Abstract
Berberine (BBR) is a natural compound with variable pharmacological effects and a broad panel of target genes. We investigated berberine’s pharmacological activities from the perspective of its nucleotide-binding ability and discovered that BBR directly regulates gene expression by targeting TATA boxes in transcriptional regulatory regions as well as the poly adenine (poly (A)) tail at the mRNA terminus. BBR inhibits gene transcription by binding the TATA boxes in the transcriptional regulatory region, but it promotes higher levels of expression by targeting the poly (A) tails of mRNAs. The present study demonstrates that TATA boxes and poly (A) tails are the first and second primary targets by which BBR regulates gene expression. The final outcome of gene regulation by BBR depends on the structure of the individual gene. This is the first study to reveal that TATA boxes and poly (A) tails are direct targets for BBR in its regulation of gene expression. Our findings provide a novel explanation for the complex activities of a small molecule compound in a biological system and a novel horizon for small molecule-compound pharmacological studies.
Similar content being viewed by others
Introduction
As a natural isoquinoline molecule, berberine (BBR) is mainly applied in intestinal infections such as bacterial enteritis and dysentery1. Modern pharmacological studies have shown that BBR has diverse biological activities, including anti-hyperlipidemia2, anti-hyperglycemia3, anti-cerebral ischemia4, prevention and treatment of neurodegeneration in Alzheimer’s disease and Parkinson’s disease5,6 and even anti-tumor effects7. However, the diversity of BBR targets that mediate these pharmacological activities are not well understood.
In the 1960s, chemists discovered BBR’s ability to bind DNA with a thymidine-adenine preference in vitro8. We previously explored BBR’s effect on TATA boxes in a biological system9. DNA TATA box elements and mRNA poly (A) tails are pivotal in the regulation of gene expression10. The fact that BBR affects both mRNA and DNA in a living cell system is an important point that has been largely ignored in the exploration of BBR’s pharmacological activities. To explain BBR’s multiple functions, we conducted a comprehensive study based on the interaction between BBR and nucleotides both in vivo and in vitro. Our findings provide novel insight into the broad pharmacological effects of BBR.
Results and discussion
Binding affinities on the nucleotides
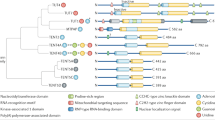
Due to BBR’s reported thymidine-adenine preference when binding to nucleotides, we first screened the binding affinities of BBR to in vitro-synthesized cis-transcriptional elements with different “TA” percentages. The binding affinity sequence, from strong to weak, was TATA box > CAAT box > GC box (Fig. 1a,b). The ability of BBR to bind TT, TA, TTAA and TATA sequences increased with increasing repetitions of the target sequences (Fig. 1c). Consistent with the “TATA” preference, BBR’s TATA binding affinity was positively correlated with the number of TATA repeats (Fig. 1d,i; Supplementary Table S1), indicating that different compositions of the “TA” variant might account for the differing effects of BBR on individual genes in vivo.
Characteristics of BBR’s affinities for DNA and RNA sequences.
(a): Controls for buffer, BBR, RNA and DNA. (b): TATA box, GC box and CAAT box. (c): Different TA compositions. (d): Different TATA boxes. In (a–d), the DNA oligos were applied at the same concentration (25 μM). (e): Poly (A). (f): Different concentration of BBR with poly (A) (20 μM). (g): Comparison of poly (A) of deoxyadenine (poly dA) (10 μM) and poly (A) of adenine (poly A) (10 μM). (h): Comparison of poly dA as a single strand (10 μM, s-30 nt) and a double strand (5 μM, d-30 bp). In (a–h), BBR was applied at the same concentration (10 μM). (i): Sequences containing TATA boxes used in (d).
BBR also binds to RNA in vitro, with a preference for adenosine (A)/uridine (U)11. BBR physically interacts with the poly (A) tail of mRNA in a concentration-dependent manner (Fig. 1e,f). BBR showed stronger binding affinity to single strand poly-adenosine than a single strand poly-deoxyadenosine (dA) (Fig. 1g; Supplementary Table S2). Nuclear magnetic resonance (NMR) spectroscopy analyses (a 2D NMR for 1H-13C cosy (HMBC)) revealed that BBR had a stronger binding affinity for single-stranded poly-adenosine, as several chemical shift changes emerged when AMP2- (monomer of poly (A)) was mixed with BBR (Supplementary Fig. S1). However, BBR had increased affinity for a complementary double strand of deoxyadenosine with deoxythymidine (dT) (Fig. 1h), indicating that BBR may bind to double-stranded DNA via a different mechanism, which might depend on the intercalation of BBR into the minor groove of the DNA double strand8. We might confirm that BBR showed the preference to a complementary double strand of deoxyadenosine with deoxythymidine and to a single stand of adenosine (poly (A)) in vitro, thus providing the targeting basis for BBR.
BBR effects on “TATA” box and poly (A) tail in wild type cells
According to the central dogma, both DNA and mRNA are important genetic materials for the expression of genetic information. The concurrence of BBR interacting with DNA double strands and mRNA poly (A) tails with a “thymidine and adenosine” preference directed us to explore the details of the mechanism by which BBR regulates gene expression12. To decipher the role of BBR’s nucleotide-binding ability in gene regulation, we used a number of genes with or without TATA boxes as models for mechanistic studies. In the wild type cells, BBR down-regulated the mRNA expression of genes containing TATA boxes in their promoter regions, including Hsp70, Il-6, Il-1β, C-fos, Pparγ and Tnfα13,14,15,16,17 (Fig. 2a–f). Conversely, BBR treatment increased the mRNA levels of genes that lack TATA boxes, including RB1, Kras, p53, Sp1, Pi3k and Akt18,19,20,21,22,23 (Fig. 2g–l). These results indicated a role for BBR binding to TATA boxes in gene promoters in the inhibition of gene transcription and indirectly suggested that BBR might also bind to poly (A) tails and consequently increase mRNA levels. When the gene transcription was inhibited by Actinomycin D, BBR protected RNA from degradation, irrespective of the presence of a TATA box (Supplementary Fig. S2). When the poly (A) tail was removed, accelerated mRNA degradation was observed, suggesting that the poly (A) tail might indeed be the target through which BBR protects mRNA from degradation (Supplementary Fig. S3).
mRNA expression of cytokines with or without a TATA box.
PC12 cells were treated with BBR for 15 h and the expression of several mRNAs was detected by qPCR. (a–f): mRNA expression of genes containing a TATA box. (g–l): mRNA expression of genes without a TATA box. The data are expressed as the mean ± S.D. from six independent experiments. *, **P < 0.05, P < 0.01 vs. control groups.
Recombination of HSP70 and Rb gene promotor to verify BBR effects
To further clarify how BBR interacts with TATA boxes and poly (A) tails, we chose as examples the HSP70 gene, which contains a TATA box24 and the Rb gene, which lacks a TATA box25. Hsp70 has a TATA box in its transcriptional regulatory region and it responds to heat stress. The expression of this gene was suppressed by BBR in vivo and in vitro (Supplementary Fig. S4). RB1 (the gene encoding Rb) is a TATA box-independent but GC box-dependent gene and it responds to ischemia-reperfusion stress. RB1 was up-regulated by BBR in vivo and in vitro, as in our previous report12.
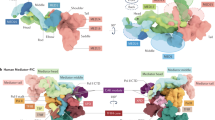
Using recombinant DNA technology, engineered plasmids containing a TATA box or a GC box were constructed (Fig. 3C; Supplementary Fig. S5). Plasmids with or without poly (A) tails were also applied to confirm the effects of BBR on poly (A) tails. The GFP gene was added to all of the plasmids to illuminate BBR’s effects without disturbing endogenous genes (Supplementary Fig. S6)26.
Plasmid construction for TATA box – GC box exchange.
(a): Fluorescence spectrum of Hsp70 with a TATA box compared with that of an Hsp70 mutant containing a “GGGCGG” sequence. BBR could clearly bind to the TATA box, increasing fluorescence, but BBR failed to increase fluorescence when the TATA sequence was replaced with GC pairs, indicating that BBR did not bind to the GC box. (b): Fluorescence spectrum of RB1 with a GC box compared with that of an RB1 mutant containing the sequence “TATAAA”. (c): Schematic of Hsp70 and RB1 promoter exchange plasmid construction. (d): Schematic of “TATAAA” and “GGGCGG” mutant plasmid construction.
Hsp70 contains both a TATA box in its promoter region and a poly (A) tail on its mRNA. The down-regulation of Hsp70 mRNA levels observed upon BBR treatment indicates that the inhibitory effect of BBR on the TATA box dominates in the presence of both a TATA box and a poly (A) tail (Fig. 4a,b). The binding affinity of BBR to the Hsp70 promoter declined when the TATA box was replaced by a GC-rich domain (Fig. 3a,b). Because of the limited distribution of BBR in cells and its lower affinity for GC-rich sequences9, it might be difficult for BBR to achieve an effective dosage to affect GC box-dependent gene transcription. In this case, we hypothesize that the activity of BBR on poly (A) tails becomes the principal effect in the absence of a TATA box, resulting in enhanced mRNA stability and up-regulation at the protein level. RB1 contains both a GC box in its transcriptional initiation region and a poly (A) tail on its mRNA. The level of RB1 mRNA increased following BBR treatment, attributable to lower mRNA degradation12. The BBR-dependent RB1 mRNA up-regulation was abolished with poly (A) depletion (Fig. 4c,d). These data collectively indicate that the TATA box is the principal target of BBR for transcriptional inhibition in vivo. However, for TATA box-independent genes, BBR can shift its primary target to the poly (A) tails of mRNAs, leading to up-regulation of protein levels through enhancement of mRNA stability.
Effect of berberine on mRNA and protein expression levels with the TATA box and GC box of Hsp70 and RB1.
(a,b): mRNA and protein expression of GFP with or without poly (A) tails transcribed by the Hsp70 promoter. (c,d): mRNA and protein expression levels of GFP with or without poly (A) tails transcribed by the RB1 promoter. (e,f): mRNA and protein expression of an Hsp70-GFP fusion protein containing the Hsp70 promoter. (g,h): mRNA and protein expression of an Hsp70-GFP fusion protein containing the RB1 promoter and a poly (A) tail. (i,j): mRNA and protein expression of an Hsp70-GFP fusion protein containing the RB1 promoter without a poly (A) tail. (k,l): mRNA and protein expression levels of a RB1-GFP fusion protein containing the Hsp70 or RB1 promoter. (m,n): mRNA and protein expression levels of a Hsp70-GFP or RB1-GFP fusion protein containing the “TATA”/“GC” sequence in the promoter. The data are expressed as the mean ± S.D. from six independent experiments. *, **P < 0.05, P < 0.01 vs. control groups. Full-length gels and blots are included in Supplementary Figure 7.
To further confirm the specificity of the TATA box target, gene cross-recombination experiments were conducted. We recombined the Hsp70 open reading frame (ORF) with the RB1 gene promoter region (containing a GC box) and simultaneously compared the effects of BBR on synthesized RNA fragments with or without a poly (A) tail. When the ORF was driven by the Hsp70 promoter (containing a TATA box), BBR treatment inhibited its expression. Conversely, when the Hsp70 promoter was replaced with the RB1 promoter, the inhibitory effect of BBR on Hsp70 expression disappeared, and, in turn, the expression of Hsp70 mRNA increased due to BBR’s effect on the poly (A) tail. When the poly (A) tail of Hsp70 (driven by the GC box-containing RB1 promoter) was removed, the increase in Hsp70 expression was abolished (Fig. 4e–j). In the opposite experiment, when the promoter region of the RB1 gene was replaced with the Hsp70 promoter region containing the TATA box, the effect of BBR on RB1 expression switched from up-regulation to inhibition because of BBR’s TATA box-mediated inhibitory effect (Fig. 4k,l).
In general, the sequence of a TATA box consists of “TATAAA” in the gene start transcription region and the sequence of a GC box consists of “GGGCGG” in the gene start transcription region. When the nucleotide sequence “TATAAA”, which is the TATA box, in the Hsp70 gene promoter region was replaced with “GGGCGG”, which is the GC box, the inhibitory effect of BBR on Hsp70 gene transcription disappeared. When the “GGGCGG” sequence in the RB1 gene promoter region was replaced with “TATAAA”, BBR acquired an inhibitory effect on RB1 gene transcription (Fig. 3d). These results demonstrate that the “TATAAA” nucleotide sequence is a functional target of BBR in its inhibition of gene transcription (Fig. 4m,n). Because of TATA box in gene transcription start site or the absence, BBR would exhibit a variety of effects.
Amplified effects of BBR on the nucleotides in pathophysiological conditions
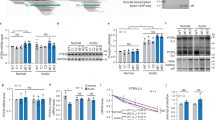
Due to the strong affinity between BBR and nucleotides, BBR might directly act to loosen packed nucleotides as soon as it enters the cell9. Furthermore, the gene structure-dependent regulatory activities of BBR might be the fundamental mechanism underlying its diverse pharmacological activities. In HepG2 cells, both GLUT2 and LDLR are GC box-dependent and poly (A) tail-containing genes27,28. BBR did not significantly influence their baseline mRNA levels under normal conditions. However, when cells suffer sugar and lipid overload, the transcriptional regions of these genes are loosened and their mRNA levels are stimulated. BBR application significantly further elevated their expression levels under these conditions. It is likely that BBR could not effectively inhibit the GC box-independent gene transcription but could prolong mRNA half-life through binding to the mRNA poly (A) tail (Fig. 5a,b), which subsequently resulted in up-regulated protein production (Fig. 5f,g). Conversely, BBR suppressed the baseline mRNA levels of TATA box-dependent and poly (A) tail-containing genes such as Hsp70, Ucp1 and Cox-229,30 under normal conditions. The application of pathophysiological models further opened the transcriptional control region of these genes and increased their expression in response to stress. Under these conditions, BBR administration further strongly inhibited the expression of these genes (Fig. 5c–e), as the TATA box-dependent inhibitory effect of BBR is its primary activity compared with its effect on mRNA. The protein production levels in response to BBR were consistent with the changes in mRNA levels (Fig. 5h–j), indicating that the translational component of gene expression might not be affected by BBR.
Expression of cytokines with or without TATA boxes in wild type cells under both physiological and pathophysiological conditions.
HepG2 cells were used for GLUT2 and LDLR, which lack TATA boxes, while PC12 cells were used for Hsp70 and Cox-2 and HIB cells were used for Ucp1, which contain TATA boxes. (a,b): mRNA expression levels of GLUT2 and LDLR. (c–e): mRNA expression of Hsp70, Ucp1 and Cox-2. (f,g): Protein expression levels of GLUT2 and LDLR. (h–j): Protein expression of HSP70, UCP1 and COX-2. (k): Differences between normal and model conditions. mRNA expression levels are displayed in green and protein expression is shown in blue. BBR had stronger effects under model conditions relative to normal conditions. BBR was applied at 0.8 μg/ml. The data are expressed as the mean ± S.D. from six independent experiments. *, **P < 0.05, P < 0.01 vs. control groups. Full-length gels and blots are included in Supplementary Figure 8.
After entering into cells, BBR could directly bind to a TATA box or a poly (A) tail. When BBR interacts with TATA boxes in gene promoters, it might hinder the reorganization of TATA-binding protein (TBP) on the TATA box, inhibiting the initiation of gene transcription. When BBR binds to poly (A) tails, it may enhance the stability of the mRNA, thus leading to a relatively higher protein level. Under pathophysiological conditions, the more loosely packed transcriptional regulatory regions allow easier access of BBR to DNA, which could lead to enhanced pharmacological activities of BBR based on its inhibitory effect on the transcription of TATA box-dependent genes. As for GC box-dependent genes, because BBR cannot reach an effective dosage to influence the transcription of these genes in living cells, the interaction between BBR and the poly (A) tail could prolong the half-life of the mRNA and subsequently maintain relatively higher protein levels (Fig. 5k), which are important for BBR’s pharmacological activity in some diseases. As an intercalator, BBR’s effects manifest as inhibited or increased gene transcription and protein production for a large number of genes. This situation occurs in pathological conditions such as venereal disease stress, indicating differences in the effects of BBR in physiological and pathological conditions.
Definition of “drug target code” of BBR
In addition to the TATA box, GC box and poly (A) tail, other gene expression-related elements composed of variant nucleotide sequences could also be targets of BBR in biological systems. Due to the limited cell-compartment distribution of BBR and the different binding affinities of BBR to adenine, cytosine, guanine and thymine, BBR has a unique activity toward each element that is determined by the individual nucleotide components. We propose that this phenomenon underlies the concept of a “drug target code”. This code is defined as the complex arrangement of drug-interacting components of a functional drug target, with the final drug activity emerging from the accumulated outcomes of all of the drug-component interactions.
Conclusion
In summary, this is the first study to comprehensively explore how BBR regulates gene expression through simultaneous direct interactions with both DNA and mRNA. BBR can interact with the TATA box transcriptional element to inhibit gene transcription and concurrently interact with mRNA in a poly (A)-dependent manner to prolong mRNA half-life, which subsequently results in relatively higher gene expression levels. The outcome of the BBR-regulated gene expression profile thus depends on the structure of the individual genes. We conclude that thymidine and adenosine are the actual targets of BBR in biological systems. Different “T/A” compositions in individual genes may function as codes that determine their accessibility to BBR, resulting in different responses to drug treatment. This evidence-supported hypothesis provides a novel understanding of the interactions between chemical compounds and biological targets, explaining the multiple activities of a single compound (such as BBR) in different physiological contexts.
Methods
Animals
The ICR mice (male, 6–8 weeks old, 20–22 g) used in this study were purchased from Vital River Laboratories (Beijing, China) and kept in an SPF (specific pathogen free) room (temperature: 25 ± 1 °C, air humidity: 50% ± 10%). Mice were fed with sterile water and the standard laboratory chow diet ad libitum. The laboratory animal facility has been accredited by AAALAC (Association for Assessment and Accreditation of Laboratory Animal Care International). All experimental procedures were approved by the IACUC (Institutional Animal Care and Use Committee) of Tsinghua University and carried out in accordance with the People’s Republic of China Legislation Regarding the Use and Care of Laboratory Animals (Approval ID: 2014-DuLJ-BBRDNRRNA).
Fluorescence measurement
The fluorescence analysis was performed in a black microplate (384-well, Corning, USA) on a Multifunctional Microplate Reader (Varioskan Flash, Thermo Scientific, USA). The excitation wavelength was 350 nm. All of the reactions were carried out in 4 mM BPES buffer (pH = 7) at room temperature, as described previously31. Blank buffer, RNA, DNA and BBR alone were also measured as background and controls. Double-stranded DNA oligos were obtained from mix of complementary oligos (equal molar concentrations) by denaturing at 95 °C for 5 min and then slow cooling to room temperature over a period of 1 h. The DNA oligonucleotides were synthesized by Sangon Biotech (Shanghai, China) and the RNA oligos with poly (A) tails by Sigma Aldrich (USA). The sequences of all DNA and RNA oligos used in these experiments are shown in Supplementary Tables 1 & 2.
Heat stress model in vivo
In the in vivo study, the mice were randomly divided into 4 groups, including two normal groups and two hot model groups. The two normal groups were kept at room temperature; one was treated with normal saline and served as a control and the other was treated with BBR (0.8 mg/kg) by intravenous administration. The two hot model groups were first injected with normal saline or BBR (0.8 mg/kg); 1 h after injection, the mice were exposed to 40 °C in a constant thermal TS-1 Incubator (Huangshi Medical Instrument, China) for 2 h32. After the end of the heat stress, the brains of the mice were immediately isolated and kept at −80 °C for RNA and protein tests.
Oxygen-glucose deprivation (OGD) in vitro
The model of OGD followed by reperfusion was used to mimic cerebral ischemia in vitro33. PC12 cells were seeded into 6-well plates and maintained in RPMI-1640 containing 10% HS and 5% FBS under 5% CO2 at 37 °C. BBR (0.8 μg/ml) was pre-treated for 15 h prior to OGD in the drug group. In the model group, the culture medium was changed to Earle’s solution without glucose and the cell plate was placed in a humidified atmosphere without oxygen (95% N2 and 5% CO2) at 37 °C for 2 h. The medium was then replaced with normal medium and the cells were maintained under normal conditions for another 6 h as reperfusion.
In vitro models of heat and cold stress
The high temperature model was used to study the cells’ heat shock responses to high temperature stress in vitro and the low temperature model was used to test responses to cold stress. PC12 cells, used for the high temperature model. And HIB cells, used for the low temperature model, were seeded into 6-well plates. BBR (0.8 μg/ml) was pre-treated for 15 h in the drug group. In the model groups, cells were exposed to a high temperature of 43 °C for 2 h or a low temperature of 4 °C for 1 h34.
In vitro models of glucose and lipid overload
The glucose and lipid overload models were, respectively used to mimic the pathological conditions of hyperglycemia and hyperlipidemia in vitro. HepG2 cells were maintained in DMEM containing 10% FBS. Cells were seeded into 6-well plates. BBR (0.8 μg/ml) was pre-treated for 2 h in the drug group. In the glucose overload model, cells were treated with 50 mg/ml glucose diluted in normal medium for 24 h. In the lipid overload model, cells were treated with 20 μg/ml Triton WR 1339 for 24 h.
Quantitative PCR
The quantitative PCR (qPCR) protocol followed that of Jiang et al.32 All primer sequences used in these analyses are shown in Supplementary Table 335,36,37,38,39,40,41,42,43,44,45,46,47,48,49.
Western blot analysis
Western blot was performed following a previously described protocol4. β-actin served as an internal control. The primary antibodies for GLUT2, COX2 and UCP1 were purchased from Bioworld (USA). Primary antibodies for HSP70, LDLR, GFP and β-actin were purchased from Abcam (UK), Proteintech (USA), Biosmart (USA) and Santa Cruz (USA), respectively. Secondary antibodies conjugated to horseradish peroxidase (HRP) were also purchased from Santa Cruz (USA).
Construction of pEGFP-N1 (ΔpolyA-)
The plasmid pEGFP-N1 was provided by Professor Ye-Guang Chen (Tsinghua University, China). The 1550–1590 poly (A) signal was deleted by amplifying nucleotides 1–1550 as Fragment 1 and nucleotides 1590–4730 as Fragment 2, double-digesting using AseI and EcoRV and then ligating into a circular plasmid, termed pEGFP-N1 (ΔpolyA−). The primer sequences were designed using Primer Premier 5.0 software as follows: Fragment 1, sense: 5′-TAGTTATTAATAGTAATCAATTACGG-3′ (AseI), antisense: 5′-ACCGATATCAGCTGCAATAAACAAGTT-3′(EcoRV); Fragment 2, sense: 5′-CATGATATCCACTGCATTCTAGTTGTG-3′ (EcoRV), antisense: 5′-GGCTATGAACTAATGACCCCGTAAT-3′. The AseI restriction site was in the PCR product, not the primer.
Construction of Hsp70/RB1-pro-Hsp70/RB1 (ΔpolyA+/−) plasmids
The Hsp70 and RB1 promoters were cloned by PCR from the genome of PC12 cells, extracted with a Genomic DNA Kit (Tiangen Biotech., China). The plasmids Hsp70-pro(ΔpolyA+) and Hsp70-pro(ΔpolyA−) were constructed by replacing the CMV promoters of pEGFP-N1 and pEGFP-N1 (ΔpolyA−) with the Hsp70 promoter, using the restriction endonucleases AseI and EcoRI. The Hsp70 and RB1 ORFs were cloned from PC12 cDNA obtained by RT-PCR. The Hsp70 and RB1 ORFs were inserted into Hsp70-pro(ΔpolyA+) using BamHI to generate the Hsp70-pro-Hsp70(ΔpolyA+) and Hsp70-pro-Rb(ΔpolyA+) plasmids. RB1-pro(ΔpolyA+), RB1-pro(ΔpolyA−), RB1-pro-Hsp70(ΔpolyA+), RB1-pro-Rb(ΔpolyA+) and RB1-pro-Hsp70(ΔpolyA−) were constructed in a similar way. Cloning primers were as follows: Hsp70 promoter sense: 5′-TGGGAGAGGAGAGTGTGTCG-3′, antisense: 5′- GGGCGGAGAAGATCTCGAAG-3′; RB1 promoter sense: 5′-TCTTTGTAGCTGGACCTGGGCCT-3′; antisense: 5′-GGGAGCCAGCGAGCTGTGGAG-3′; Hsp70 ORF sense: 5′-ATGTCGGTGGTGGGCATAGAC-3′; antisense: 5′-ATCAATGTCCATCTCAGGAAGC-3′; RB1 ORF sense: 5′-ATGCCGCCCAAAACCCCCCGAAAAACGGCC-3′; antisense: 5′-AGCATGGATACCTCAAACAAGGAAGAGAAA-3′12. 293T cells were transfected with all types of plasmids and sorted using a BD AriaIII (Becton Dickinson, USA).
Construction of Hsp70-GC-pro and RB1-TA-pro mutant plasmids
The TATA box (TATAAA) in the Hsp70 promoter was altered to a GC box (GGGCGG) and the GC box in the RB1 promoter was changed to a TATA box using overlap extension-PCR. The overlap site-direct mutagenesis was performed as previously described50. In addition to the cloning primers for the Hsp70 and RB1 promoters, the primers used for this experiment were as follows: Hsp70 site-direct mutant primer sense: 5′-ACAGTGGGCCCTGGGCGGAACTGCGGAGGGCTT-3′, antisense: 5′-AAGCCCTCCGCAGTTCCGCCCAGGGCCCACTGT-3′; RB1 site-direct mutant primer sense: 5′-GACGTGACGCCGCTATAAAAAGTGACGTTTTCCC-3′, antisense: 5′-GGGAAAACGTCACTTTTTATAGCGGCGTCACGTC-3′. The plasmids were also transfected into 293T cells and sorted by flow cytometry.
Construction of the SP64-GFP plasmid and in vitro transcription
The pSP64 polyA vector was purchased from Promega (USA). The GFP ORF was cloned from the vector pEGFP-N1 and inserted into the pSP64 polyA vector by HindIII to obtain the SP64-GFP plasmid. The primers used to clone GFP were as follows: antisense: 5′-GATCCACCGGTCGCCACCA-3′; sense: 5′-GTACAGCTCGTCCATGCCGAGAGT-3′. The plasmid SP64-GFP was separately digested with SacI or EcoRI to generate two different types of linear transcriptional templates. Two types of GFP RNA, without or with a 30-nt poly (A) tail, were obtained in a cell-free in vitro transcription system containing NTP mix, RNase inhibitor (Transgene, China) and SP64 RNA polymerase (NEB, USA). DNA templates were digested by RNase-free DNase and then RNA transcripts were purified using phenol chloroform.
Preparation of the S20 fraction from cells and RNA degradation in vitro
PC12 cells were harvested by centrifugation at 500 g for 5 min at 4 °C and washed once with ice-cold PBS. The cell pellet was suspended in ice-cold buffer A (~200 μl per 1 × 107 cells) with protease inhibitor cocktail and disrupted by beating 15 times with a sterile pestle. The nuclei were centrifuged at 2500 g for 10 min at 4 °C. The supernatant was further centrifuged at 13000 g for 30 min at 4 °C to remove cell debris. The resulting supernatant, named the S20 fraction, was stored at −80 °C51.
pEGFP-N1 and pEGFP-N1 (ΔpolyA-) were transfected and expressed in 293T cells. RNA was extracted using the RNA pre-pure Cell/Bacteria Kit (Tiangen, China) to obtain GFP mRNA with or without a poly (A) tail. The system of RNA degradation in vitro was developed as follows: 0.13 μl BBR (50 μg/ml) (substituted with water in the control group; both added before reverse transcription), 0.83 μl 100 mM ATP, 1.25 μl RNA and 4.17 μl of the PC12 S20 fraction52. The RNA degradation mixture was incubated at 37 °C for different durations (0, 15, 30 min), then terminated by adding RNase Inhibitor and placed on ice immediately. The mix was reverse-transcribed using random primers and EasyScript Reverse Transcriptase enzyme (Transgene, China). Quantitative PCR was carried out to detect mRNA decay. The degradation of GFP RNA with or without a poly (A) tail generated by in vitro transcription was also detected by this method.
NMR spectroscopy for the binding between BBR and AMP
AMP2Na (purity ≥99%) was purchased from Xiasi Biotech (Beijing, China). The NMR analyses of AMP2Na, berberine and the mixture of the two were carried out in the Analysis Center of Tsinghua University. The HMBC (Heteronuclear Multiple-Bond Correlation) spectra were obtained at 600 MHz (1H) and 150 MHz (13C) on a JNM-ECA600 spectrometer in D2O at 23.6 °C53.
Data analysis
All data were expressed as the mean ± S.D. Student’s t-tests for statistical analysis was performed using Excel Software for Office 2013 (Microsoft, U.S.). Image processing for quantitative data of protein expression was performed using Quality ONE4.62 (BIO-RAD, U.S.). P < 0.05 was considered statistically significant.
Additional Information
How to cite this article: Yuan, Z.-Y. et al. TATA boxes in gene transcription and poly (A) tails in mRNA stability: New perspective on the effects of berberine. Sci. Rep. 5, 18326; doi: 10.1038/srep18326 (2015).
References
Menees, S., Saad, R. & Chey, W. D. Agents that act luminally to treat diarrhoea and constipation. Nat. Rev. Gastroenterol. Hepatol. 9, 661–674 (2012).
Kong, W. et al. Berberine is a novel cholesterol-lowering drug working through a unique mechanism distinct from statins. Nat. Med. 10, 1344–1351 (2004).
Tang, L. Q., Wei, W., Chen, L. M. & Liu, S. Effects of berberine on diabetes induced by alloxan and a high-fat/high-cholesterol diet in rats. J. Ethnopharmacol. 108, 109–115 (2006).
Hu, J. et al. PI3K p55γ promoter activity enhancement is involved in the anti-apoptotic effect of berberine against cerebral ischemia–reperfusion. Eur. J. Pharmacol. 674, 132–142 (2012).
Durairajan, S. S. K. et al. Berberine ameliorates β-amyloid pathology, gliosis and cognitive impairment in an Alzheimer’s disease transgenic mouse model. Neurobiol. Aging 33, 2903–2919 (2012).
Kim, M. et al. Berberine prevents nigrostriatal dopaminergic neuronal loss and suppresses hippocampal apoptosis in mice with Parkinson’s disease. Int. J. Mol. Med. 33, 870–878 (2014).
Refaat, A. et al. Berberine enhances tumor necrosis factor-related apoptosis-inducing ligand-mediated apoptosis in breast cancer. Oncol. Lett. 6, 840–844 (2013).
Krey, A. K. & Hahn, F. E. Berberine: Complex with DNA. Science 166, 755–757 (1969).
Wang, Y. et al. Comprehensive study in the inhibitory effect of berberine on gene transcription, including TATA box. PLoS ONE 6, e23495 (2011).
Bae, S. H., Han, H. W. & Moon, J. Functional analysis of the molecular interactions of TATA box-containing genes and essential genes. PLoS ONE 10, e0120848 (2015).
Islam, M. M., Chowdhury, S. R. & Kumar, G. S. Spectroscopic and calorimetric studies on the binding of alkaloids berberine, palmatine and coralyne to double stranded RNA polynucleotides. J. Phys. Chem. B 113, 1210–1224 (2009).
Chai, Y. S. et al. Inhibition of retinoblastoma mRNA degradation through Poly (A) involved in the neuroprotective effect of berberine against cerebral ischemia. PLoS ONE 9, e90850 (2014).
Faggioli, L., Costanzo, C., Donadelli, M. & Palmieri, M. Activation of the interleukin-6 promoter by a dominant negative mutant of c-Jun. Biochim. Biophys. Acta 1692, 17–24 (2004).
Wang, T., Zou, J., Cunningham, C. & Secombes, C. J. Cloning and functional characterisation of the interleukin-1β1 promoter of rainbow trout (Oncorhynchus mykiss). Biochim. Biophys. Acta 1575, 108–116 (2002).
Wierstra, I. & Alves, J. FOXM1c transactivates the human c-myc promoter directly via the two TATA boxes P1 and P2. FEBS J. 273, 4645–4667 (2006).
Church, D. M. et al. Modernizing reference genome assemblies. PLoS Biol. 9, E1001091 (2011).
Zhu, W. et al. Regulation of TNF expression by multiple mitogen-activated protein kinase pathways. J. Immunol. 164, 6349–6358 (2000).
Simpson, D. J., Hibberts, N. A., McNicol, A. M., Clayton, R. N. & Farrell, W. E. Loss of pRb expression in pituitary adenomas is associated with methylation of the RB1 CpG island. Cancer Res. 60, 1211–1216 (2000).
Esa, F., Ngah, W. Z., Jamal, A. R. & Mohd Yusof, Y. A. Inhibition of beta-catenin and KRAS expressions by Piper betle in azoxymethane-induced colon cancer of male Fischer 344 rats. Anal. Quant. Cytol. Histol. 35(6), 324–334 (2013).
Thornborrow, E. C. & Manfredi, J. J. The tumor suppressor protein p53 requires a cofactor to activate transcriptionally the human BAX promoter. J. Biol. Chem. 276, 15598–15608 (2001).
Nicolás, M., Noé, V., Jensen, K. B. & Ciudad, C. J. Cloning and characterization of the 5′-flanking region of the human transcription factor Sp1 gene. J. Biol. Chem. 276, 22126–22132 (2001).
Rojo, A. I. et al. Regulation of heme oxygenase-1 gene expression through the phosphatidylinositol 3-kinase/PKC-ζ pathway and Sp1. Free Radic. Biol. Med. 41, 247–261 (2006).
Agarwal, N. K., Qu, C., Kunkulla, K., Liu, Y. & Vega, F. Transcriptional regulation of serine/threonine protein kinase (AKT) genes by glioma-associated oncogene homolog. J. Biol. Chem. 288, 15390–15401 (2013).
Wu, C. H. et al. Analysis of core promoter sequences located downstream from the TATA element in the hsp70 promoter from Drosophila melanogaster. Mol. Cell. Biol. 21, 1593–1602 (2001).
Jeon, B. N. et al. Proto-oncogene FBI-1 (Pokemon/ZBTB7A) represses transcription of the tumor suppressor Rb gene via binding competition with Sp1 and recruitment of co-repressors. J. Biol. Chem. 283, 33199–33210 (2008).
Gómez, E. et al. Construction and validation of a GFP-based vector for promoter expression analysis in the fish pathogen flavobacterium psychrophilum. Gene 497, 263–268 (2012).
Ahn, Y. H., Kim, J. W., Han, G. S., Lee, B. G. & Kim, Y. S. Cloning and characterization of rat pancreatic β-cell/liver type glucose transporter gene: A unique exon/intron organization. Arch. Biochem. Biophys. 323, 387–396 (1995).
Li, C., Kraemer, F. B., Ahlborn, T. E. & Liu, J. Induction of low density lipoprotein receptor (LDLR) transcription by oncostatin M is mediated by the extracellular signal-regulated kinase signaling pathway and the repeat 3 element of the LDLR promoter. J. Biol. Chem. 274, 6747–6753 (1999).
Kozak, L. P., Britton, J. H., Kozak, U. C. & Wells, J. M. The mitochondrial uncoupling protein gene. Correlation of exon structure to transmembrane domains. J. Biol. Chem. 263, 12274–12277 (1988).
Appleby, S. B., Ristimaki, A., Neilson, K., Narko, K. & Hla, T. Structure of the human cyclo-oxygenase-2 gene. Biochem. J. 302, 723–727 (1994).
Yadav, R. C. et al. Berberine, a strong polyriboadenylic acid binding plant alkaloid: spectroscopic, viscometric and thermodynamic study. Bioorg. Med. Chem. 13, 165–174 (2005).
Jiang, J. F. et al. Novel effect of berberine on thermoregulation in mice model induced by hot and cold environmental stimulation. PLoS ONE 8, e54234 (2013).
Li, H. et al. Comprehensive study of baicalin down-regulating NOD2 receptor expression of neurons with oxygen-glucose deprivation in vitro and cerebral ischemia-reperfusion in vivo. Eur. J. Pharmacol. 649, 92–99 (2010).
Mosser, D. D., Caron, A. W., Bourget, L., Denis-Larose, C. & Massie, B. Role of the human heat shock protein hsp70 in protection against stress-induced apoptosis. Mol. Cell. Biol. 17, 5317–5327 (1997).
Diao, L. et al. Burn Plus Lipopolysaccharide augments endoplasmic reticulum stress and NLRP3 inflammasome activation and reduces PGC-1α in liver. Shock 41, 138–144. (2014).
Ye, F. et al. Effect of topiramate on interleukin 6 expression in the hippocampus of amygdala-kindled epileptic rats. Exp. Ther. Med. 7, 223–227 (2014).
Hueston, C. M. et al. Stress-dependent changes in neuroinflammatory markers observed after common laboratory stressors are not seen following acute social defeat of the Sprague Dawley rat. Physiol. Behav. 104, 187–198 (2011).
Hou, X. et al. PPAR-γ activation by rosiglitazone suppresses angiotensin II-mediated proliferation and phenotypictransition in cardiac fibroblasts via inhibition of activation of activator protein 1. Eur. J. Pharmacol. 715, 196–203 (2013).
Liu, C. et al. The PI3K/Akt and ERK pathways elevate thyroid hormone receptor β1 and TRH receptor to decrease thyroid hormones after exposure to PCB153 and p, p′-DDE. Chemosphere 118, 229–238 (2015).
Alonso, M. et al. Anti-obesity efficacy of LH-21, a cannabinoid CB1 receptor antagonist with poor brain penetration, in diet‐induced obese rats. Br. J. Pharmacol. 165, 2274–2291 (2012).
Ramalingam, M. & Kim, S. J. Insulin involved Akt/ERK and Bcl-2/Bax pathways against oxidative damages in C6 glial cells. J. Recept. Signal Transduct. Res. 2014, e1–7 (2014). doi: 10.3109/10799893.2014.970276.
Girotti, M., Donegan, J. J. & Morilak, D. A. Chronic intermittent cold stress sensitizes neuro-immune reactivity in the rat brain. Psychoneuroendocrinology 36, 1164–1174 (2011).
Chu, C., He, W., Kuang, Y., Ren, K. & Gou, X. Celastrol protects kidney against ischemia–reperfusion-induced injury in rats. J. Surg. Res. 186, 398–407 (2014).
Zhang, N. et al. C333H ameliorated insulin resistance through selectively modulating peroxisome proliferator-activated receptor γ in brown adipose tissue of db/db Mice. Biol. Pharm. Bull. 36, 980–987 (2013).
Wu, Y. et al. Bone marrow-derived mesenchymal stem cell attenuates skin fibrosis development in mice. Int. Wound J. 11, 701–710 (2014).
Cao, X., Yang, F. Y., Xin, Z., Xie, R. R. & Yang, J. K. The ACE2/Ang-(1–7)/Mas axis can inhibit hepatic insulin resistance. Mol. Cell. Endocrinol. 393, 30–38 (2014).
Zhao, L. et al. Inflammatory stress exacerbates hepatic cholesterol accumulation via increasing cholesterol uptake and de novo synthesis. J. Gastroenterol. Hepatol. 26, 875–883 (2011).
Wang, C. Q. et al. Overexpression of Kif2a promotes the progression and metastasis of squamous cell carcinoma of the oral tongue. Oral Oncol. 46, 65–69 (2010).
Pang, Y. N. et al. Transportation of berberine into HepG2, HeLa and SY5Y cells: a correlation to its anti-cancer effect. PLoS ONE 9, e112937 (2014).
Nelson, M. D. & Fitch D. H. A. Overlap extension PCR: an efficient method for transgene construction. Molecular Methods for Evolutionary Genetics. Humana Press 459–470 (2011).
Liu, X., Kim, C. N., Yang, J., Jemmerson, R. & Wang, X. Induction of apoptotic program in cell-free extracts: Requirement for dATP and cytochrome C. Cell 86, 147–157 (1996).
Ford, L. P., Watson, J., Keene, J. D. & Wilusz, J. ELAV proteins stabilize deadenylated intermediates in a novel in vitro mRNA deadenylation/degradation system. Genes Dev. 13(2), 188–201 (1999).
Yan, C. et al. Discovery and characterization of small molecules that target the GTPase Ral. Nature 515, 443–447 (2014).
Acknowledgements
We thank all the colleagues in our laboratory. We thank Dr. M. Zhou for discussion of HMBC spectrum of AMP and berberine. This study was supported by the National Natural Science Foundation of China (81374006, 90713043and 81073092).
Author information
Authors and Affiliations
Contributions
Y.Z.Y. designed and performed experiments, analyzed data, interpreted results and wrote the paper; X.L. performed experiments. F.L. designed and performed partial experiments. Y.S.C. designed and performed experiments. Y.G.W. designed and performed partially experiments and wrote the paper. J.F.J. provided the plasmids and analyzed the data. T.S.F. performed experiments and analyzed the data. X.P.W. supplied reagents and performed experiments partially. X.Y. performed partially experiments. X.J.Y. provided the plasmid. D.M.X. supplied reagents, designed experiments and interpreted results; L.J.D. designed experiments, interpreted results and wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Yuan, ZY., Lu, X., Lei, F. et al. TATA boxes in gene transcription and poly (A) tails in mRNA stability: New perspective on the effects of berberine. Sci Rep 5, 18326 (2016). https://doi.org/10.1038/srep18326
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep18326
This article is cited by
-
Inhibitory effects of berberine on ovarian cancer: Beyond apoptosis
Medicinal Chemistry Research (2021)
-
The mitohormetic response as part of the cytoprotection mechanism of berberine
Molecular Medicine (2020)
-
Proteomic profiling of the neurons in mice with depressive-like behavior induced by corticosterone and the regulation of berberine: pivotal sites of oxidative phosphorylation
Molecular Brain (2019)
-
Study of the whole genome, methylome and transcriptome of Cordyceps militaris
Scientific Reports (2019)
-
Anti-nociceptive effect of patchouli alcohol: Involving attenuation of cyclooxygenase 2 and modulation of mu-opioid receptor
Chinese Journal of Integrative Medicine (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.