Abstract
The crab-eating frog, Fejervarya cancrivora, is the only frog that lives near seas. It tolerates increased environmental concentrations of sodium, chloride and potassium partly by raising ion and urea levels in its blood plasma. The molecular mechanism of the adaptation remains rarely documented. Herein, we analyze transcriptomes of the crab-eating frog and its closely related saline-intolerant species, F. limnocharis, to explore the molecular basis of adaptations to such extreme environmental conditions. Analyses reveal the potential genetic mechanism underlying the adaptation to salinity for the crab-eating frog. Genes in categories associated with ion transport appear to have evolved rapidly in F. cancrivora. Both positively selected and differentially expressed genes exhibit enrichment in the GO category regulation of renal sodium excretion. In this category, the positively selected sites of ANPEP and AVPR2 encode CD13 and V2 receptors, respectively; they fall precisely on conserved domains. More differentially expressed rapidly evolved genes occur in the kidney of F. cancrivora than in F. limnocharis. Four genes involved in the regulation of body fluid levels show signs of positive selection and increased expression. Significant up-regulation occurs in several genes of F. cancrivora associated with renin-angiotensin system and aldosterone-regulated sodium reabsorption pathways, which relate to osmotic regulation.
Similar content being viewed by others
Introduction
Most amphibians cannot survive in environments of greater than 10‰ salinity and no amphibian can survive permanently in seawater1. The crab-eating frog (Fejervarya cancrivora) is an exception to this tendency. It lives abundantly in coastal lowland areas and mangrove swamps of Southeast Asia where it tolerates a maximum of ~2/3 seawater and can survive in 75% or higher concentrations2. This resistance to hyperosmotic external environmental conditions generally owes to two physiological pathways: elevation of concentrations of salinity ions, especially sodium and chloride, in its blood plasma3 and accumulation of organic osmolyte-urea in tissues such as plasma and muscle via up-regulating hepatic urea synthesis2. Tadpoles of the crab-eating frog can better tolerate salinity than adults, but their osmotic regulation may be more similar to euryhaline teleosts4. The Eurasian green toad (Pseudepidalea viridis)5, the cane toad (Rhinella marinus)6 and at least one salamander (Ambystoma tigrinum)7,8,9 are also salt-tolerant. These lineages and the crab-eating frog may have functionally convergent genes10.
Amphibians use several hormonal systems to deal with salinity, including the hypothalamo–neurohypophysial, renin–angiotensin–aldosterone, adrenocorticotropic hormone–corticosteroid and natriuretic peptides systems11. Hormones of these systems can bind themselves to plasma membrane receptors by coupling with specific G proteins or cytosolic receptors to produce physiological effects11. Several amphibians increase the blood plasma parameters and concentrations of these hormones. For example, plasma osmolality in the cane toad correlates significantly with the concentration of arginine vasotocin12. Crab-eating frogs acclimated to hyperosmotic seawater elevate significantly their mean plasma concentrations of angiotensin II and aldosterone. Mean concentrations of plasma arginine vasotocin in dehydrated frogs were approximately 2.0 ~ 3.5 times higher than in control frogs13. In amphibians, aldosterone and corticosterone can increase sodium transport in organs involved in osmotic regulation, including skin, urinary bladder and kidney14,15. Thus, these hormones appear to play vital roles in amphibian osmoregulation.
Osmotic regulation of euryhaline amphibians is a complicated process that requires further exploration, especially into the molecular basis of this adaptation. Prior studies have concentrated on physiological experiments, including measuring concentrations of plasma salinity ions and different hormones1,3,13. The expression levels of some proteins also have been investigated2. Complete gene expression profiles involved in high-salinity adaptations in the crab-eating frog remain undocumented.
High-throughput expression profiling technologies (RNA-seq) have altered our visions of the extent and complexity of eukaryotic transcriptomes in species ranging from yeast to humans16. This technology facilitates investigations into the genetic bases of adaptations. It has the potential to yield insights into the genetic adaptations of the crab-eating frog to its saline environment. This approach is especially valuable for non-model species that do not have a reference genome.
Herein, we elucidate the genetic adaptations of the crab-eating frog to its saline environment. Fejervarya cancrivora and F. limnocharis are morphologically similar4 sister species17,18,19. Whereas the former is saline-tolerant the latter is saline-intolerant, as is the closely related out-group species Hoplobatrachus rugulosus17,18,19. These associations provide the basis for understanding adaptations to living in brackish water. Multi-tissue transcriptomes for these frogs yield highly confident orthologous gene pairs and produce a comprehensive landscape of expression profiles for candidate genes involved in the adaptation. Network maps of positively selected and differentially expressed genes uncover the possible genetic base of the adaptation.
Results
De novo assembly and annotation of transcriptomes
RNA-sequencing generated 982.42 million raw reads involving about 98.24 Gb of data for brain, kidney and ventral skin tissues from F. cancrivora, F. limnocharis and H. rugulosus. After quality control, ~15.79 Gb of data per species were used for the de novo assembly of transcriptomes. In total, 188565 transcripts were assembled for F. cancrivora, 180476 for F. limnocharis and 213730 for H. rugulosus. Our Contig N50s of all samples generally ranged from 955 bp to 1689 bp (Dataset 1). Thus, the quality of our de novo assemblies was similar to those of previous studies20,21.
For F. cancrivora, 188565 transcripts with the largest contig N50s were annotated by Blastx against the protein database of the draft genome of Nanorana parkeri22. In total, 65312 contigs matched 16459 genes and these covered 75.0% of the protein-coding genes (21938) of N. parkeri (Dataset 2). The transcripts were also compared to the protein database of more distantly related Xenopus tropicalis (Ensembl, JGI_4.2.75) by using Blastx. A total of 57795 contigs of F. cancrivora matched 14106 protein-coding genes (76.5%) of X. tropicalis (Dataset 3). In total, 11548 protein-coding genes (81.9%) mapped to GO categories. We identified 55 main GO categories in three levels (23 GO terms in Biological Process, 14 GO terms in Molecular Function and 18 GO terms in Cellular Component) (Fig. 1 and Dataset 4) using WEGO23, a web tool for plotting GO annotations.
Rapid evolution of coding sequences in F. cancrivora
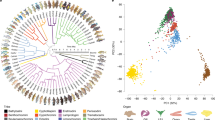
Analyses of orthologous genes suggested rapid evolution occurred in genes potentially involved in the adaptation to seawater. Among the 8698 predicted one-to-one orthologous gene pairs among the three species of frogs, 7173 were annotated as coding genes of X. tropicalis. After sequence alignment and trimming, 6959 high-confidence orthologous genes remained for estimating the evolutionary constraints acting on F. cancrivora, F. limnocharis and H. rugulosus (Fig. 2a)17,18,19. We calculated the dN/dS value for each GO term in each branch by the free ratio model implemented in PAML424. In total, 54 GO categories harbored significantly higher dN/dS values in F. cancrivora than in F. limnocharis (P < 0.05, binomial test). These included transmembrane transport (GO:0055085, P = 2.31e-32), transporter activity (GO:0005215, P = 7.05e-14), transport (GO:0006810, P = 6.45e-08), cation transport (GO:0006812, P = 1.84e-04), cellular calcium ion homeostasis (GO:0006874, P = 3.38e-4), blood vessel development (GO:0001568, P = 0.002), electron carrier activity (GO:0009055, P = 0.004), sodium ion transport (GO:0006814, P = 0.004) and ion transport (GO:0006811, P = 0.009) (Fig. 2b and Dataset 5). In contrast, 62 GO categories exhibited accelerated evolutionary rates in F. limnocharis and these mainly involved metabolism, such as fatty acid metabolic process (GO:0006631, P = 1.19e-10), fatty acid beta-oxidation (GO:0006635, P = 1.39e-06), triglyceride biosynthetic process (GO:0019432, P = 4.83e-05) and fatty acid biosynthetic process (GO:0006633, P = 9.28e-05) (Dataset 5).
Evolutionary analyses of Fejervarya cancrivora and its closely related species, F. limnocharis.
(a) The species-tree. (b) Comparison of dN/dS ratios between F. cancrivora and F. limnocharis by GO functional categories. Blue and red dots represent categories with an elevated evolutionary rates in F. limnocharis and F. cancrivora, respectively. (c) The dN/dS of differentially expressed genes in kidney of F. cancrivora compared to F. limnocharis. Red (F. cancrivora) and blue (F. limnocharis) bars denote dN/dS of differentially expressed genes and grey (F. cancrivora) and pink (F. limnocharis) bars give dN/dS of background genes. (d) Venn diagram of up-regulated genes in kidney, ventral skin and positively selected genes in F. cancrivora compared to F. limnocharis.
Positive selection in F. cancrivora
The branch-site model executed in PAML424 identified 186 candidate positively selected genes among the 6959 orthologous genes (2.7%) (Dataset 6) that possibly associated with the adaptation to high salinity. ANPEP and AVPR2 were significantly enriched in the negative regulation of renal sodium excretion (P = 2.41E-03, FDR) (Table 1 and Dataset 7). ANPEP encodes aminopeptidase N (CD13) and AVPR2 encodes the type 2 vasopressin receptor. Further, positively selected sites Q257S in ANPEP (sites 73–468, Peptidase_M1_N, IPR014782) and G178N in AVPR2 (sites1–202, GPCR_Rhodpsn_7TM, IPR017452) were located in conserved domains. Candidate genes ATP6V1F, ATP6V1G3 and ATP5A, which are involved in hydrogen-exporting ATPase activity (P = 3.707e-02, FDR), were also enriched significantly. The two v-type-H+-ATPase enzymes (Table 1) might be associated with the adaptation to high salinity.
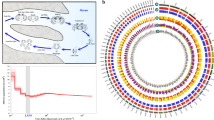
We mapped the candidate positively selected genes to the protein-protein interaction network database (InnateDB)25 to understand their biological functions. Sub-networks of more than five nodes were retained. In total, 129 seed proteins (queries) (69.4.5%) mapped to a sub-network consisting of 1756 nodes (proteins) and 2611 edges (protein-protein interactions). Positively selected XRCC6, VIM, TUFM, ALB and SNRNP70 had the strong ability to interact with other positively selected genes and non-seed proteins (degrees > 100) (Fig. 3). The protein encoded by ALB mainly functioned in the regulation of the colloidal osmotic pressure of blood; it may have played an important role in the plasma osmotic pressure of F. cancrivora.
Evolution of gene expression in F. cancrivora
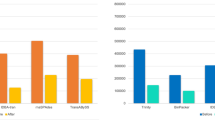
Because changes in gene expression have been considered to play more important roles in phenotypic evolution than mutations in protein-coding sequences26, we identified differentially expressed genes between F. cancrivora and its closely related species F. limnocharis. Kidney harbored far more differentially expressed genes (3239) (Dataset 8) than ventral skin (1104) (Dataset 9) and brain (35) (Fig. 4 and Dataset 10). In F. cancrivora the dN/dS values of differentially expressed genes in kidney were significantly higher than those of background genes (P = 0.0019, Mann-Whitney U test), but this pattern was not found in F. limnocharis (P = 0.5869, Mann-Whitney U test). This discovery suggested that the differentially expressed genes in F. cancrivora evolved rapidly (Fig. 2c). Further, in F. cancrivora a higher number of positively selected genes were found to harbor elevated expressions in kidney (39 positively selected genes) compared with ventral skin (24 positively selected genes) and brain (1 positively selected gene). Gene enrichment analysis of up-regulated genes in kidney found categories such as transport, regulation of urine volume, regulation of renal sodium excretion, regulation of excretion, sodium ion homeostasis and regulation of vasodilation to be over-represented (Dataset 11). Although this might be associated with adaptation to the high salinity of sea-water, the exact functions of changes in expression remained largely unclear and in need of experimental verification. Consistently, up-regulated genes in the ventral skin also showed similar enrichment. In contrast, down-regulated genes generally were involved in macromolecule metabolic processes and macromolecule modification (Dataset 12).
Visualization of differentially expressed genes in brain (a), ventral skin (b) and kidney (c) between Fejervarya cancrivora and F. limnocharis.
X-axis represents differential folds of genes (log FC) and the Y-axis shows -log P value of differentially expressed genes. Red denotes significantly up- and down-regulated genes in F. cancrivora compared to F. limnocharis. Blue denotes show genes without significant differences in levels of expressions.
The 19 positively selected genes that exhibited significantly increased levels of expression in the ventral skin and kidney of F. cancrivora were also screened (Figs 2d and 5). Genes ALB, EHHADH, CFL1and PLSCR1 involved in the regulation of body fluid levels exhibited significant enrichment (P = 0.0122, FDR) (Dataset 13). In addition, the renin-angiotensin system has long been regarded as an important regulator of systemic blood pressure and renal electrolyte homeostasis and generates a family of bioactive angiotensin peptides with varying biological activities27,28. Here, AGT, PREP, ANPEP and CTSA were found to be up-regulated in the crab-eating frog and enriched significantly in the renin-angiotensin system pathway (P = 0.0051, FDR). Further, NR3C2, PIK3CG, PDPK1 and SCNN1G showed significant up-regulation and they were included in the aldosterone-regulated sodium reabsorption pathway, which was a downstream target pathway of the renin-angiotensin system.
Expression profiles of 19 positively selected and significantly up-regulated genes in kidney (a) and in skin (b) of Fejervarya cancrivora compared to F. limnocharis.
Heatmaps show distinct expression profiles of all 19 candidate genes in kidney and ventral skin. The fpkm values were normalized by log2 (fpkm+1) and each row represents a differentially expressed gene. Blue highlights ALB, EHHADH, CFL1and PLSCR1, which associate with regulation of body fluid levels and ANPEP, which associates with negative regulation of renal sodium excretion.
Discussion
All findings for differentially expressed genes indicate that kidney might contribute more to salt adaptation than the other tested tissues. This discovery is congruent with a previous study26 in that overall differentially expressed genes may play a larger role than positively selected genes in adaptations.
Both positively selected and differentially expressed genes exhibit enrichment in the regulation of renal sodium excretion pathway. Renal sodium excretion was found to influence arterial blood pressure by affecting blood volume and electrolyte composition; inversely, blood pressure was coupled to sodium excretion through negative-feedback regulation by the renin-angiotensin-aldosterone system29. In this category, ANPEP appears to have evolved under positive selection and even experienced a significantly increased level of expression in F. cancrivora. The gene encodes aminopeptidase N (CD13), a membrane-bound protein that catalyzes the formation of natriuretic hexapeptide angiotensin IV (ANG IV) from ANG III. It was reported to be a candidate gene for handling renal salt in Rattus norvegicus30,31,32. Thus, this gene may play a primary role in the tolerance of high salinity by F. cancrivora. Candidate AVPR2 appears to have evolved under positive selection. It encodes the type 2 vasopressin receptor, which is a member of the vasopressin/oxytocin receptor subfamily of G protein-coupled receptors33. Disruption of AVPR2 causes X-linked congenital nephrogenic diabetes insipidus (NDI)34,35. Further, NDI can prevent the concentration of urine after the administration of the antidiuretic hormone arginine-vasopressin34. Because hormonal systems were shown to make important contributions to the regulation of osmotic equilibrium in amphibians13, positive selection on this gene may drive functional changes in excretion. More importantly, significantly positively selected sites of the two positively selected genes fall precisely into conserved domains. This also implies important functional changes for their protein-activities. Functional experiments can document their importance.
ATP6V1F and ATP6V1G3 associate with ion transport and in particular with hydrogen-exporting ATPase activity. Functionally, they mediate sodium transport in the mitochondria-rich (MR) cells of amphibian skin as an energizer of Na+ uptake. They are considered essential in the maintenance of amphibian body salt11,36. Thus, the evidence of positive selection on these genes suggests that the crab-eating frog possesses strong if not enhanced capacities for ion transport.
Analyses do not detect an increase in expression for 186 positively selected genes in brain, ventral skin and kidney of F. cancrivora relative to closely related F. limnocharis. This observation suggests that only a few up-regulated positively selected genes may play key roles in the adaptation of F. cancrivora to salinity. A group of associated up-regulated positively selected genes involve the osmotic organs kidney and ventral skin. The regulation of body fluid levels involves ALB, EHHADH, CFL1 and PLSCR1. They show variation in their protein-coding sequences and also harbor strong evidence of up-regulated expression. In particular, ALB encodes serum albumin, a major plasma protein that regulates osmotic pressure of blood plasma. The colloid osmotic properties of the albumin’s facilitate plasma volume expansion37,38. It appears to be an important hub-gene (degree >100) directing construction of the protein-protein interaction network. Consequently, ALB may be involved in the adaptation to high-salinity by F. cancrivora. EHHADH was shown to be highly expressed in proximal tubules and associated with energy supply in reabsorption39. Thus, synchronous changes in sequences and expression in genes that control body fluid levels may play extraordinary roles in the regulation of the osmotic pressure of blood plasma and the maintenance of electrolytic homeostasis.
The renin-angiotensin-aldosterone system plays a primary role in the hormonal osmoregulation of amphibians13. In expression levels, except for ANPEP with its changes in sequence and expression, AGT, PREP and CTSA also exhibit significant enrichment and markedly higher levels of expressions. Among these genes, AGT encodes the precursor protein angiotensinogen40, CTSA encodes a protein involved in the conversion of angiotensin I to angiotensin II41 and PREP plays a role in the degradation of angiotensin I and II42. Thus, they are important candidate genes for participating in the maintenance of osmotic equilibrium. Importantly, NR3C2, PIK3CG, PDPK1 and SCNN1G, which associate with the aldosterone-regulated sodium reabsorption pathway as the downstream target pathway of renin-angiotensin system pathway, also possess significantly elevated levels of gene expression. In particular, NR3C2 encodes MR (mineralocorticoid receptor), which can be activated by mineralocorticoid-aldosterone to regulate Na+ homeostasis, largely through multiple mechanisms that modulate the activity of the epithelial Na+ channel (ENaC)43. Thus, the candidate genes of both pathways plus the candidate positively selected genes (Fig. 6) very likely constitute adaptations of crab-eating frog to their hypertonic environment. Further functional experimentation is necessary to verify this implication.
Interactions of positively selected genes and differentially expressed genes involved in the adaptation of F. cancrivora to high salinity.
Red italics denote candidate positively selected genes, blue italics indicate significantly up-regulated genes and green italics genes experiencing positive selection and increased expression. (1) AVPR2, expressed mainly in kidney tubules, primarily serves to respond to the pituitary hormone arginine vasopressin (AVP) to maintain water homeostasis; production of ALB can facilitate blood plasma volume expansion, which serves to regulate blood pressure. (2) AGT, PREP, ANPEP and CTSA play roles in the renin-angiotenin system pathway; NR3C2, PIK3CG, PDPK1 and SCNN1G associate with aldosterone-regulated sodium reabsorption pathway. (3) EHHADH provides energy for tubule reabsorption through the oxidation of fatty acids in the proximal tubule; ATP6V1G3 and ATP6V1F are v-type-H+ ATPases and they participate in Na+ uptake as energizers.
In summary, our analyses discover positively selected and differentially expressed genes. Both variation in coding sequence and gene expression probably contribute to the adaptation of crab-eating frogs to their saltwater environment. The candidate genes and variants are promising for future studies on high salinity adaptation in poikilothermic animals. Investigations into the functions of these genes and mechanism underlying associated pathways, such as regulation of renal sodium excretion, will make valuable contributions to understanding the system.
Materials and Methods
The methods were carried out in accordance with approved guidelines.
Sample collection and RNA isolations of these tissues
Care and treatment of the frogs complied with the guidelines for the National Care and Use of Animals approved by the National Animal Research Authority (P.R. China). The experimental protocols involving live animals were approved by the Ethics and Experimental Animal Committee of Kunming Institute of Zoology, Chinese Academy of Science, China (Approval ID: SYDW-2014013).
Fejervarya cancrivora and F. limnocharis were collected in mangrove swamps, flooded rice paddies or roadside ditches in Hainan, China. Hoplobatrachus rugulosus was acquired in Guangxi, China. Frogs were euthanized by using an intraperitoneal injection of sodium pentobarbital at a dosage of 100 mg per kg body weight. After euthanasia, their brains, kidneys and ventral skin were biopsied quickly, placed in liquid nitrogen and later stored at −80 °C upon return to the laboratory. Total RNA of these tissues was extracted using TRIzol reagent (Invitrogen Corp., Carlsbad, CA). RNA purifications were performed using an RNeasy Mini Kit (Qiagen, Chatsworth, CA).
Preparation and sequencing of sample libraries and quality control of transcriptomes
Library constructions from F. cancrivora (two biological replicates), F. limnocharis (two biological replicates) and H. rugulosus (no replicates) were made using Illumina Hiseq2000 RNA sample preparation kits (Illumina, San Diego, CA). After preparing the sequencing libraries from brain, kidney and ventral skin tissues from all three species, we used a 2100-Bioanlyzer (Agilent Technologies) to assess their quality. The libraries were then sequenced on an Illumina Hiseq2000 instrument (Illumina, Inc.) as 2*101 bp. To ensure quality of sequencing, analyses used three lanes for all 15 libraries. The sequencing reads were submitted through the NCBI SRA database and can be accessed via NCBI BioProject accession SRP045588. Sequencing adaptors used for cDNA library construction were trimmed using Cutadapt (version_1.2.1)44. We employed Btrim64 (version_0.1.0)45 to delete regions with average quality scores of less than 20 and impose a minimal length equal to or greater than 20 bp. The cleaned reads were applied to de novo assembly using Trinity46 (version_2013_08_14) and its default parameters.
Annotation of transcripts and prediction of open reading frames
First, we used Blastx47 with parameters, -evalue 1e-5 and -max_target_seqs 1, (version_ ncbi-blast-2.2.27+) to align de novo contigs against the protein database of Nanorana parkeri (Anuran: Dicroglossidae)22 and protein database of Xenopus tropicalis (ftp://ftp.ensembl.org/pub/release-75/fasta/xenopus_tropicalis/pep/) to obtain a best hit (BH) of sequence alignment and to produce reliable annotations of the transcriptomes. Because the protein-coding databases of X. tropicalis and N. parkeri did not include mitochondrial genes, downstream analyses were based on nuclear coding genes only. Next, coding sequences of F. cancrivora, F. limnocharis and H. rugulosus were predicted based on the assumption that the longest open reading frame in the longest transcript per gene had a greatest chance of being a protein-coding region. GETORF in EMBOSS48 (version_6.4.0) was applied to obtain the nucleic sequences between start and stop codons and to limit the minimum size of a fragment to 120 bp. Next, transeq in EMBOSS48 (version_6.4.0) was used to obtain translated protein sequences. The longest protein sequences per gene in three species of frogs were used to identify orthologous genes. We used OrthoMCL49 (version_2.0.9) to identify orthologous genes using a Markov Cluster algorithm. Downstream analyses used orthologous genes predicted and well annotated for X. tropicalis only.
Calculation of evolutionary rate
We used the dN/dS ratio to measure the evolutionary rate along a lineage. The values of dN, dS and dN/dS ratio were estimated using the free-ratio model (Parameters: model = 1, NSsites = 0, fix_omega = 0, omega = 1) in PAML424 for each branch. Lineage-specific mean values were estimated by concatenated alignments from all orthologs. GO term data were downloaded from BioMart (Ensembl, JGI_4.2.75) and only GO categories with more than 20 orthologs were included in our analyses. GO terms experiencing relatively accelerated evolution were identified using a binomial test50.
Identification of positively selected genes
Orthologous genes were aligned by PRANK51 (Parameters: -f = fasta -F -codon -noxml -notree -nopost). We aligned the longest predicted ORFs for the longest transcript-pairs across the three species. Gblocks52,53 (version_0.91b; Parameters: -t = c -b3 = 1 -b4 = 6 -b5 = n) was employed to reduce the rate of false positive predictions due to potential sequencing errors, incorrect alignments and non-orthologous regions based on codons. After trimming, we removed fragment lengths shorter than 100 bp to obtain one-to-one orthologous genes. We calculated the ratio of nonsynonymous (Ka or dN) to synonymous (Ks or dS) substitution rate (ω = Ka/Ks or dN/dS) using CODEML in the PAML4 package24. The accepted species-tree ((F. cancrivora, F. limnocharis), H. rugulosus) was used as the guide tree17,18,19. Branch-site model24 (Parameters: Null hypothesis: model = 2, NSsites = 2, fix_omega = 1, omega = 1; Alternative hypothesis: model = 2, NSsites = 2, fix_omega = 0, omega = 1) was used to detect positively selected genes. A likelihood ratio test compared the alternative hypothesis of positive selection on the foreground branch to a null hypothesis with no positive selection on the background branch for each orthologous gene. Positively selected genes were inferred only if their P values were less than 0.05. Positively selected sites were deduced by Bayes Empirical Bayes (BEB) analyses54. Functionally conserved domains of positively selected genes were predicted by InterProsan55 (version_5.2-45.0).
Detection of differentially expressed genes between F. cancrivora and F. limnocharis
RSEM56 (version_1.2.3), an accurate transcript quantification tool for RNA-Seq data with or without a reference genome, was utilized to quantify the transcripts. Briefly, we viewed the longest orthologous transcript sequences per species of frog as the reference databases and then applied rsem-calculate-expression to map trimmed pair-end reads of brain, kidney and ventral skin onto it. For the same tissues, we used the output of RSEM as input for EdgeR57, an R software library of bioconductor (http://www.bioconductor.org), to detect differentially expressed genes between F. cancrivora and F. limnocharis. Analyses considered genes whose false discovery rate (FDR) values, using the BH approach58, were less than or equal to 0.05 to be differentially expressed. The heat maps were drawn as follows: function “heatmap.2” of the “gplots” library of R package.
Gene Ontology enrichment analyses
We identified GO categories at three levels (biological process, molecular function and cellular component) and pathways that were over-represented by positively selected genes and differentially expressed genes using the MetaCore59 pipeline. Enrichment P-values were calculated. All P-values were multi-normalized FDR using the BH approach58. Categories with a FDR of less than 0.05 were considered over-represented.
Protein-Protein Interaction (PPI) Network
We mapped the candidate positively selected genes to protein-protein interaction network database (InnateDB). Sub-network topologies were obtained by NetworkAnalyst (http://www.networkanalyst.ca/NetworkAnalyst)60. The sub-networks with node counts ≤5 were filtered and statistics of the total nodes, edges and seed proteins were obtained from the mapping overview.
Additional Information
How to cite this article: Shao, Y. et al. Transcriptomes reveal the genetic mechanisms underlying ionic regulatory adaptations to salt in the crab-eating frog. Sci. Rep. 5, 17551; doi: 10.1038/srep17551 (2015).
References
Schmidt-Nielsen, K. & Lee, P. Kidney function in the crab-eating frog (Rana cancrivora). J Exp Biol. 39, 167–177 (1962).
Wright, P. et al. The crab-eating frog, Rana cancrivora, up-regulates hepatic carbamoyl phosphate synthetase I activity and tissue osmolyte levels in response to increased salinity. J Exp Zool A Comp Exp Biol. 301, 559–568 (2004).
Gordon, M. S., Schmidt-Nielsen, K. & Kelly, H. M. Osmotic regulation in the crab-eating frog (Rana cancrivora). J Exp Biol. 38, 659–678 (1961).
Gordon, M. S. & Tucker, V. A. Osmotic regulation in the tadpoles of the crab-eating frog. J Exp Biol. 42, 437–445 (1965).
Gordon, M. S. Osmotic regulation in the green toad. J Exp Biol 39, 261–270 (1962).
Liggins, G. W. & Grigg, G. C. Osmoregulation of the cane toad, Bufo marinus, in salt water. Comp Biochem Physiol A Comp Physiol. 82, 613–619 (1985).
Kirschner, L. B., Kerstetter, T., Porter, D. & Alvarado, R. H. Adaptation of larval Ambystoma tigrinum to concentrated environments. Am J Physiol. 220, 1814–1819 (1971).
Romspert, A. P. & McClanahan, L. L. Osmoregulation of the Terrestrial Salamander, Ambystoma tigrinum, in hypersaline media. Copeia. 1981, 400–405 (1981).
Gasser, K. W. & Miller, B. T. Osmoregulation of larval blotched tiger salamanders, Ambystoma tigrinum melanostictum, in saline environments. Physiol Zool. 59, 643–648 (1986).
Gordon, M. S. & Notar, J. C. Can systems biology help to separate evolutionary analogies (convergent homoplasies) from homologies? Prog Biophys Mol Biol. 117, 19–29 (2015).
Uchiyama, M. & Konno, N. Hormonal regulation of ion and water transport in anuran amphibians. Gen Comp Endocrinol. 147, 54–61 (2006).
Konno, N., Hyodo, S., Takei, Y., Matsuda, K. & Uchiyama, M. Plasma aldosterone, angiotensin II and arginine vasotocin concentrations in the toad, Bufo marinus, following osmotic treatments. Gen Comp Endocrinol. 140, 86–93 (2005).
Uchiyama, M. et al. Changes in plasma angiotensin II, aldosterone, arginine vasotocin, corticosterone and electrolyte concentrations during acclimation to dry condition and seawater in the crab-eating frog. Gen Comp Endocrinol. 195, 40–46 (2014).
Stiffler, D. F., De, R. M. L., Hanson, P. B. & Marshall, M. Interrenal function in larval Ambystoma tigrinum. I. responses to alterations in external electrolyte concentrations. Gen Comp Endocrinol. 62, 290–297 (1986).
Van, D. W. & Zeiske, W. Ionic channels in epithelial cell membranes. Physiol Rev. 65, 833–903 (1985).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet. 10, 57–63 (2009).
Kotaki, M. et al. Molecular phylogeny of the diversified frogs of genus Fejervarya (Anura: Dicroglossidae). Zoolog Sci. 27, 386–395 (2010).
Kurabayashi, A., Kuramoto, M., Joshy, H. & Sumida, M. Molecular phylogeny of the ranid frogs from Southwest India based on the mitochondrial ribosomal RNA gene sequences. Zoolog Sci. 22, 525–534 (2005).
Ren, Z. et al. Complete nucleotide sequence and gene arrangement of the mitochondrial genome of the crab-eating frog Fejervarya cancrivora and evolutionary implications. Gene. 441, 148–155 (2009).
Looso, M. et al. A de novo assembly of the newt transcriptome combined with proteomic validation identifies new protein families expressed during tissue regeneration. Genome Biol. 14, R16 (2013).
Yang, W., Qi, Y., Bi, K. & Fu, J. Toward understanding the genetic basis of adaptation to high-elevation life in poikilothermic species: a comparative transcriptomic analysis of two ranid frogs, Rana chensinensis and R. kukunoris. BMC Genomics. 13, 588 (2012).
Sun, Y. B. et al. Whole-genome sequence of the Tibetan frog Nanorana parkeri and the comparative evolution of tetrapod genomes. Proc Natl Acad Sci USA. 112, E1257–1262 (2015).
Ye, J. et al. WEGO: a web tool for plotting GO annotations. Nucleic Acids Res. 34, W293–297 (2006).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol. 24, 1586–1591 (2007).
Breuer, K. et al. InnateDB: systems biology of innate immunity and beyond–recent updates and continuing curation. Nucleic Acids Res. 41, D1228–1233 (2013).
Wray, G. A. The evolutionary significance of cis-regulatory mutations. Nat Rev Genet. 8, 206–216 (2007).
Campbell, D. J. The renin-angiotensin and the kallikrein-kinin systems. Int J Biochem Cell Biol. 35, 784–791 (2003).
Engeli, S., Negrel, R. & Sharma, A. M. Physiology and pathophysiology of the adipose tissue renin-angiotensin system. Hypertension. 35, 1270–1277 (2000).
Toney, G. M., Vallon, V. & Stockand, J. D. Intrinsic control of sodium excretion in the distal nephron by inhibitory purinergic regulation of the epithelial Na(+) channel. Curr Opin Nephrol Hypertens. 21, 52–60 (2012).
Kotlo, K. et al. Functional polymorphism of the Anpep gene increases promoter activity in the Dahl salt-resistant rat. Hypertension. 49, 467–472 (2007).
Kotlo, K., Shukla, S., Tawar, U., Skidgel, R. A. & Danziger, R. S. Aminopeptidase N reduces basolateral Na+ -K+ -ATPase in proximal tubule cells. Am J Physiol Renal Physiol. 293, F1047–1053 (2007).
Rangel, R. et al. Impaired angiogenesis in aminopeptidase N-null mice. Proc Natl Acad Sci USA. 104, 4588–4593 (2007).
Oksche, A. & Rosenthal, W. The molecular basis of nephrogenic diabetes insipidus. J Mol Med. 76, 326–337 (1998).
Bichet, D. G. et al. X-linked nephrogenic diabetes insipidus: from the ship Hopewell to RFLP studies. Am J Hum Genet. 51, 1089–1102 (1992).
Wildin, R. S., Cogdell, D. E. & Valadez, V. AVPR2 variants and V2 vasopressin receptor function in nephrogenic diabetes insipidus. Kidney Int. 54, 1909–1922 (1998).
Ehrenfeld, J. Active proton and urea transport by amphibian skin. Comp Biochem Physiol A Mol Integr Physiol. 119, 35–45 (1998).
Convertino, V. A., Brock, P. J., Keil, L. C., Bernauer, E. M. & Greenleaf, J. E. Exercise training-induced hypervolemia: role of plasma albumin, renin and vasopressin. J Appl Physiol Respir Environ Exerc Physiol. 48, 665–669 (1980).
Gillen, C. M. et al. Plasma volume expansion in humans after a single intense exercise protocol. J Appl Physiol. 71, 1914–1920 (1991).
Klootwijk, E. D. et al. Mistargeting of peroxisomal EHHADH and inherited renal Fanconi’s syndrome. N Engl J Med. 370, 129–138 (2014).
Armesto, P. et al. Molecular characterization and transcriptional regulation of the renin-angiotensin system genes in Senegalese sole (Solea senegalensis Kaup, 1858): differential gene regulation by salinity. Comp Biochem Physiol A Mol Integr Physiol. 184, 6–19 (2015).
Miller, J. J., Changaris, D. G. & Levy, R. S. Conversion of angiotensin I to angiotensin II by cathepsin A isoenzymes of porcine kidney. Biochem Biophys Res Commun. 154, 1122–1129 (1988).
Irazusta, J., Silveira, P. F., Gil, J., Varona, A. & Casis, L. Effects of hydrosaline treatments on prolyl endopeptidase activity in rat tissues. Regul Pept. 101, 141–147 (2001).
Quinn, S., Harvey, B. J. & Thomas, W. Rapid aldosterone actions on epithelial sodium channel trafficking and cell proliferation. Steroids. 81, 43–48 (2014).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. J EMBnet. 17, 10–12 (2011).
Kong, Y. Btrim: a fast, lightweight adapter and quality trimming program for next-generation sequencing technologies. Genomics. 98, 152–153 (2011).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol. 29, 644–652 (2011).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J Mol Biol. 215, 403–410 (1990).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet. 16, 276–277 (2000).
Li, L., Stoeckert, C. J. Jr. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Consortium, C. S. a. A. Initial sequence of the chimpanzee genome and comparison with the human genome. Nature. 437, 69–87 (2005).
Loytynoja, A. & Goldman, N. webPRANK: a phylogeny-aware multiple sequence aligner with interactive alignment browser. BMC Bioinformatics. 11, 579 (2010).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 17, 540–552 (2000).
Talavera, G. & Castresana, J. Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol. 56, 564–577 (2007).
Yang, Z., Wong, W. S. & Nielsen, R. Bayes empirical Bayes inference of amino acid sites under positive selection. Mol Biol Evol. 22, 1107–1118 (2005).
Syed, A. & Upton, C. Java GUI for InterProScan (JIPS): a tool to help process multiple InterProScans and perform ortholog analysis. BMC Bioinformatics. 7, 462 (2006).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 12, 323 (2011).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 26, 139–140 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy Stat Soc, Ser B. 57, 289–300 (1995).
Bessarabova, M., Ishkin, A., JeBailey, L., Nikolskaya, T. & Nikolsky, Y. Knowledge-based analysis of proteomics data. BMC Bioinformatics. 13 Suppl 16, S13 (2012).
Xia, J., Benner, M. J. & Hancock, R. E. NetworkAnalyst–integrative approaches for protein-protein interaction network analysis and visual exploration. Nucleic Acids Res. 42, W167–174 (2014).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China [grant number31321002] and the Animal Branch of the GermplasmBank of Wild Species of Chinese Academy of Sciences (the Large Research Infrastructure Funding).We thank Wei-wei Zhou for helpful discussions and comments and Jie-Qiong Jin for sample collection and technical help in the lab.
Author information
Authors and Affiliations
Contributions
J.C., Y.P.Z. and D.D.W. conceived and designed the experiments. Y.S. analyzed the data and wrote the manuscript. L.J.W., M.L.H. and H.M.C. collected the data samples. L.Z. performed the separation, purification and sequencing of RNA. R.W.M. guided and revised this paper. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Shao, Y., Wang, LJ., Zhong, L. et al. Transcriptomes reveal the genetic mechanisms underlying ionic regulatory adaptations to salt in the crab-eating frog. Sci Rep 5, 17551 (2015). https://doi.org/10.1038/srep17551
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep17551
This article is cited by
-
Hepatic transcriptome of the freeze-tolerant Cope’s gray treefrog, Dryophytes chrysoscelis: responses to cold acclimation and freezing
BMC Genomics (2020)
-
Transcriptome analysis reveals the genetic basis underlying the development of skin appendages and immunity in hedgehog (Atelerix albiventris)
Scientific Reports (2020)
-
Transcriptome analysis reveals the genetic basis underlying the seasonal development of keratinized nuptial spines in Leptobrachium boringii
BMC Genomics (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.