Abstract
Autosomal dominant polycystic kidney disease (ADPKD) is one of the most frequently inherited renal diseases caused by mutations in PKD1 and PKD2. We performed mutational analyses of PKD genes in 49 unrelated patients using direct PCR-sequencing and multiplex ligation-dependent probe amplification (MLPA) for PKD1 and PKD2. RT-PCR analysis was also performed in a family with a novel PKD2 splicing mutation. Disease-causing mutations were identified in 44 (89.8%) of the patients: 42 (95.5%) of the patients showed mutations in PKD1 and 2 (4.5%) showed mutations in PKD2. Ten nonsense, 17 frameshift, 4 splicing and one in-frame mutation were found in 32 of the patients. Large rearrangements were found in 3 patients and missense mutations were found in 9 patients. Approximately 61.4% (27/44) of the mutations are first reported with a known mutation rate of 38.6%. RNA analysis of a novel PKD2 mutation (c.595_595 + 14delGGTAAGAGCGCGCGA) suggested monoallelic expression of the wild-type allele. Furthermore, patients with PKD1-truncating mutations reached end-stage renal disease (ESRD) earlier than patients with non-truncating mutations (47 ± 3.522 years vs. 59 ± 11.687 years, P = 0.016). The mutation screening of PKD genes in Chinese ADPKD patients will enrich our mutation database and significantly contribute to improve genetic counselling for ADPKD patients.
Similar content being viewed by others
Introduction
Autosomal dominant polycystic kidney disease (ADPKD) is one of the most frequently inherited renal diseases worldwide with an estimated incidence of 1:400 to 1:1,000 and is characterized by bi-lateral renal cysts in the liver, seminal vesicles, pancreas and arachnoid membrane, as well as extra-kidney abnormalities1. ADPKD is a common kidney disorder in China, affecting more than 1.3 million Chinese individuals. Approximately 10–20% of ADPKD children are hypertensive and the majority of adults have hypertension with normal kidney function2. Approximately 50% of individuals with ADPKD will progress to end-stage renal disease (ESRD) by the age of 60 years and these patients account for 5–8% of the renal replacement population3.
ADPKD is a heterogeneous monogenic disorder resulting from mutations in two genes: PKD1 and PKD2. The PKD1 gene (OMIM 601313), located in chromosome region 16p13.3, consists of 46 exons with an open reading frame of 12,912 bp that encodes the 4,303-amino acid peptide polycystin-1 (PC1)4. Exons 1–33 of PKD1 are duplicated approximately six times at the homologous genes (HGs), which has made the genetic analysis of PKD1 challenging5. PKD2 (MIM 173910, chromosome region 4q21-22) is a single-copy gene including 15 exons with a 2,907-bp coding sequence and is predicted to encode a 968-amino acid peptide called polycystin-2 (PC2)6. PC1 and PC2 act as flow-dependent mechanosensors in renal primary cilium that regulate the differentiated state of tubular epithelial cells7. Clinical data show that mutations in PKD1 and PKD2 account for 85% and 15% of all cases of ADPKD, respectively8. Compared to PKD2 mutations (where the average age of ESRD onset is 74 years), PKD1 mutations (where the average age of ESRD onset is 54.3 years) are associated with the more serious form of the disease9.
Diagnosis of ADPKD is performed mainly by renal ultrasound, computed tomography (CT) or magnetic resonance imaging (MRI) but cannot exclude the disease in at-risk individuals until the age of 40 years, especially in families with mutations in PKD210. Molecular diagnostics play a significant role in confirming a definite diagnosis, especially in young renal donors, patients with a negative family history, individuals with early onset ADPKD or atypical symptoms and for subjects with affected relatives11. There is no hotspot mutation for PKD1 or PKD2, indicating that mutations are usually unique to a single family and are highly variable12. The structural complexity of PKD1 and the high allelic heterogeneity of PKD genes make clinical molecular diagnostics difficult13. Although a few studies on novel mutations in the PKD1 and PKD2 genes in Chinese patients have been carried out with different methods, including PCR-single-strand conformation polymorphism (PCR-SSCP)14, denaturing high-performance liquid chromatography (DHPLC)15 and next-generation sequencing (NGS)16, direct sequencing is undoubtedly the gold standard for accurately identifying the majority of PKD mutations17. Multiplex ligation-dependent probe amplification (MLPA) was developed to detect large genomic rearrangements in PKD genes that cannot be detected by sequencing18.
Analysis of the pathogenicity of variants of an uncertain significance plays an important role in the molecular diagnosis of ADPKD because of the high level of genetic variation found in the PKD1 gene19. So far, approximately 2,322 sequence variants of PKD1 and 278 sequence variants of PKD2 have been reported in the Autosomal Dominant Polycystic Kidney Disease Mutation Database (PKDB, http://pkdb.mayo.edu; January 2015). The Human Gene Mutation Database (HGMD) has recorded 1,177 sequence variants of PKD1 and 211 sequence variants of the PKD2 gene (http://www.hgmd.cf.ac.uk). However, mutation data for PKD genes from different populations would provide a better interpretation of genetic testing results.
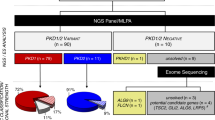
In the present study, we performed long-range PCR (LR-PCR) followed by nested PCR and MLPA of PKD1 and PKD2 in 49 Chinese patients with a definite diagnosis of ADPKD. A group of novel mutations in PKD1 and PKD2 is described in this paper. All mutation data detected will contribute to better diagnostics and genetic counselling in a clinical setting.
Results
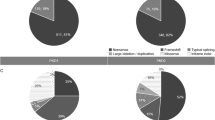
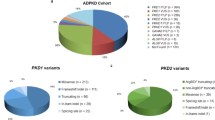
We performed complete mutational analysis by direct sequencing and MLPA analysis of PKD1 and PKD2 in 49 unrelated patients with the diagnosis of ADPKD obtained by ultrasound. Thirty-two definitely pathogenic variants and 12 likely pathogenic variants (42 variants in PKD1 and 2 in PKD2) were found in 44 patients (Table 1), resulting in a variant detection rate of 89.8% (44/49). The variants we found represent 61.4% (27/44) novel variants and 38.6% (17/44) known variants. The remaining 5 patients (10.2%) were classified with an undetermined genotype either because no variant was detected or the identified variant was considered to be nonpathogenic. Figure 1 shows the distribution of definitely pathogenic and likely pathogenic variants found in the PKD1 gene. We further confirmed that there are no variant hotspots in PKD1 gene.
Schematic representation of polycystin-1, representing the location of mutations found in this study or previous reports (adapted by permission from Macmillan Publishers Ltd: [NATURE GENETICS], Hugheset al.19954).
Nonsense mutations and frameshift mutations are indicated with arrows. The positions of in-frame deletions and splicing changes are indicated with boxes (G0576: c.8017-2-1delAG; G0649: c.8157-8159delCAC; G1410: c.10220 + 2T > C; G0053: c.10617 + 1delG). Likely pathogenic mutations are indicated with triangles (1–9: see Table 3 for details).
Definite pathogenic mutations
Definite pathogenic mutations were found in 32 of the families and included 10 nonsense mutations, 17 frameshift mutations, 2 splicing mutations and 3 large rearrangements. These disease-causing mutations are shown in Table 2. The percentage of our ADPKD patients without a family history was 12.2% (6/49). We analysed PKD1 and PKD2 in the parents of six probands with no family history of ADPKD and found that in five instances, the pathogenic mutations occurred de novo in the probands. No pathogenic mutation of PKD1 or PKD2 was found in the other patient without a family history ADPKD.
RT-PCR analysis of PKD2 mRNA in patient G0904
A novel splicing mutation in PKD2 (c.595_595 + 14delGGTAAGAGCGCGCGA) was found in patient G0904 and was predicted to affect the splice site of the gene. The predicted absence of PKD2 exon 2 would produce a premature termination codon (PTC) downstream in exon 3 and therefore could not escape nonsense-mediated mRNA decay (NMD)20,21. To evaluate the influence of the c.595_595 + 14del, RT-PCR analysis of total RNA extracted from peripheral blood mononuclear cells (PBMCs) from patient G0904 and a healthy control was performed using the primers PKD2-E1F/PKD2-E2R and PKD2-E1F/PKD2-E4R. It is worth noting that we could not detect aberrant PKD2 transcripts in patient G0904 (Fig. S1A). However, heterozygosity of rs2728118 (c.420G > A) in exon 1 of PKD2 was only found in the genomic DNA (GA) and not in the cDNA of the patient (AA) (Fig. S1C). rs2728118 was inherited from the patient’s father and the c.595_595 + 14del mutation was inherited from the patient’s mother (Fig. S1B). rs2728118 is located upstream of the c.595_595 + 14del mutation and is distributed in trans on the chromosome. Further quantitative PCR (qPCR) analysis indicated that the patient PKD2 mRNA level was reduced to 55.97 ± 1.78% (P = 5.44 × 10−5) compared to the normalized control mRNA level (Fig. S1D). These results suggested monoallelic expression of the wild-type allele, inheritance form the patient’s father and nonsense-mediated mRNA decay (NMD) of the aberrantly spliced PKD2 transcripts.
Likely pathogenic mutations
A total of 22 unclassified variants and 12 previously reported polymorphisms (Table S1) were detected in our patients. We evaluated the pathogenic potential of the unclassified variants using web-based prediction programs. The results of the evaluation are presented in Table 3. Twelve likely pathogenic mutations (11 PKD1 mutations and 1 PKD2 missense mutation) were identified in 12 patients. Among these mutations, one small novel in-frame mutation in PKD1 (c.8157-8159delCAC) showed high evolutionary conservation and two novel splicing variants in PKD1 (c.10220 + 2T > C and c.10617-1delG) were predicted to affect the splice site and were found to segregate with the disease in affected families. Therefore, these three mutations in PKD1 are highly likely to be pathogenic mutations. Two novel substitutions PKD1 (p.R3046C and p.Y3819N) coexisted in patient G0241 and were predicted in silico to be likely pathogenic and likely polymorphic, respectively. Amino acid multi-alignments demonstrated that the positions of the mutations are highly conserved across species for both p.R3046C and p.Y3819N (Fig. 2). Therefore, we speculate that p.R3046C and p.Y3819N are hypomorphic alleles. Segregation of the two variants with the disease was not validated due to the lack of blood samples from the families.
Two PKD1 missense mutations (p.R3046C and p.Y3819N) coexisted in a Chinese patient with ADPKD.
(a) The sequencing pattern of p.R3046C and p.Y3819N in patient G0241. Both sequences are in the forward direction. The bottom line corresponds to the mutated sequence; the upper line represents the wild-type sequence. (b) Multiple sequence alignments for the PKD1 variant changes p.R3046C and p.Y3819N.
Large deletion mutation
To identify large deletions or duplication mutations that cannot be detected by Sanger sequencing, we performed a copy number analysis of the probands without pathogenic mutations in the PKD genes (Fig. S2). Using MLPA, three large deletions of the PKD1 gene were found in patients G0677, G0018 and G1800. Deletion of PKD1 exon 1 (relative peak ratio 0.52) was found in patient G0677, a 45-year-old man and a similar result was found in his affected family members. Deletion of PKD1 exon 21 (relative peak ratio 0.51) was identified in patient G1800 and his affected mother. Patient G1800 is 26 years old and was found to have bilateral renal cysts at approximately 18 years of age. The renal function of patient G1800 is well controlled except for slight hypertension. DNA sequence analysis of exons 1 and 21 demonstrated the absence of single-base mutations under the oligonucleotide probe. Patient G0018, who is 20 years old and without a family history of ADPKD, showed a deletion for probes 1 to 30. Large deletion mutations segregated with the disease in all of the affected family members tested, although q-PCR confirmation was not performed at the nucleotide level due to the sequence complexity of the PKD1 locus.
Genotype/phenotype correlation
Of the 49 probands, 32 had already reached ESRD (Table 4). We therefore performed a Kaplan-Meier survival curve analysis to investigate whether the type of PKD1 mutation (non-truncating mutations including missense and small in-frame mutations vs. truncating mutations) influenced the age of ESRD onset. As shown in Fig. 3, the age of ESRD onset in patients with PKD1-truncating mutations (n = 23) was earlier than that of the patients with non-truncating PKD1 mutations (n = 9) (log-rank test, P = 0.016). The median age of ESRD onset in the patients with PKD1-truncating mutations and patients with non-truncating mutations was 47 years (95% CI, 47 ± 3.522 years) and 59 years (95% CI, 59 ± 11.687 years), respectively.
Discussion
To date, 2,322 pathogenic mutations for PKD1 and 278 for PKD2 have been reported in the PKDB. In our study, we analysed 49 ADPKD families using DNA sequencing and MLPA with a mutation detection rate of 89.8%. Among the 44 mutations we found, 61.4% (27/44) are novel mutations and 38.6% (17/44) are known mutations. The 44 pathogenic mutations we identified occurred in unrelated patients, suggesting that each patient had a unique pathogenic mutation. Our findings are consistent with previous studies showing that less than 2% of unrelated ADPKD-affected families carry the same mutation22. Our results indicate that no mutation hotspot exists in either PKD1 or PKD2. Therefore, a complete mutation analysis of PKD1 and PKD2 is needed for Chinese patients with ADPKD who require a genetic diagnosis. The 27 novel mutations we found in the Chinese population will enrich the PKD mutation database and significantly contribute to the genetic counselling of ADPKD patients.
Patients with PKD2 mutations typically develop ESRD two decades later than those with mutations in PKD19. Therefore, it is of prognostic value to determine the location of the mutation in an affected family (PKD1 or PKD2). The mutational distribution in the PKD2 gene could not be determined because only two exonic mutations were found. The mutation detection rate of PKD2 (4.1%) in our study was much lower than the average percentage (15%)8. This difference may be due to the low number of patients analysed in our study compared to previous studies by Rossetti et al.
Previous studies have shown that the mutation type and location in PKD1 may influence renal survival23. We detected a total of 44 pathogenic variations in the PKD1 gene; of these, 9 (20.9%) are located in exon 15, which is consistent with the findings recorded in the PKDB (277/1,272). This region corresponds to the junction of the PKD repeats and the REJ domain of the resultant protein. In our study, 78.6% of the mutations in PKD1 were predicted to truncate the protein, including frameshift mutations, nonsense mutations, splicing mutations and large deletions. The high frequency of these mutations is in concordance with recent results from Cornec-Le Gall et al., who showed that approximately two thirds of PKD1 mutation-positive pedigrees carry truncating mutations24. Cornec-Le Gall et al. reported that carriers of a PKD1-truncating mutation have a significantly earlier age of ESRD onset than patients with a non-truncating mutation (55 years vs. 67 years). Our data support the view that a more severe phenotype can be expected in patients with a PKD1-truncating mutation. The exception was patient G0599, who carried the known missense mutation p.C155Y but also had received a right renal transplant at 26 years old.
Two novel PKD1 mutations (p.R3046C and p.Y3819N) were found in a female patient with ADPKD and it is probable that both mutations are hypomorphic alleles. However, DNA samples from the parents of this patient were not available. Moreover, the likely pathogenic mutations identified in our study should be confirmed in future studies including additional ADPKD families.
Large genomic rearrangements account for approximately 6.8% of the pathogenic mutations in the PKD1 gene and an even smaller percentage of PKD2 gene mutations18. In our study, we identified three large deletions: one involving deletion of exon 21, one involving deletion of exons 1 to 30 and a large deletion of exon 125. Our data are consistent with previously reported data.
The high level of allelic heterogeneity in both the PKD1 and PKD2 genes and the prevalence of private mutations in ADPKD patients imply that there is a high frequency of de novo mutations in this disease. Indeed, approximately 10% of adult ADPKD patients do not have a family history of the disease26. The percentage of our ADPKD patients without a family history of disease was 12.2%, which is consistent with previously reported results. Five different pathogenic mutations occurred de novo in probands without a family history of ADPKD.
Pathogenic mutations were not found in 10.2% of our 49 unrelated patients, a result in accordance with those reported in a previous CRISP study (10.9%)8. This finding may be because mutations occur within deep intronic regions, as well as in promoters and other distantly located regulatory regions not covered by the current exon-based sequencing method. Alternatively, some of the missense mutations that are classified as nonpathogenic mutations by the prediction system may represent hypomorphic alleles. Such variants alone may result in only mild cystic disease, but two such variants in trans may cause disease27,28. Additionally, we only screened PKD1 and PKD2 mutations and therefore cannot exclude the existence of other mutations that might contribute to the cystic phenotype, such as those in HNF1b, PRKCSH, SEC63 or PKHD129. Furthermore, mosaicism may influence the genotype and phenotype of ADPKD; however, this condition is usually not detected in screenings and a significant proportion of de novo mosaic mutations may be missed30,31. The likely pathogenic mutations identified in our study need to be confirmed in future studies with additional ADPKD families.
Conclusions
In our study, 27 novel pathogenic mutations in the PKD genes were detected in 49 Chinese individuals. A novel splicing mutation in PKD2 (c.595_595 + 14delGGTAAGAGCGCGCGA) was confirmed to be definitely pathogenic. Patients carrying PKD1 mutations, especially those with truncating mutations, could have a more rapidly progressive disease than those with non-truncating mutations. Our study will enrich the mutation database of PKD and significantly contribute to the genetic counselling of ADPKD patients.
Methods
Patients
A total of 49 unrelated patients were enrolled from the Women’s Hospital School of Medicine Zhejiang University from October 2010 to December 2013. Patients were definitively diagnosed with ADPKD based on the criteria recommended by Ravine D et al.32. All patients provided written informed consent and their family and medical histories were recorded. The general clinical data of the probands are summarized in Table 5. Peripheral blood samples were collected from all probands and their family members when possible. The study was performed with the approval of the Ethics Committee of the Women’s Hospital School of Medicine Zhejiang University. The study was conducted in adherence to the Declaration of Helsinki.
Mutation analysis of PKD1 and PKD2
Genomic DNA was extracted from peripheral blood samples using the QIAGEN spin columns on a QIACube (QIAGEN GmbH) according to the manufacturer’s instructions. The mutational screening of PKD1 and PKD2 via Sanger sequencing was carried out in the 49 probands. LR-PCR followed by nested PCR was adopted for the mutational analysis of the PKD1 gene5. The duplicated region of PKD1 was amplified in eight specific long fragments by LR-PCR (exon 1, 2–7, 8–12, 13–15, 15–21, 22, 23–28 and 29–34)33 using TaKaRa LA Taq TM (TaKaRa Bio Inc.). Exons 1–34 of PKD1 were then amplified by nested PCR with these LR templates and exons 35–46 were directly PCR amplified and sequenced in both directions. Exons 1–15 of PKD2 including the adjacent 30–50 bp intronic sequence was amplified from genomic DNA according to a previous report with little modification34. PCR amplification primers for the various LR-PCR fragments are provided in Supplemental Table S1.
MLPA
MLPA analysis was performed with a SALSA MLPA KIT P351-B1/P352-B1 PKD1-PKD2 kit (MRC-Holland, Amsterdam, the Netherlands) according to the manufacturer’s instructions. The kit covers PKD1 probes for exons 37 to 46, its upstream regulatory sequences, all PKD2 exons and three exons of the TSC2 gene adjacent to the PKD1 gene35. The result of our MLPA analysis was scanned using the ABI PRISM® 3100 Genetic Analyzer (Applied Biosystems). The data were analysed using the Coffalyser MLPA analysis tool (MRC-Holland, Amsterdam, the Netherlands).
RT-PCR analysis of PKD2 mRNA in PBMCs
Total RNA from PBMCs was extracted and reverse transcribed from patient G0904 and a healthy control. RT-PCR was performed with primers PKD2-E1F/PKD2-E2R and PKD2-E1F/PKD2-E4R. qPCR was performed with primers PKD2-E13F/PKD2-E14R and the results were normalized to glyceraldehyde-3-phosphate dehydrogenase (GAPDH), according to the method described previously36. RT-PCR and qPCR primers are listed in Table S3.
Data analysis and sequence variation classification
Mutations of PKD genes were analysed using Mutation Surveyor® software. Nucleotide changes were nominated according to the NCBI reference sequences of PKD1 (NM_001009944.2) and PKD2 (NM_000297.3). The HGMD (http://www.hgmd.cf.ac.uk), the Exome Sequencing Project (http://evs.gs.washington.edu/EVS) and PKDB (http://pkdb.mayo.edu) were checked for previously reported sequence changes. Novel mutations in this study were assessed for their pathogenic potential. Nonsense or frameshift variants resulting in a PTC were identified to be definitely pathogenic. The pathogenicity of missense variants was computationally evaluated with the SIFT, PolyPhen2 and AlignGVGD prediction programs by analysing interspecies sequence variations37,38,39. The NetGene2 (http://www.cbs.dtu.dk/services/NetGene2/)40 and Human Splicing Finder (HSF) (http://www.umd.be/HSF/)41 software were used to evaluate the splice site mutations. All variations analysed by these web-based software programs were finally sorted into four categories: likely pathogenic, indeterminate, likely polymorphic and polymorphic. Only gene variations that were predicted to be damaging by SIFT, PolyPhen-2 and AlignGVGD were considered to be “likely pathogenic”, as long as no other definite mutation was found in the same patient. If a definite mutation coexisted with a damaging missense mutation in the same patient, the missense mutation was considered to be “indeterminate”. Similarly, only the variations that were scored as benign by all the software programs were considered to be “polymorphic”. Otherwise, the mutations were classified as “likely polymorphic”. Furthermore, pedigree co-segregation analysis of the potential pathogenic mutations in PKD genes was examined in all available members of the probands’ families (including healthy individuals). Likely pathogenic missense or splice site variants would segregate within the affected family.
Statistical analysis
Cumulative renal survival curves were generated using the Kaplan–Meier method and compared using the log-rank test. P < 0.05 was considered to be statistically significant. All analyses were performed using SPSS 17.0.
Additional Information
How to cite this article: Liu, B. et al. Identification of novel PKD1 and PKD2 mutations in Chinese population with autosomal dominant polycystic kidney disease. Sci. Rep. 5, 17468; doi: 10.1038/srep17468 (2015).
Change history
23 February 2016
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Torres, V. E. & Harris, P. C. Mechanisms of Disease: autosomal dominant and recessive polycystic kidney diseases. Nat. Clin. Pract. Nephrol. 2, 40–55; quiz 55 (2006).
Rossetti, S. et al. Incompletely penetrant PKD1 alleles suggest a role for gene dosage in cyst initiation in polycystic kidney disease. Kidney Int. 75, 848–855 (2009).
Braun, W. E. Autosomal dominant polycystic kidney disease: emerging concepts of pathogenesis and new treatments. Cleve. Clin. J. Med. 76, 97–104 (2009).
Hughes, J. et al. The polycystic kidney disease 1 (PKD1) gene encodes a novel protein with multiple cell recognition domains. Nature genetics 10, 151–160 (1995).
Watnick, T. J. et al. An unusual pattern of mutation in the duplicated portion of PKD1 is revealed by use of a novel strategy for mutation detection. Human Molecular Genetics 6, 1473–1481 (1997).
Mochizuki, T. et al. PKD2, a gene for polycystic kidney disease that encodes an integral membrane protein. Science 272, 1339–1342 (1996).
Tan, Y. C. Autosomal dominant polycystic kidney disease : Genetics, mutations and microRNAs. Biochimica et Biophysica Acta 1812, 1202–1212 (2011).
Rossetti, S. et al. Comprehensive molecular diagnostics in autosomal dominant polycystic kidney disease. J. Am. Soc. Nephrol. 18, 2143–2160 (2007).
Kurashige, M. et al. A comprehensive search for mutations in the PKD1 and PKD2 in Japanese subjects with autosomal dominant polycystic kidney disease. Clin. Genet. 87, 266–272 (2015).
Pei, Y. et al. Unified criteria for ultrasonographic diagnosis of ADPKD. J. Am. Soc. Nephrol. 20, 205–212 (2009).
Trujillano, D. et al. Diagnosis of autosomal dominant polycystic kidney disease using efficient PKD1 and PKD2 targeted next-generation sequencing. Mol. Genet. genomic Med. 2, 412–421 (2014).
Gout, A. M. et al. PKDB: Polycystic Kidney Disease Mutation Database—A Gene Variant Database for Autosomal Dominant Polycystic Kidney Disease. Hum. Mutat. 28, 654–659 (2007).
Tan, A. Y. et al. Molecular Diagnosis of Autosomal Dominant Polycystic Kidney Disease Using Next-Generation Sequencing. J. Mol. Diagnostics 16, 216–228 (2014).
Ding, L. et al. Novel mutations of PKD1 gene in Chinese patients with autosomal dominant polycystic kidney disease. Nephrol Dial Transplant 17, 75–80 (2002).
Yu, C. et al. Identification of novel mutations in Chinese Hans with autosomal dominant polycystic kidney disease. BMC Med. Genet. 12, 164 (2011).
Yang, T. et al. Identification of novel mutations of PKD1 gene in Chinese patients with autosomal dominant polycystic kidney disease by targeted next-generation sequencing. Clin. Chim. Acta 433, 12–19 (2014).
Liu, G. et al. Development and validation of a whole genome amplification long-range PCR sequencing method for ADPKD genotyping of low-level DNA samples. Gene 550, 131–135 (2014).
Consugar, M. B. et al. Characterization of large rearrangements in autosomal dominant polycystic kidney disease and the PKD1/TSC2 contiguous gene syndrome. Kidney Int. 74, 1468–1479 (2008).
Garcia-Gonzalez, M. A. et al. Evaluating the clinical utility of a molecular genetic test for polycystic kidney disease. Mol. Genet. Metab. 92, 160–167 (2007).
Maquat, L. E. Nonsense-mediated mRNA decay: splicing, translation and mRNP dynamics. Nat. Rev. Mol. Cell Biol. 5, 88–99 (2004).
Khajavi, M. et al. Nonsense-mediated mRNA decay modulates clinical outcome of genetic disease. Eur. J. Hum. Genet. 14, 1074–1081 (2006).
Audrézet, M. P. et al. Autosomal dominant polycystic kidney disease: comprehensive mutation analysis of PKD1 and PKD2 in 700 unrelated patients. Hum. Mutat. 33, 1239–1250 (2012).
Rossetti, S. et al. The Position of the Polycystic Kidney Disease 1 (PKD1) Gene Mutation Correlates with the Severity of Renal Disease. J. Am. Soc. Nephrol. 13, 1230–1237 (2002).
Cornec-Le Gall, E. et al. Type of PKD1 Mutation Influences Renal Outcome in ADPKD. J. Am. Soc. Nephrol. 24, 1006–1013 (2013).
Chang, M. Y. et al. Novel PKD1 and PKD2 mutations in Taiwanese patients with autosomal dominant polycystic kidney disease. J. Hum. Genet. 58, 720–727 (2013).
Neumann, H. P. et al. Adult patients with sporadic polycystic kidney disease : the importance of screening for mutations in the PKD1 and PKD2 genes. Int Urol Nephrol. 44, 1753–1762 (2012).
Rossetti, S. et al. Incompletely penetrant PKD1 alleles suggest a role for gene dosage in cyst initiation in polycystic kidney disease. Kidney Int. 75, 848–855 (2009).
Pei, Y. et al. A missense mutation in PKD1 attenuates the severity of renal disease. Kidney Int. 81, 412–417 (2012).
Obeidova, L. et al. Novel mutations of PKD genes in the Czech population with autosomal dominant polycystic kidney disease. BMC Med. Genet. 15, 41 (2014).
Reiterová, J. et al. Autosomal dominant polycystic kidney disease in a family with mosaicism and hypomorphic allele. (2013). BMC Nephrol. 14, 59 (2013).
Tan, A. Y. et al. Autosomal dominant polycystic kidney disease caused by somatic and germline mosaicism. Clin. Genet. 87, 373–377 (2014).
Ravine, D. et al. Evaluation of ultrasonographic diagnostic criteria for autosomal dominant polycystic kidney disease 1. Lancet. 343, 824–827 (1994).
Phakdeekitcharoen, B., Watnick, T. J. & Germino, G. G. Mutation Analysis of the Entire Replicated Portion of PKD1 Using Genomic DNA Samples. J. Am. Soc. Nephrol. 12, 955–963 (2001).
Tan, Y. C. et al. Novel Method for Genomic Analysis of PKD1 and PKD2 Mutations in Autosomal Dominant Polycystic Kidney Disease. Hum. Mutat. 30, 264–273 (2008).
Schouten, J. P. et al. Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acids Res. 30, e57 (2002).
Zhang, J. et al. A Functional Alternative Splicing Mutation in AIRE Gene Causes Autoimmune Polyendocrine Syndrome Type 1. PLoS One 8, e53981 (2013).
Ramensky, V. et al. Human non-synonymous SNPs : server and survey. Nucleic Acids Res. 30, 3894–3900 (2002).
Thusberg, J. et al. Performance of Mutation Pathogenicity Prediction Methods on Missense Variants. Hum. Mutat. 32, 358–368 (2011).
Pauline, C. Accounting for Human Polymorphisms Predicted to Affect Protein Function Accounting for Human Polymorphisms Predicted to Affect Protein Function. Genome Research. 12, 436–446 (2002).
Brunak, S. & Engelbrecht, J. Prediction of Human mRNA Donor and Acceptor Sites from the DNA Sequence. J. Mol. Biol. 220, 49–65 (1991).
Desmet, F. O. et al. Human Splicing Finder : an online bioinformatics tool to predict splicing signals. Nucleic Acids Res. 37, e67 (2009).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (NO. 81471506; NO. 81170620; NO. 81401219), the Shanghai Municipal Commission of Science and Technology Program (NO. 14411965100; NO. 15411966700) and the Research Project of the Chinese Ministry of Education (NO. 113038A).
Author information
Authors and Affiliations
Contributions
B.L., S.C.C., C.M.X. and H.H.F. conceived of this study and drafted the manuscript. B.L., S.C.C., J.Y.Z. and Y.T.H. carried out the LR-PCR and nested PCR assays. Y.M.Y. and K.Y. performed the MLPA analysis. Y.Q.Q. and M.Y.D. helped collect the clinical samples. F.J. participated in the design of the study. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Liu, B., Chen, SC., Yang, YM. et al. Identification of novel PKD1 and PKD2 mutations in a Chinese population with autosomal dominant polycystic kidney disease. Sci Rep 5, 17468 (2015). https://doi.org/10.1038/srep17468
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep17468
This article is cited by
-
Potential role for protein kinase D inhibitors in prostate cancer
Journal of Molecular Medicine (2023)
-
Genetic Characteristics of Korean Patients with Autosomal Dominant Polycystic Kidney Disease by Targeted Exome Sequencing
Scientific Reports (2019)
-
Novel mutations of PKD genes in Chinese patients suffering from autosomal dominant polycystic kidney disease and seeking assisted reproduction
BMC Medical Genetics (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.