Abstract
Vitellogenesis or maternal yolk formation is considered critical to the reproduction of egg-laying animals. In invertebrates, however, most of its regulatory genes are still unknown. Via a combined mapping and whole-genome sequencing strategy, we performed a forward genetic screen to isolate novel regulators of yolk production in the nematode model system Caenorhabditis elegans. In addition to isolating new alleles of rab-35, rab-10 and M04F3.2, we identified five mutant alleles corresponding to three novel regulatory genes potently suppressing the expression of a GFP-based yolk reporter. We confirmed that mutations in vrp-1, ceh-60 and lrp-2 disrupt endogenous yolk protein synthesis at the transcriptional and translational level. In contrast to current beliefs, our discovered set of mutants with strongly reduced yolk proteins did not show serious reproduction defects. This raises questions as to whether yolk proteins per se are needed for ultimate reproductive success.
Similar content being viewed by others
Introduction
In vertebrates, the hypothalamus-pituitary-gonad (HPG) axis is a regulatory system responsible for the neuroendocrine control of reproduction. One downstream process regulated by this HPG axis in oviparous females is the synthesis of egg-yolk proteins by the liver, which are required for oogenesis1,2. Yolk proteins are among the most abundant proteins in eggs3,4. They are primarily produced by the mother in a tissue outside the gonad, are heavily glycosylated and have lipid-binding properties3. As such, they provide essential nutrients to the eggs to support embryonic development4,5,6. Most invertebrate species also rely on vitellogenesis, i.e. yolk build-up in the maturing oocytes, for reproduction. To date, only scattered pieces of information are available on the (hormonal) regulation of protostome yolk protein gene expression in response to environmental conditions4,7,8,9. It can be expected that genetic control of reproduction in invertebrates will - to a certain extent - be similar to the vertebrate system, as supported by homologous gene sequences found in distinct species. Yet, many invertebrate-, clade- or species-specific factors are thought to exist as well - e.g. depending on distinct reproductive cycles. To explore genetic determinants of invertebrate yolk protein production and their evolutionary conservation, we relied on the nematode model species Caenorhabditis elegans and its powerful genetic toolbox, which readily enables the identification of new mutations in key genes and the dissection of molecular genetic pathways.
Putative orthologues of the components of the vertebrate HPG axis have previously been reported for the worm10,11,12,13, giving first indications of evolutionary conservation of at least a part of the reproductive control system. Starting from an unbiased screen for mutants defective in vit-2 (vitellogenin) gene expression as visualized by a gfp reporter gene, we here report on three molecular players (i.e. VRP-1, CEH-60 and LRP-2) previously unknown to regulate yolk protein synthesis. We further explored the transcriptional interdependency between these regulators and discuss the unexpected low impact of massive yolk protein production on overall reproductive success.
Results
Genetic ethyl methanesulfonate (EMS) screen
The C. elegans genome harbours six vit genes, all encoding yolk proteins14,15. These are subdivided into two classes - YP170 and YP88/115 - named according to their approximate molecular weight15,16,17. We created a reporter strain expressing a functional fluorescent VIT-2(YP170B)::GFP fusion protein which is, like endogenous yolk, transported from the intestine into the growing oocytes6,18,19,20,21.
Starting from this strain, roughly 7,500 EMS-mutagenized haploid genomes were manually analysed during a phenotypic selection step based on aberrant gfp expression. This ranged from complete loss of the gfp signal (five mutants) over reduced gfp expression (one mutant) to abnormal accumulation of GFP (three mutants) (Table 1, Supplementary Fig. S1). The latter three display a typical receptor-mediated endocytosis phenotype, noticeable by abnormal accumulation of fluorescence in the body cavity, outlining the internal organs, with little or no fluorescence in the oocytes and embryos6.
Mapping and allele characterization
To elucidate the phenotype-causing recessive mutations, we relied on a one-step WGS and SNP-based mapping procedure originating from a successful approach in plants22,23. We retrieved for each mutagenized sequenced genome an unambiguous map position (Supplementary Fig. S1). WGS of the starting strain allocated transgene lstIs13 [Pvit-2::vit-2::gfp] to the 5.90–5.92 Mb region on chromosome X. We next looked into all possible causative variants within each mutant’s mapping region (Supplementary Table S1). Some protein coding sequence variants - i.e. premature stops, frameshifts and splice site variants - are of special interest as a result of their allegedly higher impact on protein function.
We isolated three mutant alleles of genes previously described to be involved in yolk (receptor) endocytic trafficking24,25, consistent with our observations of their characteristic receptor-mediated endocytosis mutant phenotype. These are a novel missense mutation in rab-35 (RAB family), a novel premature stop in rab-10 and a premature stop in M04F3.2 (Table 1). The isolation of these mutants provides proof of concept of our genetic screen.
Despite the premature stop codon (W222amber) arising in oac-2 for mutant LSC652, its yolk protein mutant phenotype could not be rescued by introducing the wild-type copy of this gene. This mutant also contains an exonic missense mutation in tam-1 (tandem array expression modifier), a known transgene silencer26 (Table 1). Additional YP170B analysis (see below) confirmed that tam-1(lst538) causes reduced expression of the vit-2::gfp reporter. Similar false positives have been described for reporter-based screens before27,28,29.
In addition, we isolated five alleles comprising three genes that are so far unknown to play any role in yolk protein regulation. Novel causal protein-changing variants of two genes could be pinpointed in the corresponding mutants by rescuing vit-2::gfp expression with genomic wild-type copies of these candidates (Supplementary Fig. S1). We uncovered a premature stop allele and a splice site acceptor variant in Y54G2A.3, which we named vrp-1 (vitellogenin-regulating Caenorhabditis-specific protein) (Supplementary Fig. S1, Table 1). According to miR-abela analysis30, the first intron of the vrp-1 gene contains a potential functional RNA. We therefore also rescued the vrp-1(lst461) strain with a cDNA construct showing that its protein-coding sequence underlies the observed phenotype. Absence of similarities in BLAST searches support its Caenorhabditis-specific character. The homology-independent search tool FFPred 2.031 predicts VRP-1 to be involved in DNA-dependent regulation of transcription, further supported by its nuclear localization (see below).
Premature stop and splice site variants were similarly discovered for the ceh-60 gene (C. elegans homeobox) in two mutant strains (Supplementary Fig. S1, Table 1).
A novel nonsense mutation was unveiled in lrp-2 (low-density lipoprotein receptor related) (Supplementary Fig. S1, Table 1). For this strain, however, we could not obtain a genomic rescue, probably due to its relatively large size (~19 kbp) in combination with the strain’s ‘bag of worms’ phenotype. Because no other high impact type of variant resided in its small mapping region, we decided to use the earlier isolated lrp-2(gk556942) null mutant in addition to our lrp-2(lst464) mutant to ultimately verify the role of lrp-2 in the regulation of yolk protein production during later experiments32. This relates lrp-2 to its potential effects on yolk protein levels based on the indirect evidence for the identity of the phenotype-causing mutation in the lrp-2(lst464) strain.
Since the newly discovered alleles of vrp-1, ceh-60 and lrp-2 are of special interest, we backcrossed these strains multiple times to the premutagenized background strain and verified the molecular lesions by PCR and Sanger sequencing to fully support their robust identification. These strains - i.e. vrp-1(lst461), vrp-1(lst539), ceh-60(lst466), ceh-60(lst491), lrp-2(lst464) and lrp-2(gk556942) - will collectively be referred to as YPR (yolk protein regulating) mutant strains. YPR mutant strains were also outcrossed to wild type in order to eliminate their vit-2::gfp transgenic array.
Endogenous yolk protein and vit mRNA levels are repressed in YPR mutants
Having obtained novel regulators of vit-2 gene expression, it can now be asked whether the allelic variants emerging from our screen also have a more general impact on yolk protein production (i.e. YP170 and YP88/YP115) in C. elegans.
Compared to the control strains, endogenous YP170 yolk protein levels were strongly reduced or nearly absent in all YPR mutants throughout reproductive adulthood (Fig. 1, Supplementary Fig. S2 and S3, Supplementary Table S2). Whereas this effect was even more pronounced at the YP170B::GFP fusion protein levels (Supplementary Fig. S2 and S3), it holds for endogenous yolk levels in a wild-type background - i.e. independent of the vit-2::gfp transgene (Fig. 1, Supplementary Fig. S2, Supplementary Table S2).
Relative quantification of endogenous yolk protein and vit mRNA abundance.
YP170 yolk protein levels as analysed by SDS-PAGE were normalized against myosin levels (top row, Supplementary Fig. S2 and S3). Compared to the wild-type ( ), vit-2::gfp reporter (
), vit-2::gfp reporter ( ) and tam-1(lst538) (
) and tam-1(lst538) ( ) controls, endogenous YP170 levels are considerably lower in all YPR mutant populations, i.e. vrp-1(lst461) (
) controls, endogenous YP170 levels are considerably lower in all YPR mutant populations, i.e. vrp-1(lst461) ( ), vrp-1(lst539) (
), vrp-1(lst539) ( ), ceh-60(lst466) (
), ceh-60(lst466) ( ), ceh-60(lst491) (
), ceh-60(lst491) ( ), lrp-2(lst464) (
), lrp-2(lst464) ( ) and lrp-2(gk556942) (
) and lrp-2(gk556942) ( ). Also in a wild-type background, mutant YPR alleles suppress endogenous YP170 (
). Also in a wild-type background, mutant YPR alleles suppress endogenous YP170 ( ). The vit-2 (second row) and vit-3 (third row) mRNA expression profiles are consistent with these YP170 yolk protein data (first row). YP88 immunoblot data are expressed relative to each sample’s total protein signal (fourth row; Supplementary Table S2). In sharp contrast to both ceh-60, the vrp-1(lst461) and both lrp-2 mutants, the vrp-1(lst539) mutant still displays some YP88 immunoreactivity. The underlying vit-6 (bottom row) mRNA expression profiles again correlate well with these YP88 yolk protein data (Supplementary Table S2). For each indicated time point throughout reproductive development, the mean value of a maximum of three (mRNA) or four (protein) biologically independent measurements ± SEM is plotted and connected to assist in overall profile evaluation (see also Supplementary Table S2).
). The vit-2 (second row) and vit-3 (third row) mRNA expression profiles are consistent with these YP170 yolk protein data (first row). YP88 immunoblot data are expressed relative to each sample’s total protein signal (fourth row; Supplementary Table S2). In sharp contrast to both ceh-60, the vrp-1(lst461) and both lrp-2 mutants, the vrp-1(lst539) mutant still displays some YP88 immunoreactivity. The underlying vit-6 (bottom row) mRNA expression profiles again correlate well with these YP88 yolk protein data (Supplementary Table S2). For each indicated time point throughout reproductive development, the mean value of a maximum of three (mRNA) or four (protein) biologically independent measurements ± SEM is plotted and connected to assist in overall profile evaluation (see also Supplementary Table S2).
Endogenous YP170 levels were not reduced in tam-1(lst538), in contrast to the YP170B::GFP fusion protein levels, as can be expected for a transgene silencer mutation (Fig. 1 and S2A-C, Supplementary Table S2). We selected this mutant as an additional negative control during further analyses.
Endogenous YP170 levels in wild-type and vit-2::gfp reporter controls seem to remain stable throughout reproductive adulthood, with higher relative amounts present in wild type. However, YP170::GFP adds to the total YP170 levels of the reporter strain, which overall reaches levels similar to the wild-type endogenous YP170 pool. In addition, YP170::GFP increases throughout reproductive adulthood, an observation that has been reported by others for endogenous YP17033.
We further monitored endogenous levels of the individual YP88 yolk protein by means of an anti-vitellogenin antibody (Fig. 1 and S2D, Supplementary Table S2)3. Throughout reproductive adulthood, the YP88 pool was abundantly detected in the control strains, but severely reduced in vrp-1(lst539) and virtually absent in all other YPR mutants. Though post-translational influences cannot entirely be excluded, these results presumably also apply to YP115, since it originates from the same precursor polypeptide. YP115 cross-reactivity of the YP88 antibody supports this statement (Supplementary Fig. S2).
To verify whether the differences observed at the protein level indeed resulted from decreased vit gene expression, we measured relative expression levels of vit-2, -3 (~YP170) and -6 (=YP88/YP115) in representative YPR mutants (i.e. ceh-60(lst466), vrp-1(lst466) and lrp-2(lst464)), in comparison with the control strains. The other vit genes contributing to the YP170 pool - i.e. vit-1, -4 and -5 - were not analysed since we (Supplementary Fig. S2) and others have observed vit-1 to -5 to be expressed at similar levels33,34.
Throughout fertile adulthood, vit-2, -3 and -6 transcript levels are overall reduced in all studied YPR mutants as compared to controls (Fig. 1, Supplementary Table S2). Yet, this effect is less pronounced when comparing with the reporter strain only. This can be attributed to the much faster decrease in vit-2, -3 and -6 transcript levels towards the end of reproductive adulthood in the reporter control strain compared to wild type. Such a decline in vit-2 transcript levels has also been reported by others in a wild-type background33.
In addition, we verified the proper initiation of vit-2 expression in the reporter strain, which happens with the same timing and to the same extent as wild-type (endogenous only) vit-2 expression initiation (Fig. 2). In contrast, the vrp-1 and ceh-60 mutants fail to substantially up-regulate their vit-2 and vit-6 expression, even though it is initiated at the same time - though moderately (vrp-1(lst461)) or nearly not (ceh-60(lst466)).
vit-2 and vit-6 gene expression are not properly switched on in YPR mutants.
Light grey dotted line: wild-type lin-42 profile to assist in developmental timing evaluation111, wild type ( ), vit-2::gfp reporter control (
), vit-2::gfp reporter control ( ), vrp-1(lst461) (
), vrp-1(lst461) ( ), ceh-60(lst466) (
), ceh-60(lst466) ( ), all quantified as of the beginning of the L4 stage (profiles connect single measurements). (a) vit-2 mRNA levels of wild-type and vit-2::gfp reporter animals are heavily up-regulated upon becoming young adults, whereas this is not convincingly so in vrp-1(lst461) and nearly not at all in ceh-60(lst466) mutants. The vit-2 mRNA pool in all except wild-type animals also contains mRNA derived from the translational vit-2::gfp reporter construct. (b) In wild type, vit-6 up-regulation initiates slightly before that of vit-2 and covers an impressively larger dynamic range. vit-6 is the only YP88/YP115-providing gene, whereas the other five C. elegans vit genes can contribute to the YP170 pool. Also here, a very moderate (vrp-1(lst461)) - to no (ceh-60(lst466)) vit-6 up-regulation is observed in the selected YPR mutants. The lin-42 profile in panel b has been multiplied by 40 as compared to all other figures in this manuscript to facilitate visibility.
), all quantified as of the beginning of the L4 stage (profiles connect single measurements). (a) vit-2 mRNA levels of wild-type and vit-2::gfp reporter animals are heavily up-regulated upon becoming young adults, whereas this is not convincingly so in vrp-1(lst461) and nearly not at all in ceh-60(lst466) mutants. The vit-2 mRNA pool in all except wild-type animals also contains mRNA derived from the translational vit-2::gfp reporter construct. (b) In wild type, vit-6 up-regulation initiates slightly before that of vit-2 and covers an impressively larger dynamic range. vit-6 is the only YP88/YP115-providing gene, whereas the other five C. elegans vit genes can contribute to the YP170 pool. Also here, a very moderate (vrp-1(lst461)) - to no (ceh-60(lst466)) vit-6 up-regulation is observed in the selected YPR mutants. The lin-42 profile in panel b has been multiplied by 40 as compared to all other figures in this manuscript to facilitate visibility.
Taken together, these data show that on top of vit-2::gfp reporter gene suppression (Supplementary Fig. S2), the here isolated YPR mutants display severely reduced endogenous yolk protein and corresponding vit mRNA levels.
Intestinal VRP-1, amphid CEH-60
To better understand how VRP-1 regulates yolk protein synthesis, we sought to identify its expression pattern. Using a translational vrp-1::gfp reporter construct, we observed expression in the intestinal nuclei of both adult and larval hermaphrodites (Fig. 3). Gene expression patterns and stages of ceh-60, lrp-2 and vit-6 have been determined previously (Supplementary Table S3). Due to its intriguing site of action, we reconfirmed the ceh-60 expression pattern by confocal microscopy and DiI staining (Supplementary Fig. S4). As reported previously35, we could only observe robust ceh-60 expression in a single pair of amphids and occasionally weakly so in a second pair.
Transcriptional regulation in vit-regulatory mutants
To examine relevant periods of action during development, as well as to establish possible genetic interdependence in their control of yolk protein expression, relative expression levels of vrp-1, ceh-60 and lrp-2 genes were measured in vrp-1(lst461), ceh-60(lst491), wild-type and reporter controls. vrp-1 (Fig. 4a), ceh-60 (Fig. 4b) and lrp-2 (Supplementary Fig. S5) transcript levels are initially up-regulated during the L3-L4 wild-type moult, a transcriptional event that seems to be largely unaffected in all mutants under study. During the L4-adult moult, transcription of vrp-1 and ceh-60, but not lrp-2, is activated once more in wild-type animals. ceh-60 transcript levels are down-regulated in vrp-1(lst461), while the reverse is equally true. While vrp-1 and ceh-60 transcripts are repressed in their respective mutants, this is not true for lrp-2 (Supplementary Fig. S5).
YPR gene profiles reveal critical times during development and possible genetic pathway interdependency.
Light grey dotted line: wild-type lin-42 profile to assist in developmental timing evaluation111. (a) is vrp-1; (b) is ceh-60 expression in the following strains: wild type ( ), vit-2::gfp (
), vit-2::gfp ( ), vrp-1(lst461) (
), vrp-1(lst461) ( ) and ceh-60(lst491) (
) and ceh-60(lst491) ( ) (all profiles connect single measurements). In line with their effects on yolk levels in C. elegans, YPR genes are up-regulated in the later part of the developmental cycle (also Supplementary Fig. S5). vrp-1 levels are affected in ceh-60 mutants and vice-versa.
) (all profiles connect single measurements). In line with their effects on yolk levels in C. elegans, YPR genes are up-regulated in the later part of the developmental cycle (also Supplementary Fig. S5). vrp-1 levels are affected in ceh-60 mutants and vice-versa.
Reproduction seems unaffected by yolk depletion
Generally, vit genes are believed to act in a dose-dependent manner36. C. elegans may thus require all six active vit genes for its few intestinal cells to provide sufficient yolk proteins for the production of a large amount of eggs during its short life span18,37. However, apart from LRP-2 (whose mutants display the ‘bag of worms’ phenotype), malfunctioning of neither VRP-1 nor CEH-60 causes rigorous reproduction defects at first sight.
To evaluate the effect of a strongly impaired yolk protein pool on reproductive potential, we determined the overall brood size for all YPR mutants, as well as a more detailed progression of egg-laying for vrp-1(lst539) and ceh-60(lst466) mutants and the vit-2::gfp strain. lrp-2 mutants were not considered here, since internally hatched larvae tend to damage the mother's gonad, impeding the production of eggs. Strikingly, the total amount of viable offspring is unaffected in all YPR mutants under study (Supplementary Fig. S6). In addition, while the timing of egg-laying at best seems to be slightly delayed in vrp-1 and ceh-60 mutants, this phenotype could not be rescued by reintroducing YPR wild-type copies capable of restoring vit-2::gfp expression in these strains (Fig. 5). Thus, despite their disastrous effect on endogenous yolk protein production, the studied YPR alleles do not seem to affect egg-laying.
Progression of egg-laying in YPR mutants.
Three-hourly collected mean values ± SEM (n = number of adults evaluated) of offspring were binned per 12 hours to create egg-laying profiles for vit-2::gfp, (a) vrp-1(lst539), (b) ceh-60(lst466) and both rescued and non-rescued siblings from extrachromosomal rescue strains of each of these mutants. Compared to vit-2::gfp, both vrp-1 and ceh-60 mutants display similar viable offspring numbers (also Supplementary Fig. S6). (a+b) Potential small delays in egg-laying could not be rescued.
Survival upon starvation-induced L1 diapause
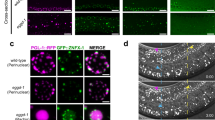
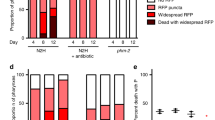
vrp-1 and ceh-60 YPR mutants cultured under standard laboratory conditions do not display overt defects in fertility nor development, despite their endogenous yolk protein pools being considerably depleted. This might be due to the presence of plentiful food in the worm’s vicinity. Yolk has been suggested to serve as an important energy source for larval survival upon L1 diapause - an alternate larval stage entered when C. elegans embryos hatch in the absence of food38,39. We found that deprived ceh-60(lst466) larvae display a convincing survival defect (Fig. 6a and Supplementary Fig. S7). At a worm density of 11 worms/μl, less than 20% of ceh-60(lst466) larvae survive their first day of starvation (p = 1.7E-09), as opposed to a median survival of 19 and 17.5 days of the wild-type and tam-1(lst538) controls. Though survival of L1 diapause for tam-1(lst538) mutants drops slightly faster than wild type, this difference is not significant (p = 0.669); supporting a similar behaviour for all control strains (Fig. 6a). vrp-1(lst539) results in a reduced survival of about 7 days compared to its control (p = 8.68E-05), although far more modest than that observed for ceh-60(lst466) (Fig. 6b). Our findings are therefore consistent with these yolk-reducing mutations having an important role for larval survival under conditions of limited food availability.
L1 diapause survival is distinctly affected in ceh-60 and vrp-1 mutants.
(a) As opposed to controls, ceh-60(lst466) ( ) individuals cannot cope with survival in absence of food (p = 1.7E-09) (also Supplementary Fig. S7). While also declining slightly earlier, tam-1(lst538) (
) individuals cannot cope with survival in absence of food (p = 1.7E-09) (also Supplementary Fig. S7). While also declining slightly earlier, tam-1(lst538) ( ) controls do not differ significantly from wild-type decline (p = 0.669). (b) Also vrp-1(lst539) (
) controls do not differ significantly from wild-type decline (p = 0.669). (b) Also vrp-1(lst539) ( ) shows a decrease in fitness under these conditions (p = 8.68E-05), though far less outspoken than the ceh-60(lst466) mutant. Because L1 diapause survival depends on culture density, panels (a) with 11 worms/μl and (b) with 18 worms/μl cannot be directly compared112.
) shows a decrease in fitness under these conditions (p = 8.68E-05), though far less outspoken than the ceh-60(lst466) mutant. Because L1 diapause survival depends on culture density, panels (a) with 11 worms/μl and (b) with 18 worms/μl cannot be directly compared112.
Discussion
Invertebrate reproduction is a process that for diverse reasons grasps the attention of many. For one, egg-laying is an easily measurable phenotypic readout, exploited to raise our fundamental understanding of biological signalling pathways40,41,42. In addition, there’s a solid industrial interest in applying new findings in this field to diversify invertebrate pest control43,44,45,46. Most importantly, if we were to better understand the molecular mechanisms acting to control invertebrate reproduction, it would help us answer important questions about evolutionary similarities and/or discrepancies10. Because in egg-laying species the embryo strongly depends on maternal yolk for its development47,48,49, we generated a C. elegans vit-2::gfp reporter strain to find novel genetic modifiers of yolk protein production. This helped us to further refine the scientific view on the control of yolk production and reproduction in a nematode, revealing some unexpected insights.
While specifically looking for mutants in which the developmental fate of yolk proteins is inappropriately executed, we identified three novel genetic components involved in the C. elegans stage-, sex- and tissue-specific vit regulatory system.
The DNA-binding CEH-60 (C. elegans homeobox) protein is homologous to the Drosophila melanogaster EXD (extradenticle) protein, encoding a HOX (homeobox) cofactor50,51 and its vertebrate counterparts, the PBX (pre-B cell leukaemia transcription factor) proteins52. PBX1 proteins are molecular mediators of sexual differentiation53,54. The here described role for ceh-60 in C. elegans yolk protein production therefore adds experimental support for a fundamental similarity between the vertebrate and invertebrate control systems of reproduction. However, its exact role within these systems might vary over species. We here confirmed previously reported expression of ceh-60 in only a few amphid neurons implicated in sensing environmental cues35,55. These findings now set the scene for a detailed search for environmental inputs that via neuronal signalling potentially modulate intestinal function related to reproduction. In mice, PBX1 proteins act in the gonadotropin-releasing hormone neurons, which further supports the potentiality for such an axis to exist56.
We isolated another putative transcription factor, which we annotated as VRP-1 (vitellogenin-regulating Caenorhabditis-specific protein) and showed it to be an intestinal player. As such, VRP-1 is well-positioned to directly influence yolk protein production, a process that occurs in the intestine only. However, in contrast to the vit genes which are only expressed in adult hermaphrodites18, vrp-1 is also expressed in larvae, albeit at much lower levels. These findings point to a broader role for vrp-1, which may from the L3 stage onwards (upon considerable up-regulation) start to serve the time- and tissue-specific regulation of vit expression.
To date, no (in)vertebrate orthologues of these two C. elegans vit regulators have been reported to be involved in yolk protein production. Moreover, all our bioinformatic searches support the Caenorhabditis-specific nature of VRP-1, indicating that in C. elegans, the regulation of yolk protein production might have at least one genus-specific aspect.
We were unable to rescue vit-2::gfp reporter expression of the lrp-2(lst464) mutant. Even though relying on tight mapping results, its annotation in our isolated strain is therefore supported by indirect evidence. Yet, the reduced vit transcript and yolk protein levels in the independently isolated lrp-2(gk556842) mutant32 indicate that LRP-2 is an actual yolk protein regulator. The LRP-2 (low-density lipoprotein (LDL) receptor related) protein is expressed in a multitude of tissues57 and is, like all known yolk receptors, a member of the LDL receptor superfamily of lipoprotein receptors58,59. Generally, such receptors are known to orchestrate (in)vertebrate cholesterol homeostasis60, a process that is crucial to reproduction since in most studies so far, all known regulatory pathways converge onto the production of steroid hormones. In C. elegans, extracellular sterols are possibly taken up by the digestive tract or through the cuticle and are of vital importance for reproduction5. LRP-2 might be involved in LDL-derived cholesterol distribution and transport to hypodermal and neuronal tissues for synthesis of the endogenous sterol-derived dafachronic acids (DAs)61,62,63,64. These hormones play a role in promoting reproductive development by bypassing entry into the alternative L3/dauer larval stage65. If the similarity to other (in)vertebrate systems would hold, these could also be functionally involved in the control of yolk protein synthesis in the adult. In a preliminary test, we were unable to rescue any of the YPR mutant phenotypes by supplementing the worms with (25S)-Δ7-DA, a C. elegans steroid hormone able to fully rescue constitutive dauer mutants66. Functional relations between the YPR proteins and DA signalling nevertheless remain possible. Providing the mutants with (25S)-Δ7-DA alone might be an insufficient or incorrect source of steroid. Because of the tight link between vitellogenesis and steroidogenesis1,2, it will be of paramount importance to further study possible interactions and dependencies in C. elegans.
lrp-2 mutants display the ‘bag of worms’ phenotype, a viviparity-like strategy. The loss of yolk production, caused by a malicious mutation in one of the regulatory genes controlling vitellogenesis, might therefore have initiated an escape mechanism, i.e. internal hatching67. Several other causes exist for this strategy, including deficient vulva development or inadequate motility of the vulval muscles. lrp-2 mutants indeed suffer from insufficient yolk protein availability, but the gene is also expressed in nervous tissue and in vulval and uterine muscles. It has been described before that LRP-1 and 2 are cooperatively required for the fibroblast growth factor EGL-17 (egg-laying defective)-dependent regulation of sex myoblast migrations during larval development68 and as such are involved in the generation of uterine and vulval musculature69. For both independent lrp-2 mutants, we were so far unable to separate their suppressed yolk protein levels from their ‘bag of worms’ phenotype. Taken together, these findings point towards a tight link between the EGL circuit and yolk production. This is further supported by compromised egg-yolk pools in egl-15 knockdown animals, as observed by others25.
Vitellogenesis encompasses an important metabolic cost and the number of genes involved in the production of vit mRNAs may be surprisingly large3. It can therefore be asked whether and how the novel genetic players retrieved from our screen may act together to influence C. elegans yolk protein production. Since we were mainly interested in the control of yolk protein synthesis, we did not specifically survey the involvement of yolk and yolk receptor endocytic trafficking components in this regard. Nevertheless, it is possible that transcriptional vit regulators also affect the level of yolk transport, e.g. through control of the earlier described factors RME-2, RAB-35, RAB-10 and M04F3.26,24,25.
Based on the transcriptional and translational data, some initial mechanistic concepts were obtained. Overall, the YP170::GFP fusion protein pool is more prone to suppression by the here identified yolk protein regulators than the endogenous YP170 yolk protein pool in the reporter background. However, also in a wild-type background - i.e. having only the endogenous yolk protein pool - YP170 levels are severely reduced for YPR mutant alleles. The vit-2::gfp reporter construct probably acts as a sink for transcription factors and their potential regulators.
Regarding LRP-2, the RNA data were not unambiguous since we detected wild-type levels of lrp-2 transcripts in the lrp-2 mutant. Due to its large size (~14.5 kbp), the lrp-2 transcript might be less vulnerable to nonsense-mediated mRNA decay70,71,72, resulting in a slower turnover and possibly explaining the higher detection levels. Furthermore, another study classified L3-to-adult mRNA profiles of thousands of transcripts as either “flat”, “rising” or “oscillating”73. Supporting our observations, all vit genes were classified as rising, vrp-1 as rising and ceh-60 as oscillating. Contrary to our data, lrp-2 was classified as flat. While these authors measured expression levels at 25 °C and therefore may have started their observations right after the here observed lrp-2 peak, this discrepancy should caution against conclusions solely based on the lrp-2 expression profile. Yet, behaving quite different from vrp-1 and ceh-60, it seems that the LRP-2 receptor regulates yolk protein production distinctly from the proposed signalling interactions described below.
Our data support that ceh-60 and vrp-1 influence each other’s expression levels, arguing for a signalling system between the amphids and the intestine characterized by bidirectional information transfer. CEH-60 is well-localized to integrate environmental information important for egg maturation. Based on its nuclear localization and bioinformatic predictions, VRP-1 may well be a transcription factor. vrp-1 expression starts to rise prior to the marked ceh-60 boost (the latter upon completion of larval development), but without this boost, its expression levels cannot rise any further. Conversely, without proper vrp-1 expression, ceh-60 levels remain low as of L4.
It has been shown that ceh-60 and all vit genes are down-regulated in egg-laying defective ets-4 mutants74. This suggests that the intestinal longevity player ETS-4 communicates with CEH-60 to integrate longevity cues in the system, which makes sense from an energetic point of view: environmental cues should generally coincide with other factors, e.g. information on nutrients available via the intestine. In D. melanogaster a mechanism has been described wherein the presence of dietary compounds in the intestine enhances, via intermediate player(s) yet to be identified, yolk protein gene expression in the fat body and thus egg production9. The initiation of vitellogenesis in Aedes aegypti and Sarcophaga bullata in response to, respectively, a blood or liver meal, is yet another well-known indicator of the existence of such a control mechanism7,75,76,77. In C. elegans, other previously identified intestinal vit regulators, such as the lipid metabolism-regulating transcription factor KLF-3 (Krüppel-like factor (zinc finger protein))78,79 and to a lesser extent RBPL-1 (retinoblastoma binding protein like)80, are also possibly involved in the integration of signals regarding the nutritional status of the intestine, i.e. the prime site of lipid metabolism81. Future experiments will be needed to reveal the interplay of these regulators with the here identified regulators of yolk protein production. Particularly KLF-3 is of interest, since the vertebrate CEH-60 homolog PBX1 is known to modulate transcriptional regulation via the KLF-3 homolog, KLF482.
The view that energetically costly investments in the production of egg-yolk are needed to enable viable offspring is - in nematodes - supported by the phenotype of rme-2 mutants. In these mutants, the slightly smaller oocytes lack yolk proteins due to a malfunctioning of yolk endocytosis, ovulation is defective and both the production of embryos as well as their viability is reduced6. While it should be noted that our particular forward genetic approach restricted the identification of yolk protein regulatory mutants to those able to reach fertile adulthood, it should still enable the isolation of mutants with a severely reduced reproduction potential. Therefore, we wanted to assess whether for the vrp-1 and ceh-60 mutants, yolk protein production would indeed be a predictive criterion for overall reproductive success.
Contrary to what would be assumed based on the abundant expression of six vit genes in wild-type adults36, YPR mutants displayed no clear reproduction defects with the exception of the ‘bag of worms’ phenotype for lrp-2 mutants. Indeed, strongly reduced yolk protein pools are not necessarily critical for successful brood development in our C. elegans experiments, as opposed to e.g. D. melanogaster83. The simplest explanation would be that in these mutants, the little remaining fuel suffices for the production of the observed amount of eggs. This could suggest that the enormous amounts of yolk serve another purpose.
Even though hermaphrodites usually self-fertilize in the wild84, they might de facto prepare for the higher offspring numbers observed when mated with a male85,86. However, we observed far greater reductions in yolk protein content in our set of mutants than can directly be correlated to this phenomenon. Alternatively, yolk proteins could hypothetically be implicated in yet unidentified, co- and post-reproductive processes in the adult87. This is somewhat supported by the observation that yolk synthesis, once induced, is not switched off again in post-reproductive adults, although this might equally well be an unwanted side-effect inherent to ageing, as explained by Blagosklonny’s hyperfunction theory88,89.
These two possibilities are nevertheless difficult to reconcile from an evolutionary point of view: why would nature have specifically selected those C. elegans that invest in immense amounts of yolk production, if it hadn’t provided them with a general hermaphrodite-specific advantage up to their reproductive period? We therefore timed the egg-laying profile of selected YPR mutants with great detail. While their overall offspring numbers are nearly similar to wild-type levels, vrp-1 and ceh-60 mutants displayed a small delay in egg-laying. In the wild, where resources are scarce, the massive C. elegans yolk protein supplies may therefore capacitate the fertile adult with a very efficient way of outcompeting others. We could however not rescue this phenotype, arguing against this hypothesis. Alternatively, it could be possible that levels capable of restoring vit-2::gfp reporter expression may yet not suffice to restore other phenotypical consequences. This can be understood from the clear preference of VIT-2::GFP production over endogenous yolk in the reporter strain. Taken together, this means that vrp-1 and ceh-60 genes at best have a mild influence on the timing of egg-laying under standard laboratory conditions. However, in the wild, these ample-food conditions are presumably rarely met.
Therefore, we attempted to demonstrate the importance of yolk for larval survival during L1 diapause emergence in YPR mutants39. When hatched in the absence of food, the survival of vrp-1(lst539) and ceh-60(lst466) yolk-depleted L1 larvae is moderately to severely compromised, respectively. These findings at least suggest that while the remaining low levels of yolk in these mutants may suffice under rich nutritional conditions, they represent a disadvantage when food is in short supply. In the adult, decisions with respect to entering the reproductive state depend on the environment’s nutritional value and are taken long before ceh-60 and vrp-1 can act on adult vit expression (i.e. the L3-L4 moult). The observed abundance of yolk under optimal conditions must therefore at least in part be a consequence of the natural selection events in favour of high yolk amounts to preserve L1 survival in the absence of food.
Our data on reproduction might also comply with a more refined escape mechanism, in which possible compensating substances preserve embryo viability - thus reproductive success - but not necessarily L1 diapause survival when yolk protein levels are low. After fertilization and cleavage, the yolk particles of control animals are approximately evenly distributed among the daughter cells and only slowly metabolized6. Considerable amounts of yolk remain in newly hatched larvae, the majority of which in intestinal cells. They continue to utilize yolk as a food source, reflected by the fast degradation of residual yolk proteins in L1 larvae. Corresponding phenomena are observed in amphibians90 and insects91,92. This observation has led to the assumption that yolk proteins must initially be present in excess19, but in fact, it could even be true that in C. elegans, embryonic development does not strongly rely on yolk. Alternative nutrients could be transported into the embryo, a process that could involve a more general transport of lipoproteins via the C. elegans yolk receptor, RME-26, in addition to yet unknown transport mechanisms. If these substances are inadequate to serve as backup under harsh conditions, this would still be in line with the observed L1 diapause defects.
In conclusion, starting from a forward genetic screen, we could identify novel genetic regulators of yolk protein production in C. elegans, hereby improving our knowledge on invertebrate control of yolk production, a process assumed to only serve reproduction. Our data support that parallel pathways are active in multiple tissues and that the proteins VRP-1, CEH-60 and LRP-2 are involved in the general regulation of C. elegans yolk protein production. Based on our data, it seems that C. elegans produces far more yolk than is needed to sustain its embryos. Enormous amounts of yolk might have been selected for in the wild, enabling not embryonic, but larval survival when resources are scarce. Yet, our findings still suggest an additional investment in yolk, opening up new research possibilities as to the why and how of this energy-costly process.
Methods
Strains, microscopy and growth conditions
Following strains were obtained via the CGC: N2 Bristol93 wild-type control, CB4856 Hawaiian94 isolate and VC40291 lrp-2(gk556942) null mutant32. The UL2612 Pceh-60::gfp strain was a kind gift of Professor I. A. Hope, University of Leeds, Leeds, UK35. For the forward genetic screen, we generated LSC276 lstIs13 [Pvit-2::vit-2::gfp] as starting strain to avoid potential non-standard unc-119 expression levels in the existing RT130 unc-199(ed3); pwIs23 [Pvit-2::vit-2::gfp, unc119(+)] strain. LSC276 was then used to obtain the following mutant strains: LSC462 rab-35(lst462); lstIs13 [Pvit-2::vit-2::gfp], LSC463 rab-10(lst463); lstIs13 [Pvit-2::vit-2::gfp], LSC537 M04F3.2(gk2294); lstIs13 [Pvit-2::vit-2::gfp], LSC648 vrp-1(lst461); lstIs13 [Pvit-2::vit-2::gfp], LSC649 ceh-60(lst491); lstIs13 [Pvit-2::vit-2::gfp], LSC650 ceh-60(lst466); lstIs13 [Pvit-2::vit-2::gfp], LSC651 vrp-1(lst539); lstIs13 [Pvit-2::vit-2::gfp], LSC652 tam-1(lst538); lstIs13 [Pvit-2::vit-2::gfp] and LSC653 lrp-2(lst464); lstIs13 [Pvit-2::vit-2::gfp]. Mutant strains were backcrossed several times (Table 1) with LSC276 (lstIs13 [Pvit-2::vit-2::gfp]) males on the basis of their aberrant yolk protein synthesis phenotype and maintained as homozygotes according to standard methods93. Similarly, the lrp-2(gk556942) mutant was backcrossed twice to N2 males based on its ‘bag of worms’ phenotype. We furthermore outcrossed vrp-1(lst539); lstIs13 [Pvit-2::vit-2::gfp], ceh-60(lst466); lstIs13 [Pvit-2::vit-2::gfp], ceh-60(lst491); lstIs13 [Pvit-2::vit-2::gfp] and lrp-2(lst464); lstIs13 [Pvit-2::vit-2::gfp] animals to N2 in order to remove the transgenic array from its background and respectively obtained strains LSC901 vrp-1(lst539), LSC897 ceh-60(lst466), LSC903 ceh-60(lst491) and LSC904 lrp-2(lst464). The causal allele of each (back- or outcrossed) mutant used during further analyses was confirmed by PCR and Sanger sequencing (Supplementary Table S4). For all these analyses, random animals from the population were always used.
All fluorescence and bright-field micrographs in this study were obtained with an Axio Imager. Z1 light microscope equipped with an AxioCam MRm camera (1388 × 1040 pixels) using the digital image processing software ZEN 2012 (Carl Zeiss Microscopy, Göttingen, Germany) and identical settings.
Amphid neurons of Pceh-60::gfp worms were stained with 1,1'-dioctadecyl-3,3,3',3'-tetramethylindocarbocyanine perchlorate (DiI; Molecular Probes, Carlsbad, California)95,96,97. Fluorescent signals were visualized by an Olympus Fluoview FV1000 (IX81) confocal microscope and confocal Z-stack projections were exported through Imaris 7.2 (Olympus, Tokyo, Japan).
All strains were maintained at 20 °C on standard nematode growth medium (NGM) agar plates seeded with Escherichia coli OP5098.
Transgenic C. elegans strains
Vitellogenin reporter
We constructed a transgenic vitellogenin reporter strain for a subsequent forward genetic screen. The plasmid V2B3 (a kind gift of Professor B. Grant, Rutgers University, New Jersey), encoding the full-length yolk protein YP170B fused to GFP99 and expressed under vit-2 promoter control, was injected at 80 ng/μl into N2 worms using standard microinjection techniques100. One stably integrated line was produced using a UV crosslinker (BLX-254; Vilber Lourmat, Marne-la-Vallée, France) following standard protocols and backcrossed ten times to N2 to obtain the lstIs13 [vit-2::gfp] reporter strain101,102,103.
YPR mutant rescues
For vrp-1(lst539) and ceh-60(lst466) strains, gDNA rescue experiments by germline transformation were executed. We PCR amplified gDNA rescue fragments using the Pfu DNA polymerase (Thermo Scientific, St. Leon-Rot, Germany) and oligonucleotides listed in Supplementary Table S5. These fragments contain the full-length gDNA, the upstream promoter and downstream regulatory sequence. Rescue strains were obtained from microinjections at 20 ng/μl for ceh-60(lst466) (LSC898) and 50 ng/μl for vrp-1(lst539) (LSC899) with an intestinal Pelt-2::mCherry selectable co-injection marker.
Similarly, microinjections at 5, 20 and/or 50 ng/μl were carried out in order to obtain the genomic rescues of the non-backcrossed mutant strains (Supplementary Fig. S1).
A rescue analysis using vrp-1 cDNA sequence was additionally carried out. Overlapping fragments of the vrp-1 promoter region (306 bp) and vrp-1 cDNA were generated using the primer sets listed in Supplementary Table S5 and subsequently fused using a PCR fusion-approach104. The obtained fragment was fused to an overlapping fragment of the 3′ UTR of vrp-1 and microinjected at 50 ng/μl.
VRP-1 reporter strain
For in vivo localization of VRP-1, overlapping fragments of full-length vrp-1 gDNA lacking its stop codon together with a 342 bp upstream regulatory region and the gfp region of pPD95.75 (Fire Lab C. elegans Vector Kit, 1995) were first amplified by PCR (oligonucleotides: see Supplementary Table S6). Subsequently, these fragments were fused with a set of nested primers to obtain a translational Pvrp-1::vrp-1::gfp fusion construct driven by a promoter region of 306 bp104. Several independent transgenic lines were obtained by germline transformation of N2 worms with 50 ng/μl of Pvrp-1::vrp-1::gfp co-injected with a pan-neuronal Prgef-1::rfp selectable marker (contained on plasmid pCB101, a kind gift of Professor M. Doitsidou and Professor O. Hobert, Colombia University, New York)105.
Yolk protein mutant isolation
For the manual screen, early L4 lstIs13 [Pvit-2::vit-2::gfp] animals were mutagenized with 50 mM of EMS according to standard protocol93. After 24 hours, 3 to 5 mutagenized gravid adults were placed in each of a total of 40 founder P0 plates. During the next 12 hours, all F1 progeny (~3,750) of the mutagenized P0 animals were singled out. gfp expression from the lstIs13 transgene of the ensuing progeny (F2 generation) was scored via microscopic observation (Leica MZ16 F; Leica Microsystems, Wetzlar, Germany). Individual mutants were picked and their phenotypic heritability confirmed in order to establish mutant lines. By first crossing mutant worms with both N2 and premutagenized lstIs13 males and then assessing fluorescence levels, we determined whether mutations were dominant or recessive, or alternatively, whether the gfp array was hit.
Sequencing of YPR mutants
WGS SNP mapping strategy
Following the guidelines previously described22, we set up crosses between ten of the isolated, homozygous recessive mutant strains (Bristol background) and CB4856 Hawaiian males. ~50 F2 recombinant worms (Supplementary Fig. S1) segregating from this cross and displaying the proper mutant phenotype were individually reselected. They were allowed to self-fertilize and their F3 and F4 progeny were pooled. For mutants displaying complete loss of gfp expression, only plates originating from F2 animals that appeared to be positive (but were not particularly homozygous) for the gfp reporter gene, as assessed by PCR, were pooled (oligonucleotides: Supplementary Table S4). Genomic C. elegans DNA samples were prepared22 and at least 3 μg of genomic DNA (gDNA) (≥30 ng/μl) from each sample was submitted to BGI Tech Solutions Co., Limited (Hong Kong, China) for short-insert library preparation and 100 bp paired-end sequencing on an Illumina HiSeq 2000 sequencer to obtain 3 Gb clean data, corresponding to a ~30-fold genome coverage across all non-gap regions. A gDNA sample of pure background strain was similarly handled to obtain 1 Gb clean data.
WGS data analysis using the CloudMap pipeline
The generated Illumina 1.5 FASTQ data files were uploaded into Galaxy and analysed using the CloudMap Hawaiian variant mapping with WGS and variant calling workflow (http://usegalaxy.org/cloudmap)106. Two pre-processing steps were run on the FASTQ files in Galaxy. First, we concatenated the FASTQs for each sample using the Concatenate Datasets tool and subsequently converted the resulting file into FASTQ Sanger quality encoding using the FASTQ Groomer tool107. Prior to the automated execution of the different implicated bioinformatics processing steps, we adapted the workflow to accept paired-end FASTQ files according to the user guide. The default tool settings further used have thoroughly been described before106.
Yolk protein analysis
The YP170 yolk protein pool was analysed for each condition as previously described33, with animals growing on plates (90 mm diameter) treated topically with 80 μl of 50 mM 2′-deoxy-5-fluorouridine (FUdR; Sigma-Aldrich, St. Louis, Missouri) to avoid offspring accumulation108. This did not result in visible changes of vit-2::gfp reporter expression of the control worms. Since yolk proteins are first expressed during L4 lethargus18, age-matched L4 stage larvae were obtained for each condition and 50 hermaphrodites were harvested in up to quadruplicate 24, 36, 48, 60, 75, 84 and 108 hours later into 15 μl M9 buffer, diluted in 15 μl 2x Laemmli Sample Buffer (Bio-Rad, Hercules, California) containing β-mercaptoethanol (Sigma-Aldrich). Worm samples were subsequently incubated during 15 min in a 70 °C water bath and vortexed every 5 min before optional storage at −80 °C. Prior to loading 15 μl on a 4–12% Bis-Tris Criterion XT precast polyacrylamide gel (Bio-Rad), protein samples were spun for 5 min to remove insoluble precipitates. According to the manufacturer’s instructions, gels were stained and destained using SimplyBlue SafeStain (Life Technologies).
To ultimately verify the identity of the Coomassie G-250-stained YP170 (~170 kDa) and YP170B::GFP (~180 kDa) protein bands, they were first manually excised using a fresh scalpel for each band and destained during 30 min in 100 μl 100 mM ammonium bicarbonate, interrupted by occasional vortexing. Next, we followed our general lab protocol for trypsin digestion109. The tryptic peptide samples were dried using a SpeedVac Concentrator (Savant) and subsequently purified to remove interfering salt contaminants using Pierce C18 Spin Columns corresponding to manufacturer’s guidelines (Thermo Scientific). The samples were dried again in the vacuum evaporator and stored at −80 °C before mass spectrometric (MS) analysis performed on an electrospray ionization quadrupole time-of-flight (ESI-QTOF) mass spectrometer micrOTOF-Q (Bruker Daltonics, Bremen, Germany) operated in reflectron positive ion mode. We identified proteins through peptide mass fingerprinting using Mascot (Matrix Science, London, UK). An error window on experimental peptide mass values of 400 ppm and on MS/MS fragment ion mass values of 600 mmu was used in all searches and we allowed one missed cleavage per peptide. Carbamidomethylation (C) and oxidation (M) were respectively chosen as fixed and variable modifications. The ~170 kDa searches were limited to C. elegans proteins and the ~180 kDa searches to all entries in the SwissProt database.
Protein bands corresponding to YP170 were quantified relative to myosin protein levels (~200 kDa) by using the Image Lab Software Version 4.1 (Bio-Rad).
In parallel, rat polyclonal antiserum anti-YP88 (a kind gift of Professor T. Blumenthal, University of Colorado Boulder, Colorado and Professor S. Strome, University of California, Santa Cruz, California)3, were applied to detect the YP88 yolk pool during immunoblot assays. Thereto, 15 μl of single fertile hermaphrodite protein samples prepared 36, 60, 84 and 108 hours post the L4 stage were separated on a 4–12% Bis-Tris Criterion XT precast polyacrylamide gel. Polyclonal rabbit anti-rat immunoglobulins conjugated to horseradish peroxidase acted as the secondary antibody (Dako, Glostrup, Denmark). Signal quantification relative to an Amersham Deep Purple total protein stain (GE Healthcare, Piscataway, New Jersey) was performed using the Image Lab Software Version 4.1.
Real-time PCR analysis
Circa three fully-grown plates (90 mm diameter) containing age-matched populations of fertile hermaphrodite adults were obtained in up to triplicate for each condition. For developmental analysis, a time series consisting of two to three fully-grown plates per sample and per condition was collected; for the adult vit expression profiles, cultures were treated with FUdR as described above. Total RNA was prepared using the RNeasy kit (Qiagen, Hilden, Germany) and subjected to DNaseI (Qiagen) treatment. Next, cDNA was synthesized in duplo from up to 500 ng of total RNA in a 100 μl-volume reaction using the PrimeScript RT Master Mix (Takara, Ohtsu, Japan), upon which technical replicates were pooled. Per biological replica, technical duplicate or triplicate 20 μl qRT-PCR reactions were set up in 96 well plates using the SYBR Green PCR Master Mix (Applied Biosystems, Foster City, California) and reactions were run on the StepOnePlus Real-Time PCR System (Applied Biosystems). The primer sets used in these reactions for transcripts vit-2, -3, -4, -5, -6, vrp-1, lrp-2, ceh-60 and lin-42 are listed in Supplementary Table S7. The relative expression level of each gene transcript in control versus mutant was assessed by a geNorm-based normalization strategy, in which cdc-42, pmp-3 and tba-1 emerged as optimal reference genes for this study110. Analysis of variance (ANOVA) statistics running post hoc Dunnett’s tests were used to obtain p-values of significance for the analysis of individual transcripts in each mutant condition.
Egg-laying
The progression of egg-laying and the overall brood size were determined by selecting synchronized L4 nematodes and placing them each on a single NGM plate with OP50 bacteria. From 18 hours until 96 hours after mid-L4, worms were repeatedly transferred every 3 hours to a freshly seeded NGM plate for a period of 12 hours, for practical reasons alternated with a single transfer after a full period of 12 hours. We counted the number of offspring when in the L4/adult stage. Incomplete offspring data due to escaping or dying mothers were omitted from the analyses. Total brood sizes were compared using ANOVA statistics running a post hoc Dunnett’s test.
L1 diapause survival assay
Starvation-induced L1 diapause survival assays were performed essentially as described previously using comparable densities for mutant and control strains39. Synchronized L1 larvae were incubated in 3 ml of sterilized S-basal buffer at 20 °C. For each condition, aliquots containing roughly 200 animals were placed in triplicate on individual seeded plates every single day during a period of 10 days starting at the first starvation day, followed by every 3 days for the consecutive period. The number of surviving animals was counted when in the L4/adult stage at 20 °C. The number of surviving animals at the first day of starvation was set at 100% in order to calculate the percentage of surviving animals at the following time points. The L1 diapause data were statistically compared using an analysis of covariance (ANCOVA) model. The rate of survival was log-transformed to accommodate the assumption of normality and data were analysed using R (The R Foundation for Statistical Computing, Vienna, Austria).
In general, statistical analyses and graphing in this manuscript were performed using Prism 6 (GraphPad Software, La Jolla, California), except for Fig. 3, S3 and S4 (made using Excel 2007).
Additional Information
How to cite this article: Van Rompay, L. et al. New genetic regulators question relevance of abundant yolk protein production in C. elegans. Sci. Rep. 5, 16381; doi: 10.1038/srep16381 (2015).
Change history
20 November 2015
The version of this Article previously published quoted an incorrect abbreviation for Liesbeth Van Rompay in the 'How to cite this article' section. This has now been corrected in both the PDF and HTML versions of the paper.
References
Tata, J. R., James, T. C., Watson, C. S., Williams, J. L. & Wolffe, A. P. Hormonal regulation and expression of vitellogenin multigene family. Ciba Found. Symp. 98, 96–110 (1983).
Awruch, C. A. Reproductive endocrinology in chondrichthyans: The present and the future. Gen. Comp. Endocrinol. 192, 60–70 (2013).
Sharrock, W. J. Cleavage of two yolk proteins from a precursor in Caenorhabditis elegans. J. Mol. Biol. 174, 419–431 (1984).
Winter, C. In Reproductive biology of invertebrates - Progress in vitellogenesis ( Raikhel, A. & Sappington, T. ) 1–27 (Plenum Press, 2002).
Matyash, V. et al. Distribution and transport of cholesterol in Caenorhabditis elegans. Mol. Biol. Cell 12, 1725–1736 (2001).
Grant, B. & Hirsh, D. Receptor-mediated endocytosis in the Caenorhabditis elegans oocyte. Mol. Biol. Cell 10, 4311–4326 (1999).
Huybrechts, R. & De Loof, A. Similarities in vitellogenin and control of vitellogenin synthesis within the genera Sarcophaga, Calliphora, Phormia and Lucilia (Diptera). Comp. Biochem. Physiol. Part B Comp. Biochem. 72, 339–344 (1982).
Bownes, M. Expression of the genes coding for vitellogenin (yolk protein). Ann. Rev Entomol. 31, 507–531 (1986).
Bownes, M., Scott, A. & Shirras, A. Dietary components modulate yolk protein gene transcription in Drosophila melanogaster. Development 103, 119–128 (1988).
Frooninckx, L. et al. Neuropeptide GPCRs in C. elegans. Front. Endocrinol. (Lausanne). 3, 167. 1–18 (2012).
Cho, S., Rogers, K. W. & Fay, D. S. The C. elegans glycopeptide hormone receptor ortholog, FSHR-1, regulates germline differentiation and survival. Curr. Biol. 17, 203–212 (2007).
Vadakkadath Meethal, S. & Atwood, C. S. Identification of a gonadotropin-releasing hormone receptor orthologue in Caenorhabditis elegans. BMC Evol. Biol. 6, 103 (2006).
Kudo, M., Chen, T., Nakabayashi, K., Hsu, S. Y. & Hsueh, A. J. The nematode leucine-rich repeat-containing, G protein-coupled receptor (LGR) protein homologous to vertebrate gonadotropin and thyrotropin receptors is constitutively active in mammalian cells. Mol. Endocrinol. 14, 272–284 (2000).
The C.elegans sequencing consortium. Genome sequence of the nematode C. elegans: a platform for investigating biology. Science 282, 2012–2018 (1998).
Blumenthal, T. et al. Cloning of a yolk protein gene family from Caenorhabditis elegans. J. Mol. Biol. 174, 1–18 (1984).
Klass, M. R., Wolf, N. & Hirsh, D. Further characterization of a temperature-sensitive transformation mutant in Caenorhabditis elegans. Dev. Biol. 69, 329–335 (1979).
Spieth, J. & Blumenthal, T. The Caenorhabditis elegans vitellogenin gene family includes a gene encoding a distantly related protein. Mol. Cell. Biol. 5, 2495–2501 (1985).
Kimble, J. & Sharrock, W. J. Tissue-specific synthesis of yolk proteins in Caenorhabditis elegans. Dev. Biol. 96, 189–196 (1983).
Sharrock, W. J. Yolk proteins of C. elegans. Dev. Biol. 96, 182–188 (1983).
Sharrock, W. J., Sutherlin, M. E., Leske, K., Cheng, T. K. & Kim, T. Y. Two distinct yolk lipoprotein complexes from Caenorhabditis elegans. J. Biol. Chem. 265, 14422–14431 (1990).
Hall, D. H. et al. Ultrastructural features of the adult hermaphrodite gonad of Caenorhabditis elegans: relations between the germ line and soma. Dev. Biol. 212, 101–123 (1999).
Doitsidou, M., Poole, R. J., Sarin, S., Bigelow, H. & Hobert, O. C. elegans mutant identification with a one-step whole-genome-sequencing and SNP mapping strategy. PLoS One 5, e15435 (2010).
Schneeberger, K. et al. SHOREmap: simultaneous mapping and mutation identification by deep sequencing. Nat. Methods 6, 550–551 (2009).
Sato, M. et al. Regulation of endocytic recycling by C. elegans Rab35 and its regulator RME-4, a coated-pit protein. EMBO J. 27, 1183–1196 (2008).
Balklava, Z., Pant, S., Fares, H. & Grant, B. D. Genome-wide analysis identifies a general requirement for polarity proteins in endocytic traffic. Nat. Cell Biol. 9, 1066–1073 (2007).
Hsieh, J. et al. The RING finger/B-box factor TAM-1 and a retinoblastoma-like protein LIN-35 modulate context-dependent gene silencing in Caenorhabditis elegans. Genes Dev. 13, 2958–2970 (1999).
Vastenhouw, N. L. et al. Gene expression: long-term gene silencing by RNAi. Nature 442, 882 (2006).
Kim, J. K. et al. Functional genomic analysis of RNA interference in C. elegans. Science 308, 1164–1167 (2005).
Wang, D. et al. Somatic misexpression of germline P granules and enhanced RNA interference in retinoblastoma pathway mutants. Nature 436, 593–597 (2005).
Sewer, A. et al. Identification of clustered microRNAs using an ab initio prediction method. BMC Bioinformatics 6, 267 (2005).
Minneci, F., Piovesan, D., Cozzetto, D. & Jones, D. T. FFPred 2.0: improved homology-independent prediction of gene ontology terms for eukaryotic protein sequences. PLoS One 8, e63754 (2013).
Thompson, O. et al. The million mutation project: a new approach to genetics in Caenorhabditis elegans. Genome Res. 23, 1749–1762 (2013).
DePina, A. S. et al. Regulation of Caenorhabditis elegans vitellogenesis by DAF-2/IIS through separable transcriptional and posttranscriptional mechanisms. BMC Physiol. 11, 11 (2011).
Halaschek-Wiener, J. et al. Analysis of long-lived C. elegans daf-2 mutants using serial analysis of gene expression. Genome Res. 15, 603–615 (2005).
Reece-Hoyes, J. S. et al. Insight into transcription factor gene duplication from Caenorhabditis elegans Promoterome-driven expression patterns. BMC Genomics 8, 27 (2007).
Brawand, D., Wahli, W. & Kaessmann, H. Loss of egg yolk genes in mammals and the origin of lactation and placentation. PLoS Biol. 6, e63 (2008).
Byrne, B. M. et al. Rudimentary phosvitin domain in a minor chicken vitellogenin gene. Biochemistry 28, 2572–2577 (1989).
Bossinger, O. & Schierenberg, E. The use of fluorescent marker dyes for studying intercellular communication in nematode embryos. Int. J. Dev. Biol. 40, 431–439 (1996).
Chotard, L., Skorobogata, O., Sylvain, M. A., Shrivastava, S. & Rocheleau, C. E. TBC-2 is required for embryonic yolk protein storage and larval survival during L1 diapause in Caenorhabditis elegans. PLoS One 5, e15662 (2010).
Rezával, C., Nojima, T., Neville, M. C., Lin, A. C. & Goodwin, S. F. Sexually dimorphic octopaminergic neurons modulate female postmating behaviors in Drosophila. Curr. Biol. 24, 725–730 (2014).
Meelkop, E. et al. PDF receptor signaling in Caenorhabditis elegans modulates locomotion and egg-laying. Mol. Cell. Endocrinol. 361, 232–240 (2012).
Jee, C. et al. CNP-1 (ARRD-17), a novel substrate of calcineurin, is critical for modulation of egg-laying and locomotion in response to food and lysine sensation in Caenorhabditis elegans. J. Mol. Biol. 417, 165–178 (2012).
Wilke, A. B. & Marrelli, M. T. Genetic control of mosquitoes: population suppression strategies. Rev. Inst. Med. Trop. Sao Paulo 54, 287–292 (2012).
Van Hiel, M. B. et al. Neuropeptide receptors as possible targets for development of insect pest control agents. Adv. Exp. Med. Biol. 692, 211–226 (2010).
Gäde, G. & Goldsworthy, G. J. Insect peptide hormones: a selective review of their physiology and potential application for pest control. Pest Manag. Sci. 59, 1063–1075 (2003).
Handler, A. M. & Beeman, R. W. United States Department of Agriculture-Agricultural Research Service: advances in the molecular genetic analysis of insects and their application to pest management. Pest Manag. Sci. 59, 728–735 (2003).
Rookledge, K. A. & Heald, P. J. Dwarf eggs and the timing of ovulation in the domestic fowl. Nature 210, 1371 (1966).
Tian, X., Gautron, J., Monget, P. & Pascal, G. What makes an egg unique? Clues from evolutionary scenarios of egg-specific genes. Biol. Reprod. 83, 893–900 (2010).
Jorgensen, P., Steen, J. A., Steen, H. & Kirschner, M. W. The mechanism and pattern of yolk consumption provide insight into embryonic nutrition in Xenopus. Development 136, 1539–1548 (2009).
Lawrence, P. A. & Morata, G. Homeobox genes: their function in Drosophila segmentation and pattern formation. Cell 78, 181–189 (1994).
McGinnis, W. & Krumlauf, R. Homeobox genes and axial patterning. Cell 68, 283–302 (1992).
Rauskolb, C. & Wieschaus, E. Coordinate regulation of downstream genes by extradenticle and the homeotic selector proteins. EMBO J. 13, 3561–3569 (1994).
Laurent, A., Bihan, R., Omilli, F., Deschamps, S. & Pellerin, I. PBX proteins: much more than Hox cofactors. Int. J. Dev. Biol. 52, 9–20 (2008).
Lichtenauer, U. D. et al. Pre-B-cell transcription factor 1 and steroidogenic factor 1 synergistically regulate adrenocortical growth and steroidogenesis. Endocrinology 148, 693–704 (2007).
Bargmann, C. I. In WormBook (The C. elegans Research Community) (2006). 10.1895/wormbook.1.123.1.
Rave-Harel, N. et al. TALE homeodomain proteins regulate gonadotropin-releasing hormone gene expression independently and via interactions with Oct-1. J. Biol. Chem. 279, 30287–97 (2004).
McKay, S. J. et al. Gene expression profiling of cells, tissues and developmental stages of the nematode C. elegans. Cold Spring Harb. Symp. Quant. Biol. 68, 159–169 (2003).
Schneider, W. J. Vitellogenin receptors: oocyte-specific members of the low-density lipoprotein receptor supergene family. Int. Rev. Cytol. 166, 103–137 (1996).
Sappington, T. W. & Raikhel, A. S. Molecular characteristics of insect vitellogenins and vitellogenin receptors. Insect Biochem. Mol. Biol. 28, 277–300 (1998).
Willnow, T. E., Christ, A. & Hammes, A. Endocytic receptor-mediated control of morphogen signaling. Development 139, 4311–4319 (2012).
Gerisch, B. & Antebi, A. Hormonal signals produced by DAF-9/cytochrome P450 regulate C. elegans dauer diapause in response to environmental cues. Development 131, 1765–1776 (2004).
Mak, H. Y. & Ruvkun, G. Intercellular signaling of reproductive development by the C. elegans DAF-9 cytochrome P450. Development 131, 1777–1786 (2004).
Jia, K., Albert, P. S. & Riddle, D. L. DAF-9, a cytochrome P450 regulating C. elegans larval development and adult longevity. Development 129, 221–231 (2002).
Gerisch, B., Weitzel, C., Kober-Eisermann, C., Rottiers, V. & Antebi, A. A hormonal signaling pathway influencing C. elegans metabolism, reproductive development and life span. Dev. Cell 1, 841–851 (2001).
Mahanti, P. et al. Comparative metabolomics reveals endogenous ligands of DAF-12, a nuclear hormone receptor, regulating C. elegans development and lifespan. Cell Metab. 19, 73–83 (2014).
Sharma, K. K. et al. Synthesis and activity of dafachronic acid ligands for the C. elegans DAF-12 nuclear hormone receptor. Mol. Endocrinol. 23, 640–648 (2009).
Chen, J. & Caswell-Chen, E. P. Facultative vivipary is a life-history trait in Caenorhabditis elegans. J. Nematol. 36, 107–113 (2004).
Kamikura, D. M. & Cooper, J. A. Lipoprotein receptors and a disabled family cytoplasmic adaptor protein regulate EGL-17/FGF export in C. elegans. Genes Dev. 17, 2798–2811 (2003).
Thomas, J. H., Stern, M. J. & Horvitz, H. R. Cell interactions coordinate the development of the C. elegans egg-laying system. Cell 62, 1041–1052 (1990).
’t Hoen, P. A. C. et al. mRNA degradation controls differentiation state-dependent differences in transcript and splice variant abundance. Nucleic Acids Res. 39, 556–566 (2011).
Sarin, S. et al. Analysis of multiple ethyl methanesulfonate-mutagenized Caenorhabditis elegans strains by whole-genome sequencing. Genetics 185, 417–430 (2010).
Keeling, K. M. et al. Leaky termination at premature stop codons antagonizes nonsense-mediated mRNA decay in S. cerevisiae. RNA 10, 691–703 (2004).
Hendriks, G. J., Gaidatzis, D., Aeschimann, F. & Großhans, H. Extensive oscillatory gene expression during C. elegans larval development. Mol. Cell 53, 380–392 (2014).
Thyagarajan, B. et al. ETS-4 is a transcriptional regulator of life span in Caenorhabditis elegans. PLoS Genet. 6, e1001125 (2010).
Clifton, M. E. & Noriega, F. G. The fate of follicles after a blood meal is dependent on previtellogenic nutrition and juvenile hormone in Aedes aegypti. J. Insect Physiol. 58, 1007–1019 (2012).
Telang, A., Li, Y., Noriega, F. G. & Brown, M. R. Effects of larval nutrition on the endocrinology of mosquito egg development. J. Exp. Biol. 209, 645–655 (2006).
Van Handel, E. & Lea, A. O. Vitellogenesis in blood-fed Aedes aegypti in the absence of the heads, thorax and ovaries. J. Insect Physiol. 30, 871–875 (1984).
Zhang, J. et al. Regulation of lipoprotein assembly, secretion and fatty acid β-oxidation by Krüppel-like transcription factor klf-3. J. Mol. Biol. 425, 2641–2655 (2013).
Zhang, J. et al. Mutation in Caenorhabditis elegans Krüppel-like factor, KLF-3 results in fat accumulation and alters fatty acid composition. Exp. Cell Res. 315, 2568–2580 (2009).
Huang, P., Ma, X., Zhao, Y. & Miao, L. The C. elegans Homolog of RBBP6 (RBPL-1) regulates fertility through controlling cell proliferation in the germline and nutrient synthesis in the intestine. PLoS One 8, e58736 (2013).
Brock, T. J., Browse, J. & Watts, J. L. Genetic regulation of unsaturated fatty acid composition in C. elegans. PLoS Genet. 2, e108 (2006).
Bjerke, G. A., Hyman-Walsh, C. & Wotton, D. Cooperative transcriptional activation by Klf4, Meis2 and Pbx1. Mol. Cell. Biol. 31, 3723–3733 (2011).
Bownes, M., Lineruth, K. & Mauchline, D. Egg production and fertility in Drosophila depend upon the number of yolk-protein gene copies. Mol. Gen. Genet. 228, 324–327 (1991).
Barrière, A. & Félix, M. A. High local genetic diversity and low outcrossing rate in Caenorhabditis elegans natural populations. Curr. Biol. 15, 1176–1184 (2005).
Hodgkin, J. Innematode C. elegans ( Wood, W. ) 243–279 (Cold Spring Harbor Laboratory Press, 1988).
Hughes, S. E., Evason, K., Xiong, C. & Kornfeld, K. Genetic and pharmacological factors that influence reproductive aging in nematodes. PLoS Genet. 3, e25 (2007).
Bownes, M. Why is there sequence similarity between insect yolk proteins and vertebrate lipases? J. Lipid Res. 33, 777–790 (1992).
Blagosklonny, M. V. Answering the ultimate question ‘what is the proximal cause of aging’? Aging (Albany. NY). 4, 861–877 (2012).
Gems, D. & de la Guardia, Y. Alternative perspectives on aging in Caenorhabditis elegans: reactive oxygen species or hyperfunction? Antioxid. Redox Signal. 19, 321–329 (2013).
Gerhart, J. C. In Biological regulation and development ( Goldberger, R. F. ) 133–316 (Plenum Press, 1980).
Anderson, D. T. In Developmental systems: insects ( Counce, S. ) 95–164 (Academic Press, 1972).
Anderson, D. T. In Developmental systems: insects ( Counce, S. ) 165–242 (Academic Press, 1972).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Hodgkin, J. & Doniach, T. Natural variation and copulatory plug formation in Caenorhabditis elegans. Genetics 146, 149–64 (1997).
Hedgecock, E. M., Culotti, J. G., Thomson, J. N. & Perkins, L. A. Axonal guidance mutants of Caenorhabditis elegans identified by filling sensory neurons with fluorescein dyes. Dev. Biol. 111, 158–170 (1985).
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 314, 1–340 (1986).
Tong, Y. G. & Bürglin, T. R. Conditions for dye-filling of sensory neurons in Caenorhabditis elegans. J. Neurosci. Methods 188, 58–61 (2010).
Sulston, J. & Hodgkin, J. In The nematode Caenorhabditis elegans (Wood, W.) 587–606 (Cold Spring Harbor Laboratory Press, 1988).
Chalfie, M., Tu, Y., Euskirchen, G., Ward, W. W. & Prasher, D. C. Green fluorescent protein as a marker for gene expression. Science 263, 802–805 (1994).
Mello, C. C., Kramer, J. M., Stinchcomb, D. & Ambros, V. Efficient gene transfer in C. elegans: extrachromosomal maintenance and integration of transforming sequences. EMBO J. 10, 3959–3970 (1991).
Evans, T. C. In WormBook (The C. elegans Research Community) (2006). 10.1895/wormbook.1.108.1.
Jin, Y. In C. elegans: A practical approach (Hope, I. A.) 69–96 (Oxford University Press, 1999).
Mitani, S. Genetic regulation of mec-3 gene expression implicated the specification of the mechanosensory neuron cell types in Caenorhabditis elegans. Dev. Growth Differ. 37, 551–557 (1995).
Hobert, O. PCR fusion-based approach to create reporter gene constructs for expression analysis in transgenic C. elegans. Biotechniques 32, 728–730 (2002).
Bénard, C., Tjoe, N., Boulin, T., Recio, J. & Hobert, O. The small, secreted immunoglobulin protein ZIG-3 maintains axon position in Caenorhabditis elegans. Genetics 183, 917–927 (2009).
Minevich, G., Park, D. S., Blankenberg, D., Poole, R. J. & Hobert, O. CloudMap: a cloud-based pipeline for analysis of mutant genome sequences. Genetics 192, 1249–1269 (2012).
Blankenberg, D. et al. Manipulation of FASTQ data with Galaxy. Bioinformatics 26, 1783–1785 (2010).
Mitchell, D. H., Stiles, J. W., Santelli, J. & Sanadi, D. R. Synchronous growth and aging of Caenorhabditis elegans in the presence of fluorodeoxyuridine. J. Gerontol. 34, 28–36 (1979).
De Haes, W. et al. Metformin promotes lifespan through mitohormesis via the peroxiredoxin PRDX-2. Proc. Natl. Acad. Sci. USA 111, E2501–2509 (2014).
Vandesompele, J. et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 3, RESEARCH0034 (2002).
Jeon, M., Gardner, H. F., Miller, E. A., Deshler, J. & Rougvie, A. E. Similarity of the C. elegans developmental timing protein LIN-42 to circadian rhythm proteins. Science 286, 1141–1146 (1999).
Artyukhin, A. B., Schroeder, F. C. & Avery, L. Density dependence in Caenorhabditis larval starvation. Sci. Rep. 3, 2777 (2013).
Flibotte, S. et al. Whole-genome profiling of mutagenesis in Caenorhabditis elegans. Genetics 185, 431–441 (2010).
Acknowledgements
We thank Professors B. Grant, O. Hobert, T. Blumenthal, S. Strome, I. A. Hope for reagents, G. Minevich (Colombia University, New York) for helpful discussions on the CloudMap output, W. De Haes (KU Leuven, Belgium) for statistical analyses and N. Suetens (KU Leuven, Belgium) for excellent technical assistance. We are especially grateful to Professor Liliane Schoofs (KU Leuven, Belgium) for critical reading of the manuscript. Some strains were provided by the Caenorhabditis Genetics Center, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440). This work was supported by the Research Foundation-Flanders (FWO-Vlaanderen; grant G.0601.11.N.10) and the University of Leuven (GOA/11/002). LT and IB are FWO postdoc fellows.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: L.V.R. assisted by L.T. Performed the experiments: L.V.R. (all), assisted by C.B. (YP170 quantification, egg-laying, L1 diapause), IB (egg-laying), L.T. (mRNA quantification). Analyzed the data: L.V.R. (all) J.C. (functional tests) L.T. (mRNA). Interpreted the data: L.V.R. and L.T. Wrote the manuscript: L.V.R. and L.T., assisted by J.C., I.B. and C.B.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Van Rompay, L., Borghgraef, C., Beets, I. et al. New genetic regulators question relevance of abundant yolk protein production in C. elegans. Sci Rep 5, 16381 (2015). https://doi.org/10.1038/srep16381
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep16381
This article is cited by
-
Vitellogenin accumulation leads to reproductive senescence by impairing lysosomal function
Science China Life Sciences (2023)
-
Chronic temperature stress inhibits reproduction and disrupts endocytosis via chaperone titration in Caenorhabditis elegans
BMC Biology (2021)
-
PQM-1 controls hypoxic survival via regulation of lipid metabolism
Nature Communications (2020)
-
Maternal age generates phenotypic variation in Caenorhabditis elegans
Nature (2017)
-
Age-associated vulval integrity is an important marker of nematode healthspan
AGE (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.