Abstract
Rice blast is one of the most devastating rice diseases and continuous resistance breeding is required to control the disease. The rice blast resistance gene Pi54 initially identified in an Indian cultivar confers broad-spectrum resistance in India. We explored the allelic diversity of the Pi54 gene among 885 Indian rice genotypes that were found resistant in our screening against field mixture of naturally existing M. oryzae strains as well as against five unique strains. These genotypes are also annotated as rice blast resistant in the International Rice Genebank database. Sequence-based allele mining was used to amplify and clone the Pi54 allelic variants. Nine new alleles of Pi54 were identified based on the nucleotide sequence comparison to the Pi54 reference sequence as well as to already known Pi54 alleles. DNA sequence analysis of the newly identified Pi54 alleles revealed several single polymorphic sites, three double deletions and an eight base pair deletion. A SNP-rich region was found between a tyrosine kinase phosphorylation site and the nucleotide binding site (NBS) domain. Together, the newly identified Pi54 alleles expand the allelic series and are candidates for rice blast resistance breeding programs.
Similar content being viewed by others
Introduction
Rice is a staple food for more than half of the world’s population and therefore important for global food security. Rice blast, which is caused by the fungus Magnaporthe oryzae, is one of the most devastating rice diseases worldwide. The rice blast infection can damage almost all parts of the plant at various growth stages1,2. Frequent epidemics and outbreaks of rice blast were reported in different major rice growing countries, with yield losses ranging from 20 to 100%1,3. Considering the severity of the disease and its significant contribution to the rice yield gap, extensive research in different laboratories is addressing various aspects essential for management of the disease, including genomics, infection mechanism, host-pathogen interactions and resistance breeding.
Deployment of host plant resistance is the most effective approach for the control of rice blast. To date nearly 100 blast resistance (R) genes and over 350 quantitative trait loci (QTLs) have been identified1, of which 21 have been cloned and characterized in detail4,5,6,7,8. Molecular marker information is facilitating marker-assisted selection (MAS) for incorporating these blast resistance genes into rice breeding programs. Examples of successful deployment of blast R genes include the introgression of Piz5 and Pi54 into an elite basmati rice restorer line ‘PRR78’9 and the development of pyramided lines with Pib and Pish10 as well as Pi1, Piz-5 and Pita combinations11 in the cultivar CO39 background. Despite the availability of several cloned R genes, rice blast remains problematic because resistance is often lost in new varieties after a few years of intensive agricultural use. The rapidly evolving blast fungus can overcome the resistance conferred by major blast resistance genes. Because of the importance of the disease, M. oryzae was the first phytopathogenic fungus whose genome was sequenced. The M. oryzae genome contains retrotransposons and abundant repetitive elements in regions carrying the avirulence (Avr) genes12. To better understand the genetic variation among M. oryzae strains, genomes of three additional M. oryzae strains were re-sequenced13,14. This together with recent studies on blast Avr genes revealed high levels of non-synonymous variations as well as frequent loss and gain of Avr genes, which explains the rapid evolving nature of the blast fungus13,15.
Pertaining to the genetic variability and pathogenicity of the rice blast fungus, genes involved in host plant resistance also co-evolved with high levels of allelic and copy number variations16. Many of the major blast resistance genes are located in complex gene clusters on chromosomal loci harbouring multiple other blast resistance genes. Several of these genes also exist in multiple copy numbers, serving as evidence for the natural selection-driven R gene diversification17,18. Moreover, minor sequence variations among blast R genes, such as the presence of a few single nucleotide polymorphisms (SNPs), lead to major differences in their function or resistance spectrum. For instance, only few amino acid changes among the predicted proteins of blast resistance genes Pi2, Pi9 and Piz-t determine their resistance specificities19, with each of them conferring broad-spectrum resistance against different rice blast isolates20. In addition to the resistance spectrum difference among individual blast resistance genes, different allelic forms of major R genes also often have varied specificities. For example, a single amino acid change differentiates the resistant and susceptible alleles of rice blast resistance genes Pita and Pid221,22. Likewise, alleles of the powdery mildew resistance Pm3 gene in wheat exhibit different resistance patterns against a range of powdery mildew isolates23,24,25. Combining such alleles of a major R gene in a single genetic background or through multiline approach is a possible strategy to improve resistance in the field26,27. The presence of multiple allelic forms might reduce the selection pressure on the pathogen, thereby increasing the durability of resistance5. In addition, intragenic pyramiding that combines the specificities of different alleles is an alternative to broaden the recognition spectrum as compared to the parental alleles25. This has been successfully demonstrated in the case of Pm3 where the specificities of Pm3d and Pm3e were combined into a single allele. Moreover, a detailed molecular analysis of such allelic R gene variants facilitated understanding of host-pathogen interactions and evolution of disease resistance genes. Therefore, exploring naturally available genetic diversity for new alleles of resistance genes is an efficient approach for broadening the resistance sources against rice blast.
Seed banks are rich sources of such natural genetic diversity considering their large collections of cultivars, traditional varieties and wild relatives of many crops. This genetic potential can be accessed for identification of novel genes and/or their functional alleles. Several agriculturally relevant genes were obtained from wild relatives and landraces, such as the bacterial blight resistance gene Xa21 from O. longistaminata28 and various blast resistance genes such as Pi9 from O. minuta29, Pid3-A4 from O. rufipogon30 and Pi-hk1 from a landrace Heikezijing31. The available information on molecular markers together with advancement in sequencing technologies can be applied to dissect the molecular diversity of any favourable trait. Sequence based allele mining is one such approach32 for exploring allelic diversity of major genes controlling important traits such as disease resistance23.
Here we report the allelic diversity of a major rice blast resistance gene Pi54 in Indian germplasm. Pi54 (formerly known as Pi-kh) was first identified in the Indian rice cultivar HR22 and was cloned from the indica type rice cultivar Tetep. The gene confers broad-spectrum resistance against Indian rice blast isolates33 and is being effectively used in several rice breeding programs in India. Pi54 belongs to the ‘Coiled Coil - Nucleotide Binding Site - Leucine Rich Repeats’ (CC-NBS-LRR) class of R genes and the protein activates several downstream defence related pathways upon pathogen attack34. The new Pi54 alleles that we have identified expand the known Pi54 allelic series and could potentially serve as a genetic resource for breeding or engineering resistance to rice blast.
Results
Selection of rice genotypes for Pi54 allele mining
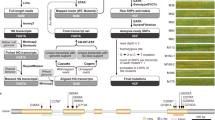
A set of 885 rice accessions originating from India and found resistant against rice blast, both under natural as well as controlled conditions, were chosen to study the allelic variation of the Pi54 gene (Fig. 1). These accessions were recorded with a resistant phenotypic score of 0-3 against a field mixture of naturally existing M. oryzae strains when screened in the uniform blast nursery (UBN). Under controlled conditions, the selected accessions were found resistant against at least one of the five tested individual blast strains35 i.e., M101-1-2-9-1, M39-1-2-21-2, JMB8401, M64-1-3-9-1 and Ca41 (Fig. 1). The 885 Indian accessions are also annotated as blast resistant in the International Rice Germplasm Collection Information System (IRGCIS) of the International Rice Genebank (IRG). The selected accessions represent three major varietal groups, indica (87.9%), javanica (8.6%) and japonica (3.5%). They were further screened for the presence of Pi54 using the ‘Pi54 MAS’ marker36, which is a functional co-dominant marker that amplifies a 216 bp fragment if a Pi54 allele is present or a 359 bp fragment if a Pi54 allele is interrupted by a 144 bp insert (often associated with the susceptible phenotype)36 (Supplementary Fig. S1). A total of 329 accessions showed the presence of the amplified 216 bp product and were selected as the candidates for sequence-based allele mining of Pi54 (Fig. 1, Supplementary Table S1).
Selection of rice accessions for Pi54 allele mining.
The 885 rice accessions that were resistant with a phenotypic score between 0 and 3 in the uniform blast nursery (UBN) as well as against any of the five pure blast isolates tested35 were screened using the Pi54MAS marker. The 329 rice accessions identified as molecular positives were selected as candidates for Pi54 allele mining. The data on number of accessions resistant (score 0–3) against each of the five blast isolates is also presented.
Isolation of new allelic forms of Pi54
Following the confirmation using the diagnostic marker ‘Pi54 MAS’, the full length coding region of the gene was amplified from all 329 accessions. The primers flanking the start and stop codons as annotated for Pi54 allele from the cultivar Tetep37 (hereafter, referred as Pi54_Tetep) were used for the sequence based amplification of Pi54 alleles. Detailed analysis of the sequences obtained from these 329 accessions identified eleven alleles of Pi54, based on the comparison with Pi54_Tetep37 which was also used as a reference sequence to assign domains. The obtained sequences were further compared with already reported Pi54 allelic sequences4,38,39,40 specifically for the ORF region as reported for Pi54_Tetep. The allele Pi54_3708 and the previously reported Pi54 alleles from a landrace Satti and the wild rice O. nivara were found identical. Similarly Pi54_40636 was found identical to alleles in the haplogroup-H2 from recently reported Pi54 alleles39. The nine new alleles we identified are unique due to the presence of unique SNPs/deletions and/or combination of shared SNPs observed in these alleles (Supplementary Table S2). Figure 2 shows the schematic representation of the sequence alignment of the identified Pi54 alleles to the reference allele Pi54_Tetep. Except for the three alleles Pi54_42439, Pi54_22419 and Pi54_3708 that were exclusively identified in single accessions, each of the remaining eight alleles was identified in several rice accessions (Table 1, Supplementary Table S2). Pi54_40996 was identified in 110 rice accessions and therefore is the most widespread Pi54 allele. The alleles Pi54_13758, Pi54_22126, Pi54_22597 and Pi54_40636 were also relatively widespread and present in the range of 34 to 53 rice accessions. In addition to the new alleles, 33 accessions have the Pi54_Tetep allele (Supplementary Table S2).
Schematic representation of sequence alignment of newly identified Pi54 alleles.
The DNA sequence alignment of 11 allelic forms of Pi54 isolated from the studied Indian accessions together with Pi54_Tetep as reference is shown. The alleles Pi54_3708 and Pi54_40636 were found identical to recently reported alleles of Pi5439 but the remaining nine alleles are unique. The domain regions of the Pi54 alleles are illustrated at the bottom as grey boxes (CC, NBS and LRR) and the brown box within LRR1 represents a zinc finger motif. The unit scale indicates the nucleotide position. The black lines on the bars as well as the numbers on the right indicate nucleotide polymorphisms compared to Pi54_Tetep. Numbers within brackets indicate the non-synonymous SNPs. The gap in the bars for certain alleles indicates 2 bp (Pi54_22419, Pi54_10202 and Pi54_13758) and 8 bp (Pi54_42439 and Pi54_40996) deletions. * indicates the alleles for which non-synonymous SNPs were not calculated due to the presence of deletions causing premature stop codons or a change in the ORF.
Sequence analysis of Pi54 alleles
The three major CC, NBS and LRR domains are conserved in the eleven Pi54 alleles identified in the Indian accessions. Six alleles (Pi54_22126, Pi54_22597, Pi54_40636, Pi54_41818, Pi54_52694 and Pi54_3708) have a complete open reading frame (ORF) similar to that of Pi54_Tetep. The remaining 5 alleles (Pi54_40996, Pi54_13758, Pi54_10202, Pi54_42439 and Pi54_22419) appear to have shorter ORFs of variable sizes due to predicted premature stop codon(s) introduced by deletions of either 2 bp (Pi54_22419, Pi54_10202 and Pi54_13758) or 8 bp (Pi54_42439 and Pi54_40996) (Fig. 2, Table 1). These eleven alleles differed from Pi54_Tetep by a total of 46 nucleotide polymorphisms, three double deletions and a stretch of 8 bp deletions present either uniquely or shared among the different alleles. Pi54_13758 is the only allele with a double deletion in the CC domain. A ‘SNP-rich region’ with nearly half of the total polymorphisms is found between nucleotides 232 and 429 (Fig. 3a). Five SNPs are present in the NBS domain of which three are synonymous changes (Fig. 2). In case of the LRR domains, one synonymous mutation was found in the Zn finger motif of the LRR1 domain and two non-synonymous mutations were found in both LRR domains, one in each (Fig. 2). All Pi54 alleles have an intact and highly conserved Zinc finger motif with only a single synonymous mutation at position 774. Similarly, the tyrosine kinase phosphorylation site is highly conserved, suggesting the significance of these motifs for Pi54 function. In addition, several post-translational modification (PTM) sites such as 13 casein kinase II (CK2) phosphorylation sites, four N-glycosylation sites, four N-myristoylation sites and three protein kinase C (PKC) phosphorylation sites were predicted in all Pi54 alleles41, but their number varied among the alleles because of SNPs and deletions (Table 2). Twenty-nine of the 46 SNPs are shared among all of the alleles except Pi54_3708, which differs from Pi54_Tetep by only 2 SNPs at positions 591 and 795. One of the two SNPs in Pi54_3708 is a synonymous mutation at position 591 that is shared with all other identified Pi54 alleles (i.e., from AAG to AAA; coding for Lysine). Seven alleles have at least one unique SNP (four of them non-synonymous) or deletions specific to that allele but absent in the other Pi54 alleles (Table 1). The remaining four alleles share the SNPs with the other alleles but in varied combinations that still makes them unique Pi54 alleles. When compared to the recently reported Pi54 alleles, 84% of the polymorphisms observed in these eleven alleles were found to be shared with several other Pi54 alleles, the majority of which were identified in cultivars and landraces reported to be resistant against the diagnostic blast isolate Mo-nwi-37-139. Additionally, several large insertions/deletions were found in recently reported Pi54 alleles39, particularly in those identified from wild rice species, but none of these INDELs were present in the eleven alleles we identified. The presence of these polymorphic yet shared nucleotide changes between alleles indicate frequent sequence exchange between the alleles, possibly by gene conversion.
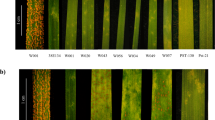
SNP rich region in the new Pi54 alleles.
(a) Selection window of the SNP-rich region (between nucleotide position 232 and 429) from the sequence alignment of the eleven studied Pi54 alleles. Dots represent the nucleotides that are identical to the Pi54-Tetep reference and the polymorphisms are represented by single letter code for nucleotides in different colours A (green), T (red), G (black) and C (blue) respectively. The domains and PTM sites are illustrated on top of the reference sequence. The unit scale on top indicates the nucleotide position. (b) Sliding-window analysis of nucleotide diversity (π) observed in the identified Pi54 alleles. The domains are illustrated below the unit scale that represents nucleotide position.
Nucleotide polymorphism analyses for the eleven Pi54 alleles were performed in DnaSP42. The overall nucleotide diversity (π) of the identified Pi54 alleles was calculated as 0.01551. The sliding window analysis of nucleotide diversity of Pi54 alleles showed a high rate of diversity within the SNP-rich region described above (Fig. 3b). The number of mutations (η-46) and the number of segregations sites (S-46) were the same, suggesting their positive selection. The negative Tajima’s D test value (−0.02431) also suggests that the alleles are under positive selection. However, the ratio of non-synonymous to synonymous substitutions per respective site (ka/ks value = 0.257) does not indicate a strong positive selection.
Analysis of predicted Pi54 proteins
The alleles with a complete ORF based on the Pi54_Tetep reference allele were subjected to protein prediction and analyses. Pi54_Tetep has a NBS domain of 21 amino acids (AA) and two non-synonymous mutations in the NBS domain were found among the studied Pi54 alleles. One of these non-synonymous mutations replaces tyrosine with phenylalanine at position 155 in all predicted proteins except Pi54_3708. The second non-synonymous mutation changes a candidate serine casein kinase II phosphorylation site at position 158 to threonine in Pi54_41818, Pi54_22126, Pi54_40996 and Pi54_42439. Pi54_Tetep has two LRR domains, each 23 AA in length. The only non-synonymous mutation in LRR1 that replaces leucine with phenylalanine at position 265 is observed in Pi54_3708 exclusively (Supplementary Table S3). The non-synonymous mutation in LRR2 changes threonine to alanine at position 272 and is shared among seven predicted Pi54 protein sequences (Pi54_10202, Pi54_40636, Pi54_22597, Pi54_41818, Pi54_22126, Pi54_40996 and Pi54_42439). The allele Pi54_22419 also has this LRR2-specific mutation but the predicted protein was not analysed because of a two bp deletion in its nucleotide sequence. The ‘PredictProtein’ server43 was used to analyse the Pi54 protein sequences of alleles with complete ORFs for secondary structure elements, disordered regions, as well as exposed and buried regions. All analysed Pi54 proteins have predicted disordered regions within their CC domains from position 15 to 36 (Fig. 4a). To predict the surface accessibility of polymorphic AA residues, the Pi54 proteins were subjected to NetSurfP analysis44. The majority of the polymorphic residues are exposed and more than half of the predicted exposed residues are located within a PTM site and/or within a specific domain of Pi54 (Fig. 4b) suggesting that these AA changes are likely critical for interaction with M. oryzae effector proteins. A single SNP that replaces methionine with leucine at AA position 214 in 10 out of 11 alleles (except Pi54_3708) results in a predicted imperfect leucine zipper-like pattern (LxxxxxxLxxxxxxLxxxxxxL) but the predicted protein structures of various alleles retain a typical horse shoe-shaped conformation (Supplementary Fig. S2).
Schematic representation of secondary structure elements, protein-binding sites, exposed/buried and disordered regions within Pi54 proteins.
(a) Protein sequences of alleles with complete ORFs as that of reference sequence Pi54_Tetep were analyzed using ‘PredictProtein’ server. (b) Surface accessibility of AA residues of Pi54 proteins predicted using NetSurfP server. ‘X’ represents polymorphic AA residues identified at least in one of the analysed Pi54 proteins. The conserved AA residues are represented by their respective single letter code. The polymorphic residues predicted as solvent exposed are highlighted in green and the ones predicted as buried are in red. The posttranslational modification sites and domain regions are illustrated.
Phylogeny of new Pi54 alleles and their distribution among rice subspecies
Five of the eleven Pi54 alleles are unique to either indica or javanica subspecies, whereas the remaining alleles (Pi54_40996, Pi54_13758, Pi54_22126, Pi54_22597 and Pi54_40636) are distributed among all three subspecies (Table 2). Pi54_3708 identified in an indica accession in the present study was previously isolated from a landrace Satti as well as from O. nivara. Pi54_10202, Pi54_41818, Pi54_52694 and Pi54_42439 were the alleles identified only in the indica subgroup. Pi54_22419 is the only allele found exclusively in javanica while no allele sequence specific to japonica was identified. The reference allele Pi54_Tetep is widespread among indica accessions and is also identified in some of the javanica accessions.
To understand the genetic relatedness among the Pi54 alleles, phylogeny of the eleven alleles was analyzed together with the previously reported allelic forms of Pi54, including the alleles from eight wild rice species4,38,40 and the recently reported Pi54 allelic sequences from Indian landraces and cultivated varieties39. A phylogenetic tree was constructed using the nucleotide sequences corresponding to the complete ORF reported for Pi54_Tetep. Four major clusters were observed and the Pi54 alleles from most of the wild rice accessions formed a separate sub-cluster within cluster II (Fig. 5), but the alleles from O. nivara, O. rufipogon and O. minuta clustered together with alleles from cultivated rice. Pi54_3708 clustered together with alleles from wild rice O. nivara, landrace Satti and other cultivars including Pi54_Tetep (Fig. 5) within cluster II. The allele Pi54_40636 clustered with alleles from landraces and other cultivated rice accessions within cluster I. Except for Pi54_3708 and Pi54_40636, all the other alleles identified in this study formed two clear sub-clusters within cluster I and III. The five alleles in cluster I are closely related to O. minuta while the four alleles falling into cluster III show relatedness to different landraces and cultivars.
Phylogenetic relationship among Pi54 alleles.
Analysis was performed using Pi54 sequences identified in our study material together with previously reported Pi54 sequences from wild rice species and cultivars. The ORF for all the sequences (including the previously reported sequences) were extracted from NCBI database based on the reported ORF for Pi54_Tetep reference sequence. Bootstrap values (1000 replications) are mentioned at the branch nodes. The alleles identified in our study material are labelled in yellow and the reference allele in green. The leaf nodes are coloured purple for wild rice, pink for landraces, green for cultivars, black for unknown status or unknown + breeding lines and blue for landraces + unknown/breeding line/cultivar, respectively.
Discussion
Considering the narrow genetic base of agriculturally grown varieties, exploration of natural genetic diversity is critically important to broaden and diversify our resistance sources against detrimental crop diseases, including rice blast. Allelic variants of an R gene may vary in their Avr recognition specificities, thereby conferring race-specific or broad spectrum resistances. For example, assessment of allelic diversity of the barley powdery mildew resistance gene Mla45 and flax rust resistance gene L46,47 revealed different functional allelic forms carrying several sites of positive selection. The wheat powdery mildew resistance gene Pm3 currently has 17 identified functional alleles, representing one of the largest allelic series characterized among plant R genes24. Such allelic variants with SNPs of additive effects are sometimes more rewarding as they might confer broad-spectrum resistance to pathogen isolates from different geographical locations. For example, the orthologue of the rice blast resistance gene Pid3 isolated from the Indian cultivar Kasalath confers broad-spectrum resistance against Chinese blast isolates (20 of 23 tested isolates) compared to Pid3 orthologues from other cultivated and wild rice varieties6. Similarly, identification of new allelic forms of cloned rice blast R genes revealed a unique spectrum of resistance among these alleles, including alleles of Pi355, alleles of Pi54 from O. rhizomatis (Pi54rh) and O. officinalis (Pi54of)4,38 and various alleles at the AC134922 locus18. Notably, Pi35 that confers quantitative broad-spectrum resistance is an allele of Pish, which confers race-specific resistance against rice blast5. Here, we investigated the allelic diversity of Pi54, a major blast resistance gene conferring resistance against rice blast in India. We identified nine new alleles of Pi54 that significantly extend the known Pi54 allelic series. Among the investigated Indian accessions, the distribution of new Pi54 alleles appeared to be enriched in indica as compared to japonica and javanica accessions.
Pi54 has a NBS-LRR structure with a CC domain at the N terminus48. The NBS domain is reported to function as a molecular switch in disease signalling and the LRR region is known to be involved in physical interactions with the AVR proteins49. Recent studies also revealed interactions of non-LRR and non-NBS regions of resistance proteins, such as CC domains, with their corresponding AVR proteins50,51,52. The CC domain of the potato Rx protein, which confers resistance against potato virus X, contains disordered regions that engage in intramolecular interaction with the NB-LRR region and intermolecular interaction with the Rx cofactor53. Our analysis of Pi54 proteins revealed disordered regions in the CC domain of all alleles, suggesting an important role of the CC domain for the functional structure of Pi54 and in intramolecular and/or intermolecular interactions. The proteins encoded by other allelic forms of Pi54 (Pi54, Pi54rh, Pi54of) interact with AVR-Pi54 using various domain as well as non-domain regions4. These studies together with our own results signify the role of non-domain regions in specificity determination. We have identified only a few polymorphic sites in the NBS and LRR domains of the new Pi54 alleles, while the majority of the sequence polymorphisms are in the region between the tyrosine kinase phosphorylation site and the NBS domain, which we refer to as the ‘SNP-rich region’. The SNP-rich region as well as the overall nucleotide polymorphism of the alleles identified in this study is shared among the fifty Pi54 haplotypes that were recently reported from Indian rice varieties39. Despite the presence of large InDels in some alleles, such highly conserved polymorphic nucleotides suggest frequent re-shuffling between Pi54 alleles by gene conversion and recombination.
We found that 67% of the amino acid changes among the minor motifs/patterns and PTM sites are also in the SNP-rich region of the gene. The majority of the polymorphisms at the PTM sites does not result in their loss but could influence the accessibility of acceptor residue(s) within a PTM site for the modifying enzymes. More than half of the polymorphic AA residues predicted as exposed are located within a PTM site. It has been suggested that the difference in number and distribution of phosphorylation motifs between the resistant Tetep and susceptible Nipponbare alleles of Pi54 are responsible for the resistance phenotype in Tetep54. We therefore expect that the observed changes in the PTM sites together with the shared and unique SNPs observed (in domain and non-domain regions) among the identified alleles might influence the protein structure, their interactions with effector proteins and downstream defence signalling. This in turn could result in novel resistance specificities. The varied patterns of resistance/susceptibility among the rice accessions containing the new Pi54 alleles to different rice blast strains further indicate a possibility of novel and/or altered resistance spectrum as compared to the known Pi54 alleles (Supplementary Table S1). However, this would need to be functionally tested using approaches such as gene silencing and/or complementation assays. Although the presented alleles were isolated from selected rice accessions that are annotated as blast resistant in IRGCIS as well as in our own screening experiments under natural and controlled conditions, the possibility for resistance in these lines due to the presence of other major R gene(s) and/or QTLs cannot be ruled out. Once functionally validated, these alleles could be used in various combinations with previously reported allelic forms of Pi54 or with other rice blast resistance genes in order to enhance the durability of blast resistance. Furthermore, detailed understanding of polymorphic sites controlling resistance will facilitate intragenic allele pyramiding to combine resistance specificities of different alleles25.
Materials and Methods
Plant material and molecular screening for Pi54
The rice germplasm material for this study was obtained from the International Rice Genebank (IRG) of the International Rice Research Institute (IRRI), Philippines. The accessions were screened for rice blast resistance against field mix-inoculum and five pure blast isolates35. Based on the standard scoring system for leaf blast55 (scale 0–9), accessions that were resistant with a phenotypic score of 0-3 against field mix-inoculum and against any of the five isolates were selected for molecular screening. Molecular screening was carried out to identify the presence of Pi54 using a functional co-dominant marker ‘Pi54 MAS’36 that differentiates Pi54 alleles with and without an insertion of 144 bp, the former being correlated with the susceptible allele. Genomic DNA extracted from Pi54 monogenic line was used as the positive control and that from susceptible rice cultivar LTH was used as the negative control for the Polymerase chain reaction (PCR), in addition to a water control.
Isolation and cloning of Pi54 alleles
Forward primer 5′-TACCTGATGGTTCTTTAAAATTGGG (designed for this study) and reverse primer 5′-CATAAGCTAGACCTTGAAGGATGTC38 were used for isolation of the full-length coding region of Pi54 alleles as annotated for Pi54 allele from cultivar Tetep37. PCR was performed with an initial denaturation at 95 °C for 5 minutes; followed by additional denaturation at 98 °C for 20 seconds, annealing at 61 °C for 20 seconds, extension at 72 °C for 1 minute (these three steps repeated for 35 cycles); followed by final extension at 72 °C for 2 minutes, using KAPA HiFi HotStart DNA polymerase (high fidelity proof-reading enzyme). The amplified products were cloned using pJET1.2 blunt end cloning vector.
Sequencing and sequence analysis
Forward primer 5′-GTTAGGCCTTCAGGAATGGAGTGC was used as internal sequence primer in addition to the standard forward and reverse primers from the pJET1.2 cloning vector for the complete sequence coverage. The primers for isolation and sequencing of Pi54 alleles were designed in DNASTAR-SeqBuilder, using Pi54_Tetep as reference (Accession Nr. AY914077). DNA sequencing was performed using Applied Biosystems Capillary Sequencer 3730. Sequence assembly, consensus development and alignments were done using ‘CLC Genomics Workbench, version-7.5′. Multiple sequence alignment was performed to identify the Single Nucleotide Polymorphisms (SNPs). Sequence polymorphism analyses such as, number of segregation sites and mutations, sliding window analysis of nucleotide diversity, ka/ks ratio and Tajima’s D test were performed using DnaSP-v542. Alleles with unique SNPs or InDels (insertion or deletion) were re-sequenced and in addition, re-amplified (from genomic DNA of respective accessions), re-cloned and re-sequenced for SNP/InDel confirmation. For the allelic forms identified in more than four accessions (thereby representing multiple independent amplification and cloning events), no repetitions were made.
Phylogenetic analysis
Phylogenetic analysis was performed using DNA sequences of the newly identified Pi54 alleles together with previously reported Pi54 alleles from cultivated rice, landraces and wild rice species to analyse their genetic relatedness4,38,39,40. The nucleotide sequences of previously reported Pi54 alleles i.e., from start to stop codon (including introns, if any) corresponding to that of Pi54_Tetep were obtained from NCBI. The sequence alignments were performed in CLC genomics workbench. The tree was constructed using ‘Neighbor-joining’ method and ‘Jukes-Cantor’ for nucleotide distance measure and bootstrap performed with 1000 replications.
Protein prediction and analysis
The protein sequences of Pi54 alleles were obtained using the EXPASY translation tool (http://web.expasy.org/translate/). The major domains such as NBS and LRR regions were assigned as previously reported for Pi5437,40. The CC domain was predicted using the COILS server (http://www.ch.embnet.org/software/COILS_form.html). The patterns and post-translational modification sites were predicted using ScanProsite server41. The secondary structure, solvent accessibility, disorders and flexibility regions of the predicted proteins were analysed using the PredictProtein server43. The surface accessibility of AA residues of Pi54 proteins were predicted in NetSurfP server44. Phyre2 server56 was used for three-dimensional protein structure predictions of proteins of the alleles with complete ORFs based on the Pi54_Tetep reference allele. The prediction parameter was set to ‘normal’ mode for all the proteins analysed. PyMOL-version 1.3 was used to visualize, analyse and edit the protein structures for their secondary structure elements and orientation.
Additional Information
Accession codes: The newly identified Pi54 sequences have been submitted to the GenBank database with accession numbers, Pi54_40996 = KR052922, Pi54_13758 = KR052923, Pi54_22126 = KR052924, Pi54_22597 = KR052925, Pi54_40636 = KR052926, Pi54_10202 = KR052927, Pi54_41818 = KR052928, Pi54_52694 = KR052929, Pi54_42439 = KR052930, Pi54_22419 = KR052931 and Pi54_3708 = KR052932.
How to cite this article: Vasudevan, K. et al. Identification of novel alleles of the rice blast resistance gene Pi54. Sci. Rep. 5, 15678; doi: 10.1038/srep15678 (2015).
Change history
06 January 2016
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Sharma, T. R. et al. Rice Blast Management Through Host-Plant Resistance: Retrospect and Prospects. Agric. Res. 1, 37–52 (2012).
Skamnioti, P. & Gurr, S. J. Against the grain: safeguarding rice from rice blast disease. Trends Biotechnol. 27, 141–50 (2009).
Khush, G. S. & Jena, K. K. Advances in Genetics, Genomics and Control of Rice Blast Disease. (Springer: Netherlands,, 2009). 10.1007/978-1-4020-9500-9
Devanna, N. B., Vijayan, J. & Sharma, T. R. The blast resistance gene Pi54of cloned from Oryza officinalis interacts with Avr-Pi54 through its novel non-LRR domains. PLoS One 9, e104840 (2014). 10.1371/journal.pone.0104840
Fukuoka, S. et al. Multiple functional polymorphisms in a single disease resistance gene in rice enhance durable resistance to blast. Sci. Rep. 4, 1–7 (2014).
Xu, X. et al. Excavation of Pid3 orthologs with differential resistance spectra to Magnaporthe oryzae in rice resource. PLoS One 9, 1–10 (2014).
Zhai, C. et al. Function and interaction of the coupled genes responsible for Pik-h encoded rice blast resistance. PLoS One 9, e98067 (2014).
Ma, J. et al. Pi64, Encoding a Novel CC-NBS-LRR Protein, Confers Resistance to Leaf and Neck Blast in Rice. MPMI 28, 558–568 (2015).
Singh, V. K. et al. Marker-assisted simultaneous but stepwise backcross breeding for pyramiding blast resistance genes Piz5 and Pi54 into an elite Basmati rice restorer line ‘PRR78’. Plant Breed. 495, 486–495 (2013).
Koide, Y. et al. Development of pyramided lines with two resistance genes, Pish and Pib, for blast disease (Magnaporthe oryzae B. Couch) in rice (Oryza sativa L.). Plant Breed. 129, 670–675 (2010).
Hittalmani, S., Parco, A., Mew, T. V., Zeigler, R. S. & Huang, N. Fine mapping and DNA marker-assisted pyramiding of the three major genes for blast resistance in rice. TAG Theor. Appl. Genet. 100, 1121–1128 (2000).
Dean, R. a et al. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 434, 980–6 (2005).
Xue, M. et al. Comparative analysis of the genomes of two field isolates of the rice blast fungus Magnaporthe oryzae. PLoS Genet. 8, e1002869 (2012).
Yoshida, K. et al. Association genetics reveals three novel avirulence genes from the rice blast fungal pathogen Magnaporthe oryzae. Plant Cell 21, 1573–91 (2009).
Huang, J., Si, W., Deng, Q., Li, P. & Yang, S. Rapid evolution of avirulence genes in rice blast fungus Magnaporthe oryzae. BMC Genet. 15, 45 (2014).
Jacob, F., Vernaldi, S. & Maekawa, T. Evolution and conservation of plant NLR functions. Front. Immunol. 4, 1–16 (2013).
Kato, H. et al. Structural diversity and evolution of the Rf-1 locus in the genus Oryza. Heredity (Edinb). 99, 516–24 (2007).
Wang, D. et al. Allele-mining of rice blast resistance genes at AC134922 locus. Biochem. Biophys. Res. Commun. 446, 1085–1090 (2014).
Zhou, B. et al. The eight amino-acid differences within three leucine-rich repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to Magnaporthe grisea. Mol. Plant. Microbe. Interact. 19, 1216–28 (2006).
Wu, K., Xu, T., Guo, C., Zhang, X. & Yang, S. Heterogeneous evolutionary rates of Pi2/9 homologs in rice. BMC Genet. 13, 73 (2012).
Bryan, G. T. et al. A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta. Plant Cell 12, 2033–46 (2000).
Chen, X. et al. A B-lectin receptor kinase gene conferring rice blast resistance. Plant J. 46, 794–804 (2006).
Bhullar, N. K., Street, K., Mackay, M., Yahiaoui, N. & Keller, B. Unlocking wheat genetic resources for the molecular identification of previously undescribed functional alleles at the Pm3 resistance locus. Proc. Natl. Acad. Sci. USA 106, 9519–24 (2009).
Bhullar, N. K., Zhang, Z., Wicker, T. & Keller, B. Wheat gene bank accessions as a source of new alleles of the powdery mildew resistance gene Pm3: a large scale allele mining project. BMC Plant Biol. 10, 1–13 (2010).
Brunner, S. et al. Intragenic allele pyramiding combines different specificities of wheat Pm3 resistance alleles. Plant J. 64, 433–45 (2010).
Chen, Y., Singh, S., Rashid, K., Dribnenki, P. & Green, A. Pyramiding of alleles with different rust resistance specificities in Linum usitatissimum L. Mol. Breed. 21, 419–430 (2007).
Brunner, S. et al. Transgenic Pm3 multilines of wheat show increased powdery mildew resistance in the field. Plant Biotechnol. J. 10, 398–409 (2012).
Ronald, P., Albano, B. & Tabien, R. Genetic and physical analysis of the rice bacterial blight disease resistance locus, Xa21. Mol. Gen. Genet. MGG 236, 113–120 (1992).
Qu, S. et al. The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rich repeat protein and is a member of a multigene family in rice. Genetics 172, 1901–14 (2006).
Lv, Q. et al. Functional analysis of Pid3-A4, an ortholog of rice blast resistance gene Pid3 revealed by allele mining in common wild rice. Phytopathology 103, 594–9 (2013).
Wu, Y. et al. Fine mapping and identification of blast resistance gene Pi-hk1 in a broad-spectrum resistant japonica rice landrace. Phytopathology 103, 1162–8 (2013).
Kumar, G. R. et al. Allele mining in crops: prospects and potentials. Biotechnol. Adv. 28, 451–61 (2010).
Rai, A. K. et al. Functional complementation of rice blast resistance gene Pi-k h (Pi54) conferring resistance to diverse strains of Magnaporthe oryzae. J. Plant Biochem. Biotechnol. 20, 55–65 (2011).
Gupta, S. K. et al. The single functional blast resistance gene Pi54 activates a complex defence mechanism in rice. J. Exp. Bot. 63, 757–72 (2012).
Vasudevan, K., Vera Cruz, C. M., Gruissem, W. & Bhullar, N. K. Large scale germplasm screening for identification of novel rice blast resistance sources. Front. Plant Sci. 5, 1–9 (2014).
Ramkumar, G. et al. Development and validation of functional marker targeting an InDel in the major rice blast disease resistance gene Pi54 (Pik h). Mol. Breed. 27, 129–135 (2011).
Sharma, T. R. et al. High-resolution mapping, cloning and molecular characterization of the Pi-k (h) gene of rice, which confers resistance to Magnaporthe grisea. Mol. Genet. Genomics 274, 569–78 (2005).
Das, A. et al. A novel blast resistance gene, Pi54rh cloned from wild species of rice, Oryza rhizomatis confers broad spectrum resistance to Magnaporthe oryzae. Funct. Integr. Genomics 12, 215–28 (2012).
Thakur, S. et al. Extensive sequence variation in rice blast resistance gene Pi54 makes it broad spectrum in nature. Front. Plant Sci. 6, 1–14 (2015).
Kumari, A. et al. Mining of rice blast resistance gene Pi54 shows effect of single nucleotide polymorphisms on phenotypic expression of the alleles. Eur. J. Plant Pathol. 137, 55–65 (2013).
De Castro, E. et al. ScanProsite: Detection of PROSITE signature matches and ProRule-associated functional and structural residues in proteins. Nucleic Acids Res. 34, 362–365 (2006).
Librado, P. & Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Yachdav, G. et al. PredictProtein-an open resource for online prediction of protein structural and functional features. Nucleic Acids Res. 42, W337–43 (2014).
Petersen, B., Petersen, T. N., Andersen, P., Nielsen, M. & Lundegaard, C. A generic method for assignment of reliability scores applied to solvent accessibility predictions. BMC Struct. Biol. 9, 51 (2009).
Seeholzer, S. et al. Diversity at the Mla powdery mildew resistance locus from cultivated barley reveals sites of positive selection. Mol. Plant. Microbe. Interact. 23, 497–509 (2010).
Ellis, J. G., Lawrence, G. J., Luck, J. E. & Dodds, P. N. Identification of regions in alleles of the flax rust resistance gene L that determine differences in gene-for-gene specificity. Plant Cell 11, 495–506 (1999).
Ravensdale, M. et al. Intramolecular interaction influences binding of the Flax L5 and L6 resistance proteins to their AvrL567 ligands. PLoS Pathog. 8, e1003004 (2012).
Young, N. D. The genetic architecture of resistance. Curr. Opin. Plant Biol. 3, 285–290 (2000).
McHale, L., Tan, X., Koehl, P. & Michelmore, R. W. Plant NBS-LRR proteins: adaptable guards. Genome Biol. 7, 212 (2006).
Cesari, S. et al. The rice resistance protein pair RGA4/RGA5 recognizes the Magnaporthe oryzae effectors AVR-Pia and AVR1-CO39 by direct binding. Plant Cell 25, 1463–81 (2013).
Kanzaki, H. et al. Arms race co-evolution of Magnaporthe oryzae AVR-Pik and rice Pik genes driven by their physical interactions. Plant J. 894–907 (2012). 10.1111/j.1365-313X.2012.05110.x
Césari, S. et al. The NB-LRR proteins RGA4 and RGA5 interact functionally and physically to confer disease resistance. EMBO J. 33, 1–19 (2014).
Hao, W., Collier, S. M., Moffett, P. & Chai, J. Structural basis for the interaction between the potato virus X resistance protein (Rx) and its cofactor ran GTPase-activating protein 2 (RanGAP2). J. Biol. Chem. 288, 35868–35876 (2013).
Kumar, S. P., Dalal, V., Singh, N. K. & Sharma, T. R. Comparative Analysis of the 100 kb Region Containing the Pi-k h Locus Between indica and japonica Rice Lines Gene prediction and annotation. Genomics. Proteomics Bioinformatics 5, 35–44 (2007).
IRRI. Standard Evaluation System for Rice 4th edn, (Manila, INGER Genetic Resource Center, 1996).
Kelley, L. A. & Sternberg, M. J. E. Protein structure prediction on the Web: a case study using the Phyre server. Nat. Protoc. 4, 363–71 (2009).
Acknowledgements
We thank the International Rice Genebank of the International Rice Research Institute (IRRI), Philippines, for providing seed material. Kind support of Dr. Casiana Vera Cruz, IRRI, for obtaining the seeds and our previously reported germplasm screening against rice blast is greatly appreciated. We acknowledge the help from Dr. Thomas Wicker for generating the schematic representation of alleles presented in Fig. 2. DNA sequencing was performed at the Genetic Diversity Centre (GDC) of ETH Zurich. We thank Dr. T.R. Sharma for time-to-time discussions regarding Pi54 alleles. The research was supported by funds from ETH Zurich to W.G and from ETH Global (previously North-South Centre) to N.K.B.
Author information
Authors and Affiliations
Contributions
N.K.B conceived and designed the experiment, K.V. carried out the experiments, K.V., W.G. and N.K.B. discussed the data, K.V. and N.K.B. wrote the manuscript, W.G. and N.K.B. edited the final manuscript. All authors have read the manuscript and agree with its content.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Com-mons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Vasudevan, K., Gruissem, W. & Bhullar, N. Identification of novel alleles of the rice blast resistance gene Pi54. Sci Rep 5, 15678 (2015). https://doi.org/10.1038/srep15678
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep15678
This article is cited by
-
Identification of Novel Alleles of the Rice Blast-Resistance Gene Pi9 through Sequence-Based Allele Mining
Rice (2020)
-
Genome wide association studies for japonica rice resistance to blast in field and controlled conditions
Rice (2020)
-
Blast resistance gene Pi54 over-expressed in rice to understand its cellular and sub-cellular localization and response to different pathogens
Scientific Reports (2020)
-
N-Glycoproteome Reveals That N-Glycosylation Plays Crucial Roles in Photosynthesis and Carbon Metabolism in Young Rice Leaves
Journal of Plant Biology (2020)
-
An update on molecular mechanism of disease resistance genes and their application for genetic improvement of rice
Molecular Breeding (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.