Abstract
The maintenance of stability during perturbations is essential for living organisms and cellular networks organize multiple pathways to enable elements to remain connected and communicate, even when some pathways are broken. Here, we evaluated the biconnectivity of the metabolic networks of 506 species in terms of the clustering coefficients and the largest biconnected components (LBCs), wherein a biconnected component (BC) indicates a set of nodes in which every pair is connected by more than one path. Via comparison with the rewired networks, we illustrated how biconnectivity in cellular metabolism is achieved on small and large scales. Defining the biconnectivity of individual metabolic compounds by counting the number of species in which the compound belonged to the LBC, we demonstrated that biconnectivity is significantly correlated with the evolutionary age and functional importance of a compound. The prevalence of diseases associated with each metabolic compound quantifies the compounds vulnerability, i.e., the likelihood that it will cause a metabolic disorder. Moreover, the vulnerability depends on both the biconnectivity and the lethality of the compound. This fact can be used in drug discovery and medical treatments.
Similar content being viewed by others
Introduction
Cellular metabolism enables the execution of numerous cellular functions on the basis of the genetic information contained within living organisms. Metabolic reactions and compounds are intricately wired and their organization may change during evolution in fluctuating environments over long periods of time1,2,3,4,5. The structures of various cellular networks have been studied for the past decade6,7,8,9,10,11,12,13,14,15 and are quite different from the structures of random networks16,17,18; they are characterized by enhanced modularity19,20 and a power-law distribution of the number of links (degree) of each node21,22. Efforts have been made to understand the origin and implications of the anomalous yet universal features of biological networks. For example, the gene duplications and mutations that accompany the inheritance of interaction links have been proposed as the mechanisms that result in broad degree distributions in cellular networks5. The topology of human cellular networks has also proven to be relevant to the pathogenesis and prevalence of clinical disorders23,24,25,26,27.
In a networked system, a path is necessary for the communication and interaction of two elements. A connected component, which is a set of nodes wherein every pair of nodes is connected by one or more paths, can act as a functional module, which comprises smaller modules in charge of different roles. The largest connected component (LCC) contains the core part of a given network28. Theoretical studies have demonstrated that networks with power-law degree distributions, referred to as scale-free (SF) networks, are capable of forming an LCC that is comparable in size to the total number of nodes, even with only a small number of links. This finding suggests that SF networks can reliably execute their core functions, even with the loss of a significant fraction of links29.
Small perturbations are often more common than large ones. Cellular networks need to maintain their regularity during small but frequent perturbations; therefore, a single path between two elements may be insufficient for the robust maintenance of their normal states. If more than one disjoint path exists between two nodes, the nodes can remain connected and therefore function normally, even with some loss in one path. The impact of multiple pathways has recently been investigated in various contexts, including the epistatic interactions of different enzymes in yeast metabolism30, the pair-wise interactions of drugs31 and the synthetic lethality of bacterial metabolism32,33.
The reaction pathways of metabolism are organized for the survival and reproduction of a species34. If there are alternatives for many pathways, the robustness under perturbation is increased, thus endowing a species with a high degree of resilience. However, the establishment of backup pathways may have a biochemical and entropic cost; the establishment of a pathway can require biochemical resources and it can take substantial time for a mutant equipped with a backup pathway of interest to appear. Therefore, the organization of alternative or backup pathways can indicate the ways in which living organisms remodel their cellular networks towards high resilience during evolution. In this work, we analyzed the structures of the metabolic networks of hundreds of living species to determine the organization of backup pathways. We consulted the BioCyc database version 13.18 to construct the bipartite metabolic networks of 506 species and to numerically determine their clustering coefficients and biconnected components (BCs). The clustering coefficient quantifies the abundance of diamond-shaped subgraphs that involve two distinct reactions that process two metabolites. In graph theory, two nodes are considered to be biconnected if there are two disjoint paths connecting them. A BC is a subset of nodes and links in which every pair of nodes is connected by two or more disjoint paths and therefore is expected to be robust against perturbations because most pairs of nodes would remain connected even if some pathways break.
We demonstrated that real networks have, compared with randomly wired networks that preserve degree distributions of both metabolites and reactions, greater clustering coefficient but smaller largest biconnected component(LBC) and indicating the scale-dependent organization of pathways for the maintenance of order. This feature is preserved even after coarse graining of nodes into modules. We subsequently defined the biconnectivity of individual compounds by counting the number of species in which the compound belonged to the LBCs. The establishment of a detour pathway is determined by the competition in terms of cost and benefit. Evolutionarily old compounds, identified here with those present in many species, were demonstrated to have high biconnectivity because they are associated with many backup pathways. Additionally, the important compounds, which may be lethal if their concentrations reach abnormal levels and which were identified with compounds located close to the biomass components, exhibited high biconnectivity, thus suggesting that abundant backup pathways are associated with compounds of functional importance. The biconnectivity of each compound is related to the maintainability of its normal state. Therefore, the likelihood of disease occurrence related to a compound is expected to depend on the compounds biconnectivity. By identifying human diseases, the pathogenesis of which could potentially involve the abnormality of each metabolic compound8,24,25,35,36,37, we demonstrated that the prevalence of the associated diseases was negatively correlated with biconnectivity. Disease prevalence should also depend on the lethality of the compound and we demonstrated that the correlation between the biconnectivity and the lethality of metabolic compounds facilitates an understanding of the disease prevalence patterns.
The work we present here is an extensive and quantitative analysis of pathway organization across hundreds of species. The obtained results offer insights into the evolutionary pressure imposed on the topology of metabolic networks that should be stable while interacting ceaselessly with fluctuating environments. We discuss the implications of our findings for human diseases and drug target searches.
Results
Biconnectivity of the metabolic network of each species
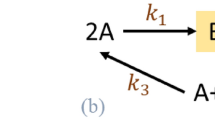
In the bipartite metabolic network for each species, a reaction is connected by undirected links to all its substrates and products [Fig. 1(a–c)]. Although the reversibility of individual reactions may provide important information, some irreversible reactions may be forced in the reverse direction under certain physiological conditions of temperature or metabolite concentrations38,39,40. The numbers of metabolic reactions, Nr, compounds, Nc and links, L, are not broadly distributed over species [Fig. 1(d)]. In contrast, the number of reactions that process a compound is highly heterogeneous. The distribution of the degree k of a compound follows a power-law  , with the exponent γ, which ranges between 2 and 3 [Fig. 1(e)]6.
, with the exponent γ, which ranges between 2 and 3 [Fig. 1(e)]6.
Properties of the metabolic networks of the 506 species investigated.
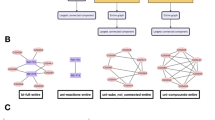
(a–c) Bipartite networks of compounds and reactions of (a) Ashbya gossypii ATCC 10895 (Nc = 46 and Nr = 24), which is known to have the smallest genome among eukaryotes60. (b) Burkholderia sp. (Nc = 1951 and Nr = 1888) and (c) human (Nc = 1513 and Nr = 1451). Here, Nc and Nr represent the numbers of compounds (red squares) and reactions (blue circles). Among the 506 species considered in this work, Burkholderia sp. has the largest metabolism and Ashbya gossypii has the smallest. (d) Distribution of Nc and Nr. Inset: Distribution of the mean degree  with
with  . (e) Distribution of the exponent γ, which characterizes the degree distribution for compounds
. (e) Distribution of the exponent γ, which characterizes the degree distribution for compounds  .
.
In this study, we applied the compound-centric view in considering the biconnectivity of the metabolic networks, because the stable production and consumption of metabolites is the most important function of metabolism. Moreover, the presence of two distinct paths is more meaningful for a pair of compounds than for a pair of reactions. When two metabolites are connected by two pathways, the production of one metabolite from the other is conducted along two pathways and can occur even if one pathway is broken. However, two pathways that connect two reactions do not guarantee the stability of both reactions.
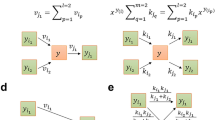
The clustering coefficient of each metabolic network was computed, as described in Methods. Using the algorithm in ref. 41, we obtained the connected components and the BCs of each metabolic network [Fig. 2(a)]. A metabolic network typically consists of a single LCC and many small connected components. The LBC lies inside the LCC and the fraction of the nodes that belong to the LBC,  , with S2 the number of nodes in the LBC and
, with S2 the number of nodes in the LBC and  , characterizes the biconnectivity of a network on the largest scale. Except for the LBC, other BCs are quite small in a given metabolic network [Fig. 2(b)].
, characterizes the biconnectivity of a network on the largest scale. Except for the LBC, other BCs are quite small in a given metabolic network [Fig. 2(b)].
Biconnected components (BC).
(a) A network of 5 compound nodes (red squares) and 5 reaction nodes (blue circles) has two BCs. BC 1, of size 5, is the LBC; thus, m2 of this network is  . (b) Distribution of the relative size
. (b) Distribution of the relative size  of a BC in the 506 species considered. Note that the fraction of nodes in each BC other than the LBC is less than 0.05.
of a BC in the 506 species considered. Note that the fraction of nodes in each BC other than the LBC is less than 0.05.
For comparison, we generated three ensembles of rewired networks; they are obtained by relocating of randomly selected links completely randomly (‘rewired-all’), by changing the end reactions of the links to randomly selected reactions, which preserves the degree of every compound (‘rewired-c’) and by exchanging the neighbor reactions of two metabolites via crosslinking, which preserves the degree distributions of both compounds and reactions (‘rewired-cr’).
The clustering coefficients, C and the fractions of the LBC, m2, were plotted as a function of the mean degree  for the real and the rewired networks of the 506 species in Fig. 3(a,b). On average, 75% of the nodes of the real networks belonged to the LBC, i.e.,
for the real and the rewired networks of the 506 species in Fig. 3(a,b). On average, 75% of the nodes of the real networks belonged to the LBC, i.e.,  and
and  . The clustering coefficient was greater than any of the considered rewired networks, thus suggesting enhanced biconnectivity on a small scale. The fraction of the LBC was smaller than the rewired-all and the rewired-cr networks; however, it was larger than the rewired-c networks.
. The clustering coefficient was greater than any of the considered rewired networks, thus suggesting enhanced biconnectivity on a small scale. The fraction of the LBC was smaller than the rewired-all and the rewired-cr networks; however, it was larger than the rewired-c networks.
Biconnectivity of the metabolic networks.
(a) Plot of the clustering coefficient C versus the mean degree  . Each of the 506 data points grouped as ‘rewired-all’ (‘rewired-c’, ‘rewired-cr’) is the average over 100 rewired-all (rewired-c, rewired-cr) networks generated for each species. Inset: Both
. Each of the 506 data points grouped as ‘rewired-all’ (‘rewired-c’, ‘rewired-cr’) is the average over 100 rewired-all (rewired-c, rewired-cr) networks generated for each species. Inset: Both  and
and  are positive, which indicates that C(real) is larger than the rewired networks. (b) Plot of the fraction of the LBC, m2 versus the mean degree
are positive, which indicates that C(real) is larger than the rewired networks. (b) Plot of the fraction of the LBC, m2 versus the mean degree  . Inset:
. Inset:  is negative, which indicates that the m2 of the real networks is smaller than the rewired-cr networks. (c) Distribution of the size of a module Nintra. P(Nintra) is broader for the real networks than the rewired networks. Inset: Plot of the number of modules Nmod versus
is negative, which indicates that the m2 of the real networks is smaller than the rewired-cr networks. (c) Distribution of the size of a module Nintra. P(Nintra) is broader for the real networks than the rewired networks. Inset: Plot of the number of modules Nmod versus  . (d) Plot of the clustering coefficient Cmod versus the mean degree
. (d) Plot of the clustering coefficient Cmod versus the mean degree  for the modular networks. The modular network is a unipartite network; thus, Cmod is evaluated following the conventional definition,
for the modular networks. The modular network is a unipartite network; thus, Cmod is evaluated following the conventional definition,  where Aij is the adjacency matrix of the modular network. (e) Plot of m2,mod versus the mean degree
where Aij is the adjacency matrix of the modular network. (e) Plot of m2,mod versus the mean degree  for the modular networks.
for the modular networks.
It has been shown42,43 that m2 depends on  and the large-k behavior of the degree distribution p(k). When
and the large-k behavior of the degree distribution p(k). When  is not too small, the random SF networks have smaller LBCs than the Erdös-Rény (ER) networks, which are constructed by randomly assigning links and thus have narrow Poisson degree distributions. Therefore, the order of the m2s of the real, rewired-all and rewired-c networks can be understood by noting that the rewired-all networks have narrower degree distributions for both reactions and compounds compared with the real networks, whereas the rewired-c networks have broader degree distributions for reactions than the real networks. It should be noted, however, that in large-scale failures in which a significant number of links are lost and
is not too small, the random SF networks have smaller LBCs than the Erdös-Rény (ER) networks, which are constructed by randomly assigning links and thus have narrow Poisson degree distributions. Therefore, the order of the m2s of the real, rewired-all and rewired-c networks can be understood by noting that the rewired-all networks have narrower degree distributions for both reactions and compounds compared with the real networks, whereas the rewired-c networks have broader degree distributions for reactions than the real networks. It should be noted, however, that in large-scale failures in which a significant number of links are lost and  becomes quite small, hub nodes, the abundance of which is responsible for broad degree distributions, play a dominant role as seeds for building connected components and BCs, leading to larger LCCs and LBCs than without hubs43,44. Therefore, the abundance of hub nodes facilitates large-scale failure survival.
becomes quite small, hub nodes, the abundance of which is responsible for broad degree distributions, play a dominant role as seeds for building connected components and BCs, leading to larger LCCs and LBCs than without hubs43,44. Therefore, the abundance of hub nodes facilitates large-scale failure survival.
The topology of metabolic networks exhibits various features, such as modularity19,20, which may be responsible for the difference between the real networks and the rewired-cr networks. Cellular metabolism consists of biochemical pathways, such as glycolysis, the TCA cycle and fatty acid pathways45 and the graph-theoretical approaches enable us to computationally identify the modules that exhibit high intraconnectivity and low interconnectivity20,46. In the application of the algorithm utilizing the maximization of modularity47, one can obtain the modules, which provide a coarse-grained view of the cellular metabolism in each species. We identified a larger number of modules Nmod in the real networks than any ensemble of the rewired networks. These modules were non-uniform in size compared with the rewired networks [Fig. 3(c)]. The networks of Nmod nodes and Lmod links, in which two nodes (modules) are connected if a link connects the nodes that belong to them in the original network, exhibit clustering coefficients and LBCs larger than the modular networks extracted from the ER networks [blue lines in Fig. 3(d,e)] for a given mean degree  . In the real networks, compared with the modular networks from the rewired-cr networks, the modular clustering coefficients were similar; however, the modular LBC, m2,mod, was substantially larger in the rewired-cr networks than the real networks, similar to m2. Given the prominent modularity of metabolic networks19,20, the smaller values of m2, a measure of the long-range biconnectivity, of the real networks than the rewired-cr networks can be related to the larger number of modules and their low biconnectivity. The backup pathways are subsequently expected to be selectively established for pathways that require a high degree of stability.
. In the real networks, compared with the modular networks from the rewired-cr networks, the modular clustering coefficients were similar; however, the modular LBC, m2,mod, was substantially larger in the rewired-cr networks than the real networks, similar to m2. Given the prominent modularity of metabolic networks19,20, the smaller values of m2, a measure of the long-range biconnectivity, of the real networks than the rewired-cr networks can be related to the larger number of modules and their low biconnectivity. The backup pathways are subsequently expected to be selectively established for pathways that require a high degree of stability.
Although the specific modules in a given network may differ depending on the algorithm used, we demonstrated that our results were not changed even when we used a different algorithm, such as the algorithm in48 (Fig. S2). To further confirm the robustness of our results, we performed the same analysis after removal of the highly abundant compounds that are involved in numerous reactions, such as protons, water, adenosine triphosphate (ATP) and carbon dioxide49, in the metabolic networks of the studied species and our results remained valid (Fig. S3). For example, even without currency metabolites49, the fraction of the LBC is only mildly changed  . We also analyzed the well-curated metabolic reconstructions of two strains of Escherichia coli and Helicobacter pylori, Pseudomonas putida, Staphylococcus aureus, Methanosarcina barkeri, Mycobacterium tuberculosis, Saccharomyces cerevisiae and Homo sapiens from the BiGG database50; our results did not qualitatively change, although some trends were difficult to identify because of the small number of data points [Fig. S4].
. We also analyzed the well-curated metabolic reconstructions of two strains of Escherichia coli and Helicobacter pylori, Pseudomonas putida, Staphylococcus aureus, Methanosarcina barkeri, Mycobacterium tuberculosis, Saccharomyces cerevisiae and Homo sapiens from the BiGG database50; our results did not qualitatively change, although some trends were difficult to identify because of the small number of data points [Fig. S4].
The evolution of species, including speciation, may have occurred along with the recruitment of new pathways, which enhanced stability, or the removal of those that were merely redundant, thus leading to the gradual variation of the biconnectivity of metabolism1,2. Using the National Center for Biotechnology Information (NCBI) taxonomy database for species classification according to morphological studies and molecular information51,52, we computed the network distance, dij, which represents the length of the shortest path in the phylogenetic tree, between every pair of species i and j to define as the evolutionary distance. Considering the normalized difference of m2 between two species i and j, defined as  , we demonstrated that
, we demonstrated that  increased with the evolutionary distance dij; the Pearson correlation coefficient (PCC) was 0.017 and the P-value (P) was smaller than 10−8. The exponent 1/2 was introduced to allow the small values of
increased with the evolutionary distance dij; the Pearson correlation coefficient (PCC) was 0.017 and the P-value (P) was smaller than 10−8. The exponent 1/2 was introduced to allow the small values of  to significantly contribute to the average over species pairs.
to significantly contribute to the average over species pairs.
Biconnectivity of individual metabolic compounds
The cross-species analysis of the BCs of the metabolic networks indicates the heterogeneity of the metabolic compounds that participate in cellular metabolism. Let us equate the ratio of the number of species that have a compound i in metabolism to the total number of all considered species by the popularity fi53. Compounds with a large f and the enzymes that bind to them, may have been introduced to metabolism in the early stages of evolution and inherited by many contemporary species. Therefore, compound popularity can be a measure of evolutionary age. The popularity is broadly distributed and follows a power-law distribution:  , where
, where  .
.
If a compound belongs to the LBC, it can maintain a connection to most core compounds; thus, its concentration can be readily stabilized, even when some pathways are inactive. We define the biconnectivity, bi, of a compound i, as the ratio of the number of species that have compound i in the LBC to the number of all species having it. The compound biconnectivity is expected to be related to the maintainability of the normal state against perturbations by utilizing alternative pathways. We demonstrated that among 6058 compounds 3071 compounds had b = 0, which indicates that they were not protected by backup pathways in their communication with the core compounds in any species. Many of these unprotected compounds were not isolated; they belong to the LCC. Defining the (single) connectivity gi as the ratio of the number of species that have a compound i in the LCC to the number of all species that have it, we demonstrated that 5097 compounds had g = 1 and 2545 compounds had g = 1 and b = 0. We hypothesize that the abnormal concentration of a compound with low biconnectivity is not sufficiently lethal to the cells overall function as to overcome the entropic and biological costs of establishing backup pathways. In contrast, 971 compounds, approximately 16% overall, belonged to the LBC in all species. Chlorides, 3-beta-hydroxysterols, brassinosteroids, trans-polyisoprenyls, amino acids, quinones, folates and uridine diphosphate glucose were the first few types of compounds that exhibited high biconnectivity. However, organosulfur, antibiotics and sugar-alcohols exhibited quite low biconnectivity. The biconnectivity distribution P(b) peaked both at b = 0 and b = 1, whereas the connectivity distribution P(g) peaked at g = 1 [Fig. 4(a)].
The cross-species biconnectivity of the metabolic compounds.
(a) The average (single-) connectivity  and the biconnectivity
and the biconnectivity  of the compounds of given popularity f. The correlation between g and f was PCC = 0.102 and between b and f was PCC = 0.358. Inset: The distributions of g and b. (b) Plot of the average biconnectivity
of the compounds of given popularity f. The correlation between g and f was PCC = 0.102 and between b and f was PCC = 0.358. Inset: The distributions of g and b. (b) Plot of the average biconnectivity  of the compounds of given closeness c. The correlation between b and c is PCC = 0.389. Inset: (left) Distribution of the closeness c. (right) Plot of
of the compounds of given closeness c. The correlation between b and c is PCC = 0.389. Inset: (left) Distribution of the closeness c. (right) Plot of  versus c with the log-binned data used. A crossover was identified from
versus c with the log-binned data used. A crossover was identified from  with
with  (solid line) to
(solid line) to  (dot-dashed line).
(dot-dashed line).
We also evaluated the biconnectivity of each of the 924 pathways, as listed in the BioCyc database, via the mean biconnectivity of the involved compounds. When the analysis was restricted to the pathways consisting of more than 10 reactions, glycolysis had the largest biconnectivity b = 0.999, which indicates that the compounds involved in these pathways had that value of biconnectivity on average. Following the glycolysis pathways were L-histidine biosynthesis (b = 0.991), L-lysine biosynthesis (b = 0.989), adenosylcobalamin biosynthesis (b = 0.973), mixed acid fermentation (b = 0.962) and folate biosynthesis (b = 0.953). In contrast, nicotine degradation (b = 0.324) had the smallest biconnectivity, followed by the tRNA charging pathway (b = 0.382), the gibberellin inactivation pathway (b = 0.436), the phospholipase pathways (b = 0.524) and mycolate biosynthesis (b = 0.557).
Evolutionarily old compounds may have had more chances to acquire alternative pathways than young compounds over time, which effectively reduced the cost of establishing backup pathways. Relating the popularity f to the evolutionary age of each compound, one can expect a positive correlation between the popularity f and the biconnectivity b, which was confirmed by our analysis shown in Fig. 4(a). The correlation between the connectivity g and f was substantially weaker.
The metabolic compounds occupy different locations in each metabolic network and play different roles of varying importance in metabolism. On average, the compounds located close to other compounds may be lethal because their abnormality can quickly spread to other compounds. The deprivation of essential enzymes can halt the production of the biomass components, as well as cell growth54. For each compound i, we evaluated the average of the inverse of the network distance to the biomass components and denoted it by the closeness ci to represent its potential lethality. We used the union of the biomass components identified for several well-studied species55, which may deviate from the true list of biomass components of each species. In ref. 55, the biomass components of B. subtilis, E. coli, H. pylori, S. aureus, M. barkeri and S. cerevisiae have been experimentally determined. Here, we consider the lethality of a compound as the probability that local perturbation around the compound can threaten the most important function of the metabolic network that generates the biomass components; furthermore, the compounds’ lethality may arise in a context other than their impact on biomass components. Figure 4(b) indicates that the biconnectivity b increases with the closeness c. This correlation implies that the metabolic networks have many loops around the biomass components and the backup pathways are not equally distributed but concentrated around functionally important (lethal) compounds. Interestingly, we identified a crossover behavior b as a function of c:  , with
, with  for small c and
for small c and  for large c, which indicates that the evolutionary pressure for backup pathways is substantially stronger for the compounds with high lethality.
for large c, which indicates that the evolutionary pressure for backup pathways is substantially stronger for the compounds with high lethality.
Implications of biconnectivity on the prevalence of human diseases
The malfunction of an enzyme accompanied by perturbation in the concentrations of related compounds is not confined and may spread in metabolism, which could result in a disorder on a large scale. Backup pathways help suppress these cascading failures. Human diseases, particularly metabolic diseases, may arise from cellular-level failures; thus, the prevalence pattern of various diseases may reflect the organization of backup pathways.
Inherited disorders are associated with the mutations of specific gene(s) and have been archived, e.g., in the Online Mendelian Inheritance in Man (OMIM) database35. In cases in which a disease gene generates enzymatic proteins that catalyze certain reactions, the perturbations of the compounds processed by the reactions may be involved in the pathogenesis of the associated diseases24,37. We consulted the OMIM database35 and the BioCyc database8 to identify the diseases potentially associated with each metabolic compound [Fig. 5(a)]. To assess the likelihood that a compound’s fluctuation would eventually lead to a human disease, we computed the mean prevalence of the associated disease. Here, the prevalence is the fraction of the patients diagnosed with a given disease among all patients in the Medicare dataset24,25,36,37. For each compound i, we took the mean prevalence of the associated diseases as its vulnerability vi.
The vulnerability and the biconnectivity of the metabolic compounds.
(a) The diseases associated with glycerate-2P. The compound (red square) is processed by three reactions (blue ellipses) denoted by the Enzyme Commission (EC) number. The genes (triangles) that generate the enzymatic proteins catalyzing the reactions are also shown, some of which are involved in the pathogenesis of three inherited diseases (trapezoids). The vulnerability of glycerate-2P is given by the mean of the prevalence values of the three diseases,  . (b) Plots of the average vulnerability
. (b) Plots of the average vulnerability  of the metabolic compounds in the LBC
of the metabolic compounds in the LBC  LBC), outside the LBC but in the LCC
LBC), outside the LBC but in the LCC  LBC,
LBC,  LCC) and outside the LCC
LCC) and outside the LCC  LCC) of the human metabolic network. (c) Distribution of the biconnectivity b for the three groups of the compounds considered in (b). (d) The average vulnerability
LCC) of the human metabolic network. (c) Distribution of the biconnectivity b for the three groups of the compounds considered in (b). (d) The average vulnerability  of the compounds of a given biconnectivity b and closeness c. The correlation between v and b was negative,
of the compounds of a given biconnectivity b and closeness c. The correlation between v and b was negative,  . In contrast, the correlation between v and c was not significant,
. In contrast, the correlation between v and c was not significant,  . (e) Plot of
. (e) Plot of  versus b for the compounds in the LBC. (f) Plot of
versus b for the compounds in the LBC. (f) Plot of  versus b for the compounds outside the LBC and in the LCC. (g) The vulnerability
versus b for the compounds outside the LBC and in the LCC. (g) The vulnerability  as a function of biconnectivity b and closeness c.
as a function of biconnectivity b and closeness c.
The human metabolic network has 1513 compounds that are processed by 1451 reactions and only 754 compounds are associated with 477 diseases. The compounds that have multiple pathways to the core compounds are less likely to cause diseases than the compounds without multiple pathways. In Fig. 5(b), the disease compounds in the LCC but outside the LBC of the human metabolic network have an average vulnerability  , which is larger than
, which is larger than  of the human LBC (P = 0.002). In contrast, the disease compounds located outside the LCC have
of the human LBC (P = 0.002). In contrast, the disease compounds located outside the LCC have  , which is smaller than the compounds in the LCC because local fluctuations in the small components do not affect the LCC in which most elements reside.
, which is smaller than the compounds in the LCC because local fluctuations in the small components do not affect the LCC in which most elements reside.
An example of a disease that is associated with low-b and a high prevalence is depressive disorder with an accelerated response to antidepressant drug treatment, with  . It is associated with peptidyl-proline, in which b = 0. Hyperproteinemia associated with angiotensinogen in which b = 0 has
. It is associated with peptidyl-proline, in which b = 0. Hyperproteinemia associated with angiotensinogen in which b = 0 has  . Renal carcinoma associated with 12 compounds that have
. Renal carcinoma associated with 12 compounds that have  has
has  . Other examples include renal tubular dysgenesis (b = 0,
. Other examples include renal tubular dysgenesis (b = 0,  and dyskeratosis congenita (b = 0.0046 and
and dyskeratosis congenita (b = 0.0046 and  .
.
Whether a disease compound belongs to the LBC and the LCC of the human metabolic network depends on the cross-species biconnectivity b. The disease compounds outside the human LBC have b = 0, with few exceptions [Fig. 5(c)]. Many disease compounds in the human LBC have b = 1, whereas some have b < 1. Diverse features of the human metabolic networks may contribute to preventing local perturbations from spreading and the compound biconnectivity may provide the relevant information. The compounds with high biconnectivity are less vulnerable than the compounds that exhibit low biconnectivity. In Fig. 5(d), the vulnerability v is shown to decrease with the biconnectivity b  . This correlation was stronger when we disregarded the currency metabolites in each metabolic network
. This correlation was stronger when we disregarded the currency metabolites in each metabolic network  [Supplementary Material]. The vulnerability of the 498 compounds in the human LBC exhibited a negative correlation with their biconnectivity
[Supplementary Material]. The vulnerability of the 498 compounds in the human LBC exhibited a negative correlation with their biconnectivity  , thus implying that the compounds with high biconnectivity are stabilized better than the compounds with low biconnectivity, while they are all in the human LBC.
, thus implying that the compounds with high biconnectivity are stabilized better than the compounds with low biconnectivity, while they are all in the human LBC.
The vulnerability of a compound depends on its lethality and its capacity to maintain its normal state, which can be roughly represented as follows:

The closeness ci of a compound can quantify its lethality and the biconnectivity bi represents a measure of the maintainability of the normal state under perturbations. The vulnerability v is expected to increase with the closeness c if other conditions are identical. However, the lethality and the maintainability are not independent of each other; evolutionary pressure is imposed to enhance the maintainability of lethal compounds, as supported by the correlation between the closeness c and the biconnectivity b shown in Fig. 4(b). The correlation between b and c makes it difficult to identify their independent contributions to vulnerability, as indicated in the weak correlation between v and c in Fig. 5(d). Interestingly, for compounds outside the human LBC, a higher biconnectivity is associated with greater vulnerability [Fig. 5(f)], which contrasts the negative correlation identified for the disease compounds in the human LBC. To understand the reason for this difference, let us assume that the lethality and the maintainability are proportional to the closeness c and the biconnectivity b, respectively, in Eq. (1). They are correlated with each other as shown in Fig. 4(b), in which  with
with  for small c
for small c  and
and  for large c
for large c  . Inserting
. Inserting  into
into  of Eq. (1), we find that
of Eq. (1), we find that  . Therefore, v decreases with b if α > 1, which may be the case for the compounds in the human LBC that have a large b. However, v increases with b if α < 1, which is valid for the compounds outside the LBC that have a small b. Figure 5 (g) shows the plot of the vulnerability as a function of closeness c and biconnectivity b. Both plots in Fig. 5(e,f) indicate that v peaks at intermediate values of b, which results from v increasing with b for small b and decreasing with b for large b.
. Therefore, v decreases with b if α > 1, which may be the case for the compounds in the human LBC that have a large b. However, v increases with b if α < 1, which is valid for the compounds outside the LBC that have a small b. Figure 5 (g) shows the plot of the vulnerability as a function of closeness c and biconnectivity b. Both plots in Fig. 5(e,f) indicate that v peaks at intermediate values of b, which results from v increasing with b for small b and decreasing with b for large b.
These findings suggest that the network properties of the metabolic compounds in other species can be helpful for understanding human diseases. The cross-species features reflect the generic features of each compound identified in diverse metabolic networks that have evolved in different ways; thus, they carry information that could not be identified by only analyzing the human metabolic network. Our findings could be useful in the search for drug targets and medical treatments. For example, if we must choose one of two candidate target compounds to set up alternative pathways and thereby suppress the spread of perturbations initiated somewhere in metabolism, we can first select the compound with increased biconnectivity. We expect this approach to establish the additional pathway and make it stable relatively easily.
Discussion
Our study illustrates a novel use of the cross-species cellular network data that are rapidly accumulating in this post-genomic era. Metabolic pathway organization is essential to allow living organisms to be resilient to perturbations; thus, we performed an extensive quantitative analysis of the biconnectivity of the cellular metabolism in hundreds of species. As a result, we were able to understand how biconnectivity is achieved on small and large scales in real metabolic networks in the presence of other structural features, such as modularity and heterogeneous connectivity patterns. The idea that biconnectivity serves for resilience to perturbations is supported by the negative correlation between the prevalence of associated diseases and the biconnectivity of compounds. Furthermore, we demonstrated that the pattern of disease prevalence can be better understood by considering the dependence of the compound vulnerability on both the lethality and the biconnectivity, suggesting its potential use in drug design and medical treatments.
Our study does have several limitations. We used the biomass components known in the well-curated metabolic reconstructions55,56 as the biomass components of the species studied in our work as long as they were present in the metabolic network of a given species, which must be curated by plugging the species-specific biomass components. The development of a biconnectivity measure that incorporates the direction of chemical reactions could better elucidate pathway organization in metabolism. Although we focused on individual compounds, the properties of pairs of compounds in terms of their multiple paths deserve investigation. The present work focused on the structure of metabolic networks; however, an understanding of the dynamics, e.g., flux-balance modeling, would allow us to identify different sets of pathways that are active in wild-type and perturbation conditions, which would thereby facilitate an understanding of homeostatic mechanisms in living organisms. It would also be interesting to investigate structural biconnectivity and dynamical flux-coupling relations. The idea that biconnected compounds are resilient to perturbations can be confirmed and better supported by in-depth studies regarding the stoichiometry of specific compounds.
Methods
Database
We used the BioCyc database version 13.18, which included 506 species, to construct the bipartite metabolic networks of 453 bacteria, 34 archaea and 19 eukarya. The latest version is 19.1, which covers 5700 metabolic pathways as of 2015. The same compounds in different compartments were considered to reflect different nodes. The transport reactions between different compartments were included.
Clustering coefficient of a bipartite network
A diamond-shaped subgraph (a quad) that involves two metabolic compounds and two reactions involves two disjoint pathways that connect the two compounds. For a bipartite metabolic network of the adjacency matrix Acr, we defined the clustering coefficient as follows:

which is the ratio of the total number of quads that involve distinct pairs of compounds to the total number of reactions that involve pairs of compounds. If two compounds had k distinct reactions processing them, the number of quads would be k2. Therefore, C would be larger than 1 if all pairs of compounds had more than one reaction processing them. If all pairs of compounds were involved in only one reaction, C would be zero. This definition of the clustering coefficient is rather different from the previous definitions for bipartite networks57,58,59.
Popularity, biconnectivity and closeness
Let  be the set of the 506 species considered in this work. Using
be the set of the 506 species considered in this work. Using  or 0 to represent whether a species
or 0 to represent whether a species  has a compound i or not, respectively and
has a compound i or not, respectively and  or 0 to represent whether a species
or 0 to represent whether a species  has the compound i in the LBC or not, respectively, we can represent the popularity and the biconnectivity as
has the compound i in the LBC or not, respectively, we can represent the popularity and the biconnectivity as  and
and  , respectively. Considering the set
, respectively. Considering the set  of the 129 biomass components identified in at least one species, i.e., B. subtilis, E. coli, H. pylori, S. aureus, M. barkeri and S. cerevisiae55,56, we first made the set of biomass components
of the 129 biomass components identified in at least one species, i.e., B. subtilis, E. coli, H. pylori, S. aureus, M. barkeri and S. cerevisiae55,56, we first made the set of biomass components  of a species
of a species  . We subsequently computed the network distance
. We subsequently computed the network distance  between a compound i and each biomass component
between a compound i and each biomass component  in each species
in each species  and evaluated the closeness as
and evaluated the closeness as  .
.
Additional Information
How to cite this article: Kim, P. et al. Biconnectivity of the cellular metabolism: A cross-species study and its implication for human diseases. Sci. Rep. 5, 15567; doi: 10.1038/srep15567 (2015).
References
Horowitz, N. H. On the evolution of biochemical syntheses. Proc. Natl. Acad. Sci. USA 31, 153 (1945).
Jensen, R. A. Enzyme recruitment in evolution of new function. Annu. Rev. Microbiol. 30, 409 (1976).
Yčas, M. On earlier states of the biochemical system. J. theor. Biol. 44, 145 (1974).
Light, S. & Kraulis, P. Network analysis of metabolic enzyme evolution in Escherichia coli. BMC Bioinformatics 5, 15 (2004).
Yamada, T. & Bork, P. Evolution of biomolecular networks - lessons from metabolic and protein interactions. Nat. Rev. Mol. Cell Biol. 10, 791 (2009).
Jeong, H., Tombor, B., Albert, R., Oltvai, Z. N. & Barabási, A. L. The large-scale organization of metabolic networks. Nature 407, 651 (2000).
Fell, D. A. & Wagner, A. The small world of metabolism. Nat. Biotech. 18, 1121 (2000).
Karp, P. D. et al. Expansion of the BioCyc collection of pathway/genome databases to 160 genomes. Nucleic Acids Res. 33, 6083 (2005).
Duarte, N. C. et al. Global reconstruction of the human metabolic network based on genomic and bibliomic data. Proc. Natl. Acad. Sci. USA 104, 1777 (2007).
Lee, T. I. et al. Transcriptional Regulatory Networks in Saccharomyces cerevisiae. Science 298, 799 (2002).
Babu, M. M., Luscombe, N. M., Aravind, L., Gerstein, M. & Teichmann, S. A. Structure and evolution of transcriptional regulatory networks. Curr. Opin. Struct. Biol. 14, 283 (2004).
Uetz, P. et al. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature 403, 623 (2000).
Xenarios, I. et al. DIP, the Database of Interacting Proteins: a research tool for studying cellular networks of protein interactions. Nucleic Acids Res. 30, 303 (2002).
Rual, J.-F. et al. Towards a proteome-scale map of the human protein-protein interaction network. Nature 437, 1173 (2005).
Krogan, N. J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637 (2006).
Barabási, A.-L. & Oltvai, Z. N. Network biology: understanding the cell’s functional organization. Nat. Rev. Genet. 5, 101 (2004).
Albert, R. Scale-free networks in cell biology. J. Cell Sci. 118, 4947 (2005).
Alon, U. An Introduction to Systems Biology: Design Principles of Biological Circuits (Chapman and Hall/CRC, 2006).
Shen-Orr, S. S., Milo, R., Mangan, S. & Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet. 31, 64 (2002).
Ravasz, E., Somera, A., Mongru, D., Oltvai, Z. & Barabási, A.-L. Hierarchical Organization of Modularity in Metabolic Networks. Science 297, 1551 (2002).
Barabási, A.-L. & Albert, R. Emergence of Scaling in Random Networks. Science 286, 509 (1999).
Albert, R. & Barabási, A.-L. Statistical mechanics of complex networks. Rev. Mod. Phys. 74, 47 (2002).
Goh, K.-I. et al. The human disease network. Proc. Natl. Acad. Sci. USA 104, 8685 (2007).
Lee, D.-S. et al. The implications of human metabolic network topology for disease comorbidity. Proc. Natl. Acad. Sci. USA 105, 9880 (2008).
Park, J., Lee, D.-S., Christakis, N. A. & Barabási, A.-L. The impact of cellular networks on disease comorbidity. Mol. Sys. Biol. 5, 262(2009).
Barabasi, A.-L., Gulbahce, N. & Loscalzo, J. Network medicine: a network-based approach to human disease. Nat. Rev. Genet. 12, 56 (2011).
Vidal, M., Cusick, M. E. & Barabási, A.-L. Interactome Networks and Human Disease. Cell 144, 986 (2011).
Almaas, E., Oltvai, Z. N. & Barabási, A.-L. The Activity Reaction Core and Plasticity of Metabolic Networks. PLoS Comput. Biol. 1, e68 (2005).
Albert, R., Jeong, H. & Barabási, A.-L. Error and attack tolerance of complex networks. Nature 406, 378 (2000).
Segre, D., DeLuna, A., Church, G. M. & Kishony, R. Modular epistasis in yeast metabolism. Nat. Genet. 37, 77 (2005).
Yeh, P., Tschumi, A. I. & Kishony, R. Functional classification of drugs by properties of their pairwise interactions. Nat. Genet. 38, 489 (2006).
Ghim, C.-M., Goh, K.-I. & Kahng, B. Lethality and synthetic lethality in the genome-wide metabolic network of Escherichia coli. J. theor. Biol. 237, 401 (2005).
Shen, Y. et al. Blueprint for antimicrobial hit discovery targeting metabolic networks. Proc. Natl. Acad. Sci. USA 107, 1082 (2010).
Schmidt, S., Sunyaev, S., Bork, P. & Dandekar, T. Metabolites: a helping hand for pathway evolution? Trends Biochem. Sci. 28, 336 (2003).
Online mendelian inheritance in man, omim®. McKusick-Nathans Institute of Genetic Medicine, Johns Hopkins Univer- sity (Baltimore, MD), World Wide Web URL: http://omim.org/ (Date of access : 21/02/2014).
Hidalgo, C. A., Blumm, N., Barabási, A.-L. & Christakis, N. A. A Dynamic Network Approach for the Study of Human Phenotypes. PLoS Comput. Biol. 5, e1000353 (2009).
Lee, D.-S. Interconnectivity of human cellular metabolism and disease prevalence. J. Stat. Mech.: Theor. Exp. 2010, P12015 (2010).
Faust, K., Dupont, P., Callut, J. & van Helden, J. Pathway discovery in metabolic networks by subgraph extraction. Bioinformatics 26, 1211 (2010).
Croes, D., Couche, F., Wodak, S.J. & van Helden, J. Metabolic PathFinding: inferring relevant pathways in biochemical networks. Nucleic Acids Res. 33, W326 (2005).
Patil, K. R. & Nielsen, J. Uncovering transcriptional regulation of metabolism by using metabolic network topology. Proc. Natl. Acad. Sci. USA 102, 2685 (2005).
Hopcroft, J. & Tarjan, R. Algorithm 447: efficient algorithms for graph manipulation. Comm. ACM 16, 372 (1973).
Newman, M. & Ghosal, G. Bicomponents and the Robustness of Networks to Failure. Phys. Rev. Lett. 100, 138701 (2008).
Kim, P., Lee, D.-S. & Kahng, B. Phase transition in the biconnectivity of scale-free networks. Phys. Rev. E 87, 022804 (2013).
Lee, D.-S., Goh, K.-I., Kahng, B. & Kim, D. Evolution of scale-free random graphs: Potts model formulation. Nucl. Phys. B 696, 351 (2004).
Michal, G. Biochemical Pathways: An Atlas of Biochemistry and Molecular Biology 2nd edn (eds Michal, G. et al.) (Wi- ley, 1999).
Girvan, M. & Newman, M. E. J. Community structure in social and biological networks. Proc. Natl. Acad. Sci. USA 99, 7821 (2002).
Clauset, A., Newman, M. E. J. & Moore, C. Finding community structure in very large networks. Phys. Rev. E 70, 066111 (2004).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech.: Theor. Exp. 2008, P10008 (2008).
Holme, P. & Huss, M. Currency metabolites and network representations of metabolism. arXiv:0806.2763v1 (2008).
Schellenberger, J., Park, J.O., Conrad, T.M. & Palsson, B. Ø. BiGG: a Biochemical Genetic and Genomic knowledgebase of large scale metabolic reconstructions. BMC Bioinformatics 11, 213 (2010).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 37, D5 (2009).
Benson, D. A., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J. & Sayers, E. W. GenBank. Nucleic Acids Res. 37, D26 (2009).
Bernhardsson, S., Gerlee, P. & Lizana, L. Structural correlations in bacterial metabolic networks. BMC Evol. Biol. 11, 20 (2011).
Palsson, B. Ø. Systems Biology: Properties of Reconstructed Networks (Cambridge University Press, 2006).
Systems biology research group: Downloads, University of California (San Diego, CA), World Wide Web URL: http://systemsbiology.ucsd.edu/Downloads (Date of access : 31/12/2013).
Han, K. Classification of organisms using backup reactions in metabolic networks. Master thesis, Seoul National University (2010).
Lind, P. G., González, M. C. & Herrmann, H. J. Cycles and clustering in bipartite networks. Phys. Rev. E 72, 056127 (2005).
Opsahl, T. Triadic closure in two-mode networks: Redefining the global and local clustering coefficients. arXiv:1006.0887 (2010).
Kitsak, M. & Krioukov, D. Hidden variables in bipartite networks. Phys. Rev. E 84, 026114 (2011).
Dietrich, F. S. et al. The Ashbya gossypii Genome as a Tool for Mapping the Ancient Saccharomyces cerevisiae Genome. Science 304, 304 (2004).
Acknowledgements
This work was supported by National Research Foundation of Korea (NRF) grants funded by the Korean Government (MSIP and NCRI) (No. 2014-069005 (BK) and Nos 2012R1A1A2005252 and 2013R1A1A201068845 (DSL)).
Author information
Authors and Affiliations
Contributions
D.-S.L. and B.K. conceived the project; P.K. performed the simulations; all authors interpreted the results and contributed to the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kim, P., Lee, DS. & Kahng, B. Biconnectivity of the cellular metabolism: A cross-species study and its implication for human diseases. Sci Rep 5, 15567 (2015). https://doi.org/10.1038/srep15567
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep15567
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.