Abstract
Previous publications on stapled peptide inhibitors against Mdm2/Mdm4-p53 interactions have established that this new class of drugs have the potential to be easily optimised to attain high binding affinity and specificity, but the mechanisms controlling their cellular uptake and target engagement remain elusive and controversial. To aid in understanding the rules of peptide and staple design and to enable rapid optimisation, we employed the newly-developed cellular thermal shift assay (CETSA). CETSA was able to validate stapled peptide binding to Mdm2 and Mdm4 and the method was also used to determine the extent of cellular uptake, cellular availability and intracellular binding of the endogenous target proteins in its native environment. Our data suggest that while the stapled peptides engage their targets intracellularly, more work is needed to improve their cellular entry and target engagement efficiency in vivo. CETSA now provides a valuable tool to optimize such in vivo properties of stapled peptides.
Similar content being viewed by others
Introduction
Commonly dubbed the ‘guardian of the genome’, the tumour suppressor p53 has been found to be mutated in half of all cancers1,2 and with more than 70 000 publications describing the gene, it is undoubtedly one of the most important and well-studied cancer-related genes. About 50% of cancers lack p53 activity due to mutations within the coding regions while the other 50% have aberrations that, in one way or another, reduce the function of the wild type p53 pathway3. In either situation, the transcriptional role of p53 is perturbed, resulting in the inability to activate or repress p53 target genes that govern cell cycle arrest or apoptosis. In additional, transcription-independent roles of p53 are disrupted. Ultimately, the lack of p53 activity results in unfettered cell division, genomic instability and cellular transformation.
Wild type p53 can be reactivated by inhibiting its structurally similar negative regulators Mdm2 and Mdm4. Blocking the interactions between p53 and Mdm2/Mdm4 prevents the occlusion of the p53 transactivation domain, as well as preventing p53 nuclear export, ubiquitination and proteasomal degradation4,5. This results in the accumulation of p53, allowing both p53 transcription-dependent and independent activity, fulfilling its role as a tumour suppressor. Small molecules, like nutlin-36 or MI-2197, have been developed that mimic the p53 N-terminal region that interfaces with Mdm2 (reviewed in Brown et al. 20098). More recently, the Mdm2 binding motif of p53 has been used to engineer peptides that bind to Mdm2 and Mdm49,10,11. To maintain peptide helicity and to allow additional contacts with Mdm2/Mdm4, hydrocarbon staples have been added, increasing the affinity and activity of some of these stapled peptides12. Furthermore, hydrocarbon stapling vastly improves the helical stability of the peptides, promotes their cellular uptake and protects them from protease degradation.
The therapeutic effects of modern oncology drugs are obtained by the specific interaction of drugs with their molecular targets, often simply described as “targeted therapy” to distinguish these approaches from more generic cytotoxic drugs and radiation therapy. Therefore, in drug design and optimisation, the knowledge of how well a drug binds to its target is of utmost importance in determining drug efficacy and understanding potentially deleterious side effects. There are several methods to assay for drug-target engagement and one of the most direct methods is through the use of thermal shift assays13,14,15,16. These assays utilise the principle that protein conformation is stabilised by interactions with ligands and the proteins therefore undergo denaturation at higher temperatures. The amount of thermal shift in the presence of ligands correlates well with ligand affinity17 and can be used to approximate binding affinities between drugs and their target proteins. Traditionally, these assays require the use of purified proteins, representing a challenge in the case of proteins that are difficult to isolate. Recently, cellular thermal shift assays have been developed to overcome this requirement, using crude cell lysates or even intact cells and tissues rather than purified proteins18. Instead of fluorescent dyes that bind to hydrophobic protein cores, thermal denaturation is measured by separating the soluble proteins with centrifugation, followed by Western blotting to quantitate the protein of interest. The added benefit is that CETSA allows the observation of thermal shifts of proteins in their cellular environments, which is more representative of the true target environment, taking account of the presence of other cellular proteins, lipids and biomolecules at high concentrations. CETSA can be also used to assay for drug entry and target engagement within cells18.
There is a demand in establishing rapid and reliable assays to deconvolute the complexity of targeting intracellular protein-protein interactions in therapeutics19. With increasing evidence that targeting the Mdm2/Mdm4/p53 axis is crucial in the treatment of several types of malignancies, the dual inhibition of both Mdm2 and Mdm4 is potentially a key strategy in therapeutics20,21. In this study, we utilised CETSA to assay for the efficacy of Mdm2/Mdm4 inhibitory stapled peptides, both in terms of Mdm2/Mdm4 engagement in cells and cell lysates, as well as for entry and availability in cells. The stapled peptides were found to induce large thermal shifts in the stability of their target proteins in cell lysates largely in concordance to their affinities estimated using other techniques with purified proteins, though discrepancies may arise from the use of different approaches to estimate binding affinities. In CETSAs involving whole cells, the thermal shifts were repressed compared to CETSAs involving cell lysates, indicating the poorer efficiency of the stapled peptides in reaching their targets within cells. This study provided evidence that CETSA is an immensely useful tool for drug design and optimisation, not just in target affinity, but also in cell entry and availability and is capable of dealing with novel drug classes such as larger peptide molecules.
Materials and Methods
Peptide synthesis and purification
Materials
Ramage Chemmatrix resin was obtained from PCAS-Biomatrix (Quebec, Canada). L-amino acids were obtained from Advanced Chemtech (Louisville, KY). Fmoc-threonine, serine, glutamic acid and tyrosine were t-butyl protected and Fmoc-tryptophan was used without BOC protection. Unnatural alkenyl amino acids, (S)-N-Fmoc-2-(4’-pentenyl)alanine and (R)-N-Fmoc-2-(7’-octenyl)alanine denoted as Xs and Xr respectively on the peptide sequences were purchased from OKeanos (Beijing, China). 5-FAM was obtained from GL Biochem (Shanghai, China). All other solvents and reagents were obtained from Sigma-Aldrich. 1, 2-dichloroethane (DCE) was dried overnight over activated molecular sieves and purged with Argon for 30 min prior to use. All other reagents were used as received.
Peptide synthesis
The stapled peptides were synthesized manually by Fmoc chemistry at the 0.1 mmol scale using Ramage Chemmatrix resin (0.53 mmol/g). The dry resin was swelled with 1-methyl-2-pyrrolidinone (NMP) before use. The Fmoc protecting group was removed by treatment with 20% piperidine in NMP (15 min). Solutions of the Fmoc-protected amino acids (5 equiv.), diisopropylcarbodiimide (DIC, 5 equiv.) and 1-aza benzotriazole (HOAt, 5 equiv.) in NMP (0.5 M) were activated for 7 min and added to the resin. The coupling time was 1 hour for all amino acids except for (S)-N-Fmoc-2-(4’-pentenyl)alanine and (R)-N-Fmoc-2-(7’-octenyl)alanine, denoted Xs and Xr, respectively. Those alkenyl amino acids (4 equiv.) were coupled to the peptide resin for two hours. Following deprotection of the final Fmoc group, the peptides were acetylated using a mixture of acetic anhydride/diisopropylethylamine/DMF (2/2/1) for 60 min. After each coupling, deprotection and acetylation reaction, the resin was thoroughly washed with NMP, methanol and diethyl ether.
Ring-closing metathesis of resin-bound, N-acetylated peptides was performed manually using a 5 mg/mL solution of Grubbs I catalyst (20 mol%) in dry 1,2-dichloroethane (DCE) at room temperature under an atmosphere of inert argon (3 × 2 h treatments). After the reaction, the solution was drained, the resin washed with DCE (3 × 1 min), DMSO (1 × 2 h) and MeOH (3 × 1 min) then dried in vacuo overnight. Cleavage of the peptide from the resin was achieved using 8 mL of TFA cocktail consisting trifluoroacetic acid/triisopropylsilane/water (95/2.5/2.5) for 2 h followed by filtration and precipitation with diethyl ether. The precipitate was collected by centrifugation, dried and redissolved in a 3:2 mixture of acetonitrile and water.
The 5-FAM was incorporated to the peptide resin after the Fmoc deproctection of the cyclized peptide. 5-FAM (7 equiv) was dissolved in a mixture of DMSO/ DMF 2:1 preactivated with DIC/HOAt (7 equiv) and added to the resin overnight.
The pure peptides (>90% purity) were obtained by purification using a preparative HPLC system (Agilent) on a Jupiter C12 reversed-phase preparative column (Phenomenex, 4 μm, Proteo 90 Å, 250 × 10 mm). The structures of the stapled peptides are depicted in Supplementary Figures 1 and 2.
Cell culture and drug treatments
OCI/AML-2 cells (DMSZ) were cultured in MEM Alpha supplemented with 20% foetal bovine serum (FBS) and penicillin/streptomycin, in a 37 °C humidified incubator with 5% CO2 atmosphere. 24 hours prior to drug treatment, cells were transferred to MEM Alpha containing 10% FBS. Cells were pre-treated with 10 μM MG132 for one hour, before harvesting for cell lysate CETSA or treatment for whole cell CETSA. For whole cell CETSAs, cells were treated with the indicated amounts of nutlin-3 or stapled peptides for four hours before collection. The total amount of DMSO, used to dissolve the compounds, was fixed at 0.35% during incubation.
T22 p53 β-galactosidase based reporter assay
The T22 p53 β-galactosidase based reporter assay was performed as described by Brown et al.10. T22 cells, which were stably transfected with a p53 responsive β-galactosidase reporter, were seeded into 96-well plate at a cell density of 8000 cells per well22. Cells were maintained in Dulbecco’s Minimal Eagle Medium (DMEM) with 10% FBS and penicillin/streptomycin and incubated for 24 hours before treatment with the stapled peptides for 18 hours. β-galactosidase activity was assayed using the FluoReporter LacZ/Galactosidase Quantitation kit (Invitrogen) as per manufacturer’s instructions. Measurements were made using an Envision plate reader (Perkin Elmer). Experiments were carried out independently thrice.
Lysate CETSA
Cells pre-treated with 10 μM MG132 for one hour were lysed in kinase buffer (25 mM Tris pH 7.4, 10 mM MgCl2, 2 mM DTT, 1x protease (Roche) and phosphatase inhibitor cocktails (Thermo Scientific)) by dounce homogenisation with a 2 ml Wheaton mortar and pestle. Cell lysates were cleared by centrifugation and aliquoted for treatment with nutlin-3 or stapled peptides. Following a 15 min incubation at room temperature with the drugs, the cell lysates were individually heated to the indicated temperatures for 2 min, cool at room temperature for 2 min and placed on ice. Insoluble proteins were separated by centrifugation and the soluble proteins were used for SDS PAGE and Western blotting for the indicated proteins. Three independent replicates were performed.
Whole cell CETSA
Cells were pre-treated with 10 μM MG132 for one hour before a four hour treatment with the indicated drugs. They were then aliquoted for heating to different temperatures for 2 min, cooled at room temperature for 2 min and placed on ice. Kinase buffer was added to the cells, followed by lysis with three freeze/thaw cycles in liquid nitrogen. Insoluble proteins were separated by centrifugation and the soluble proteins were used for SDS PAGE and Western blotting. Three independent experiments were performed.
Confocal microscopy
For confocal microscopy, HCT116 cells were grown in McCoy’s 5A media supplemented with 10% foetal bovine serum and penicillin/streptomycin in a 37 °C humidified incubator with 5% CO2 atmosphere. The cells were treated with 25 μM FAM-labelled stapled peptides for four hours before fixation by 3.7% formaldehyde. Nuclei were stained with DAPI-containing mounting media (Olink Bioscience) prior to microscopy. Micrographs were obtained with a Zeiss LSM 510 scanning laser confocal microscope with 63 × /1.4 oil objective.
Data analysis
The signal intensity of each protein band on the Western blots were normalised with the signal intensity at 37 °C (Supplementary Figure 3). The relative signal intensities were plotted against temperature using Graphpad (Prism) and a variable slope sigmoidal curve was fitted by least squares for each data set using the following equation (1):

Initial values for top and bottom plateaus were set at 1.0 and 0 respectively for curve fitting, while initial values for Tm50 and Hill Slope were automatically chosen. As the asymmetry of errors is small, errors were assumed to be symmetrical for statistical analyses. Thermal shifts were calculated using ΔTm50, which is the difference in temperatures between the control and treatment, at which 50% signal intensity (protein denaturation) was observed.
Results
Stapled peptides bind to Mdm2 and Mdm4 at high affinity in CETSA
Stapled peptides based on the N-terminal Mdm2-binding domain on p53 have been shown to successfully inhibit Mdm2 and Mdm4 interacting with p53 in in vitro thermofluor assays, as well function in cell based p53 reporter based activation assays. Moreover, similar to the small molecule Mdm2 antagonist nutlin-3, these inhibitors elicit cell cycle arrest and cell death responses in cellular assays from previous studies9,10,11. Because Mdm2/Mdm4 antagonistic stapled peptides are well-studied despite being relatively new and represent a new potential drug class, they are ideal molecules to test the CETSA method’s ability to assay for affinities, cellular uptake and target engagement in vivo. The CETSA data would be invaluable in determining the type of optimisations required to improve the various properties to the stapled peptides, allowing them to become effective therapeutic agents against cancers with wild type p53.
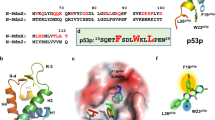
The stapled peptides used in this study have previously been reported to bind strongly to Mdm2 and Mdm4. Table 1 summarises their amino acid sequence, together with their respective IC50s reported in the literature. Also included is the IC50s for nutlin-36, which will be used as a positive control for Mdm2 thermal stabilisation. These stapled peptides have been shown to specifically inhibit the interaction between p53 and Mdm2/Mdm410,11. It is important to note that though the derivation of IC50s differ among studies, the implication remains that these compounds exhibit high affinities to Mdm2 and/or Mdm4. The physiological effects of these peptides in activating p53 were recapitulated using a p53 reporter assay (Fig. 2a in Brown et al., 2013). Stapled peptides, or nutlin-3, were added to T22 cells, which have been stably transfected with a p53-responsive β-galactosidase construct23 and p53 activation was measured by β-galactosidase activity. Cells treated with increasing concentrations of Staplin, Staplin-2 or ATSP-7041 demonstrated correspondingly increasing p53 activity and plateaued off at maxima, while p53 activation by nutlin-3 exhibited a sharp decline at higher concentrations due to the non-specific effects of high concentrations of nutlin-3. These readouts demonstrated that the stapled peptides were able to activate p53 in cellular assays.
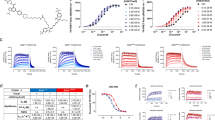
The stapled peptides (and nutlin) were analysed for the thermal stabilisation of Mdm2 and Mdm4 in AML2 cell lysates (Fig. 1a,b). Representative Western blots for the stabilisation of Mdm2 are shown in Supplementary Figure 3. Without the addition of Mdm2 specific-antagonists, Mdm2 has a Tm50 (the temperature at which 50% of proteins are precipitated by thermal denaturation) of 53.8 °C. The addition of nutlin-3 increased its Tm50 to 55.9 °C. In other words, nutlin-3 resulted in a change of 2.09 °C in Tm50 over the control, denoted by the ΔTm50. In general, the more strongly a ligand bound to a protein, the greater the thermal stabilisation observed24. Therefore, the ΔTm50 serves as an indication of how strongly the stapled peptides and nutlin-3 is bound to Mdm2, their affinities, in the presence of a multitude of other cellular proteins. In summary, the CETSA results suggest that Staplin and Staplin-2 bound Mdm2 and Mdm4 in cell extracts with high affinities, with ΔTm50 of 6.76 °C (Mdm2) and 5.48 °C for (Mdm4) for Staplin and 6.58 °C (Mdm2) and 5.44 °C (Mdm4) for Staplin 2, resulting in the highest thermal shifts in Mdm2 and Mdm4 denaturation in the panel tested, despite a ten-fold smaller IC50 reported for nutlin-3. Cellular thermal shifts therefore represent a better indication of efficacy in a cellular environment than IC50 values, which only gives an indication of binding affinities in a system containing only purified proteins.
Lysate CETSA demonstrated that stapled peptides based on Mdm2-binding site of p53 bind to both Mdm2 and Mdm4.
a,b. Thermal shift plots of lysate CETSA of nutlin-3 and stapled peptide interactions with Mdm2 (a) and Mdm4 (b) and their corresponding ΔTm50 values with 95% confidence intervals, calculated assuming symmetric errors of Tm50s. R2 values of curve fitting are also included. Nutlin-3 and peptides were added to 5 mg/ml AML2 cell lysates at 10 μM.
Assessment of Mdm2/Mdm4 binding by CETSA following modification of stapled peptides
As CETSA was developed as a quick way to assay for drugs during optimisation, we were interested in how robust the technique was in assessing target binding following modification of the stapled peptides. FAM (carboxyfluorescein) moieties were conjugated to Staplin-2 and ATSP-7041 and their contribution to Mdm2 and Mdm4 thermal shifts were analysed with CETSA on AML2 cell extracts. Figure 2a depicts the differences between unlabelled and labelled Staplin-2, with the modified Staplin-2 attributing thermal shifts 2.71 °C and 2.99 °C less for Mdm2 and Mdm4 respectively, compared to unlabelled Staplin-2 (Fig. 2c). Interestingly, FAM-labelled ATSP-7041 exhibited increased Mdm2 binding, resulting in a thermal shift of 4.21 °C compared to 1.52 °C of unlabelled ATSP-7041, though its affinity to Mdm4 was weakened, showing a 0.4 °C decrease in thermal shift (Fig. 2b,c). T22 p53 reporter assays using both labelled and unlabelled versions of Staplin-2 and ATSP-7041 showed congruent data, recapitulating the CETSA observations that FAM labelling Staplin-2 negatively affected its activity on Mdm2, while FAM labelling ATSP-7041 enhanced its activity (Fig. 2d). Taken together, these results demonstrated that CETSA can be used to quickly assay for differences in binding affinities during drug modification.
Lysate CETSA can be used to assay for the effect of stapled peptide modification on binding affinities.
a. Modification of stapling-2 with FAM reduced the binding affinity to Mdm2 and Mdm4. b. FAM labelling on ATSP-7041 increased its affinity to Mdm2, but not Mdm4. c. Differences in Tm50 between labelled and unlabelled stapled peptides. d. T22 p53 reporter assay with the labelled and unlabelled stapled peptides in the presence of 10% FCS.
Whole cell CETSA as a tool to determine cellular entry and target engagement
CETSA can also be used to assay for drug-protein interaction within cells. In this variant, cells were exposed to stapled peptides or nutlin-3 for 4 hours, when cellular uptake of stapled peptides, p53 activity and accumulation of p53 and Mdm2 was observed, but without significant apoptosis or cell death. The treated cells were collected and aliquoted to be heated to increasing temperatures. A simple propidium iodide exclusion assay was performed to determine the maximum permissible temperature of 62 °C before cell membrane failure, well exceeding the temperatures at which thermal stabilisation of Mdm2 and Mdm4 was observed (Supplementary Figure 4).
As the drugs inhibit Mdm2-p53 interaction and therefore degradation, the accumulated p53 would upregulate Mdm2 expression, raising the levels of Mdm2 in the drug-treated cells well above that of the untreated cells. In order to obtain comparable levels of Mdm2 in both the treated and control groups, cells were pre-treated with the proteasome inhibitor MG132 before introducing stapled peptides or nutlin-3 for whole cell CETSAs, or before lysis for cell lysate CETSAs. The accumulation of ubiquitinated proteins may confound the actual denaturation temperatures of Mdm2 and Mdm4, but the addition of proteasome inhibitor is not a variable in this experiment, allowing comparison between techniques and treatment groups.
The whole cell CETSA with nutlin-3 produced a thermal shift for Mdm2 that did not differ significantly to that observed in the cell lysate CETSA (Fig. 3a, ΔΔTm50 = 0.24 ± 1.42 °C). The mean thermal shift from the whole cell CETSA was 89% of the mean thermal shift for the cell lysate CETSA, indicating that nutlin-3 was able to enter cells and bind to Mdm2 efficiently. On the other hand, despite treatment with a higher peptide concentration, the thermal shifts observed for Mdm2 and Mdm4 for Staplin-2 in the whole cell CETSAs was notably less than recorded for the equivalent cell lysate CETSAs (Fig. 3b, ΔΔTm50 = 4.01 °C for Mdm2; ΔΔTm50 = 3.58 °C for Mdm4). ). The mean thermal shifts from the whole cell CETSA were 40% and 33% of the mean thermal shift for the cell lysate CETSA for Mdm2 and Mdm4 respectively. This suggests that Staplin-2 cellular entry and target engagement was impaired and therefore unable to recapitulate the thermal shifts in the cell lysate CETSAs. This is reinforced by the observation of high concentrations of FAM-labelled peptides within vesicles in cells, showing that the availability of stapled peptides for Mdm2/Mdm4 binding was impaired (Fig. 3c).
Comparisons between CETSA on whole cells and cell lysates can be used to determine drug cellular entry and target engagement.
a. The thermal stabilisation of Mdm2 by 10 μM nutlin-3 was assayed by CETSA with cell lysates and whole cells. In both cases, nutlin-3 resulted in very similar thermal shifts, as determined by the ΔΔTm50. b. The thermal shift of CETSA with whole cells with 25 μM Staplin-2 showed differences in thermal shifts. c. Confocal microscopy of HCT116 cells treated with Staplin-2FAM prior to fixation showed vesicular accumulation of the FAM-labelled peptides.
Comparing the differences in thermal shifts between the cell lysate and the whole cell CETSAs generates an index that correlates with the cellular availability of a drug. The combination of the two assays therefore serve as a powerful tool to enable the optimisation of stapled peptide-based drugs not only in terms of target affinity, but also its ability to enter cells to reach their intended targets.
Discussion
The efficacy of a drug for therapeutics was commonly assessed by observing for phenotypic changes, such as the shrinking of a tumour mass or lowering of blood pressure. Such an approach to drug discovery led to issues with the activation or inhibition of other cellular pathways, typically resulting in deleterious side effects. This was especially the case for the activation of p53—a multitude of stimuli can drive the activation of p53 and induce downstream phenotypic changes. As our knowledge of biology increases, we came to recognise the importance of designing drugs that target a particular pathway specifically and drugs designed or discovered to have such properties are, unsurprisingly, better tolerated and generally have better efficacy.
While the concept of stapling peptides is not new, the technology has yet to reach maturity and disparate views have started to appear in the past few years. For instance, the efficacy of Bax-activating stapled peptides has been found to be highly context-dependent, showing enhanced apoptotic activity in some systems but not others25,26,27. Clearly, stapling alone does not enhance a peptide’s efficacy and until we fully understand the rules for peptide therapeutics, copious optimisation is required to select the ones with the best activities. The process presented in this study aims to provide an effective method to aid in the optimisation of this new class of drugs.
Possibly the most critical way to determine if a drug is specific for a particular pathway is to measure its direct binding to its intended molecular target. Some drugs may be modified to carry affinity tags, which allows direct observations of target engagement though pulldown experiments. However, for many small molecules, the inclusion of an affinity tag adversely affects their binding to their targets, thus indirect measurements of target binding is required.
Several techniques have been developed to measure protein-ligand interactions using spectroscopic and calorimetric analysis13,28. As described earlier, these methods generally require proteins of very high purity and in many instances the major bottleneck in drug design and development is the inability to produce purified protein. The CETSA method described by Molina et al. was developed to overcome this major obstacle, allow the analysis of protein-ligand interaction in multicomponent solutions—whole cells and tissues and cell extracts. This method has since been used to validate the binding of several drugs to their protein targets, such as MTH129, Bcl-2/Bcl-xL30 and PARP131. Here, we validated the ability of stapled peptide to bind Mdm2 and Mdm4, both in cell extracts, as well as in intact cells.
Besides the main advantage of circumventing the need for purified proteins, CETSA allows the monitoring of drug transport, availability and target engagement within cells. The stapled peptides described in this study have been reported to bind Mdm2 and Mdm4 with very high, low-nanomolar affinities and this was recapitulated in the cell lysate CETSAs, where massive thermal stabilisation was observed with the addition of these peptides. However, the main disadvantage of these peptides is their relative difficulty in entering cells and reaching their intended targets, as shown by the relatively poor thermal stabilisation in the whole cell CETSA experiments. This may be overcome by altering the staple used to enforce peptide helicity, either by the use of other linkers such as “double-click“ staples32, disulphides33, thioethers34, lactams35 or oximes36, or by adding functional groups to the alkene hydrocarbon staple to aid in cellular availability. Nonetheless, this study provides strong evidence that CETSA can be employed to investigate drug cellular entry and availability.
The relative ease of the CETSA technique would be invaluable for rapid optimisation of drug design, not just in terms of absolute binding affinities, but also in terms of cellular uptake and target accessibility. The technique is also applicable on samples from animal models and patients18. Most importantly, these measurements would be performed on endogenous proteins in their native cellular environments, greatly reducing multiple confounding factors that are associated with similar assays.
Additional Information
How to cite this article: Tan, B. X. et al. Assessing the Efficacy of Mdm2/Mdm4-Inhibiting Stapled Peptides Using Cellular Thermal Shift Assays. Sci. Rep. 5, 12116; doi: 10.1038/srep12116 (2015).
References
Hollstein, M., Sidransky, D., Vogelstein, B. & Harris, C. C. p53 mutations in human cancers. Science 253, 49–53 (1991).
Vogelstein, B., Lane, D. & Levine, A. J. Surfing the p53 network. Nature 408, 307–310 (2000).
Momand, J., Jung, D., Wilczynski, S. & Niland, J. The MDM2 gene amplification database. Nucleic Acids Res. 26, 3453–3459 (1998).
Huang, L. et al. The p53 inhibitors MDM2/MDMX complex is required for control of p53 activity in vivo. Proc. Natl. Acad. Sci. 108, 12001–12006 (2011).
Michael, D. & Oren, M. The p53–Mdm2 module and the ubiquitin system. Semin. Cancer Biol. 13, 49–58 (2003).
Vassilev, L. T. et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303, 844–848 (2004).
Shangary, S. et al. Temporal activation of p53 by a specific MDM2 inhibitor is selectively toxic to tumors and leads to complete tumor growth inhibition. Proc. Natl. Acad. Sci. 105, 3933–3938 (2008).
Brown, C. J., Lain, S., Verma, C. S., Fersht, A. R. & Lane, D. P. Awakening guardian angels: drugging the p53 pathway. Nat. Rev. Cancer 9, 862–873 (2009).
Bernal, F. et al. A Stapled p53 Helix Overcomes HDMX-Mediated Suppression of p53. Cancer Cell 18, 411–422 (2010).
Brown, C. J. et al. Stapled Peptides with Improved Potency and Specificity That Activate p53. ACS Chem. Biol. 8, 506–512 (2013).
Chang, Y. S. et al. Stapled α−helical peptide drug development: A potent dual inhibitor of MDM2 and MDMX for p53-dependent cancer therapy. Proc. Natl. Acad. Sci. 201303002 (2013). doi: 10.1073/pnas.1303002110.
Kim, Y.-W., Grossmann, T. N. & Verdine, G. L. Synthesis of all-hydrocarbon stapled α-helical peptides by ring-closing olefin metathesis. Nat. Protoc. 6, 761–771 (2011).
Pantoliano, M. W. et al. High-density miniaturized thermal shift assays as a general strategy for drug discovery. J. Biomol. Screen. 6, 429–440 (2001).
Pierce, M. M., Raman, C. S. & Nall, B. T. Isothermal Titration Calorimetry of Protein–Protein Interactions. Methods 19, 213–221 (1999).
Vedadi, M. et al. Chemical screening methods to identify ligands that promote protein stability, protein crystallization and structure determination. Proc. Natl. Acad. Sci. 103, 15835–15840 (2006).
Ericsson, U. B., Hallberg, B. M., DeTitta, G. T., Dekker, N. & Nordlund, P. Thermofluor-based high-throughput stability optimization of proteins for structural studies. Anal. Biochem. 357, 289–298 (2006).
Fedorov, O. et al. A systematic interaction map of validated kinase inhibitors with Ser/Thr kinases. Proc. Natl. Acad. Sci. 104, 20523–20528 (2007).
Molina, D. M. et al. Monitoring Drug Target Engagement in Cells and Tissues Using the Cellular Thermal Shift Assay. Science 341, 84–87 (2013).
Khoo, K. H., Verma, C. S. & Lane, D. P. Drugging the p53 pathway: understanding the route to clinical efficacy. Nat. Rev. Drug Discov. 13, 217–236 (2014).
Gembarska, A. et al. MDM4 is a key therapeutic target in cutaneous melanoma. Nat. Med. 18, 1239–1247 (2012).
Tan, B. X., Khoo, K. H., Lim, T. M. & Lane, D. P. High Mdm4 levels suppress p53 activity and enhance its half-life in acute myeloid leukaemia. Oncotarget 5, 933–943 (2013).
Lu, X., Burbidge, S. A., Griffin, S. & Smith, H. M. Discordance between accumulated p53 protein level and its transcriptional activity in response to u.v. radiation. Oncogene 13, 413–418 (1996).
Hupp, T. R., Sparks, A. & Lane, D. P. Small peptides activate the latent sequence-specific DNA binding function of p53. Cell 83, 237–245 (1995).
Savitski, M. M. et al. Tracking cancer drugs in living cells by thermal profiling of the proteome. Science 346, 1255784 (2014).
Gavathiotis, E. et al. BAX activation is initiated at a novel interaction site. Nature 455, 1076–1081 (2008).
Okamoto, T. et al. Stabilizing the Pro-Apoptotic BimBH3 Helix (BimSAHB) Does Not Necessarily Enhance Affinity or Biological Activity. ACS Chem. Biol. 8, 297–302 (2013).
LaBelle, J. L. et al. A stapled BIM peptide overcomes apoptotic resistance in hematologic cancers. J. Clin. Invest. 122, 2018–2031 (2012).
Sigurskjold, B. W. Exact Analysis of Competition Ligand Binding by Displacement Isothermal Titration Calorimetry. Anal. Biochem. 277, 260–266 (2000).
Huber, K. V. M. et al. Stereospecific targeting of MTH1 by (S)-crizotinib as an anticancer strategy. Nature 508, 222–227 (2014).
Bai, L. et al. BM-1197: A Novel and Specific Bcl-2/Bcl-xL Inhibitor Inducing Complete and Long-Lasting Tumor Regression In Vivo. PLoS ONE 9, e99404 (2014).
Na, Z. et al. A Small-Molecule Protein–Protein Interaction Inhibitor of PARP1 That Targets Its BRCT Domain. Angew. Chem. 127, 2545–2549 (2015).
Lau, Y. H. et al. Investigating peptide sequence variations for ‘double-click’ stapled p53 peptides. Org. Biomol. Chem. 12, 4074–4077 (2014).
Jackson, D. Y., King, D. S., Chmielewski, J., Singh, S. & Schultz, P. G. General approach to the synthesis of short alpha.-helical peptides. J. Am. Chem. Soc. 113, 9391–9392 (1991).
Brunel, F. M. & Dawson, P. E. Synthesis of constrained helical peptides by thioether ligation: application to analogs of gp41. Chem. Commun. Camb. Engl. 2552–2554 (2005). doi: 10.1039/b419015g.
Taylor, J. W. The synthesis and study of side-chain lactam-bridged peptides. Biopolymers 66, 49–75 (2002).
Pallin, T. D. & Tam, J. P. Cyclisation of totally unprotected peptides in aqueous solution by oxime formation. J. Chem. Soc. Chem. Commun. 2021–2022 (1995). doi: 10.1039/C39950002021.
Author information
Authors and Affiliations
Contributions
B.X.T., C.J.B. and D.P.L. designed the experiments, F.J.F., T.Y.Y., C.J.B. and Q.S.T. synthesised the peptides, B.X.T. and B.H.C. performed the experiments, A.E.J. and H.L.T. assisted with troubleshooting and B.X.T., D.P.L. and P.N. analysed the data and wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Tan, B., Brown, C., Ferrer, F. et al. Assessing the Efficacy of Mdm2/Mdm4-Inhibiting Stapled Peptides Using Cellular Thermal Shift Assays. Sci Rep 5, 12116 (2015). https://doi.org/10.1038/srep12116
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep12116
This article is cited by
-
Experimental study of proteome halophilicity using nanoDSF: a proof of concept
Extremophiles (2022)
-
Therapeutic targeting of p53: all mutants are equal, but some mutants are more equal than others
Nature Reviews Clinical Oncology (2018)
-
Icaritin enhances mESC self-renewal through upregulating core pluripotency transcription factors mediated by ERα
Scientific Reports (2017)
-
Peptidomimetic inhibitors of APC–Asef interaction block colorectal cancer migration
Nature Chemical Biology (2017)
-
Mechanistic validation of a clinical lead stapled peptide that reactivates p53 by dual HDM2 and HDMX targeting
Oncogene (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.