Abstract
Hybrid seeds are used for stimulated crop production, as they harness heterosis. The achievement of complete male-sterility in the female-parent and the restored-fertility in F1-hybrids are the major bottlenecks in the commercial hybrid seed production. Here, we report a male sterility–fertility restoration system by engineering the inmost nutritive anther wall layer tapetum of female and male parents. In the female parent, high–level and stringent expression of Arabidopsis autophagy–related gene BECLIN1 was achieved in the tapetum, which altered the tapetal degeneration program, leading to male sterility. This works on our previously demonstrated expression cassette based on functional complementation of TATA-box mutant (TGTA) promoter and TATA-binding protein mutant3 (TBPm3), with modification by conjugating Long Hypocotyle in Far-Red1 fragment (HFR1NT131) with TBPm3 (HFR1NT131-TBPm3) to exercise regulatory control over it. In the male parent, tapetum–specific Constitutive photo-morphogenesis1 (COP1) was expressed. The F1 obtained by crossing these engineered parents showed decreased BECLIN1 expression, which was further completely abolished when COP1-mutant (COP1L105A) was used as a male parent, leading to normal tapetal development and restored fertility. The system works on COP1-HFR1 interaction and COP1–mediated degradation of TBPm3 pool (HFR1NT131-TBPm3). The system can be deployed for hybrid seed production in agricultural crops.
Similar content being viewed by others
Introduction
Hybrid crops have been contributing to the substantial global rise in agricultural output over the past few decades as they harness heterosis (hybrid vigor), a phenomenon of outperformance of F1 hybrid progeny compared with their parents in terms of yield, biotic and abiotic resistance1. Their utilization offers a 20% to more than 50% yield increase2 and contributes to more than half of the production of the major crops3. Precise control over pollen fertility in the female parent and the fertility restoration in F1 hybrids are the prerequisites in the commercial production of the F1 hybrid in self-pollinating crops4. Restoration of male fertility in hybrids is especially important in crops where the desired agricultural products are seeds, such as cereals, pulses and so on.
Several approaches have been explored to limit self-fertilization in the female parent line for the development of an effective hybrid seed production system, such as emasculation (manual removal of male reproductive organs), chemical-induced male sterility, cytoplasmic and nuclear male sterility and biotechnological approaches for pollen abortion. Emasculation involves labor-intensive and time-consuming processes for large-scale hybrid seed production. The use of chemicals is limited by the issues related to bio-safety, variable effects, optimum dose and cost effectiveness5. The cytoplasmic and nuclear male sterility is limited by maintenance of multiple lines, partial and unstable male sterility and limited fertility restoring gene sources, all of which restrict the economic benefits of the hybrid6. A number of biotechnological strategies have been deployed to restrict self–fertilization in plants7. Several of the transgenic systems possess the male fertility–restoring constituent8,9,10,11,12,13,14,15,16,17. However, the only commercialized transgenic male sterility method is SeedLinkTM, which relies on the expression of bacterial cytotoxic ribonuclease (Barnase) in the male reproductive organ of the female parent line and fertility restoration by ribonuclease inhibitor (Barstar) delivered by the male parent18,19. Barnase-barstar system is tested in many crops20; however, issues such as leaky expression of the barnase gene and difficulty in obtaining restoration lines of the barstar gene9,20,21,22,23 and biosafety concerns associated with the use of the bacterial cytotoxic gene in food crops are the key challenges associated with its applicability. Hence, it is desirable to develop a hybrid seed system that is equipped with capabilities of complete pollen abortion in a biologically safe and tightly controlled manner, as well as efficient male fertility restoration in the F1 hybrid.
Tapetum is the sporophytic tissue and innermost wall layer of the micro-sporangia in the angiosperm plants24. It plays an important role in the development of male gametophyte (microspore) by providing enzymes, nutrients and wall material, first by secretion and eventually by degeneration25. Tapetal degeneration is a programmed cell death (PCD) event26,27 with typical cytological features of cell shrinkage, mitochondria and cytoskeleton degeneration, nuclear condensation, oligonucleosomal cleavage of DNA, vacuole rupture and endoplasmic reticular swelling26,28,29,30,31. Tapetal PCD at a specific developmental stage is crucial for pollen fertility and disruption of the timing of PCD, either early or delayed, results in pollen abortion or male sterility. Several transgenic approaches have been developed to generate male sterile plants either through early tapetum degeneration by expressing BARNASE18, RNase -119, DIPTHERIA TOXIN-A32, RIBOSOME INACTIVATING PROTEIN33 and BAX28, or by delaying tapetal PCD through BAX INHIBITOR28, ethylene receptor gene Cm-ETR1/H69A34, cystein protease BoCysP1 and BoCP335.
Here, we established a transcription regulation system for male sterility–fertility restoration in plants. It includes two tapetum-specific expression cassettes; one expressed in the female parent and the other expressed in the male parent of the desired F1 progeny. The female expression cassette has the potential to attain high-level, stringent expression of a desired gene limited to tapetal cells, but the expression switches to abolition in F1 when regulated by the male component. We successfully deployed this system to express the desired gene (we earlier demonstrated for male sterility) Arabidopsis BECLIN115, which generates the complete male sterile parent by altering the tapetal degeneration program. The male fertility of F1 progeny was completely restored due to the abolished expression of BECLIN1,> as the tapetal degeneration program was sustained to be normal. The female component works on the functional complementation of mutated TATA-box (TGTA) and TBPm336 with aided HFR1NT131 fused to TBPm3 (HFR1NT131-TBPm3). The abolition of BECLIN1 expression in F1 was achieved by limiting TBPm3 through COP1 (male component) –mediated degradation37,38,39 of the fusion protein HFR1NT131-TBPm3. Nicotiana tabacum has been used for the proof of principle presented here, but the essential elements of the technology are generic and possibly will work in other crops.
Results
High-level, stringent and spatio-temporal expression of the desired gene in postmeiotic tapetum
We have previously utilized TATA box binding protein mutant3 (TBPm3) and TATA-box mutation TGTA complementation system36 for high-level and stringently regulated expression in anther tapetum15. The expression through promoters with TATA-box mutated to TGTA was not driven by a native TBP protein because of its altered affinity40,41,42, however, three amino-acid substitution mutant (Ile152 to Phe152, Val161 to Thr161, and Leu163 to Val163) of TBP (TBPm3) complement the TGTA-mutation and drive the expression from such promoters43,44. The high-level expression of the TGTA-TBPm3 complementation system was achieved due to a dedicated pool of TBPm3 protein to TGTA-containing promoters15,36,45. In this work, we seek transcriptional control over the TGTA-TBPm3 complemented system by fusing TBPm3 with the 131 amino-acid N-terminal domain of HFR1 (HFR1NT131) and specifically targeting it for degradation by COP1, thus limiting TBPm3 protein in the cell. The limited cellular availability of TBPm3 protein leads to the abolition of expression of the transgene in tapetum cells (Fig. 1a–c). Tapetum-specific promoter TA29 (PTA29)46 was utilized to construct a TGTA-TBPm3 complemented system to attain high-level and stringent expression, specific to the postmeiotic tapetum as previously described15, with a modification of fusing HFR1NT131 at the N-terminus of TBPm3 to excercise further control over it (Supplementary Fig. S1-a-I). The transgenic lines of Nicotiana tabacum cv. Petit Havana (NTPH) harboring this expression cassette 1370 (carrying TGTA-driven gusA gene and HFR1NT131:TBPm3 fusion in a single T-DNA cassette; Supplementary Fig. S1-a-I) were normal during growth and development with a similar range of pollen viability, germination and seed setting when compared with NTPH plants (Supplementary Fig. S1-b and c). The anther development has been divided into seven premeiotic (−7 to −1; negative sign represents stages before the meiosis) and twelve postmeiotic stages (1 to 12) in tobacco46. The expression of gusA protein was examined in the anthers of stages 2 and 3 in 1370 transgenic lines. The gusA expression in the stage 2 anther ranges from 11.45 nmoles mg−1min−1 to 47.57 nmoles m−1min−1 (Supplementary Fig. S2-a), with an average expression of 22.00 ± 5.15 nmoles mg−1min−1 (Fig. 2b); at stage 3, the expression ranges from 10.45 nmoles mg−1min−1 to 45.65 nmoles mg−1min−1(Supplementary Fig. S2-b), with average expression being 20.43 ± 5.74 nmoles mg−1min−1(Fig. 2b) in the anthers of transgenic lines evaluated. Histochemical GUS staining confirmed that the expression of gusA protein was limited to tapetal cells in anthers (Fig. 2d). Expression was not detected in other plant organs such as leaves, emasculated flower buds, stems and roots (data not shown). Taken together, our results showed that high-level and stringent expression was achieved in the postmeiotic (stages 2 and 3) tapetum, using the expression vector 1370.
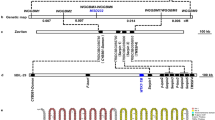
Experimental design and underlying mechanism of the male sterility–fertility restoration system in plants for hybrid seed production.
(a) Generation of male-sterile female parent for hybrid breeding. The female expression cassette (1370/1371) contains two transcriptional units: expression component (PTA29(TGTA)-gusA/BECLIN1-Tnos) and the regulatory component (PPcec-HFR1NT131-TBPm3-Tnos). The regulatory component express conjugated protein HFR1NT131-TBPm3, which binds to TATA-box mutated promoter (PTA29(TGTA)) of expression component and resulted transcriptional pre-initiation complex (PIC) formation. The high-level and tapetum-specific expression of gusA or BECLIN1 was achieved, as TGTA-mutated promoter was functionally complemented by TBPm3 (or HFR1NT131-TBPm3) pool. This expression cassette is useful in achieving tapetum-specific, high-level expression of the male-sterility gene (we expressed BECLIN1). (b) The male expression cassette (1372/1373) expresses COP1 (or COP1L105A) by using the tapetum-specific promoter A9, giving normal male-fertile plants. (c) Fertility restoration mechanism of F1. F1 is obtained by crossing with plants expressing female (male-sterile) and male (male-fertile) expression cassettes. Regulatory component express HFR1NT131-TBPm3 and male-component express COP1 (or COP1L105A) proteins in the F1-tapetal cell. The COP1 physically interacts with the HFR1NT131 fragment of the conjugated protein (HFR1NT131-TBPm3) and sequentially degrades it, resulting in the unavailability of TBPm3, hence no PIC formation on the TGTA-mutated TA29 promoter, leading to expression abolition of gusA or BECLIN1. BECLIN1 abolition resulted normal tapetal degeneration program and restored fertility of F1-progeny.[TA29(TGTA) = Tapetum-specific promoter from tobacco with mutated TATA box to TGTA, gusA= β-glucuronidase, Pcec= artificial promoter48, HFRINT131 = N-terminus 131 amino acid fragment of Long hypocotyle in far red 1 (HFR1), TBPm3 = TATA-binding protein with three amino acid substitution (Ile152 to Phe152, Val161 to Thr161, and Leu163 to Val163), Tnos= transcriptional terminator, A9= tapetum-specific promoter from Arabidopsis, COP1= Constitutive photomorphogenic1, COP1L105A= COP1-mutant with increased nuclear localization38]
Analysis of reversible expression system.
(a) The strategy of crossing; the development of F1s, transgenic plants of 1370 was taken as ♀-parent and was crossed with 1372 and 1373 as ♂-parent, resulting in F1 (1370(♀) × 1372(♂)) and F’1 (1370(♀) × 1373(♂)) progeny, respectively. (b) GUS expression analysis in ♂-parent (1370), F1 and F’1 hybrids at anther developmental stages 2 and 3; the expression values are normalized with the control (NTPH). The error bar represents SD of n = 10 independent line. (c–f) Gus staining in the stage 2 anther of the control (c), 1370 transgenic line (d) and F1 (e) and F’1 (f) lines. Bar = 100 μm. (g) Transgenic plant of 1373 (PA9:COP1L105A) showing normal growth and development. (h) Inflorescence of 1373 transgenic lines showed normal flowering and flower development.(i) Pollen viability using FDA-PI method; green fluorescence of FDA shows viable pollens whereas red fluorescence of PI indicates aborted pollens. Bar = 100 μm. (FDA= fluorescein diacetate, PI= propedium iodide). (j) In-vitro pollen germination and staining with FDA and PI, germinated pollens with pollen tube whereas abortive pollen with PI staining and no germination. Bar = 20 μm. (k) Bar graph showing the relative expression of COP1L105A in developing anthers of stages 1–6. Total RNA was isolated from anthers of the transgenic lines (1373) for qRT-PCR assay (n = 3 independent biological repeats). UBIQ10 was taken an internal control. Error bars indicate standard deviation (SD).
Transcriptional abolition of TGTA-TBPm3 system based on HFR1NT131-COP1L105A–mediated light signalling
To evaluate whether limiting cellular protein levels of TBPm3 can abolish gusA expression, we developed transgenic tobacco lines expressing COP1 under the control of Arabidopsis tapetum-specific promoter A9 (PA9)47. In construct 1370, the desired gene (gusA in the case of 1370 cassette) was expressed under the control of the TGTA-mutated tapetum-specific promoter and the translationally fused HFR1NT131-TBPm3 complex was under the regulation of the Pcec promoter48. HFR1NT131 is a truncated form (NT131: N-terminus 131-amino-acid fragment) of Long Hypocotyle in Far Red1 (HFR1), a basic helix-loop-helix (bHLH) involved in the seedling response to far-red light. The NT131 amino-acid fragment loses its bioactivity but retains its interaction ability with another protein COP1 (Fig.1a–c). COP1 is a repressor of light-mediated photo-morphogenesis that physically interacts with HFR1 (and HFR1NT131) and mediates its targeted degradation through the proteosome37,39,49,50. We, therefore, postulated that the fusion protein HFR1NT131:TBPm3 will also be degraded by COP1 and this was exploited to exercise control over the TGTA-TBPm3 complementation system (construct 1370). To achieve this, a regulatory expression cassette (1372) was designed, in which the Arabidopsis COP1 gene was placed under the control of the Arabidopsis tapetum-specific promoter A9 (PA9)47 (Supplementary Fig. S1-a-III). The tobacco transgenic lines of 1372 were normal during growth and development with normal flowering, pollen viability, in-vitro pollen germination and seed setting similar to those of the control (NTPH) plants (Supplementary Fig. S1b-c and S3a-d). The relative expression of COP1 was evaluated using qRT-PCR in anther developmental stages 1 to 6. The highest expression was found in the stage 1 anthers (Supplementary Fig. S3-e). To screen the best expressing transgenic lines, relative expression of COP1 was compared using qRT-PCR in the stage 1 anthers in 19 transgenic lines. The transgenic lines 13 and 33 were selected for further studies based on their higher expression (Supplementary Fig. S4-a). To validate our hypothesis of achieving transcriptional control over the TGTA-TBPm3 system by COP1-mediated degradation of HFR1NT131-TBPm3, crossing was performed between female 1370 lines (L3 and L5; Supplementary Fig. S2-a) and male 1372 plants (T13 and T33; Supplementary Fig. S4-a), raising F1 progeny. The F1 seeds were harvested, subjected to double antibiotic selection of kanamycine and hygromycine, considering the selectable marker genes of the constructs 1370 (NPTII) and 1372 (HPTII) and further confirmed by PCR (Supplementary Fig. S5a–c). The fluorimetric GUS analysis revealed a 4–6–fold reduced GUS activity in the anthers of F1 plants during stages s2 and s3 as compared to 1370 plants (Fig. 2b, Supplementary Fig. S2-c and d). Histochemical GUS staining of F1 plants revealed that the GUS expression was stringent to tapetal cells but diminished when compared with 1370 (Fig. 2e) Thus, COP1 imparts its control over the TGTA-TBPm3 system that is mediated by HFR1NT131- linked TBPm3 protein (HFR1NT131-TBPm3) degradation in F1 plants, albeit the expression was not completely abolished. The one reason for incomplete control of COP1 could be its light-responsive nuclear and cytoplasmic localiszation.
To develop a fertility restoration system, it is necessary to completely abolish the expression of the male sterility gene (here, BECLIN1) in the F1 hybrid. We postulate that increased nuclear localization of COP1 should cause complete degradation of HFR1NT131-TBPm3, resulting in its unavailability to form the transcriptional pre-initiation complex (PIC) on the TGTA-mutated TATA-box of the PTA29 promoter, leading to expression abolition in F1 progeny. To achieve it, COP1 was mutated to COP1L105A. The mutation in COP1 (COP1L105A) was reported to increase nuclear localization but it retained its normal functioning38. The expression cassette 1373 (Supplementary Fig. S1-a-iv) was designed to express mutated COP1 (COP1L105A) using PA9. The transgenic plants expressing COP1L105Awere found to be normal during growth, development and fertility (Fig. 2g–j, Supplementary Fig. S1a-b). The highest COP1L105A expression was found at the s1 anther development stage (Fig. 2k). Fifteen transgenic lines were compared for relative expression; lines 3 and 5 were selected for the crossing experiment based on their higher expression (Supplementary Fig. S4-b). The cross was made between female 1370 (emasculated) and male 1373 to raise F’1 seeds. The F’1 seeds were screened on double antibiotic selection medium (Kanamycin and Hygromycin) and standard PCR. The fluorimetric and histochemical GUS analysis of anthers of F’1 plants at developmental stages s2 and s3 showed completely abolished expression of gusA protein (Fig. 2b,f and Supplementary Fig. S2e,f). The results indicated that the nuclear localization of COP1L105A imparts its complete control over the TGTA-TBPm3 system through HFR1NT131-mediated degradation of TBPm3.
Complete male sterility by high-level expression of BECLIN1 in tapetum cells
We previously demonstrated that high-level expression of Arabidopsis ATG6/BECLIN1 in the tapetum under control of the TGTA-TBPm3 complementation system leads to complete male sterility in tobacco15. To develop a male sterility–fertility restoration system, we cloned Arabidopsis ATG6/BECLIN1 into the 1370 vector replacing gusA, thus developing the construct 1371 (Supplementary Fig. S1-a-ii). The tobacco transgenic lines expressing 1371 showed no morphological anomaly, with normal growth and development comparable to wild plants except male sterility (Fig. 3a,b). The flowers of 1371 transgenics showed healthy stigma and other accessory whorls, whereas pollen grains laden to the anther were very low (Fig. 3c,d). The flowers failed to self-fertilize because of the formation of non-viable pollen, which eventually dries out and drops from the floral axis (Fig. 3d arrow shows point of detachment and drying); whereas when fertilized with the viable pollen of wild plants, a normal seed setting was observed (data not shown). The results suggest that stigma receptivity and female reproductive organs of 1371 transgenics were normal and the drop of flowers was due to the development of aborted pollen and failure in fertilization.
Analysis of BECLIN1 expressing transgenic lines.
(a) Control (wild type) and (b) 1371 transgenic plants. Inflorescence, flower and seed setting in control (c) and 1371 transgenic plants (d). The control and 1371 plants showed normal growth and development except male sterility in 1371 transgenics (arrows indicate the dying and detachment of flower that are unsuccessful in fertilization due to male sterility). (e) qRT-PCR for BECLIN1 expression in 1371 transgenic lines of anther at stages 2 to 6, highest expression was found in stage 2 and 3 of anther development. Error bars represent SD of three (n = 3) biological replicates. Transmission electron micrograph (TEM) of pollen of anther stage 1 to 6 in control (f–k) and in 1371 transgenic plants (l–q). Bar = 5 μm.
The relative expression of Arabidopsis BECLIN1 was compared using qRT-PCR at a different anther developmental stage of the 1371 transgenic lines; the highest expression was found in stages 2 and 3 (Fig. 3e). To understand the effect of postmeiotic, tapetum-specific and high-level expression of BECLIN1 on the pollen development, transmission electron microscopy (TEM) of developing pollens was performed. No obvious differences were found till stage 1, between 1371 transgenics and control (NTPH) as meiosis was normal and microspore tetrad was surrounded by the callose in both control (Fig. 3f, Supplementary Fig. S6-b) and in transgenic (Fig. 3l, Supplementary Fig. S6-i) plants. At stage 2, microspores became free due to callose decomposition, the nucleus was displaced aside due to a single large vacuole and development of the orderly microspore exine structure was seen in the control (Fig. 3g, Supplementary Fig. S6-c); whereas in 1371 transgenics the tetrad was not fully separated due to remnants of the callose deposition and deformation had started in microspores (Fig. 3m, Supplementary Fig. S6-j). The vacuole gets enlarged with demarked wall formation in stage 3 microspores in case of control (Fig. 3h, Supplementary Fig. S6-d), whereas deformation increases in 1371 transgenics (Fig. 3n, Supplementary Fig. S6-k). At stage 4, the generative cell was attached to the wall with a prominent vegetative nucleus, cytoplasm and normal exine and intine development in the control (Fig. 3i, Supplementary Fig.S6-e); whereas in 1371 transgenics, the cytoplasm was collapsed and pollen wall deformations were visible (Fig. 3o, Supplementary Fig. S6-e). The microspore shape became nearly circular and was filled with dense cytoplasm and small vesicles; the pollen wall also differentiated into well-developed exine and intine at stages 5,6 in control pollens (Fig. 3j,k, Supplementary Fig. S6f-g); whereas in case of pollens of 1371 transgenics, the cytoplasm collapsed and severe deformity was observed in the pollen wall (Fig. 3p,q, Supplementary Fig. S6m-n).
We next examined the pollen viability, in-vitro pollen germination and seed setting in wild and 1371 transgenics. The 1371 transgenic lines showed considerably low pollen viability (Fig. 4a,b,u) with nil germination (Fig. 4m,n,u) as compared to the control (wild type). In control plants, bagging of a young inflorescence before anthesis results in normal seed setting (Fig. 4q,v), unlike in male sterile plants in which all the bulbs were seedless (Fig. 4r,v). Further, the scanning electron micrograph (SEM) of stage 7 pollen of control showed normal pollen morphology (Fig. 4e,f and Supplementary Fig. S7a-b); whereas deformed pollens were seen in the 1371 transgenic lines (Fig. 4g,h and Supplementary Fig. S7c-d). The results obtained by the new system are in agreement with our previous finding15 that overexpression of BECLIN1 in the anther tapetum resulted in complete male sterility in the tobacco transgenic plants. In addition, the male sterile female lines are equipped with additional regulatory control over the TGTA-TBPm3 system operated through the male component.
Transgenic male sterility and fertility restoration in tobacco.
[a–d] Pollen viability assay using FDA-PI method in control (a) 1371 transgenic (b) F1 obtained by crossing (1371(♀) × 1372(♂)) (c) and F’1 (1371(♀) × 1373(♂)) (d) lines. Bar = 100 μm, inset bar = 20 μm. [e–l] Scanning electron microscopic (SEM) analysis in control (e and f), 1371 transgenic (g and h), F1 (i and j) and F’1 lines (k and l). Bar = 50 μm in e, g, i and k and 10 μm in f,h,j and l. [m–p] In-vitro pollen germination assay in control (m), 1371 transgenic (n), F1 (o) and F’1 (p) lines pollens. Bar = 100 μm. [q–t] Seed setting in control (q) 1371 transgenic (r) F1 hybrid (s) and F’1 hybrid (t) lines. [u] Quantitative analysis (in %) of pollen viability and in-vitro pollen germination in control, 1371 transgenic, F1 hybrid and F’1 hybrid lines. Error bar represents SD of n = 10 lines. [v] Quantitative analysis (in mg/pod) of seed setting in control, 1371 transgenic, F1 hybrid and F’1 hybrid lines. Error bar represents SD of n = 10 lines.
Complete restoration of male fertility in F1 hybrid
Restoration of pollen fertility in F1 progeny is a prerequisite for hybrid seed production. The male sterile 1371 transgenic female lines were crossed with the selected male lines expressing either COP1 (1372) or COP1L105A (1373) to raise F1 and F’1 progeny, respectively (Supplementary Fig. S5). In F1 [1371(♀) × 1372(♂)] plants, pollen viability was recorded ~15.14% (Fig. 4c,u) as compared with ~77.71% of control; whereas in-vitro pollen germination was ~18.5% (Fig. 4o,u) in F1 as compared with ~70% in case of control plants. The pollen viability of F1 was low, but the seed setting was found to be comparable to the control plants (Fig. 4s,v). The SEM of stage 7 pollens showed a high population of aborted and deformed pollens with a few normal architecture pollens in F1 plants (Fig. 4i,j, Supplementary Fig. S7 e-f).The relative expression of BECLIN1 and COP1 was quantified in s1-s6 anther developmental stages of F1 plants. We found higher accumulation of AtBECLIN1 transcript in s1, s2 and s3 stages. The COP1 expressed at considerably high level in the s1 stage. We next compared the expression of BECLIN1 in s2 stage in F1 with that of 1371 transgenic male sterile lines and found about ~21-fold decrease in the expression (Fig. 5a,c,d). The results, thus, indicate that COP1 (1372) mediates partial restoration of fertility in F1 plants by reducing the expression of BECLIN1. In F’1 [1371(♀) × 1373(♂)] progeny, the pollen viability and pollen germination was found to be comparable to that of control plants (Fig. 4d,p,u). The SEM of stage 7 pollen showed pollen architecture similar to control pollen (Fig. 4k,l and Supplementary Fig. S7g-h). The seed setting was normal with seed weight ~78.89 mg/pod, comparable to control ~74.23 mg/pod (Fig. 4t,v). The expression of BECLIN1 was completely abolished, while the expression of COP1 was found to be the highest in the s1 anther stage in F’1 (Fig. 5b–d). The results established complete restoration of male fertility in F’1 plants by utilizing the principle of COP1-mediated degradation of HFR1NT131-TBPm3 and by abolishing the expression of BECLIN1.
Expression analysis.
[a] Bar diagram showing the expression of BECLIN1 and COP1 in the anther stages 1 to 6 in F1 lines (1371(♀) × 1372(♂)). Total RNA was isolated from the anther of stages 1 to 6 for qRT-PCR assay (n = 3 independent biological repeats). UBIQ10 was used as an internal control. BECLIN1 expression value at stage 1 was set as 1 and the relative gene expression levels were calculated. Error bars indicate SD. [b] Bar diagram showing the expression of BECLIN1 and COP1L105A in the anther stages 1–6 in the F’1 lines (1371(♀) × 1373(♂)). Total RNA was isolated from the anther of stages 1–6 for qRT-PCR assay (n = 3 independent biological repeats). UBIQ10 was used as an internal control. BECLIN1 expression value at stage 1 was set as 1 and the relative gene expression levels were calculated. Error bars indicate SD. [c] Bar diagram showing the expression of BECLIN1 in the anther stage 3 of 1371, F1 and F’1. [d] Bar diagram showing the expression of COP1 and COP1L105A in the anther stage 1 of 1372, 1373, F1 and F’1 plants.
BECLIN1 expression resulted in abnormal tapetal growth and delayed degeneration
To determine the cause of pollen abortion in BECLIN1 expressing transgenics (1371), we examined cross-sections of anther from control (NTPH) and 1371 transgenic lines at different anther developmental stages46. At stage −1, the tapetum was well developed enclosing microspore mother cells (MMC) which were in the stage of meiotic division in the control (Fig. 6a). At stage 1, the meiotic division occurs, forming dyad and tetrad with distinct tapetum in the control (Fig. 6b). However, no obvious differences were observed in the 1371 transgenic anther in stages −1 and 1, when compared with the control (Fig. 6h,i). The morphological defects started surfacing at stage 2; at this stage, well-developed tapetal layer enclosing nascent-free microspores were observed in control anther (Fig. 6c), whereas in 1371 anther, tapetal cells showed abnormal growth and the entire pollen sac was filled with debris and unstructured cell matter (Fig. 6j). At stage 3, control tapetal cells shrank and microspores were clearly visible (Fig. 6d); whereas in 1371, the abnormal tapetal structure, debris and unstructured cell matter filled the pollen sac and microspores were not clearly visible (Fig. 6k). At stage 4, the control tapetum was degenerating and only epidermis and endothecial layers were visible as surrounding filled mature pollens (Fig. 6e). Interestingly in 1371 anther, tapetal layer remained intact with empty pollens in the pollen sac (Fig. 6l). At stages 5,6, the control tapetum was completely degenerated and pollens were mature with a darkly stained dense cytoplasm (Fig. 6f,g); whereas in 1371 transgenics, the tapetum was still visible as enclosing empty aborted pollens (Fig. 6m,n). To ensure that the abnormal tapetal growth and delayed degeneration in the male sterile parent was because of the expression of BECLIN1, the anther cross-sections of the F’1 were observed, as we had already observed complete abolition of BECLIN1 expression in F’1. The cross-section of F’1 showed normal development of tapetum and pollen similar to the control (Fig. 6o–u), confirming that the tapetal abnormality and pollen abortion in 1371 were due to the expression of BECLIN1.
Transverse anatomical comparison of anther development in wild type (NTPH), Beclin1 transgenic (1371) and F’1 Semi-thin cross sections of control
([a] to [g]), 1371 transgenic ([h] to [n]) and F’1 ([o] to [u]) at anther stages from −1 to stage 6. E, epidermis; En, endothecium; ML, middle layer; T, tapetum; TDR, tetrads; Msp, microspore; Ps, pollen sac; C-T, connective tapetum, MMC, microspore mother cell; aMsp, aborted microspore. Bars = 100 μm.
Discussion
In this study, we have developed a male sterility–fertility restoration system for heterosis breeding in plants. In the female parent, tapetum-specific, high-level and postmeiotic expression of the Arabidopsis BECLIN1/ATG6 gene led to complete male sterility. The tapetum–specific expression was completely abolished and male fertility was restored in the F1 hybrid.
The male sterility system (construct 1371; Supplementary Fig. S1a-I) is the modification of our previously reported two component system15 based on the principle of expression restoration of a TGTA-mutated promoter by providing a complementing TBPm3, which binds specifically to the TGTA36. The system gave several-fold enhancement of expression over the native tapetum-specific promoter (Fig. 1a), in agreement with our previous report15. In this study, we included the light-regulated transcription factor HFR1 that exhibits its COP1-mediated degradation37,49,50 to limit TBPm3 and abolish expression of the desired gene. HFR1 is a transcription factor (bHLH) consisting of 292 amino acids, of which the N-terminus 131 amino acid interacts with COP1 and the C-terminus 161 amino acid has a functional role to bind DNA and promote photomorphogenesis39,49. We fused the HFR1NT131 fragment to the N-terminus of TBPm3, thus making the fusion protein HFR1NT131-TBPm3. This fusion did not affect the functionality of TBPm3, resulting in a high-level expression of the desired gene (gusA or BECLIN1) (Fig. 1a). The transgenic plants expressing gusA were normal in growth and development and the strength and stringency of the expression system were not compromised (Fig. 2b,c–f and Supplementary Fig. S1-b and c). To ensure the transcriptional abolition of tapetum-specific expression in F1 progeny, it was crossed with the male parent expressing COP1 under the regulation of the Arabidopsis PA9 promoter47 (Fig. 1b). The expression of COP1 ensures its availability to bind with HFR1NT131 and to degrade it in the F1 tapetal cell. The degradation of TBPm3 was also achieved, as it was conjugated HFR1NT131-TBPm3 (Fig. 1c). However, COP1 facilitates only partial abolition of the TGTA-TBPm3 complementation system in F1 (Fig. 2b,c–f). The COP1 protein is known to shuttle between the cytoplasm and nucleus39 and stoichiometrically insufficient nuclear concentration of COP1 might be a reason for partial expression reversion. Therefore, we made use of an alternative COP1-mutant (COP1L105A)38. The mutation retained dimerization and functional activity of the COP1 but increased its nuclear abundance. The expression of the reporter gene (gusA) was completely abolished in F’1 progeny when COP1L105A lines were used as a male parent, instead of native COP1 (Fig. 2b,c–f). Thus, a tapetum-specific reversible expression system was established.
The male sterility and fertility restoration system developed by us are generic in nature and hence can be used with other reported genes for male sterility. Thus, the female expression cassette (1371) can be used to generate the male-sterile parent by expressing any known gene reported for the male sterility such as BARNASE18 , BAX28 and so on. The major advantage with our system that there is no specific requirement of the fertility restoration gene for example BARSTAR in BARNASE18,19. The male expression cassette (Construct 1373) offers restoration through abolishing transcription of the male sterility gene. We used the Arabidopsis BECLIN1/ATG615 gene to raise complete male sterile transgenic plants. Genetic engineering of male-sterility and the fertility-restoration system have emerged as tangible options for hybrid seed production. Several restoration systems have been reported to redeem male fertility by inactivating the male sterility protein18,19, degrading transcripts of the male sterility gene14,17,51 and site-specific recombination in the male sterility gene8,10. However, efficient restoration of fertility has been discussed as one of the limiting factors in some of the systems14,21,51. The present system offers a system equipped with complete male sterility and fertility restoration in F1-progeny (Figs 4,1a–c; proposed model) with a novel approach.
The restoration of fertility of F1 hybrid is prerequisite, especially when the economic product is seed. The barnase/barstar system18,19 was deployed for the commercial hybrid production but the identification of efficient restorer (barstar) line was proven to be difficult in Brassica juncea; one in 54 cross-recombination between barnase (male sterile) × barstar (restorer) was adequately restored male fertility in the F1 hybrid21,23. Tapetum-specific promoter TA29 driven barstar (restorer) restore 65.6% male fertility (in terms of pollen viability), it was further improved to 78–90% when PA9 and chimeric system were used to express barstar (restorer)9,21. Barnase weakly expressed in vegetative tissue resulted yield penalty in the plants23. The other barnase based systems; the Cre/loxp-mediated site-specific recombination system10, two-component system22 and split-gene system4,11 claimed 100% restoration of F1 fertility, however, use of toxic gene of trans-origin limited the acceptability due to biosefty concern in some countries. Pathogenesis-related (PR) β-1,3-glucanase gene based male sterility was only partially restored by pA9-driven sense and antisense PR glucanase fragments51. The temperature-sensitive DIPTHERIA TOXIN-A (DTAts) confer conditional-male-sterility (18 °C male sterility, 26 °C restored fertility)32 and reversible male sterility in egg plant16 claimed complete restoration but works on ethanol inducible method which limit its practical applicability. In our system, female expression cassette (1371) expressing plants generate complete male sterility; when compared with control (pollen viability (%): 77.7 ± 1.3 (100%), pollen germination (%): 70 ± 3.6 (100%) and seed-setting (mg/pod): 74.23 ± 5 (100%)), 10 randomly selected BECLIN1 expressing transgenic lines showed pollen viability (%): 0.76 ± 0.78 (~0.96%), pollen germination (%): 0.76 ± 0.73 (~2.3%) and nil seed setting (Fig. 4u,v). The fertility restoration of F1 progeny works on transcription abolition of male sterility gene (BECLIN1) regulated through male parent. When COP1 expressing lines (1372) were used as male parent, F1 showed ~21-fold reduction in BECLIN1 expression (Fig. 5c), which restored pollen viability (%): 15.14 ± 1.8 (20%), pollen germination (%): 18.5 ± 3.3 (26%) that is sufficient for optimal seed setting (mg/pod) 60.2 ± 15 (81%) (Fig. 4u,v). We observed further improvement in fertility restoration when COP1-mutant (COP1L105A) lines (1373) were taken as male parent, which completely abolished the expression of BECLIN1 resulting complete restoration of pollen fertility in F’1 with pollen viability (%): 74.58 ± 1.2 (96%), pollen germination (%): 69 ± 4.6 (~97.14% ) and seed setting (mg/pod) 71.9 ± 6.1 (97%) (Fig. 4u,v) comparable to the untransformed control plants.
Maintaining the male-sterile female lines is a prerequisite for future commercial application of this technique and the genetic design of the male-sterile female line (construct 1371, Fig. S1a-ii) provides this opportunity. Crossing the heterozygous male-sterile female parent (BECLIN1/−) with its wild type (−/−) results in ~50% of the male-sterile progeny (1:1 ratio, (BECLIN1/−) and (−/−)). In future approaches, linking the herbicide resistance gene in the construct 1371 (as in SeedLinkTM) will enable the selection of male-sterile female parents. However, this requires overplanting and eliminating half of the sown plants by applying herbicide to obtain pure male-sterile female parents.
In conclusion, we have developed a system for tapetum-specific, high-level expression of the desired gene to achieve complete male-sterility, along with a system for transcriptional control over the expression system for fertility-restoration in the F1-hybrid. The tapetum-specific expression of the BECLIN1/ATG6 gene facilitated complete male sterility and COP1-mediated HFR1 degradation system was used for repression of transcription of BECLIN1 followed by fertility restoration in the F1 hybrid. The proposed male sterility-fertility restoration system described here will be a valuable future contribution for exploiting hybrid vigor and commercial production of hybrid seed.
Methods
DNA Constructs, Transformation and transgenic development
The four expression cassettes were constructed in the plant expression vector pBI101 and modified pCAMBIA1300 (Schematically presented in Supplementary Fig. S1A). All the vector constructs that were transformed in Agrobacterium and transgenic tobacco plants were developed as previously described15.
Site-directed mutagenesis
TGTATATG mutation was introduced into the TATATATG box region of the TA29 promoter using Quik Change XL Kit (Stratagene, http://www.stratagene.com) as previously described15. In COP1, the L105A mutation38 was introduced using site directed mutagenesis PCR was performed according to the manufacturer’s instructions. 5′TTCGCGGCCGATAAGGCAGCGAAG 3′ mutation was introduced into the 5′ TTCTTGCTCGATAAGCTATTGAAG 3′ region of the COP1 gene by using two sets of primers COPM_f1 5′gct tta ccc taa ttt cgc ggc ccg ata agc tat tga aga aaa ctt c 3′, COPM_r1 5′gtt ttc ttc aat agc tta tcg gcc gcg aaa tta ggg taa agc tg 3′ and COPM_f2 5′ taa ttt ctt gct cga taa ggc agc gaa gaa aac ttc agc tcg gc 3′, COPM_r2 5′ cga gct gaa gtt ttc ttc gct gcc tta tcg agc aag aaa tta gg 3′ primers, Clones were screened by DNA sequencing for the desired mutation using T3 and T7 primers.
Fluorimetric and histochemical GUS assay
Fluorimetric GUS assay was performed as previously described by36. All results are an average of 10 independent lines with three independent experiments of T1 lines. Histochemical GUS analysis was performed as described by15.
Pollen viability and germination assay
Pollen from the blooming stage of flowers was collected and incubated in Fluorescein Diacetate (FDA)—Propidium Iodide (PI) solution for [2 mg/ml (FDA) in acetone and diluted by 10% sucrose drop by drop until turning milky with 1 mg/ml PI (PBS) added to a final concentration] for 5 min. The pollens were centrifuged at 5000 rpm for 1 min, the supernatant was discarded, 3 washings were done in phosphate-buffered saline (PBS) and finally suspended in 50ul PBS. It was mounted on slowfade@antifade (Molecular probes) and observed under a confocal microscope (LSM510META, CarlZeiss) with FDA (green) and PI (red) filters. The percentage of pollen viability that was counted was an average of ten transgenic (n = 10) with three independent experiments. An in-vitro pollen germination test was performed using artificial liquid medium (10% Sucrose, 0.1 mg/ml Boric acid, 0.3 mg/ml calcium nitrate, 0.2 mg/ml magnesium sulfate and 0.1 mg/ml potassium nitrate). Pollen germination images were taken on Leica microscope. Pollen germination (%) was an average of 10 lines (n = 10) with three replicates of the experiment.
Transmission electron microscopy (TEM) and Scanning electron microscopy (SEM)
Anther samples of developmental stages from −5 to +6 stage as described by46 were washed with 1xPBS (pH 7.2) before fixing in 2.5% glutaraldehyde prepared in .1 M sodium cacodylate (Ladd Research) buffer (pH 7.2) for 2 h at 4 °C. Samples were washed thrice, with 0.1 M sodium cacodylate buffer and post-fixed in 1% osmium tetraoxide for 2 h. Further, samples were washed with sodium cacodylate, dehydrated in acetone series (15–100%) and embedded in araldite-DDSA mixture (Ladd Research Industries, USA). After baking at 60 °C, blocks were cut (60–80 nm thick) by an ultra-microtome (Leica EM UC7) and sections were stained by uranyl acetate and lead citrate. Analysis of sections was done under FEI Tecnai G2spirit twin transmission electron microscope equipped with Gatan digital CCD camera (Netherland) at 60 or 80 kV.
For SEM analysis stage 7 anthers of control (NTPH), 1371 transgenic and F1 and F’1 were washed twice in 0.1 M Sodium cacodylate buffer and fixed in 2.5% glutaraldehyde and 4% paraformaldehyde overnight 4 °C. Samples were washed thrice in 0.1 M Sodium cacodylate, 20 min each and transferred to Osmium tetraoxide overnight; further, two washings in 0.1 M sodium cacodylate buffer were done, dehydration was conducted in acetone series, 15%, 30%, 60% and 90% and 3 changes were made in 100% for 20 min each. Further samples were dehydrated till they reach critical point (CPD), anthers were ruptured and pollens were taken on the stub adhesive, coated with gold particles (2 coating) and observed under the scanning electron microscope (FEG450 Quanta, Netherland).
RNA extraction and quantitative real-time RT-PCR
Total RNA was isolated using Plant Spectrum Total RNA isolation kit according to manual instructions (Sigma –Aldrich, http://www.sigmaaldrich.com). After DNaseI treatment (Ambion Inc, Austin TX USA), RNA integrity was checked by electrophoresis and quantified by using using a NanoDrop® ND-1000 UV-Vis spectrophotometer. 2 μg RNA was reverse transcribed using oligo(dT) primers and Superscript II RT (Invitrogen, Rockville, MD, USA) into first-strand c-DNA in a 20-μL reaction as per manual instructions. Quantitative real-time PCR was performed by Express SYBR®Green ER™ qPCR SuperMix Universal (Invitrogen) using the ABI 7500 Fast Real-Time PCR Detection System (Applied Biosystems). The gene-specific primers listed in Table S1 were utilized. UBQ10 was taken as an internal control. Relative expression was calculated from threshold cycle values52). Three independent qRT-PCR reactions were performed on different cDNA samples.
Microtome and light microscopy
Tobacco anthers of developmental stage from −1 to +646 were immediately fixed in Poly/LEM fixative (Polysciences, Inc. Cat# 16864) overnight at 4 °C. Fixed anthers were dehydrated in ethanol series 50%, 70%, 85%, 95% and 100% for 1h each. Infiltration was done in 2:1, 1:1 and 1:0 of (ethanol: infiltration solution) for 1h each and embedded in resin (JB-4 Embedding Kit, Polysciences Inc. Eppelheim, Germany). A 4-μM-thick section were cut by using Leica microtome, stained with 0.1% toluidine blue O’ (Sigma Aldrich) and pictures were captured using a Nikon microscope. Pictures were adjusted using adobe paint (NET) software.
Crossing strategies and F1 screening
The best expressing lines of transgenic 1370 were taken as the female parent and were crossed by the two best performing lines of 1372 and 1373 which were taken as male parents; F1 and F’1 were generated. Similarly 1371 transgenics were taken as the female parent and were crossed with the best performing 1372 and 1373 pollens; F1 and F’1 were generated. The F1/F’1 seeds were germinated on medium containing kanamycin (300 mg/litre) and hygromycin (50 mg/litre) on a petri-dish for 3 weeks. Plantlets of selected crosses were transferred into pots in a transgenic house.
Additional Information
How to cite this article: Singh, S. P. et al. A novel male sterility-fertility restoration system in plants for hybrid seed production. Sci. Rep. 5, 11274; doi: 10.1038/srep11274 (2015).
References
Schnable, P. S. & Springer, N. M. Progress toward understanding heterosis in crop plants. Annu Rev Plant Biol 64, 71–88 (2013).
Tester, M. & Langridge, P. Breeding technologies to increase crop production in a changing world. Science 327, 818–822 (2010).
Shaoqing Li DYaYZ . Characterization and Use of male sterility in hybrid rice breeding. Journal of Integrative Plant Biology 49, 791–804 (2007).
Kempe, K., Rubtsova, M. & Gils, M. Split-gene system for hybrid wheat seed production. Proc Natl Acad Sci U.S.A. 111, 9097–9102 (2014).
Parodi, P. C. & Gaju, M. D., Male sterility induced by the chemical hybridizing agent clofencet on wheat, Triticum aestivum and T. turgidum var. durum. Ciencia e investigación agraria 36, 267–276 (2009).
Chen, L. & Liu, Y. G. Male sterility and fertility restoration in crops. Annu Rev Plant Biol 65, 579–606 (2014).
Katja Kempe, M. G. Pollination control technologies for hybrid breeding. Molecular Breeding 27, 417–437 (2011).
Bayer, M. H. D. Restoring full pollen fertility in transgenic male-sterile tobacco (Nicotiana tabacum L.) by Cre-mediated site-specific recombination. Mol Breed 15, 193–203 (2005).
Bisht, N. C., Jagannath, A., Burma, P. K., Pradhan, A. K. & Pental, D. Retransformation of a male sterile barnase line with the barstar gene as an efficient alternative method to identify male sterile-restorer combinations for heterosis breeding. Plant Cell Rep 26, 727–733 (2007).
Cao, B., Huang, Z., Chen, G. & Lei, J. Restoring pollen fertility in transgenic male-sterile eggplant by Cre/loxp-mediated site-specific recombination system. Genet Mol Biol 33, 298–307 (2010).
Gils, M. et al. A novel hybrid seed system for plants. Plant Biotechnol J 6, 226–235 (2008).
Li, S. F., Iacuone, S. & Parish, R. W. Suppression and restoration of male fertility using a transcription factor. Plant Biotechnol J 5, 297–312 (2007).
Nizampatnam, N. R. & Dinesh Kumar, V. Intron hairpin and transitive RNAi mediated silencing of orfH522 transcripts restores male fertility in transgenic male sterile tobacco plants expressing orfH522. Plant Mol Biol 76, 557–573 (2010).
Schmulling, T., Rohrig, H., Pilz, S., Walden, R. & Schell, J. Restoration of fertility by antisense RNA in genetically engineered male sterile tobacco plants. Mol Gen Genet 237, 385–394 (1993).
Singh, S. P. et al. BECLIN1 from Arabidopsis thaliana under the generic control of regulated expression systems, a strategy for developing male sterile plants. Plant Biotechnol J 8, 1005–1022 (2010).
Toppino, L. et al. Reversible male sterility in eggplant (Solanum melongena L.) by artificial microRNA-mediated silencing of general transcription factor genes. Plant Biotechnol J 9, 684–692 (2010).
Zabaleta, E., Mouras, A., Hernould, M. & Suharsono, Araya A. Transgenic male-sterile plant induced by an unedited atp9 gene is restored to fertility by inhibiting its expression with antisense RNA. Proc Natl Acad Sci U.S.A. 93, 11259–11263 (1996).
Mariani, C., Beuckeleer, M. D., Truettner, J., Leemans, J. & Goldberg, R. B. Induction of male sterility in plants by a chimaeric ribonuclease gene. Nature 347, 737–741 (1990).
Mariani, C. G. V. et al. A chimaeric ribonuclease-inhibitor gene restores fertility to male sterile plants. Nature 357, 384–387 (1992).
Banga, S. K. B. P & Banga, S. S. Genetically engineered systems of male sterility. J Oilseeds Res 23, 1–7 (2006).
Bisht, N. C. J. A, Gupta, V, Burma, P. K & Pental, D. A two gene—two promoter system for enhanced expression of a restorer gene (barstar) and development of improved fertility restorer lines for hybrid seed production in crop plants. Mol Breed 14, 129–144 (2004).
Burgess, D. G. et al. A novel, two-component system for cell lethality and its use in engineering nuclear male-sterility in plants. Plant J 31, 113–125 (2002).
Jagannath, A. A. N., Gupta, V., Pradhan, A. K., Burma, P. K. & Pental, D. Development of transgenic barstar lines and identifi- cation of a male sterile (barnase)/restorer (barstar) combination for heterosis breeding in Indian oilseed mustard (Brassica juncea). Curr Sci 82, 46–52 (2002).
Goldberg, R. B., Beals, T. P. & Sanders, P. M. Anther development: basic principles and practical applications. Plant Cell 5, 1217–1229 (1993).
Zhu, J. et al. Defective in Tapetal development and function 1 is essential for anther development and tapetal function for microspore maturation in Arabidopsis. Plant J 55, 266–277 (2008).
Papini, A. & Brighigna, S. M. L. Programmed-cell-death events during tapetum development of angiosperms. Protoplasma 207, 213–221 (1999).
Wu, H. M. & Cheun, A. Y. Programmed cell death in plant reproduction. Plant Mol Biol 44, 267–281 (2000).
Kawanabe, T., Ariizumi, T., Kawai-Yamada, M., Uchimiya, H. & Toriyama, K. Abolition of the tapetum suicide program ruins microsporogenesis. Plant Cell Physiol 47, 784–787 (2006).
Li, N. et al. The rice tapetum degeneration retardation gene is required for tapetum degradation and anther development. Plant Cell 18, 2999–3014 (2006).
Luo, D. et al. A detrimental mitochondrial-nuclear interaction causes cytoplasmic male sterility in rice. Nat Genet 45, 573–577.
Varnier, A. L., Mazeyrat-Gourbeyre, F., Sangwan, R. S. & Clement, C. Programmed cell death progressively models the development of anther sporophytic tissues from the tapetum and is triggered in pollen grains during maturation. J Struct Biol 152, 118–128 (2005).
Guerineau, F., Sorensen, A. M., Fenby, N. & Scott, R. J. Temperature sensitive diphtheria toxin confers conditional male-sterility in Arabidopsis thaliana. Plant Biotechnol J 1, 33–42 (2003).
Cho, H. J., Kim, S., Kim, M. & Kim, B. D. Production of transgenic male sterile tobacco plants with the cDNA encoding a ribosome inactivating protein in Dianthus sinensis L. Mol Cells 11, 326–333 (2001).
Takada, K., Ishimaru, K., Minamisawa, K., Kamada, H. & Ezura, H. Expression of a mutated melon ethylene receptor gene Cm-ETR1/H69A affects stamen development in Nicotiana tabacum. Plant Sci 169, 935–942. (2005).
Konagaya, K., Ando, S., Kamachi, S., Tsuda, M. & Tabei, Y. Efficient production of genetically engineered, male-sterile Arabidopsis thaliana using anther-specific promoters and genes derived from Brassica oleracea and B. rapa. Plant Cell Rep 27, 1741–1754 (2008).
Chaturvedi, C. P. et al. Mutated TATA-box/TATA binding protein complementation system for regulated transgene expression in tobacco. Plant J 50, 917–925 (2007).
Duek, P. D., Elmer, M. V., van Oosten, V. R. & Fankhauser, C. The degradation of HFR1, a putative bHLH class transcription factor involved in light signaling, is regulated by phosphorylation and requires COP1. Curr Biol 14, 2296–2301 (2004).
Subramanian, C. et al. The Arabidopsis repressor of light signaling, COP1, is regulated by nuclear exclusion: mutational analysis by bioluminescence resonance energy transfer. Proc Natl Acad Sci U.S.A. 101, 6798–6802 (2004).
Yang, J. et al. Light regulates COP1-mediated degradation of HFR1, a transcription factor essential for light signaling in Arabidopsis. Plant Cell 17, 804–821 (2005).
Chen, W. & Struhl, K. Saturation mutagenesis of a yeast his3 “TATA element”: genetic evidence for a specific TATA-binding protein. Proc Natl Acad Sci U.S.A. 85, 2691–2695 (1988).
Harbury, P. A. & Struhl, K. Functional distinctions between yeast TATA elements. Mol Cell Biol 9, 5298–5304 (1989).
Wobbe, C. R. & Struhl, K. Yeast and human TATA-binding proteins have nearly identical DNA sequence requirements for transcription in vitro. Mol Cell Biol 10, 3859–3867 (1990).
Reddy, P. & Hahn, S. Dominant negative mutations in yeast TFIID define a bipartite DNA-binding region. Cell 65, 349–357 (1991).
Strubin, M. & Struhl, K. Yeast and human TFIID with altered DNA-binding specificity for TATA elements. Cell 68, 721–730 (1992).
Pan, S., Czarnecka-Verner, E. & Gurley, W. B. Role of the TATA binding protein-transcription factor IIB interaction in supporting basal and activated transcription in plant cells. Plant Cell 12, 125–136 (2000).
Koltunow, A. M., Truettner, J., Cox, K. H., Wallroth, M. & Goldberg, R. B. Different temporal and spatial gene expression patterns occur during anther development. Plant Cell 2, 1201–1224 (1990).
Paul, W., Hodge, R., Smartt, S., Draper, J. & Scott, R. The isolation and characterisation of the tapetum-specific Arabidopsis thaliana A9 gene. Plant Mol Biol 19, 611–622 (1992).
Sawant, S., Singh, P. K., Madanala, R. & Tuli, R. Designing of an artificial expression cassette for the high level expression of transgenes in plants. Theoretical and Applied Genetics 102, 635–644 (2001).
Jang, I. C., Yang, J. Y., Seo, H. S. & Chua, N. H. HFR1 is targeted by COP1 E3 ligase for post-translational proteolysis during phytochrome A signaling. Genes Dev 19, 593–602 (2005).
Kim, Y. M., Woo, J. C., Song, P. S. & Soh, M. S. HFR1, a phytochrome A-signalling component, acts in a separate pathway from HY5, downstream of COP1 in Arabidopsis thaliana. Plant J 30, 711–719 (2002).
Hird, D. L., Paul, W., Hollyoak, J. S. & Scott, R. J. The restoration of fertility in male sterile tobacco demonstrates that transgene silencing can be mediated by T-DNA that has no DNA homology to the silenced transgene. Transgenic Res 9, 91–102 (2000).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Acknowledgements
The authors are grateful to the Department of Biotechnology (DBT), the Government of India, for funding the research project under Sr.IYBA. They are also thankful to the Council of Scientific and Industrial Research (CSIR) for fellowship funding support. They also acknowledge the support of Dr. Manu Agarwal, Delhi University, India for critical reading and the important suggestions in their article.
Author information
Authors and Affiliations
Contributions
S.V.S., S.P.S.1 and S.P.S.2 designed the study. S.P.S.1 and S.V.S. analyzed the data. All the experiments are performed by S.P.S.1. S.P.S.2 and TP helped in designing and cloning few constructs. S.V.S., S.P.S.1, S.P.S.2 and R.R.S. contributed towards the preparation of this article.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Singh, S., Singh, S., Pandey, T. et al. A novel male sterility-fertility restoration system in plants for hybrid seed production. Sci Rep 5, 11274 (2015). https://doi.org/10.1038/srep11274
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep11274
This article is cited by
-
Comparative transcriptome analysis in Chinese cabbage (Brassica rapa ssp. pekinesis) for DEGs of Ogura-, Polima-CMS and their shared maintainer
Physiology and Molecular Biology of Plants (2020)
-
Evolvement of transgenic male-sterility and fertility-restoration system in rice for production of hybrid varieties
Plant Molecular Biology (2018)
-
Molecular Approaches for Manipulating Male Sterility and Strategies for Fertility Restoration in Plants
Molecular Biotechnology (2017)
-
Cytoplasmic male sterility (CMS) in hybrid breeding in field crops
Plant Cell Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.