Abstract
The evolution of multicellularity required novel mechanisms for intercellular communication, but their origin is unclear. Dictyostelium cells exchange signals to position specialized cell types in multicellular spore-bearing structures. These signals activate complex pathways that converge on activation of cAMP-dependent protein kinase (PKA). Genes controlling PKA were detected in the Dictyostelid unicellular ancestors, which like most protists form dormant cysts when experiencing environmental stress. We deleted PKA and the adenylate cyclases AcrA and AcgA, which synthesize cAMP for PKA activation, in the intermediate species Polysphondylium, which can develop into either cysts or into multicellular structures. Loss of PKA prevented multicellular development, but also completely blocked encystation. Loss of AcrA and AcgA, both essential for sporulation in Dictyostelium, did not affect Polysphondylium sporulation, but prevented encystation. We conclude that multicellular cAMP signalling was co-opted from PKA regulation of protist encystation with progressive refunctionalization of pathway components.
Similar content being viewed by others
Introduction
The transition from uni- to multicellularity occurred at least eight times independently and enabled the evolution of complex macroscopic life forms. Multicellularity allows division of labour between cells and the construction of multi-layered tissues and organs, in which specialized cells perform different functions. However, multicellularity also requires novel mechanisms for intercellular communication that allow the specialized cell types to differentiate in well-regulated proportions and at appropiate locations. Most unicellular eukaryotes or protists have a simple life cycle in which feeding cells differentiate into a dormant cyst, when faced with starvation or other forms of stress1. The physiology and differentiation of protists are therefore mainly regulated by environmental signals. Upon transition to multicellularity their sensory systems must have adapted to enable communication between cells. Because the signal processing systems of protists have been little investigated and the early multicellular forms are long extinct, the early stages in the evolution of multicellularity are not understood.
Dictyostelid social amoebas aggregate to form migrating sorogens, which transform into fruiting bodies, containing up to a million cells. These cells differentiate into spores and four different cell types to carry the spore mass aloft. In the genetic model system Dictyostelium discoideum, the mechanisms controlling multicellular development have been intensively investigated, highlighting a dominant role for cAMP throughout the developmental programme. D. discoideum aggregation is mediated by secreted cAMP pulses that are produced by the adenylate cyclase ACA and detected by surface cAMP receptors (cARs). Once aggregated, a second adenylate cyclase, ACG, is post-transcriptionally upregulated in the posterior region of the sorogen. cAMP, produced by ACG, acts on both cARs and PKA to induce the differentiation of prespore cells2. A third adenylate cyclase, ACR, acts later on PKA to trigger the maturation of spores and assist in the maturation of stalk cells3. ACG acting on PKA has a second role in the mature fruiting body, where it mediates inhibition of spore germination by ambient high osmolyte levels4. The cAMP phosphodiesterase RegA also plays a major role in regulating cAMP levels during cell differentiation and spore germination. RegA activity is regulated by signals that are exchanged between the maturing spore and stalk cells. These signals bind to sensor histidine kinases/phosphatases which regulate the phosphorylation state of the intrinsic response regulator of RegA5,6,7.
D. discoideum resides in group 4 of the four major groupings of Dictyostelia8, which together are members of Amoebozoa, a kingdom that consists mainly of unicellular amoebas that encyst individually. While D. discoideum development is strictly multicellular, many dictyostelids in groups 1–3, have retained the ancestral process of encystation, in addition to fruiting body formation and are thus ideally suited to investigate how cellular communication systems adapted during the transition to multicellularity. One of these species, Polysphondylium pallidum, is like D.discoideum amenable to both forward and reverse genetic approaches. In addition to the D. discoideum genome9, the genomes of P.pallidum, residing in group 2, D. fasciculatum, residing in group 1 and D. lacteum, residing in group 3 were recently sequenced by ourselves and colleagues10(Schaap, P. and Gloeckner, G. in preparation). Genome sequence of the strictly unicellular amoebozoan Acanthamoeba castellani is also available11. This information allows us to retrace conservation and change in known developmental signalling genes. We have developed procedures for successive disruption of multiple genes in Polysphondylium pallidum, which allows us to assess gene function in both unicellular and multicellular development12. In this work we test the hypothesis that multicellular sporulation is evolutionary derived from unicellular encystation by investigating the roles of the catalytic subunit of PKA (PkaC) and the adenylate cyclases ACG and ACR in the uni- and multicellular life cycles of P. pallidum.
Results
Conservation of PkaC, ACG and ACR in Amoebozoa
Social amoebas can be subdivided into four major groups, which together are members of Amoebozoa, a kingdom of mainly unicellular amoebas8. Genomes, representative of the four groups and three genomes of unicellular Amoebozoa have recently become available9,10,11,13 (http://sacgb.fli-leibniz.de/; http://www.physarum-blast.ovgu.de/). We investigated the presence of genes encoding ACR, ACG and PkaC, the catalytic subunit of PKA in these genomes. BLAST queries detected homologs of PkaC and ACR in all Amoebozoan genomes, except that of the obligatory parasite Entamoeba histolytica, while ACG was only conserved in the Dictyostelid genomes (Figure 1). Because the PkaC sequence is similar to that of many other protein kinases, we also included the closest homologs of PkaC (PkgB and PkgD) in our query. Phylogenetic inference showed that our initally selected PkaC orthologs grouped together with D. discoideum PkaC, while hits to the PKA homologs grouped with either PkgB or PkgD (Supplementary Figure 1).
Conservation of PkaC, ACG and ACR in Amoebozoa.
(A–C). PkaC, ACG and ACR sequences were retrieved from Dictybase (http://dictybase.org/) or by BLASTp or tBLASTn query of sequenced genomes using D. discoideum PkaC, ACG and ACR as bait. Sequences were aligned using Clustal Omega30 and phylogenetic relationships were determined with MrBayes31. The posterior probabilities of tree nodes are indicated by colored dots and the trees are annotated with the functional domain architecture of the proteins, as determined with SMART32. Gene identifiers are color-coded to reflect species names as outlined in panel D. (D). Phylogeny of Amoebozoa inferred from 32 aligned protein sequences33. Numbers between brackets denote the taxon group in which the Dictyostelid species reside.
All ACG proteins consist of a CHASE domain flanked by one or two transmembrane domains and a cytosolic adenylate cyclase domain (Figure 1). The ACR homologs also display a similar functional domain architecture as D. discoideum ACR with 6–7 transmembrane domains, a HATPase-c domain, two receiver domains and a cyclase catalytic domain. The HisKA autophosphorylation/dimerization domain that is usually located at the N-terminus of the HATPase-c domain is missing in the Amoebozoan ACRs. The closest homologs to ACG or ACR are prokaryote ACs, while a fungal PkaC is closest to the amoebozoan PkaCs.
Disruption of P.pallidum PkaC by homologous recombination
To assess the function of PkaC in uni- and multicellular development, we disrupted P. pallidum PkaC by replacing a PkaC internal segment with a floxed G418 cassette14 (Supplementary Figure 2). The pkac- amoebas and a control random integrant (RI) proliferated normally, but completely lost the ability to aggregate and form fruiting bodies. After 48 h of starvation, when RI cells had formed fruiting bodies, the lawn of starving pkac- cells had somewhat contracted, but aggregates were never formed (Figure 2A). Strikingly, unlike RI controls, the pkac- amoebas also could not form cysts when starved at high osmolarity (Figure 2B), a condition that triggers encystation of unicellular amoebas15.
Phenotype of P.pallidum pkac- and pkac-/PkaC mutants.
(A). Fruiting body formation. Amoebas of the P. pallidum random integrant (RI) control, pkac- mutant and pkac- complemented with PkaC and its 5′ intergenic region (see Supplementary figure 2 for generation of mutants), were freed from bacteria, incubated for 48 h on NN agar and photographed. Bar: 1 mM. (B). Encystation. RI control, pkac- and pkac-/PkaC amoebas were incubated in 0.4 M sorbitol for 48 h, stained with 0.001% Calcofluor and photographed under phase contrast and UV illumination (right panels). Bar: 10 μM.
After removal of the G418 selection cassette by transformation with cre-recombinase, the pkac- cells were transformed with the PkaC coding sequence fused to its own promoter. The resulting pkac-/PkaC cells fully regained both development into fruiting bodies and encapsulation of single amoebas into cysts (Figs. 2A,B), thus demonstrating that PkaC is essential for both multicellular development and encystation.
Disruption of AcgA and AcrA in P.pallidum
ACG has an overlapping role with ACR in prespore differentiation in D. discoideum and mediates inhibition of spore germination by high ambient osmolarity in the spore head2,4,16. To investigate whether ACG has essential roles in P. pallidum multicellular development and encystation, we disrupted the ACG gene AcgA in P. pallidum (Supplementary figure 3). The acga- mutant formed normal aggregates and fruiting bodies (Figure 3A). Surprisingly, the fruiting bodies contained viable detergent resistant spores, that showed normal inhibition of spore germination by high osmolarity (Figure 4D). The acga- mutant also showed normal encystation (Figure 3C).
Phenotype of acga- and acra- mutants.
(A). Fruiting body formation. P. pallidum wild-type, acga- and acra- amoebas, freed from bacteria, were incubated on NN agar for 48 h under ambient room lighting and photographed. Bar: 100 μM. (B). Spores. Wild type and acra- spores were harvested from fruiting bodies, stained with 0.001% Calcofluor and photographed under phase contrast and UV illumination (right panels). Bar: 10 μM. (C). Encystation. Wild-type, acga- and acra- amoebas, freed from bacteria, were incubated in 0.4 M sorbitol for 48 h, stained with Calcofluor and photographed as above. Bar: 10 μM.
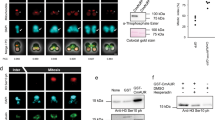
Phenotype of a P.pallidum acra-/acga- mutant.
(A). Fruiting body morphology. The AcgA gene was deleted in an acra- mutant (see Supplementary figure 3) and the resulting acra-acga- cells were developed for 48 h on NN agar. Bar: 100 μm. (B). Stalk and spores. Wild-type and acra-acga- fruiting bodies were picked up with a needle, deposited in 0.001% Calcofluor on a slide glass and photographed under phase contrast and UV. Bar: Bar 100 μm. (C). Cysts. Wild-type, acra-acga- and acra-acga- cells transformed with P. pallidum AcrA and its 5′intergenic region were incubated for 48 h in 0.4 M sorbitol and stained with Calcofluor. Bar: 10 μm. (D). Spore germination. Detergent treated spores of wild-type P. pallidum, acra-, acga-, acra-acga- and acra-acga-/AcrA were plated with E.coli on LP agar or LP agar with 0.4 M sorbitol at 200 spores per plate (143 cm2). Emerging plaques were counted after 4–6 days at 22°C. Means and SD of 3 experiments with duplicate plates for each strain are presented. (E). Cyst germination. Detergent treated cysts of the above cell lines, except acra-acga-, which does not form cysts, were plated in the presence and absence of sorbitol as described for spores. In addition to the wild-type, a random integrant of the AcrA knock-out construct in wild-type was included as a control. Means and SD of 4 experiments with duplicate plates are presented. (F). Encystation time course. Amoebas of the cell lines mentioned above were incubated in KK2 with 0.4 M sorbitol. At the indicated time points, samples were stained with Calcofluor and the numbers of fluorescent cysts and unstained amoebas were counted. Means and SD of 4 experiments, performed in duplicate.
ACR is essential for spore maturation in D. discoideum3 and to assess its role in P. pallidum, we next ablated the P. pallidum AcrA gene (Supplementary Figure 4). Surprisingly, unlike D. discoideum acra-, the P. pallidum acra- mutant displayed normal spore formation in fruiting bodes (Figure 3A,B). The acra- mutants also encysted when submerged at high osmolarity, but encystation was less efficient than in wild-type P. pallidum (Figs. 3B, 4F). To assess whether ACG and ACR are functionally redundant, we created a double acra-acga- mutant. P. pallidum acra- cells were transformed with cre-recombinase to excise the neomycin selection cassette and G418 sensitive clones were selected, which were then transformed with the AcgA knock-out construct. Strikingly, the acra-acga- mutant was also not defective in fruiting body morphology or sporulation (Fig. 4A,B). However, the ability to encyst was completely lost (Fig. 4C). To validate that this was due to loss of cAMP synthesis, the neomycin cassette was excised from the acra-acga- mutant once more and cells were transformed with a plasmid that contained a P. pallidum genomic fragment, encompassing the full length AcrA coding and 5′intergenic region. The resulting acra-acga-/AcrA mutant regained the ability to encyst (Fig. 4C), confirming that lack of encystation was due to the combined loss of AcgA and AcrA.
Roles for ACG and ACR in regulation of encystation and spore germination by osmolytes
When comparing P. pallidum to D. discoideum, a striking difference is the absence of an obvious role for ACR or ACG in P. pallidum spore differentiation. It is also remarkable that ACR can complement ACG in osmolyte-induced encystation, a role more suited for ACG with its intramolecular osmosensor17. In D. discoideum, high osmolarity prevents spores from germinating prematurely, while still in the sorus. We first investigated whether ACG or ACR still regulated spore germination in P. pallidum. Mature spores from wild-type and mutant fruiting bodies were incubated with detergent to lyse unencapsulated cells and plated clonally on E.coli lawns in the presence and absence of 0.4 M of the osmolyte sorbitol. In the absence of osmolyte, wild-type, acra-, acga-, acra-acga- and acra-acga-/AcrA spores germinated with equal efficiency (Figure 4D). In the presence of osmolyte, germination of wild-type, acga- and acra-acga-/AcrA spores was about 85% reduced and of acra- spores about 30%. Only acra-acga- spores still germinated at full efficiency. These data show that ACG and ACR both mediate inhibition of spore germination by high osmolarity, with ACR playing the more dominant role.
Cyst germination on E.coli lawns is only 30% reduced by 0.4 M sorbitol in wild-type P. pallidum and by about 50% in a random integrant of the AcrA KO construct (Fig. 4E). Inhibition of cyst germination by osmolyte was completely lost in the acra- mutant, but not in the acga- mutant. Remarkably, cyst germination was completely inhibited when acra- cells were complemented with AcrA expressed from its own promoter. This could be a consequence of AcrA overexpression, caused by integration of multiple copies of the expression vector in the P. pallidum genome.
A time course of the progression of encystation in wild type P. pallidum and all adenylate cyclase mutants showed that 60% of wild-type and RI control amoebas have encysted after 72 h of incubation with osmolyte (Fig. 4F). The acra-acga- cells have completely lost encystation, while acra- cells encyst more slowly. Unexpectedly, both the acga- cells and the acra-acga-/AcrA cells reached 70–80% encystation within 24 hours. Because acra-acga-/AcrA also showed more efficient inhibition of cyst germination by high osmolarity, this may simply imply that the increased ACR levels facilitated encystation. Why ACG should be required for encystation in an acra- background, but simultaneously reduce encystation in a wild-type background is less clear. It suggests a scenario in which ACR is the main player, with its expression down-regulated by ACG. Negative regulation of AcrA expression by ACG was previously observed in D.discoideum slugs2.
Discussion
ACR and ACG have overlapping roles in inducing prespore differentiation and spore maturation in D.discoideum, but appeared to be dispensible for the same processes in P.pallidum spore differentiation. However, the P. pallidum genome contains three copies of the gene encoding the adenylate cyclase ACA10, which is mainly involved in chemotactic signalling in D.discoideum7. One of the Ppal copies, ACA2, is expressed in prespore cells (Kawabe, Y., in preparation) and likely provides cAMP for spore differentiation.
The most striking outcome of this work is that deletion of PkaC and combined deletion of AcrA and AcgA completely blocks the process of encystation in P. pallidum. PkaC and ACR, but not ACG, are conserved in other Amoebozoa, such as Acanthamoeba castellani and Physarum polycephalum (Figure 1). Other components of the cAMP signalling pathway such as PkaR and the cAMP phosphodiesterase RegA are also conserved in these amoebozoan genomes and, remarkably, also in the amoeboflagellate Naegleria gruberi, which resides in the kingdom Excavata11,18,19.
RegA plays a central role in regulating cAMP levels during D. discoideum multicellular development, where its activity is controlled by intercellular signalling. The signals bind to sensor histidine kinases/phosphatases (SHKPs), which regulate the phosphorylation state of the intrinsic response regulator of RegA and thereby the attached cAMP phosphodiesterase activity7. The genomes of Naegleria and all sequenced Amoebozoa contain a large number of SHKPs. In non-dictyostelid amoebas, the roles of SHKPs have not yet been explored, but in D. discoideum their main target is RegA. About 7 out of the 15 Dictyostelium SHKPs mediate effects of secreted factors that control the developmental programme. However the stimuli detected by the other 8 SHKPs and those of the unicellular amoebas are likely to be of environmental origin.
Loss of RegA causes precocious encystation in P. pallidum and A.castellani19, as expected for a protein that inhibits PKA function. Combined with results obtained in this work, this allows us to propose a universal mechanism for control of amoebozoan and excavate encystation (Figure 5). In this scheme, cAMP activation of PKA induces encystation and prevents excystation. cAMP is synthesized by ACR in Amoebozoa, while in both Amoebozoa and Excavata, cAMP is hydrolysed by RegA. RegA activity is under both positive and negative regulation from external stimuli that activate a sensor histidine kinase or a sensor histidine phosphatase, respectively. For the former, such stimuli could be stress factors that cause encystation and for the latter, the stimuli could signal the presence of food to cause cyst germination.
Pathway for encystation in Amoebozoa and Excavata.
The choice between feeding trophozoite and dormant cyst stage is controlled by cAMP binding to PkaR causing dissociation from and activation of PkaC. cAMP is synthesized by adenylate cyclases, which may be activated directly by stress. cAMP is hydrolyzed by the cAMP phosphodiesterase RegA, which is respectively activated by sensor histidine kinases that sense food, or inhibited by sensor histidine phosphatases that sense stress. Violet: components that are conserved in Amoebozoa and Excavata, blue: conserved in Amoebozoa, Teal or pink, conserved in only dictyostelia or Excavata, respectively. Arrow: stimulates; crossbar: inhibits.
Researchers have been intrigued for many years by the fact that PkaC regulates so many aspects of D. discoideum development, starting with the transition from growth to development20, the regulation of chemotactic signalling21, the differentiation of prespore cells22, the maturation of spores and stalk cells23,24,25 and the germination of spores4. We show here that these roles are likely to have emerged from an original role of PKA in mediating stress-induced encystation and stress-inhibited excystation. The trigger for multicellular development is also nutrient stress and its end-point is the formation of two encapsulated cell types, the stalk cells and spores. It therefore appears that PKA retained its ancestral role in encapsulation, when Dictyostelia started to adopt a multicellular survival strategy, but additionally acquired novel roles to assist the organism to proceed through its multicellular differentiation programme in a well-regulated manner. These novel roles required novel mechanisms to regulate cellular cAMP levels. The differences in ACG, ACR and ACA function that we noted in this work between the group 2 species P.pallidum and the group 4 species D.discoideum may reflect different evolutionary trajectories towards effective cell signalling between these groups.
In conclusion, we propose that the range and complexity of the cAMP-mediated signalling cascades that control almost all stages of multicellular development in modern Dictyostelia have been co-opted from an ancestrol role for cAMP in stress-induced dormancy. The emergence of complex cell-cell communication from basic environmental sensing in Dictyostelia may prove to be a paradigm for the evolution of multicellularity in other eukaryote lineages.
Methods
Cell growth, development, encystation and sporulation
Growth and development
Polysphondylium pallidum strain PN500, was routinely grown in association with Escherichia coli on lactose peptone (LP) agar. For multicellular development, P. pallidum cells were harvested in 20 mM KH2PO4/K2HPO4, pH 6.5 (KK2), washed free from bacteria and incubated on non-nutrient (NN) agar at 106 cells/cm2 and 22°C.
Encystation
For quantification of encystation, P. pallidum cells were grown in a suspension of autoclaved Klebsiella aerogenes in KK2, until cell density reached 5 × 106–107 cells. Cells were washed, resuspend in KK2 with 0.4 M sorbitol at 107 cells/ml and shaken at 180 rpm and 21°C. Aliquots of 0.1 ml were sampled at regular intervals and supplemented with 1 μl 0.1% Calcofluor (which reacts to cellulose in the cyst wall). Total amoeba and cyst numbers were determined by counting cells in a haemocytometer under phase contrast and UV illumination, respectively. 100–500 cells were counted for each time point.
Cyst and spore germination
P. pallidum spores were harvested from 5-day old fruiting bodies. For cysts, P. pallidum cells were grown in K.aerogenes suspension as described above, washed and incubated with 0.4 M sorbitol for 3 days to allow mature cysts to form. Cysts or spores were treated for 10 min with 0.1% Triton-X100 to lyse amoeboid cells, washed with KK2, counted and clonally plated on LP with E. coli. Emerging P. pallidum plaques were counted after 4–6 days.
Gene disruption and complementation
Disruption of P. pallidum PkaC
For disruption of PkaC, two fragments of P. pallidum gDNA were amplified using primer pairs PKACI5′/PKACI3′ and PKACII5′/PKACII3′ (Supplementary table 1). The fragments were digested with XbaI/BamHI or XhoI/KpnI, using restriction sites incorporated in primer design and sequentially inserted into inserted into XbaI/BamHI and XhoI/KpnI digested plasmid pLox-NeoIII14 to flank the LoxP-neo selection cassette (Supplementary figure 2A). P. pallidum cells were transformed by electroporation with the XbaI/KpnI vector insert, followed by selection of transformants on autoclaved K.aerogenes at 300 μg/ml G41826. Genomic DNA was isolated from G418 resistant clones and screened by 2 PCR reactions (Supplementary figure 2B) and Southern blot (Supplementary figure 2C) to diagnose PkaC disruption by homologous recombination. Two knock-out (KO) and two random integrant (RI) clones were identified.
Complementation of pkac- with PkaC
To remove the A6neo cassette, KO (pkac-) cells were transformed with pA15NLS.Cre27 for transient expression of Cre recombinase. Transformed clones were replica-plated onto autoclaved K.aerogenes on LP agar with and without 200 μg/ml G418 for negative selection. To generate a PkaC expression construct, the 0.8 kb PkaC 5′intergenic region was amplified from gDNA using primers P-PKAC p3 and P-PKAC P4r (Supplementary table 1), which contain NheI and BglII restriction sites, respectively and ligated into NheI/BglII digested vector pDdCGFP28. Next, the PkaC coding sequence was amplified using primers palPKAC p9 and palPKAC p11r (Supplementary table 1), which harbour BglII and XbaI sites, respectively and ligated into the BamHI and XbaI sites of the newly created vector. This fuses PkaC N-terminally to its own 5′intergenic region and C-terminally to green fluorescent protein (GFP)(see Supplementary figure 2A). The construct was transformed into pkac- cells and G418 resistant cells were selected as described above.
Disruption of P. pallidum AcgA
To disrupt P. pallidum AcgA, two AcgA fragments of 2.8 and 1.1 kb, respectively, were amplified from P. pallidum gDNA using primer pairs Pp-ACG-P5/Pp-ACG-P3 and Pp-ACG-53H/Pp-ACG-53K (Supplementary table 1). The first fragment was reduced to 2.4 kb by XbaI/BglII digest and inserted into XbaI/BamHII digested vector pLox-NeoIII14. The second fragment was digested with HindIII/KpnI, using restriction sites introduced in the primers and inserted into the HindIII/KpnI sites of the newly generated vector pAcgA-KO (Supplementary figure 3). P. pallidum cells were transformed by electroporation with the linearized vector pAcgA-KO, followed by selection of transformants and diagnosis of gene knock-out as outlined in Supplementary figure 3.
Disruption of P. pallidum AcrA
For P. pallidum AcrA disruption, two DNA fragments of ~2.0 kb were amplified by PCR from P. pallidum gDNA, using primer pairs AcrA5′-fw/AcrA5′-rev and AcrA3′-fw/AcrA3′-rev (Supplementary table 1), which harbour KpnI/BamHI and HindIII restriction sites, respectively. The two fragments were sequentially cloned into the KpnI/BamHI and HindIII restriction sites of plasmid pLoxP-NeoI29, yielding vector pAcrA-KO. Correct orientation of the HindIII fragment was verified by digest with SalI and XbaI and DNA sequencing. The insert containing the two AcrA KO fragments flanking the floxed neomycin resistance cassette (Supplementary figure 4A) was introduced into P. pallidum cells and gene knock-outs were identified by PCR and Southern blot (Supplementary figure 4B,C).
Creation of a double acra-acga- cell line
The A6neo cassette was removed from AcrA KO cells (acra-) as described above. A G418 sensitive acra- clone was transformed with linearized pAcgA-KO to delete AcgA. The AcgA gene disruption was confirmed by PCR and Southern blot (Supplementary figure 3C).
Complementation of acra-acga- with AcrA
To express AcrA from its own promoter, a fosmid, used for P. pallidum genome mapping10 and containing AcrA, was digested whith SpeI. This released an 8.65 kb fragment, which contained the entire AcrA 5′intergenic region and coding sequence. This fragment was cloned into vector pLox-NeoIII and introduced into acra-acga- cells.
References
Schilde, C. & Schaap, P. The Amoebozoa. Methods in molecular biology 983, 1–15, 10.1007/978-1-62703-302-2_1 (2013).
Alvarez-Curto, E. et al. cAMP production by adenylyl cyclase G induces prespore differentiation in Dictyostelium slugs. Development 134, 959–966 (2007).
Soderbom, F., Anjard, C., Iranfar, N., Fuller, D. & Loomis, W. F. An adenylyl cyclase that functions during late development of Dictyostelium. Development 126, 5463–5471 (1999).
Van Es, S. et al. Adenylyl cyclase G, an osmosensor controlling germination of Dictyostelium spores. J. Biol. Chem. 271, 23623–23625 (1996).
Shaulsky, G., Fuller, D. & Loomis, W. F. A cAMP-phosphodiesterase controls PKA-dependent differentiation. Development 125, 691–699 (1998).
Thomason, P. A. et al. An intersection of the cAMP/PKA and two-component signal transduction systems in Dictyostelium. EMBO J. 17, 2838–2845 (1998).
Loomis, W. F. Cell signaling during development of Dictyostelium. Developmental Biology 391, 1–16, 10.1016/j.ydbio.2014.04.001 (2014).
Schaap, P. et al. Molecular phylogeny and evolution of morphology in the social amoebas. Science 314, 661–663 (2006).
Eichinger, L. et al. The genome of the social amoeba Dictyostelium discoideum. Nature 435, 43–57 (2005).
Heidel, A. et al. Phylogeny-wide analysis of social amoeba genomes highlights ancient origins for complex intercellular communication. Genome Res., 1882–1891, 10.1101/gr.121137.111 (2011).
Clarke, M. et al. Genome of Acanthamoeba castellanii highlights extensive lateral gene transfer and early evolution of tyrosine kinase signaling. Genome Biol. 14, R11, 10.1186/gb-2013-14-2-r11 (2013).
Du, Q. & Schaap, P. The Social Amoeba Polysphondylium pallidum Loses Encystation and Sporulation, but Can Still Erect Fruiting Bodies in the Absence of Cellulose. Protist 165, 569–579, 10.1016/j.protis.2014.07.003 (2014).
Sucgang, R. et al. Comparative genomics of the social amoebae Dictyostelium discoideum and Dictyostelium purpureum. Genome Biol. 12, R20, 10.1186/gb-2011-12-2-r20 (2011).
Kawabe, Y., Weening, K. E., Marquay-Markiewicz, J. & Schaap, P. Evolution of self-organisation in Dictyostelia by adaptation of a non-selective phosphodiesterase and a matrix component for regulated cAMP degradation. Development 139, 1336–1345, 10.1242/dev.077099 (2012).
Toama, M. A. & Raper, K. B. Microcysts of the cellular slime mold Polysphondylium pallidum. I. Factors influencing microcyst formation. J Bacteriol 94, 1143–1149 (1967).
Virdy, K. J. et al. High cAMP in spores of Dictyostelium discoideum: association with spore dormancy and inhibition of germination. Microbiology 145, 1883–1890 (1999).
Saran, S. & Schaap, P. Adenylyl cyclase G is activated by an intramolecular osmosensor. Mol. Biol. Cell 15, 1479–1486 (2004).
Fritz-Laylin, L. K. et al. The genome of Naegleria gruberi illuminates early eukaryotic versatility. Cell 140, 631–642, 10.1016/j.cell.2010.01.032 (2010).
Du, Q. et al. The cyclic AMP phosphodiesterase RegA critically regulates encystation in social and pathogenic amoebas. Cellular Signalling 26, 453–459, 10.1016/j.cellsig.2013.10.008 (2014).
Schulkes, C. & Schaap, P. cAMP-dependent protein kinase activity is essential for preaggegative gene expression in Dictyostelium. FEBS Lett. 368, 381–384. (1995).
Maeda, M. et al. Periodic signaling controlled by an oscillatory circuit that includes protein kinases ERK2 and PKA. Science 304, 875–878 (2004).
Hopper, N. A., Harwood, A. J., Bouzid, S., Véron, M. & Williams, J. G. Activation of the prespore and spore cell pathway of Dictyostelium differentiation by cAMP-dependent protein kinase and evidence for its upstream regulation by ammonia. EMBO J. 12, 2459–2466 (1993).
Harwood, A. J. et al. Culmination in Dictyostelium is regulated by the cAMP-dependent protein kinase. Cell 69, 615–624 (1992).
Hopper, N. A., Anjard, C., Reymond, C. D. & Williams, J. G. Induction of terminal differentiation of Dictyostelium by cAMP- dependent protein kinase and opposing effects of intracellular and extracellular cAMP on stalk cell differentiation. Development 119, 147–154 (1993).
Mann, S. K. O., Richardson, D. L., Lee, S., Kimmel, A. R. & Firtel, R. A. Expression of cAMP-dependent protein kinase in prespore cells is sufficient to induce spore cell differentiation in Dictyostelium. Proc. Natl. Acad. Sci. USA 91, 10561–10565 (1994).
Kawabe, Y., Enomoto, T., Morio, T., Urushihara, H. & Tanaka, Y. LbrA, a protein predicted to have a role in vesicle trafficking, is necessary for normal morphogenesis in Polysphondylium pallidum. Gene 239, 75–79 (1999).
Faix, J., Kreppel, L., Shaulsky, G., Schleicher, M. & Kimmel, A. R. A rapid and efficient method to generate multiple gene disruptions in Dictyostelium discoideum using a single selectable marker and the Cre-loxP system. Nucleic Acids Res 32, e143 (2004).
Meima, M. E., Weening, K. E. & Schaap, P. Vectors for expression of proteins with single or combinatorial fluorescent protein and tandem affinity purification tags in Dictyostelium. Protein Expr Purif 53, 283–288 (2007).
Kawabe, Y. et al. Activated cAMP receptors switch encystation into sporulation. Proc Natl Acad Sci USA 106, 7089–7094, 10.1073/pnas.0901617106 (2009).
Sievers, F. & Higgins, D. G. Clustal omega, accurate alignment of very large numbers of sequences. Methods in molecular biology 1079, 105–116, 10.1007/978-1-62703-646-7_6 (2014).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003).
Schultz, J., Milpetz, F., Bork, P. & Ponting, C. P. SMART, a simple modular architecture research tool: identification of signaling domains. Proc. Natl. Acad. Sci. USA 95, 5857–5864 (1998).
Romeralo, M. et al. Analysis of phenotypic evolution in Dictyostelia highlights developmental plasticity as a likely consequence of colonial multicellularity. Proc Biol Sci 280, 20130976, 10.1098/rspb.2013.0976 (2013).
Acknowledgements
This work was funded by grants 090276 and 100293Z/12/Z from the Wellcome Trust and grant BB/K000799/1 from the BBSRC.
Author information
Authors and Affiliations
Contributions
Y.K. prepared the acga-, acra-acga and acra-acga-/AcrA mutants, C.S. the acra- mutant and Q.D. the pkac- and pkac-/PkaC mutants. All three examined the phenotypes of their mutants and wrote sections of the manuscript. P.S. designed the study and finalized the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Kawabe, Y., Schilde, C., Du, Q. et al. A Conserved Signalling Pathway for Amoebozoan Encystation that was Co-Opted for Multicellular Development. Sci Rep 5, 9644 (2015). https://doi.org/10.1038/srep09644
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep09644
This article is cited by
-
Emerging roles for diguanylate cyclase during the evolution of soma in dictyostelia
BMC Ecology and Evolution (2023)
-
Adenylate cyclase A amplification and functional diversification during Polyspondylium pallidum development
EvoDevo (2022)
-
Glycogen synthase kinase 3 promotes multicellular development over unicellular encystation in encysting Dictyostelia
EvoDevo (2018)
-
The transcription factor Spores Absent A is a PKA dependent inducer of Dictyostelium sporulation
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.