Abstract
Pluripotent cells can be subdivided into two distinct states, the naïve and the primed state, the latter being further advanced on the path of differentiation. There are substantial differences in the regulation of pluripotency between human and mouse and in humans only stem cells that resemble the primed state in mouse are readily available. Reprogramming of human stem cells into a more naïve-like state is an important research focus. Here, we developed a pipeline to reanalyze transcriptomics data sets that describe both states, naïve and primed pluripotency, in human and mouse. The pipeline consists of identifying regulated start-ups/shut-downs in terms of molecular interactions, followed by functional annotation of the genes involved and aggregation of results across conditions, yielding sets of mechanisms that are consistently regulated in transitions towards similar states of pluripotency. Our results suggest that one published protocol for naïve human cells gave rise to human cells that indeed share putative mechanisms with the prototypical naïve mouse pluripotent cells, such as DNA damage response and histone acetylation. However, cellular response and differentiation-related mechanisms are similar between the naïve human state and the primed mouse state, so the naïve human state did not fully reflect the naïve mouse state.
Similar content being viewed by others
Introduction
Embryonic stem cells (ESCs) were first isolated from mouse embryos some thirty years ago1,2. Since then, they have been characterized in many other mammalian species, most notably in human3. ESCs are pluripotent, i.e. they can self-renew and give rise to all cell types of the adult body, which makes them attractive for therapeutic research. Stem cell research has gained additional momentum with the possibility of reprogramming somatic cells into induced pluripotent stem cells (iPSCs)4, enabling the generation of patient specific pluripotent stem cells. As reviewed by De Los Angeles et al. [2012]5, two distinct states of pluripotency are usually distinguished. Mouse ESCs (mESCs) are isolated from the inner cell mass (ICM) of mouse preimplantation embryos in vitro and resemble ICM cells at embryonic day (E) 4.5. These cells have specific properties, for which they have been termed naïve pluripotent stem cells. Most notably, they are able to generate chimeric animals with high efficiency when introduced into blastocysts5. Maintenance of mESCs depends on Lif/Jak/Stat3 and Bmp4/Smad1/5/8. mESCs are also insensitive to single cell dissociation and display dome shape colony morphology. Furthermore, female mESCs have both of their X chromosomes in an active state (XaXa). On the other hand, mouse epiblast stem cells (mEpiSCs) are isolated from early post-implantation embryos (E5.5). These cells are rarely, if at all, able to generate chimeras6 and have one of their X chromosomes inactivated in female lines (XaXi). Further, mEpiSCs are bFGF and ActivinA-Nodal/Smad2/3 dependent. In contrast to mESCs they are sensitive to single cell dissociation and display flat colony morphology. Because they represent a more developed state, they have been termed primed pluripotent stem cells.
Human embryonic stem cells (hESCs) have much more in common with mEpiSCs than with mESCs, although mESCs and hESCs are derived from the inner cells mass of pre-implantation embryos. This includes growth factor requirements and X chromosome activation state. Therefore, mEpiSCs and hESCs are thought to represent the primed state as opposed to the naïve mESCs5,7. Efforts have been devoted to convert primed into naïve pluripotent stem cells. In mouse, this has been achieved for both permissive and non-permissive strains by Hanna et al. 20098. In the latter case, however, a more complex experimental protocol was required, yielding metastable (i.e. transgene-dependent) naïve pluripotent stem cells. In the case of human ESC and iPS cells, conversion into the naïve state has been attempted9,10. Although the cells that were cultured according to the protocol by ref. 9 show some features in common with naïve mESCs, such as dome-shaped colonies, XaXa status, Lif dependence and Activin independence, these cells disintegrate after 15–20 passages, which is an impairment of the self-renewal aspect of the pluripotent state in vitro. We were interested in the molecular characterization of these “naïve-like human pluripotent stem cells” in order to elucidate which barriers are preventing their full conversion to the naïve state. We analyzed the transcriptomics data of pluripotent stem cells from different species and states, such as human versus mouse, and, naïve versus primed. From a comparative analysis of high-throughput data, for example of gene expression (transcriptomics) assays, we expected a deeper understanding of similarities and differences of stem cell regulation. Usually, differentially expressed genes (DEGs) are identified for further analysis and it is assumed that gene expression is a proxy for the expression of the protein products. Due to noise, such data often feature false positive as well as false negative findings, however. Functional interaction networks allow noise reduction by allowing consideration of interactions between the genes. Specifically, such network knowledge has successfully been employed in classification tasks11. There are many different ways in which genes (and their protein products) can interact, like dimer/multimer formation, phosphorylation or transcriptional activation and repression. These types of interactions are reflected to different extents in transcriptomics data; this has to be taken into account when interpreting such data in the context of functional networks. Noise may also be reduced by mapping the genes to functional categories such as Gene Ontology (GO,12) terms or Kyoto Encyclopedia of Genes and Genomes (KEGG,13) pathways. This abstraction is successful if general regulatory mechanisms governing stem cell identity are conserved between different species although the precise genes that take part in those mechanisms are not.
Many functional interaction networks have been published so far and they vary greatly in scope and size. Small scale expert networks with hundreds of genes and interactions, all implicated in pluripotency, have been presented by Müller et al. [2008]14 (employing machine learning on a large scale network augmented with additional literature knowledge), Som et al. [2010]15 (with a focus on careful manual literature curation), Xu et al. [2013]16 (integrating large scale experimental evidence, complemented by manual curation) and most recently by Dunn et al. [2014]17 (reconstruction of a Boolean network of transcription factors based on co-regulation). Large all-purpose functional interaction networks of tens of thousands of genes, e.g. STRING18 lack the quality that can be provided by manual curation. Yet, they may be more suitable for the analysis of high-throughput data. As expert networks invariably cover only a small amount of the genes that are interrogated in a high-throughput study, large networks are more amenable to the application of GO/KEGG abstraction, because, featuring many more genes, they are less biased with respect to specific terms. The STRING network contains both direct (physical) and indirect (functional) interactions between genes.
We recently published ExprEssence [Warsow et al. 2010]19 to analyse differential data in the context of networks. ExprEssence computes a LinkScore for every link (interaction) in a functional network and every pair of conditions described by a high-throughput data set. The LinkScore reflects the amount of concerted change that two linked genes or proteins experience during the transition between two conditions. It takes directionality into account, if such information is available, considering the expected impact of the change of the amount of a stimulator or inhibitor on the genes it regulates. The links with the lowest and highest LinkScores (the tails of the distribution) are likely to represent the major mechanistic changes that occur during the transition.
In this paper, we used ExprEssence and an all-purpose functional interaction network to study the transitions between the naïve and the primed state of pluripotency in ES and iPS cells from human and mouse. We obtained the transcriptomics data sets accompanying Hanna et al. [2009]8 and Hanna et al. [2010]9. For all pairings of species and pluripotency states, we filtered the major mechanistic changes based on the links in the tails of the respective LinkScore distribution and investigated their GO term and KEGG pathway enrichment. Finally, we aggregated data of similar transitions. Thus, our workflow employs links in a network, GO/KEGG enrichment and transition relatedness as three layers of abstraction, which collaboratively reduce the noise that is inherent to heterogeneous data. Our results suggest that a published protocol for generation of naïve human pluripotent stem cells [Hanna et al. 2010]9 gave rise to human cells that, indeed, share important mechanisms with the prototypical naïve pluripotent cells from mouse, while their primed counterparts also show biologically meaningful similarities between human and mouse. However, we also found mechanisms in the state claimed to be naïve human, which show similarities to the primed mouse state. We also investigated a more recent protocol for naïve human pluripotent stem cells by Gafni et al. [2013]10. Again, we conclude that although these cells display many similarities to the naïve mouse cells, they also display differentiation mechanisms shared with primed mouse cells. These very similarities to the primed state may be amendable to experimental intervention, in order to obtain naïve cells as in mouse.
Results
Analyzing cross-condition and cross-species pluripotency-related gene expression
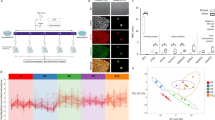
We set out to describe the transition between the naïve and the primed state of pluripotent stem cells in two mammalian species, human and mouse. The comparison is based on biological mechanisms that we define by the joint analysis of transcriptomics and network data. A roadmap to our approach is presented in Figure 1; see Methods for details.
Roadmap of the work presented in this paper.
We start by integrating several input data sets, namely one large-scale network of direct (physical) and indirect (functional) interactions derived from STRING and featuring mostly expert-curated data and two high-throughput transcriptomics studies that describe the transition between naïve and primed pluripotency in human and mouse. On this multidimensional data set, we sequentially apply computational methods (grey elliptical boxes), obtaining data in a more and more abstract form, from genes to mechanisms to gene ontology terms. Each step is intended to remove noise from the data. The result of each step is shown in a blue box. We employed several analysis and visualization methods to obtain insights into the data at each step (beige boxes).
First, we obtained two previously published transcriptomics data sets8,9. Hanna et al. [2009]8 analyzed the reprogramming of primed mouse pluripotent stem cells to the naïve pluripotent state. For this purpose, they generated microarray data describing gene expression profiles of naïve pluripotent and primed pluripotent stem cells from the NOD strain, which is generally non-permissive for the naive state and the permissive strain 129 (GSE15603). Hanna et al. [2010]9 studied the reprogramming of human pluripotent stem cells to naïve pluripotency (GSE21222). Both data sets were downloaded and processed separately as described in Methods.
Expression profiles from human and mouse stem cells were combined into one data set (see Methods) and we will refer to this and the analysis based thereon as the Hanna/Hanna data and the Hanna/Hanna analysis. Our preprocessing protocol resulted in a combined data set comprising 25 samples and approximately 11,000 genes. Samples belong to one of four conditions: naïve human (NH, n = 6), primed human (PH, n = 5), naïve mouse (NM, n = 6) and primed mouse (PM, n = 8). Hierarchical clustering of this data set separates the samples according to their pluripotency state (Figure 2A), in agreement with ref. 9. A principal component analysis (PCA) supports the conclusion drawn from the clustering (Figure 2B). However, the second principal component (PC) indicates that there may be some similarity between NH and PM, on which we will follow up upon later.
Analysis of the combined gene expression data set of Hanna et al (2009, 2010).
The Hanna/Hanna data set consists of 25 microarray samples that correspond to the four conditions naïve human (NH), primed human (PH), naïve mouse (NM) and primed mouse (PM). For the sample preprocessing and construction of the gene expression matrix, see text. A: Hierarchical clustering dendrogram of the samples (Spearman's rank correlation, complete linkage). The colour bar below the dendrogram provides information about the species (upper bar) and the pluripotency state (lower bar) of the single samples. The samples are labelled with their GEO identifiers; see data sets GSE21222 and GSE15603 for further details. B: Principal component analysis (PCA) of the data set. Four distinct clusters are identifiable, which correspond to the four conditions. The species of origin of the samples is indicated by the colour of the single symbols, while the colour of the ellipses enclosing the four clusters indicates the pluripotency state of the samples inside.
Stepwise aggregation of gene expression data
To enhance signal and remove noise in the combined data set, we processed and subsequently aggregated the data, in multiple ways as described in Figure 1.
1. Mapping of gene expression data to a functional network
The first kind of noise reduction was accomplished by the joint analysis of genes known to be related to each other. Gene expression profiles from the combined data set were thus mapped onto a functional network as described in Methods. Focusing on specificity rather than sensitivity, our network is based on experimentally validated interactions and genes that have at least one annotation in the biological process division of GO (GOBP, see Methods). The resulting network contains 5220 genes connected by 17171 interactions.
Next, for each link (interaction) in the network and each possible pair of “source” and “target” conditions, we computed the ExprEssence LinkScore (see Methods), which describes the magnitude of concerted change for the two linked genes, occurring in the transition from source to target condition. A positive LinkScore suggests simultaneous upregulation and it suggests that the corresponding interaction is more pronounced in the target condition than in the source condition. Conversely, an interaction with a negative LinkScore suggests simultaneous downregulation. Notably, false positive and false negative findings are possible, e.g. because an interaction may be context-dependent. Pairs of concerted upregulated or downregulated genes could also be identified from single-gene analyses of gene expression data or be constructed by machine learning as features. However, such approaches lack the biological focus specificity given by an underlying functional network, highlighting only pairs of co-regulated genes that are a priori known to interact, resulting in what we call highlighted mechanistic changes.
For each possible pair of source and target conditions (Figure 3), we selected the links with LinkScores at or above the 99.75 percentile, yielding twelve link sets. We then extracted from each link set the genes that are connected by those links, yielding twelve gene sets, hypothesizing that these genes are involved in the major mechanisms characterizing the transition to the respective target condition.
All pairwise comparisons of conditions and aggregates thereof.
A comparison has a source and a target condition. For example, for comparison #1, NH is the source and NM is the target condition. Aggregates of comparisons are defined as follows. (a) by target condition (NH, NM, PH, PM) and (b) block-wise. The latter are subdivided into the natural blocks naïve (N, green) and primed (P, red) and the mixed ones NHPM (blue) and NMPH (cyan), see also Supplementary Table S1. Target aggregates are indicated by arrows below the columns; all comparisons within a column constitute a single target aggregate. Block aggregates are indicated by enclosing coloured circles and rounded rectangles. All comparisons that are enclosed within a shape of a given colour are part of the aggregate that is associated with that colour; a comparison may be part of multiple aggregates. For example, N (green) aggregates comparisons with target naïve (regardless of species, but excluding naïve as the source), NH aggregates all comparisons with target NH and NHPM allows checking the hypothesis that NH and PM have properties in common. LinkScore calculations for a comparison estimate which links between genes start up during the transition from source to target. For each comparison, the top-scoring links form its link set, which in turn gives rise to a corresponding gene set.
2. Mapping of genes involved in high-scoring network links to GOBP terms
In order to facilitate interpretation of the GOBP enrichment analysis presented next, we classified the twelve comparisons into several aggregates, namely target aggregates and block aggregates (see Figure 3). The latter were further subdivided into natural and mixed block aggregates. This allowed us to identify mechanisms that are consistently enriched in transitions leading to the same conditions (target aggregates) or the same pluripotency states such as naïve or primed (block aggregates), see Figure 3. The same aggregates will be later used for a third and final noise reduction step.
In the second noise reduction step, the gene sets that we defined for the comparisons in the first step were subjected to a functional analysis based on their GOBP annotation (see Methods). We constructed a heatmap showing enriched and depleted GOBP terms for each of the twelve gene sets corresponding to the twelve comparisons (Figure 4). In contrast to the primed block aggregate, the naïve one features a larger number of depleted GOBP terms. This is also the case for the comparison #1 between NH (source) and NM (target), which thus captures “naïvity” in mouse. This enrichment pattern supports a model in which there is an upregulation of a multitude of pathways in the primed (but not naive) pluripotent stem cells. Next, we subjected the GOBP matrix (Figure 4) to PCA, using the GOBP terms as variable names (Figure 5). The first two PCs explain ~52% and ~14%, respectively, of the total variance. The first PC clearly separates the natural block aggregates primed and naïve, making the distinction between naïve and primed pluripotency the most outstanding feature of our data set.
Heatmap of evidence for associations between comparisons and GOBP terms.
Twelve gene sets were derived from the comparisons between source and target conditions, see Figure 3 and Supplementary Table S1, by selecting, using a STRING functional network, the genes involved in the links with the highest LinkScore. These gene sets were functionally annotated based on GO. For each gene set the -log p-values of the GO biological process (GOBP) term enrichment or depletion are given. Rows are scaled. In the annotation bar on top of the matrix, the colouring identifies the membership of the twelve gene sets in the naïve (green) or primed (red) block aggregate.
Principal component analysis (PCA) of evidence for associations between comparisons and GOBP terms.
PCA was performed on the matrix shown in Figure 4, taking the comparisons as samples and the GOBP terms as variable names. The comparisons are numbered, with the numbers matching those assigned in Figure 3 and Supplementary Table S1. Colours are used to reflect sample membership in the block aggregates naïve and primed, as defined in Figure 3, with grey indicating samples that are in neither aggregate.
3. Aggregation of gene sets of comparisons that describe similar transitions
We performed a third noise reduction and data aggregation step and combined the GOBP enrichment data. More specifically, we summarized the evidence for enrichment of GOBP terms for each target condition (see Methods). The resulting matrix of aggregated evidence was again subjected to a number of analyses. First, we generated a heatmap of the matrix (Figure 6). Clustering of the target aggregates (columns) results in two clusters, one of them containing the target aggregates that describe the naïve pluripotent state in human and mouse, while the other one comprises the corresponding aggregates for the primed state. This representation of the data therefore strengthens the case for a similarity between the prototypical naïve mouse and the naïve human states, as proposed by ref. 9. This clustering is restricted to the GOBP terms that were found to be specific for any of the block aggregates, as detailed below.
Heatmap of evidence for enrichment of GOBP terms in target aggregates.
Aggregation of the twelve comparisons into target aggregates was performed as described in the text. The heatmap includes only GOBP terms for which a significant enrichment in one of the block aggregates was determined using our statistical assessment. This aggregate is colour-coded to the left of the heatmap (green: naïve, red: primed, blue: NHPM).
Applying a PCA to the matrix of aggregated evidence highlights the relationships between the target aggregates as well as the relative importance (loadings) of the single GOBP terms for the target aggregates (Figure 7). Here, the first PC clearly separates the target aggregates PM and PH (which have positive scores) from the target aggregates NM and NH (negative scores), with both groups having a lower intragroup than intergroup distance. Again, we conclude that there are functional similarities between the naïve states in mouse (represented by the target aggregate NM) and human (NH). Moreover, since the first PC accounts for 74% of the variance, these similarities are substantial. However, Figure 4 indicated that the naïve block aggregate is characterized by depletion of GOBP terms that are enriched elsewhere; yet the aggregation procedure that underlies the PCA considers enriched GOBP terms only. We investigated this issue further using a statistical approach, determining how many and which GOBP terms and, hence, biological processes are in fact common between NM and NH (see Figure 8 below). While the first PC in Figure 7 seems to capture functional and species-independent characteristics of naïve and primed pluripotency, the second PC, explaining roughly 17% of total variance, has a different interpretation. Here, the target aggregates NH and PM have strikingly similar scores, hinting at some biological process similarity between the claimed naïve state in human and the primed state in mouse. This finding prompted us to define the (additional) block aggregate NHPM (see Figure 3). In the next section we will use this aggregate to decipher the GOBP terms that might underlie this finding and that calls for improvements of protocols for human naïvity.
Principal component analysis (PCA) of evidence for enrichment of GOBP terms in target aggregates.
Aggregation of the twelve comparisons into target aggregates was performed as described in the text; the matrix of log-transformed p-values (Figure 6) was then analysed by PCA. In this biplot, two scatterplots are overlaid: one for the scores of the four target aggregates, represented by their respective abbreviations (see Figure 3, denoted NH, PH, NM and PM, for the four pairs of a species and a state) and one for the relative importance (or loadings) of the GOBP terms, plotted as grey or coloured dot symbols. The symbol colour indicates for which, if any, block aggregate (as defined in Figure 3, denoted N, P, NHPM and NMPH) the term was found to be significant (see text). The axes below and to the left of the plot belong to the scores plot, the ones on top and to the right to the loadings plot.
Distribution of the Fisher statistic of GOBP terms.
Selection of GOBP terms was based on specificity; only terms that were found to be specific for one of the block aggregates are included. Fisher statistics of association p-values were computed to evaluate the enrichment of GOBP terms in block aggregates. Background distributions of Fisher statistics (i.e. its distribution for all possible 4-tupels of comparisons, see text for details) are shown in a series of boxplots. Overlaid on each boxplot are the Fisher statistics that were computed for the four block aggregates. The terms are grouped based on the block aggregate for which they were found to be specifically significant. Panels A and B contain terms that are significant for the naïve and primed block aggregate, respectively. Terms in panel C, in turn, support the mixed block aggregate NHPM. In each panel the terms are ordered by the median of the background distribution.
4. Exploration of single GOBP terms and calculation of statistical significance
Finally, we explored which single GOBP terms are of highest importance for each block aggregate, in terms of being specifically enriched in the link sets that characterize each block aggregate. In Figure 7, the GOBP terms located close to a given target aggregate feature low p-values (i.e. are highly significant) for this target aggregate. Closest GOBP terms are thus considered specific for 'their' target aggregate. Moreover, terms that are situated between two target aggregates, but more distant to the other two target aggregates, may be specific for a block aggregate. For example, a term between the aggregates NM and NH is likely to support the block aggregate naïve. Calculating statistical significance of enrichment of any such GOBP term in any given block aggregate is possible by applying Fisher's method. However, since the comparisons and the link sets and gene sets derived from them share some information (e.g. the target condition, possibly single genes etc.), single p-values are not only lacking the correction for multiple testing, but they also are not obtained in a strictly independent manner and the overall p-values should be interpreted conservatively. To account for this, for a given GOBP term we tested the Fisher scores obtained for the block aggregates against the background distribution for this GOBP term across all comparisons (see Methods). GOBP terms considered significantly enriched in a block aggregate and thus specific for it, were annotated by color in Figure 7, indicating the respective block aggregate. In Figure 8, their Fisher scores are displayed, including a boxplot for the background distribution, demonstrating that most of them are specific for one single block aggregate. Specifically, for most of the selected GOBP terms, the runner-up block aggregate has a score that lies within the box, i.e. below the 75% percentile, so the selected GOBP terms are indeed specific for their respective block aggregate.
Investigation of GOBP terms specific for the natural block aggregates
The GOBP terms that were found to be specific for any of the block aggregates by our statistical approach can be classified into two broad categories. The first one comprises biological processes that are specific for any one of the two natural block aggregates, namely naïve and primed. Such terms are found in panels A and B of Figure 8. Further, they are colored in the PCA plot of Figure 7. (The second category concerns the mixed aggregates, see discussion.)
Panel A comprises terms that are enriched in the naïve block aggregate. These terms represent processes that are overrepresented in the naïve versus the primed state. Notably, many of these GOBP terms refer to epigenetics, such as histone acetylation, which is a chromatin modification that generally correlates with active transcription. Interestingly, early embryonic cells are characterized by a hyperdynamic chromatin architecture, suggesting that heterochromatic and largely inaccessible domains are mostly formed upon differentiation. Higher activity of histone acetylation pathways suggests that the euchromatic state may be more prominent in naïve cells. This is consistent with several studies that highlight the need for open chromatin structures for successful reprogramming. For example, knockdown of Mbd3, part of the NuRD repressor complex with histone deacetylase activity, enhanced reprogramming efficiency to near 100%20. Further, culturing of naïve mouse mESCs in the presence of Lif and two inhibitors, Glycogen-synthase kinase β inhibitor and Mek inhibitor, (called 2i), or Lif and serum, causes a loss of repressive modifications21.
Further examples for terms that are commonly enriched towards naïve pluripotency are signal transduction in response to DNA damage and related terms. Maintaining genomic integrity is of critical importance for any stem cell, especially for ESCs, since DNA damage compromises the daughter cells22. Indeed, DNA repair mechanisms are very active in mouse as well as in human ESCs23,24 and reduced DNA repair responses contribute to stem cell ageing25. Notably, naïve pluripotent stem cells are situated very early in the course of development, rendering genomic integrity of utmost importance. The aspect of genomic integrity may also explain why terms related to metabolism, e.g. primary metabolic process, are characteristic of naïve pluripotency. The metabolism of primed ESCs relies on anaerobic glycolysis rather than oxidative phosphorylation (OXPHOS), presumably, amongst other reasons, because OXPHOS is associated with increased formation of reactive oxygen species (ROS), which can lead to increased genomic damage26,27,28. In the naïve ground state of pluripotency, glycolysis is further activated21. Progressive oxidation of metabolites is required and essential for differentiation; indeed, inhibition of differentiation can be achieved by inhibiting oxidization29. Other primary metabolic processes like lipid and carbohydrate metabolism are enhanced in the ground naïve state21.
Panel B in Figure 8, on the other hand, comprises terms that are enriched in the natural block aggregate primed. In analogy to the aforementioned terms, these terms represent processes that are enriched in primed cells but are depleted and/or suppressed in human and mouse cells upon induction of the naïve state. Many of these terms are related to developmental and differentiation processes. These include general terms like cell morphogenesis and regulation of cell development, as well as more specific ones like neuron differentiation. Such findings are consistent with, e.g., ref 30, who showed that differentiating ESCs are committed to a neural fate in the absence of factors that enforce alternative differentiation pathways31,32,33.
Investigation of GOBP terms specific for the naive-human/primed-mouse (NHPM) block aggregate
While panel A and B of Figure 8 likely represent commonalities between naïve and primed pluripotency, respectively, panel C comprises terms that are shared between naïve human and primed mouse cells. Such terms are responsible for the striking similarity between NH and PM in terms of the second PC, as shown in Figure 7 and these terms are discussed below.
We finally analyzed the single genes whose presence in the selected gene sets gave rise to the identification of the enriched GOBP terms. Supplementary Table S2 provides, for each given GOBP term that was left over after filtering (see Methods) and for each gene set, the list of genes within the gene set that were annotated with that GOBP term. Genes that occur in several (or even all) of the gene sets that make up one of the block aggregates are specifically important. For example, the gene PBX1 was found in all four gene sets contributing to the block aggregate primed. Indeed, PBX1 has been described as a transcriptional regulator of the neural marker gene SOX334 supporting that neural differentiation is a default differentiation pathway. (SOX3 was not part of our combined data set.)
Investigation of specific KEGG pathways
In order to compare our findings to results based on a pathway-centric compendium of functional gene annotations, we repeated our analysis pipeline using the KEGG pathway database, instead of GOBP. Figure 9 summarizes the results, on the same analysis stage as Figure 6, featuring a heatmap of KEGG pathway enrichment evidence in target aggregates. The clustering of the target aggregates confirms the purported closeness of human and mouse naïvity. This clustering considers only significant KEGG pathways, i.e. pathways that were found to be specifically enriched in any of the four block aggregates, with a p-value threshold of 0.05. Yet, a number of KEGG terms resonate well with our previous findings. Base excision repair was identified as specific for the naïve block aggregate, corroborating the GOBP term signal transduction in response to DNA damage found above. Furthermore, glycolysis and valine, leucine and isoleucine biosynthesis were among the naïve-specific terms. The former is consistent with ref. 21. With respect to the latter, it was shown that the metabolomes of hESCs and human embryonal carcinoma cells (hECCs), which are the malignant counterpart to the hESCs, share common signatures comprising amongst others also amino acid metabolism28. Further, mESCs cultured in 2i + Lif in comparison to Serum + Lif upregulate amino acid related metabolism21. However, the precise role of different amino acid pathways in pluripotency remains to be determined in detail. As for pathways that are shared between the target aggregates PH and PM, we found axon guidance, supporting our conclusion about the importance of neuron differentiation for the primed aggregate. Finally, the NHPM block aggregate features various signalling pathways involved in differentiation such as those related to TGF-beta, NOD-like and Jak/STAT, consistent with the GOBP analysis. In interpreting the results of the KEGG analysis, however, it should be noted that fewer genes could be annotated with KEGG pathways, compared to those with GOBP annotations.
Heatmap of evidence for enrichment of KEGG pathways in target aggregates.
The heatmap includes only KEGG pathways for which a significant enrichment in one of the block aggregates was determined using our statistical assessment. This aggregate is colour-coded to the left of the heatmap (green: naïve, red: primed, blue: NHPM, cyan: NMPH).
Application of the pipeline on another data set on naïve human pluripotency
Most recently, Hanna and co-workers published a follow-up paper10 on the topic of naïve human cells. Compared to their 2010 results, they used a refined protocol to establish human cells that more closely resemble the naïve mouse state of pluripotency.
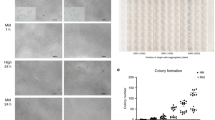
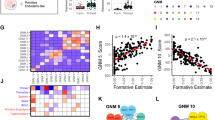
We therefore compiled a combined data set from the human gene expression data that accompanied Gafni et al. [2013]10 and the mouse gene expression data from Hanna et al. [2009]8; we will refer to this and the analysis based thereon as the Hanna/Gafni data and the Hanna/Gafni analysis. In order to ensure comparability of the analyses, we restricted the Hanna/Gafni data set to the genes that were also part of the Hanna/Hanna data set. First, we performed hierarchical clustering and PCA, both visualized in panels A and B of Figure 10; Figure 2 features the equivalent analyses for the Hanna/Hanna data. Hierarchical clustering identified two major clusters in the combined data set, which correspond to the two states of pluripotency, i.e. naïve and primed, again supporting the similarity of NH and NM cells. The PCA, however, is less clear in this regard. While the first PC clearly distinguishes the PM samples from the NM samples, the human samples form a cloud around the origin of the coordinate system. This observation suggests that important gene expression changes that distinguish naïve from primed pluripotent samples in mouse are not recapitulated in human. To gain further insights into this matter, we ran our aggregation-based analysis pipeline on the combined data set. As is evident from the column-clustering in the heatmap of GOBP enrichment shown in Figure 11 (representing the same analysis stage as Figure 4; the subsequent PCA analysis is shown in Supplementary Figure S3), three of the four single comparisons making up the block aggregate naïve, cluster closely together. Indeed, their functional signatures appear to be more consistent as compared to Figure 4. Interestingly, within the list of GOBP terms that are consistently enriched in these three comparisons we find terms like stem cell maintenance, telomere maintenance, histone modification and cell cycle process, all of them known to be important for stem cell identity. Regarding the latter, pluripotent stem cells are characterized by an accelerated cell cycle, which is slowed down upon differentiation35. The mechanisms invoked in naïve human are thus similar to the naïve mouse state and they are related to stemness. However, the fourth member of the naïve block aggregate, which corresponds to the within-species comparison of PH as source and NH as target (comparison 8), has a strikingly different biological process signature, as evident by the clustering. Since this comparison is part of the target aggregate NH, this might explain the considerable spatial distance between NH and the target aggregate NM in the PCA plot shown in Figure 12 (representing the same analysis stage as Figure 7; the underlying heatmap, taking only the significant (i.e. colored) terms of Figure 12, is shown in Supplementary Figure S4), which further points to considerable differences between NH and NM in the Hanna/Gafni analysis. On the other hand, NH and PM are located quite close to each other in the PCA plot, suggesting similarity between these, just as was the case in the Hanna/Hanna analysis (Figure 8). In line with this, our statistical assessment method again identified numerous GOBP terms that are significant for the block aggregate NHPM (Figure 13, panel C; note that panel C is truncated for the sake of readability, Supplementary Figure S2 provides an untruncated version). These terms are discussed below. Then again, there are also a number of GOBP terms that are specific for the block aggregates naïve and primed (panels A and B, respectively, of Figure 13). Among the terms for the naïve block aggregate are two terms directly related to cell cycle control, the importance of which for the pluripotent state35 was already pointed out; the respective terms are G1/S transition of mitotic cell cycle, regulation of cell cycle and associated metabolic processes. Conversely, terms related to differentiation were found for the block aggregate primed. Prominent among them were terms related to formation of ectoderm structures, such as axonogenesis and eye morphogenesis. Ectodermal differentiation-related processes were also identified as specific for the primed aggregate in the Hanna/Hanna analysis. This indicates that depletion of terms related to ectoderm differentiation is triggered by various protocols for induction of naïve pluripotency in a cross-species manner. Furthermore, within this GOBP term group are also terms that relate to mesoderm formation, like skeletal system development and kidney development, which were not observed in our Hanna/Hanna analysis (Figure 8). We take this as indication for a broader activity of differentiation processes in the primed state and, consequently, repression of these processes in the naïve state as a result of the protocol of Gafni et al. [2013]10. This is supported further by general differentiation related terms such as extracellular matrix organization and biological adhesion that are also found among the terms characterizing the block aggregate primed in the Hanna/Gafni data.
Analysis of the combined gene expression data set of Hanna et al. [2009]8 and Gafni et al. [2013]10.
The Hanna/Gafni data set consists of 26 microarray samples that correspond to four conditions: NH, PH, NM and PM. For the sample preprocessing and construction of the gene expression matrix, see text. A: Hierarchical clustering dendrogram of the samples (Spearman's rank correlation, complete linkage). The colour bar below the dendrogram provides information about the species (upper bar) and pluripotency state (lower bar) of the single samples. The samples are labelled with their GEO identifiers; see data sets GSE46872 and GSE15603 for details. B: Principal component analysis (PCA) of the data set. The colour of the symbols indicates the pluripotency state of the respective samples, while labels next to the symbols indicate the species.
Heatmap of evidence for association of comparisons and GOBP terms, for the Hanna/Gafni data set.
See Figure 4 for further details.
Principal component analysis (PCA) of evidence for enrichment of GOBP terms in target aggregates, for the Hanna/Gafni data set.
See Figure 7 for further details.
Distribution of the Fisher statistic of GOBP terms, for the Hanna/Gafni data set.
See Figure 8 for further details. Panel C features GOBP terms that are specific for the block aggregate NHPM or NMPH and it is truncated with respect to the number of terms; Supplementary Figure S2 is the untruncated version.
Discussion
When analyzing transcriptomics data, as done here, one must always be aware that the amount of mRNA in a cell is not necessarily predictive of the amount of the corresponding protein, due to posttranscriptional regulative mechanisms36,37. Hence, the success of cell programming efforts has to be further confirmed experimentally by cell morphology, growth factor requirements, functional assays like contribution to chimeras and, as a new tool, cell surface proteomics38. Moreover, the functional network and the gene ontology annotations we used contain false positive and false negative assertions, that is, they entail both incorrect and missing information. Finally, our aggregation of data for similar transitions treats species differences and developmental state differences on equal footing, calling for a scheme that may assign different weight to species differences and state differences.
Notably, species similarities and developmental state similarities are considered together in a single pipeline, designed to highlight similarities in terms of GO terms and pathways to the degree that these are robustly identifiable, i.e. after several steps of noise filtering. If the molecular mechanisms would not be ‘similar enough’ across species or states, our pipeline could not identify significant findings, many of which having high biological plausibility. Furthermore, we note that we investigate a transition from one state of pluripotency to another and several groups (ours included) have found that the genes and pathways involved in these are conserved at least to some degree, see e.g. ref. 39 and references cited therein. Along the same lines, we mixed the comparisons between conditions together into aggregates, with the purpose to detect similarity between these if there is one. Thus, from the very outset, we were open to find any robust similarity that is there. Block and target aggregates together enable to find similarity in the two naive conditions, in the two primed conditions and also between in the naive human and the primed mouse condition. These three cases are indeed present; our approach was designed to find each of these similarities if they exist. These similarities are not mutually exclusive: If mechanisms are shared between conditions A & B, other mechanisms may be shared between conditions A & C, etc., even though the conditions themselves are distinct.
Our perhaps most surprising finding is the similarity between the conditions naive human and primed mouse, captured by the Gene Ontology (GOBP) terms for the block aggregate NHPM consisting of these conditions. We interpret these terms as representative of processes that were either wrongly induced by the treatment of the human cells or, alternatively, that failed to be repressed by it. In both cases, they drive the human cells away from the prototypical state of mouse naïvity rather than bringing them closer. Inspecting Figure 8, we note that among the terms with that role are (cellular) response to (endogenous) stimulus, cell proliferation, tube morphogenesis, extracellular matrix organization, blood vessel morphogenesis and osteoblast differentiation. Human and mouse ESCs and iPSCs are cultured in media of different composition, explaining the enrichment in cellular response to stimulus and to organic substance in particular. The other biological processes are all related to differentiation pathways. Unlike neurons, which are of ectodermal origin, blood vessels and bone derive from mesoderm, however. Regarding pluripotency, one theory put forward recently proposes that in pluripotent stem cells a multitude of differentiation pathways leading to the formation of different germ layers are active simultaneously, the effectors of which compete with each other. Only if one pathway dominates, differentiation will occur accordingly. One example is SMAD2/3 (triggered by  ) and SMAD1/5/8 (triggered by BMP4) competing for SMAD440,41. Thus, some of these multiple differentiation pathways may indeed be activated in the transition of naïve to primed pluripotency in the mouse and similarly in the unstable transition of primed to naïve pluripotency in human. In the Hanna/Gafni analysis the NHPM block aggregate is again dominated by cellular response pathways such as response to organic stimulus and response to stress, as well as differentiation pathways, such as tissue development, ossification and tissue remodelling. Ossification points to mesodermal processes as in the Hanna/Hanna analysis.
) and SMAD1/5/8 (triggered by BMP4) competing for SMAD440,41. Thus, some of these multiple differentiation pathways may indeed be activated in the transition of naïve to primed pluripotency in the mouse and similarly in the unstable transition of primed to naïve pluripotency in human. In the Hanna/Gafni analysis the NHPM block aggregate is again dominated by cellular response pathways such as response to organic stimulus and response to stress, as well as differentiation pathways, such as tissue development, ossification and tissue remodelling. Ossification points to mesodermal processes as in the Hanna/Hanna analysis.
In summary, we designed a workflow that allows the (re-)analysis of noisy gene expression data, employing several layers of abstraction. We re-analyzed data sets from naïve mouse and primed as well as naïve and primed human pluripotent stem cells and characterized the various cell types by enrichment analyses. We found that cells claimed to be naïve human display an overlapping gene expression signature with the naïve mouse cells, explaining their naïve properties. However, we also found similarities to the primed mouse state. This raises the question of what caused the incomplete induction of the naïve state in hESCs. Closely related are the biological processes causing hESCs to transit from the naïve into the primed state during their isolation, as they were naïve in the inner cell mass in the first place. As summarized in Figure 14, our study suggests that there are residual biological processes typically found in primed mouse pluripotency that hinder complete induction of true human naïvity. These processes include response to endogenous stimulus and differentiation-related biological processes, which may also be at work in the defaulting of hESCs into the primed state. Inhibiting these may enable us to come even closer to the naïve human pluripotent state.
Methods
Sample selection and preprocessing: human
Human microarray data for primed and naïve stem cells of both embryonic and iPS origin were obtained from Gene Expression Omnibus (GEO)42, accession number GSE212229, performed on the experimental platform Hgu133plus2 from Affymetrix. After a preliminary analysis of the whole GSE set (see Supplementary Figure S1), the subset with GSM numbers 530613 to 530618 was taken to represent the naïve state, while GSM numbers 530608 to 530612 formed the group of primed samples. On the reduced set of samples we applied the mas5calls() function (part of the affy package of BioConductor) to obtain a list of probesets that were flagged as absent in one or more samples. The data set was then processed using the MAS5 algorithm, which encompasses background correction, normalization and summarization of the single probes of each probeset. This was followed by log2-transformation and filtering-out of all probes that did not have at least 5 present flags across all samples. Using the annotation library hgu133plus2.db, the probesets were mapped to their respective EntrezGene identifiers and probesets annotated to the same EntrezGene identifier were aggregated by their median value.
In order to reanalyze the data from Gafni et al.10 we obtained from GEO the gene expression data set GSE46872, which was run on the Affymetrix-platform HuGene1.0ST. Out of the twelve samples contained therein, all were used in this study and assigned to the naïve and primed group according to the sample metadata. Data set processing was performed with the RMA algorithm43. Using the annotation library hugene10sttranscriptcluster.db, the probesets were mapped to their respective EntrezGene identifier and probesets annotated to the same EntrezGene identifier were aggregated by their mean value. Processing the human dataset GSE21222 using RMA resulted in minimal to non-existent differences, see Supplementary Figure S5.
Sample selection and preprocessing: mouse
Mouse microarray data for primed and naïve stem cells of both embryonic and iPS origin were obtained from GEO, accession number GSE156038. The samples within the data set were run on the Agilent Whole Mouse Genome Microarray 4x44K platform. No method for assessing the rate of present/absent calls per individual probeset, comparable to the mas5calls() method, was available in this case; thus we downloaded the data in already preprocessed and normalized form via an interface provided by the R package GEOquery44. Preprocessing and normalization of the data was performed by the original authors using the Limma package. From this data set we removed two samples, namely GSM390184 and GSM390186, because they were cultured under challenging conditions, which carried the possibility of confounding our analysis. Agilent spot identifiers were mapped to mouse EntrezGene identifiers; in a second step the mouse EntrezGene identifiers were mapped to human EntrezGene identifiers using homology information from HomoloGene45. See Supplementary Text S1 for a discussion of some problems in identifier mapping. Probesets annotated to the same human EntrezGene identifier were aggregated by their median value. We used this mouse data set to compare against the human data from Hanna et al.9 as well as Gafni et al.10, giving rise to what we call the Hanna/Hanna and Hanna/Gafni data and analyses, respectively. Aggregating multiple probesets by mean or median resulted in results that were practically not distinguishable, see Supplementary Figure S6.
Data intersection
The processing of the human and mouse expression data sets both yielded an expression matrix with the samples in the columns and the genes, represented by human EntrezGene identifiers, in the rows. To enable comparisons across the matrices, in each matrix we centered each gene on its mean across the matrix, thus removing information on and possible bias in, the magnitudes of gene expression. Hanna et al.9 performed a similar processing step. Subsequently, the intersection of the genes within the human and the mouse data set was formed and a joint expression matrix containing the centered gene expression values, restricted to the intersection, was constructed. Finally, for each gene its mean and variance across the four conditions, i.e. NH, PH, NM and PM (see Figure 3), were computed.
STRING network
A comprehensive network for human was obtained from STRING 9.018. We restricted the network to links (interactions) that featured confidence scores for experimental validation equal to or greater than 600. We mapped the computed expression means and variances for each gene onto the network as node attributes. On subsets of genes derived from this data set, we applied functional analyses based on GO12, employing the biological process (GOBP) subdivision and KEGG13. Genes not annotated within this subdivision did not contribute to, nor confound, our analyses; we filtered the network retaining only genes with GOBP annotations. For the Hanna/Hanna analysis based on human data by Hanna et al.9 we thus obtained a network containing 5220 genes and 17171 interactions. The network that we obtained in the course of analyzing the Hanna/Gafni data was restricted to the genes that were also used in the Hanna/Hanna analysis; this move was intended to facilitate comparability with the latter. This resulted in a network containing 5185 genes and 17020 interactions. Network analysis was performed using ExprEssence19, which runs as a plugin in the popular network analysis tool Cytoscape v2.846, as described next.
Pairwise comparison of conditions and highlighting of mechanistic changes
The four conditions (NH, PH, NM and PM) gave rise to twelve comparisons of conditions as described in Figure 3. Any comparison is directed from source to target condition. Let A and B be two genes, with expression measurements, that are connected by a link in the network. Let μA;source, σ2A;source, nA;source and μA;target, σ2A;target, nA;target be A's mean, variance and sample size under the source and target condition, respectively. Let the values for B be defined likewise. The LinkScore L is then computed as the sum of two T statistics, one for each gene, as follows19.

A gene that is highly expressed in the target condition, but less so in the source condition will thus contribute a high positive value to the LinkScore of any given interaction that this gene is part of. An interaction with a positive overall LinkScore then experiences a “start-up” towards the target condition, according to terminology established19. For each comparison (Figure 3) we employed LinkScore calculations to identify the interactions with the highest LinkScores, i.e. the most pronounced start-ups and we defined one link set and one gene set as follows. We set a cutoff value at the 99.75 percentile of the corresponding LinkScore distribution and selected all links (referred to as link sets) with LinkScores that were equal to or greater than the cutoff, yielding a link set. The genes that are connected by the interactions within this link set were then extracted, yielding the corresponding gene set.
Functional annotation based on GOBP and KEGG
Gene sets (corresponding to the comparisons) were subjected to functional analysis, employing the R package GOstats47 to test whether terms from the biological process division of GO (GOBP,12) were over- or underrepresented, respectively. Every GOBP term was tested separately on each gene set. If a gene annotated by a GOBP term is found in a gene list, it is called a hit. GOstats uses the hypergeometric distribution to assess for a given GOBP term the significance of the deviation, between the number of actual hits and the expected number of hits, given the frequency of the term's annotation to the background gene set (i.e. the gene universe); this significance is then reported as a p-value. The gene universe was defined as the set of genes that have measurements in both species, are present in the network and are annotated with at least one GOBP term. The single p-values were log-transformed and assembled into a matrix of dimension n × m, with n being the number of GOBP terms and m being the number of gene sets (that is, twelve). Because of the way they were computed, these p-values are one-tailed, one p-value corresponding to the statement “term is overrepresented” and the other to the statement “term is underrepresented”. For each pair of GOBP term and gene set, both p-values were calculated and the appropriate one, depending on whether the number of hits was greater or less than the number expected by chance, was entered into the matrix. To reflect this, the p-values were direction-signed, where a “-” denotes underrepresentation. Multiple filtering steps were then applied to the resulting matrix. First, only GOBP terms with a significant p-value in at least two of the twelve gene sets were considered further. In addition, only GOBP terms with a minimum of five hits in at least one of the twelve gene sets were retained.
In a similar fashion we studied associations between gene sets and KEGG pathways, for which GOstats also provides an interface. The protocol resembled the one outlined above, with the exception that we did not apply the filter on the minimum number of hits; doing so would have been too restrictive with respect to the numbers of retained KEGG pathways.
Aggregation of comparisons
The twelve comparisons were aggregated based on their target condition (i.e. the name of the corresponding column in Figure 3, see also Supplementary Table S1), into one of four eponymous target aggregates, with each target aggregate consisting of 3 comparisons. Moreover, the comparisons that share a given pluripotency state (i.e. either naïve or primed) in their target condition, irrespective of species, but with the additional requirement that this state is not part of their source condition, were merged into natural block aggregates. Finally, two mixed block aggregates were created, each merging a given pluripotency state from one species with the opposite one from the other, as described in Figure 3. Following their definition, each block aggregate consists of 4 comparisons. For each pair of aggregate and GOBP term, we thus considered a k-tuple of log-transformed association p-values, with k = 3 for target aggregates and k = 4 for block aggregates. To obtain a measure of how well the k single p-values collectively support the hypothesis of enrichment, the k-tuples for a given GOBP term were aggregated into a single p-value using the Fisher method. Let p be a vector of p-values of length k. Then, the aggregation statistic F is computed as

and the corresponding p-value for each pair of a given GOBP term and condition were computed based on the chi-square distribution with k*2 degrees of freedom, log-transformed (base 10) and then assembled into an n × m matrix, where n is the number of GOBP terms and m is the number of aggregates; for both target and block aggregates, m = 4. The significance of each GOBP term with respect to the block aggregates as defined in Figure 3 was assessed in a non-parametrical way. For each GOBP term, the distribution of the Fisher statistic was computed for all possible 495 4-tuples of the twelve comparisons. A GOBP term was called enriched for a given block aggregate if less than 1% of all possible statistics equaled or exceeded the value of the statistic for this aggregate.
References
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
Martin, G. R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc Natl Acad Sci U S A 78, 7634–7638 (1981).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676, 10.1016/j.cell.2006.07.024 (2006).
De Los Angeles, A., Loh, Y.-H., Tesar, P. J. & Daley, G. Q. Accessing naïve human pluripotency. Curr Opin Genet Dev, 10.1016/j.gde.2012.03.001 (2012).
Huang, Y., Osorno, R., Tsakiridis, A. & Wilson, V. In Vivo differentiation potential of epiblast stem cells revealed by chimeric embryo formation. Cell Rep 2, 1571–1578, 10.1016/j.celrep.2012.10.022 (2012).
Zhang, B., Krawetz, R. & Rancourt, D. E. Would the real human embryonic stem cell please stand up? Bioessays 35, 632–638, 10.1002/bies.201200162 (2013).
Hanna, J. et al. Metastable pluripotent states in NOD-mouse-derived ESCs. Cell Stem Cell 4, 513–524, 10.1016/j.stem.2009.04.015 (2009).
Hanna, J. et al. Human embryonic stem cells with biological and epigenetic characteristics similar to those of mouse ESCs. Proc Natl Acad Sci U S A 107, 9222–9227, 10.1073/pnas.1004584107 (2010).
Gafni, O. et al. Derivation of novel human ground state naive pluripotent stem cells. Nature, 10.1038/nature12745 (2013).
Taylor, I. W. et al. Dynamic modularity in protein interaction networks predicts breast cancer outcome. Nat Biotechnol 27, 199–204, 10.1038/nbt.1522 (2009).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25, 25–29 (2000).
Ogata, H. et al. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res 27, 29–34 (1999).
Müller, F.-J. et al. Regulatory networks define phenotypic classes of human stem cell lines. Nature 455, 401–405, 10.1038/nature07213 (2008).
Som, A. et al. The PluriNetWork: an electronic representation of the network underlying pluripotency in mouse and its applications. PLoS One 5, e15165, 10.1371/journal.pone.0015165 (2010).
Xu, H. et al. ESCAPE: database for integrating high-content published data collected from human and mouse embryonic stem cells. Database (Oxford) 2013, bat045, 10.1093/database (2013).
Dunn, S.-J., Martello, G., Yordanov, B., Emmott, S. & Smith, A. G. Defining an essential transcription factor program for naïve pluripotency. Science 344, 1156–1160, 10.1126/science.1248882 (2014).
Szklarczyk, D. et al. The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 39, D561-D568 10.1093/nar (2011).
Warsow, G. et al. ExprEssence-revealing the essence of differential experimental data in the context of an interaction/regulation net-work. BMC Syst Biol 4, 164, 10.1186/1752-0509-4-164 (2010).
Rais, Y. et al. Deterministic direct reprogramming of somatic cells to pluripotency. Nature, 10.1038/nature12587 (2013).
Marks, H. et al. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 149, 590–604, 10.1016/j.cell.2012.03.026 (2012).
Saretzki, G. et al. Downregulation of multiple stress defense mechanisms during differentiation of human embryonic stem cells. Stem Cells 26, 455–464, 10.1634/stemcells.2007–0628 (2008).
Nagaria, P., Robert, C. & Rassool, F. DNA double strand break response in stem cells: Mechanisms to maintain genomic integrity. Biochim Biophys Acta, 10.1016/j.bbagen.2012.09.001 (2012).
Rocha, C. R. R., Lerner, L. K., Okamoto, O. K., Marchetto, M. C. & Menck, C. F. M. The role of DNA repair in the pluripotency and differentiation of human stem cells. Mutat Res 752, 25–35, 10.1016/j.mrrev.2012.09.001 (2013).
Fan, G.-C. Role of heat shock proteins in stem cell behavior. Prog Mol Biol Transl Sci 111, 305–322, 10.1016/b978-0-12-398459-3.00014-9 (2012).
Cho, Y. M. et al. Dynamic changes in mitochondrial biogenesis and antioxidant enzymes during the spontaneous differentiation of human embryonic stem cells. Biochem Biophys Res Commun 348, 1472–1478, 10.1016/j.bbrc.2006.08.020 (2006).
Chung, S. et al. Mitochondrial oxidative metabolism is required for the cardiac differentiation of stem cells. Nat Clin Pract Cardiovasc Med 4 Suppl 1 S60-S67 10.1038/ncpcardio0766 (2007).
Abu Dawud, R., Schreiber, K., Schomburg, D. & Adjaye, J. Human embryonic stem cells and embryonal carcinoma cells have overlapping and distinct metabolic signatures. PLoS One 7, e39896, 10.1371/journal.pone.0039896 (2012).
Yanes, O. et al. Metabolic oxidation regulates embryonic stem cell differentiation. Nat Chem Biol 6, 411–417, 10.1038/nchembio.364 (2010).
Ying, Q.-L., Stavridis, M., Griffiths, D., Li, M. & Smith, A. Conversion of embryonic stem cells into neuroectodermal precursors in adherent monoculture. Nat Biotechnol 21, 183–186, 10.1038/nbt780 (2003).
Masaki, H., Nishida, T., Kitajima, S., Asahina, K. & Teraoka, H. Developmental pluripotency-associated 4 (DPPA4) localized in active chromatin inhibits mouse embryonic stem cell differentiation into a primitive ectoderm lineage. J Biol Chem 282, 33034–33042, 10.1074/jbc.M703245200 (2007).
Abranches, E. et al. Neural differentiation of embryonic stem cells in vitro: a road map to neurogenesis in the embryo. PLoS One 4, e6286, 10.1371/journal.pone.0006286 (2009).
Aiba, K. et al. Defining developmental potency and cell lineage trajectories by expression profiling of differentiating mouse embryonic stem cells. DNA Res 16, 73–80, 10.1093/dnares (2009).
Mojsin, M. & Stevanovic, M. PBX1 and MEIS1 up-regulate SOX3 gene expression by direct interaction with a consensus binding site within the basal promoter region. Biochem J 425, 107–116, 10.1042/bj20090694 (2010).
Hindley, C. & Philpott, A. The cell cycle and pluripotency. Biochem J 451, 135–143, 10.1042/bj20121627 (2013).
Schwanhauesser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342, 10.1038/nature10098 (2011).
Li, J. J., Bickel, P. J. & Biggin, M. D. System wide analyses have underestimated protein abundances and the importance of transcription in mammals. Peer J 2, e270, 10.7717/peerj.270 (2014).
Rugg-Gunn, P. J. et al. Cell-surface proteomics identifies lineage-specific markers of embryo-derived stem cells. Dev Cell 22, 887–901, 10.1016/j.devcel.2012.01.005 (2012).
Som, A., Lustrek, M., Singh, N. K. & Fuellen, G. Derivation of an interaction/regulation network describing pluripotency in human. Gene 502, 99–107, 10.1016/j.gene.2012.04.025 (2012).
Loh, K. M. & Lim, B. A precarious balance: pluripotency factors as lineage specifiers. Cell Stem Cell 8, 363–369, 10.1016/j.stem.2011.03.013 (2011).
Montserrat, N. et al. Reprogramming of Human Fibroblasts to Pluripotency with Lineage Specifiers. Cell Stem Cell, 10.1016/j.stem.2013.06.019 (2013).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets-update. Nucleic Acids Res 41, D991-D995 10.1093/nar (2013).
Irizarry, R. A. et al. Exploration, normalization and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264, 10.1093/biostatistics (2003).
Davis, S. & Meltzer, P. S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics 23, 1846–1847, 10.1093/bioinformatics/btm254 (2007).
NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 42, D7-17 10.1093/nar/gkt1146 (2014).
Smoot, M. E., Ono, K., Ruscheinski, J., Wang, P.-L. & Ideker, T. Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27, 431–432, 10.1093/bioinformatics (2011).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258, 10.1093/bioinformatics (2007).
Acknowledgements
We thank Kenjiro Adachi (MPI Münster) for valuable suggestions regarding sample selection and advice on interpreting the data. This work was supported in part by the German Research Foundation (DFG, Priority Program Pluripotency and Cellular Reprogramming, FUE 583/2-2) and the German Federal Ministry of Education and Research (BMBF, Validierung des Innovationspotenzials wissenschaftlicher Forschung - VIP, 03 V0396).
Author information
Authors and Affiliations
Contributions
G.F. and M.E. conceived the study, conducted the analyses and interpreted the results. G.F., M.E. and R.A.D. wrote the main manuscript text. A.K., G.S. and L.T. critically revised the manuscript and contributed further interpretation of results and discussion. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplement
Supplementary Information
Suppl Table 2
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Ernst, M., Dawud, R., Kurtz, A. et al. Comparative computational analysis of pluripotency in human and mouse stem cells. Sci Rep 5, 7927 (2015). https://doi.org/10.1038/srep07927
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07927
This article is cited by
-
Bioinformatics Approaches to Stem Cell Research
Current Pharmacology Reports (2018)
-
Direct comparison of distinct naive pluripotent states in human embryonic stem cells
Nature Communications (2017)
-
Dissecting microRNA-mediated regulation of stemness, reprogramming, and pluripotency
Cell Regeneration (2016)
-
Ageing with elegans: a research proposal to map healthspan pathways
Biogerontology (2016)
-
Techniques of Human Embryonic Stem Cell and Induced Pluripotent Stem Cell Derivation
Archivum Immunologiae et Therapiae Experimentalis (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.