Abstract
Ferredoxins are iron-sulfur proteins that play important roles in electron transport and redox homeostasis. Yeast Apd1p is a novel member of the family of thioredoxin-like ferredoxins. In this study, we characterized the hydroxyurea (HU)-hypersensitive phenotype of apd1Δ cells. HU is an inhibitor of DNA synthesis, a cellular stressor and an anticancer agent. Although the loss of APD1 did not influence cell proliferation or cell cycle progression, it resulted in HU sensitivity. This sensitivity was reverted in the presence of antioxidant N-acetyl-cysteine, implicating a role for intracellular redox. Mutation of the iron-binding motifs in Apd1p abrogated its ability to rescue HU sensitivity in apd1Δ cells. The iron-binding activity of Apd1p was verified by a color assay. By mass spectrometry two irons were found to be incorporated into one Apd1p protein molecule. Surprisingly, ribonucleotide reductase genes were not induced in apd1Δ cells and the HU sensitivity was unaffected when dNTP production was boosted. A suppressor screen was performed and the expression of stress-regulated transcription factor Yap1p was found to effectively rescue the HU sensitivity in apd1Δ cells. Taken together, our work identified Apd1p as a new ferredoxin which serves critical roles in cellular defense against HU.
Similar content being viewed by others
Introduction
Yeast APD1 gene is generally uncharacterized for function. It was implicated in cellular resistance to sodium ions, hydrogen peroxide, elesclomol, dithiothreitol, diamide and cumol hydroperoxide in several genome-wide studies1,2,3. In another global protein localization study, Apd1p was found in both the cytoplasm and the nucleus4. The interaction between Tsa1p and Apd1p was initially suggested in a large-scale mass spectrometric analysis of protein complexes in yeast5. Tsa1p is a principal peroxiredoxin and a strong suppressor of genome instability6,7,8. Peroxiredoxins are peroxidases that function as redox-sensitive molecular switches and tumor suppressors9,10,11. TSA1-null cells have elevated levels of dNTPs and are sensitive to hydroxyurea (HU), a dNTP depleting agent8. HU inhibits the activity of ribonucleotide reductase (RNR) by quenching the tyrosyl radical at its iron-containing active site12. RNR catalyzes the synthesis of dNTPs, the building blocks of DNA13. In addition, HU also perturbs DNA repair and elevates intracellular cGMP levels12. HU can activate several arms of cellular stress response and is commonly used as an anti-cancer agent14. As an antioxidant protein, Tsa1p requires efficient electron transfer relay to support and regenerate its functional capacity. Thioredoxins and glutaredoxins are relevant reductants to regenerate peroxiredoxins9,15,16,17.
Apd1p belongs to a family of thioredoxin-like ferredoxins, which has more than 100 members found in organisms ranging from bacteria to metazoan18. Ferredoxins are iron-sulfur (Fe-S) proteins that play important roles in biology19. The thioredoxin fold is also commonly found in oxidoreductases and chaperones20. Although the function of most thioredoxin-like ferredoxins remains unknown, the presence of thioredoxin fold and ferredoxin domain in these proteins is suggestive of a role in redox sensing and electron transfer. Like thioredoxins, glutaredoxins and peroxiredoxins, thioredoxin-like ferredoxins may function as thiol-based molecular switches by modulating disulfide bond formation in their target proteins16,21. In prokaryotes, thioredoxin-like ferredoxin YfaE has been suggested to be involved in diferric-tyrosyl radical maintenance in E. coli RNR22. More recently, yeast thioredoxin-like ferredoxins Grx3/4p and Dre2p have been characterized for their roles in supporting in vivo diferric tyrosyl radical formation in RNR23. That is to say, at least some thioredoxin-like ferredoxins might play a role in redox regulation of RNR activity. It will therefore be of interest to see whether Apd1p might also serve as a redox sensor and electron carrier for RNR and/or Tsa1p.
Yap1p transcription factor and its target ATR1 are critically involved in cellular response to oxidative stress and other cytotoxic agents24,25,26,27,28,29,30. Yap1p is an AP-1-like bZIP protein that controls a large and specialized stress regulon31,32. Atr1p is a multidrug resistance transporter with multiple transmembrane segments33. Both Yap1p and Atr1p are high-copy-number suppressors of cellular sensitivity to stressors such as metals, boron, reactive oxygen species, DNA damage and metabolic inhibitors24,25,26,27,28,29,30. Particularly, Yap1p is thought to be influential in cellular response to HU34. However, the exact roles of Yap1p and Atr1p in HU sensitivity are not known.
Characterizing Apd1p might shed light on the biological function of a family of thioredoxin-like ferredoxins. In this study we set out to perform phenotypic characterization of apd1Δ yeast cells and found that they are hypersensitive to HU. The iron binding capability of Apd1p was validated and shown to be important to HU resistance. The HU sensitivity in apd1Δ cells was suppressed by YAP1 and ATR1. Our findings revealed a protective role of Apd1p in cellular defense against HU.
Results
APD1 is not essential for cell growth or cell cycle progression
Genome-wide analyses revealed that loss of APD1 in budding yeast results in cellular sensitivity to hydrogen peroxide and other stressors1,2,3. However, the exact biological function of APD1 remains to be characterized. Thus, we constructed apd1Δ cells to study how the loss of APD1 might affect cell growth and proliferation.
We first examined the cell growth pattern of apd1Δ cells. The proliferation of apd1Δ cells was not different from the WT BY4741 cells (Fig. 1A). Loss of APD1 did not affect the lag phase to log phase transition or the subsequent exponential growth, suggesting that APD1 is non-essential to cellular growth. When we analyzed cell cycle profile by flow cytometric measurement of the DNA content of cells in asynchronized log-phase culture, both WT and apd1Δ cells displayed a similar DNA profile (Fig. 1B). This indicated that APD1 is not influential on cell cycle progression. Next, we further examined the growth characteristics of WT and apd1Δ cells in different strain backgrounds. As shown in Fig. 1C, their growth rates and patterns were comparable in different strain backgrounds (i.e. W303-1a and BY4741) and different carbon sources (i.e. glucose, acetate, ethanol and glycerol). Thus, Apd1p has minimal influence on cell growth and cell cycle progression.
Growth and cell cycle profile of apd1Δ cells.
(A) Growth curve. Logarithmically growing yeast cells, BY4741 wild-type (WT) and apd1Δ, were inoculated in YPD medium. Cell density was determined by OD600. (B) Cell cycle profiling. Asynchronized cells of BY4741 WT and apd1Δ were grown to mid-log phase in YPD. Cells were fixed, treated with RNase A and stained with propidium iodide (PI). DNA content of cells was reflected in PI intensity determined by flow cytometry. (C) Spot test in different carbon sources. Ten-fold serial dilutions of strains W303-1a WT, W303-1a apd1Δ, BY4741 WT and BY4741 apd1Δ were spotted on YP medium containing glucose, acetate, ethanol or glycerol at 2%. Plates were incubated for 3–4 days at 30°C.
Deletion of APD1 sensitizes cells to hydroxyurea
In a global protein interactome study, Apd1p was fished out by Tsa1p, a principal peroxiredoxin, suggesting a possible physical interaction and functional intersection5. From our previous work, we know that loss of TSA1 results in elevation of dNTP levels and hypersensitivity to HU, a dNTP depleting agent8. Tsa1p is also required for the maintenance of genome stability6,7,8,35. Although we were unable to detect physical or genetic interaction between Apd1p and Tsa1p (data not shown), we found that loss of APD1 also confers HU sensitivity.
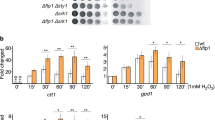
We performed spot assays on agar plates containing HU and H2O2 using APD1-proficient and -deficient cells, together with TSA1-null cells as positive controls. Intriguingly, deletion of APD1 sensitized cells to HU challenge (Fig. 2A, row 2 compared to 3). This sensitivity can be fully complemented by the re-expression of APD1 gene (Fig. 2B, row 3 compared to 1 and 2), indicating that this phenotype was due specifically to the loss of APD1. As a positive control, tsa1Δ cells were also sensitive to HU as previously described8. To our surprise but generally consistent with the lack of interaction between APD1 and TSA1, apd1Δ cells was not sensitive to H2O2 (Fig. 2A, row 2 compared to 3), while tsa1Δ cells became largely inviable in the presence of H2O2 (Fig. 2A, row 2 compared to 1). Thus, APD1 and TSA1 might play distinct roles in antioxidant defense.
Loss of APD1 sensitizes cells to HU.
(A-B) HU sensitivity in apd1Δ cells. Ten-fold serial dilutions of strains BY4741 WT, BY4741 apd1Δ and BY4741 tsa1Δ (A) or BY4741 WT pRS415, BY4741 apd1Δ pRS415 and BY4741 apd1Δ pAPD1 (B) were spotted on YPD or SC-L medium containing the indicated dose of HU or H2O2. Plates were incubated for 3–4 days at 30°C. (C-D) Influence of antioxidant on HU sensitivity of haploid and diploid apd1Δ cells. Ten-fold serial dilutions of strains BY4741 WT, BY4741 apd1Δ, BY4742 WT, BY4742 apd1Δ, BY4743 WT and BY4743 apd1Δ (C) or BY4741 WT pRS415, BY4741 WT pAPD1, BY4741 apd1Δ pRS415 and BY4741 apd1Δ pAPD1 (D) were spotted on YPD or SC-L medium containing the indicated dose and combination of HU and NAC. Plates were incubated for 3–4 days at 30°C. (E) Cells were treated as in D except that cells in the (NAC ~ HU) group were treated with NAC for 12 hours and washed extensively before spotting on HU-containing plate.
As a thioredoxin-like ferredoxin18, Apd1p is thought to be involved in redox regulation. Thus, we next asked whether the HU sensitivity of apd1Δ cells might be attributed to redox. To address this, spot assay was performed in the presence or absence of N-acetyl-cysteine (NAC), a potent antioxidant. The antioxidant property of NAC was first verified. When NAC was added to H2O2-treated BY4741 cells, the intracellular level of reactive oxygen species, as measured by fluorescence microscopy with the fluorescent probe 2′,7′-dichlorofluorescein diacetate (DCF), was reduced (Supplementary Fig. 1, panel 5 compared to 4). In contrast, treatment with HU did not significantly affect intracellular redox (Supplementary Fig. 1, panel 6 compared to 4). Both haploid (BY4741 and BY4742) and diploid (BY4743) apd1Δ strains were constructed and tested (Fig. 2C, row 2 compared to 1, row 4 compared to 3 and row 6 compared to 5). Sensitivity of apd1Δ cells with APD1 re-expression was also examined in the same setting (Fig. 2D, row 4 compared to 3). The HU sensitivity in apd1Δ cells was largely rescued by the addition of NAC (Fig. 2D, row 3 compared to 1). To rule out the possibility that NAC directly neutralizes HU in the culture medium, BY4741 cells were treated with NAC first and washed extensively before spotting them on HU-containing agar. Similar rescuing effect was observed in these cells treated sequentially with NAC and HU (Fig. 2E, row 3 compared to 1). These results suggested that APD1 serves as a physiological suppressor of HU sensitivity plausibly by regulating intracellular redox.
Iron-binding motifs are critical to HU resistance
Apd1p is a thioredoxin-like ferredoxin containing a CX3C motif and a C-terminal thioredoxin-like domain, which harbors a HX3H motif (Fig. 3A). These are iron-binding motifs resembling those found in well-characterized Fe-S clusters19.
Iron-binding motifs in Apd1p are crucial to HU resistance.
(A) Primary structure of Apd1p protein. The amino acid sequence of Apd1p was retrieved from Saccharomyces Genome Database (SGD) with the iron-binding motifs underlined and key amino acid residues highlighted in grey. (B) Influence of iron-binding motifs of Apd1p on HU sensitivity of haploid apd1Δ cells. Ten-fold serial dilutions of strains WT pRS415, WT pAPD1_WT, apd1Δ pRS415, apd1Δ pAPD1_WT, apd1Δ pAPD1_CX3C and apd1Δ pAPD1_HX3H were spotted on SC-L medium containing the indicated dose of HU. Plates were incubated for 3–4 days at 30°C. Strains in the background of haploid BY4741 were used. (C) Semi-quantitative RT-PCR analysis of APD1 transcript. Logarithmically growing cells, as indicated in A, in SC-L were harvested. Total RNA was extracted and used for cDNA synthesis. PCR was performed to assess the levels of APD1 and ANB1 transcripts. The housekeeping gene ANB1 was used as an internal loading control of duplex PCR. As a negative control (-ve), no reverse transcriptase was added to the reaction in lane 1. (D) Western blot analysis of Apd1-Mycp proteins. Western blotting was performed with mouse anti-Myc (Roche) and mouse anti-Pgk1p (Invitrogen) antibodies.
Given that the iron-binding motifs of Apd1p might be important for function, we asked whether disrupting these motifs could affect the ability of Apd1p to rescue HU sensitivity in APD1-null cells. Through site-directed mutagenesis, we constructed mutants in which critical residues of the two iron-binding motifs CX3C and HX3H were altered. Plasmids expressing WT and mutant APD1 were transformed into apd1Δ cells and spot assay was performed (Fig. 3B). Re-expression of APD1 was verified at mRNA level by RT-PCR (Fig. 3C, lanes 5–7 compared to 4) and at protein level by Western blotting (Fig. 3D, lanes 4–6 compared to 3). Notably, the CX3C mutant of Apd1p could minimally rescue the HU sensitivity (Fig. 3B, row 5 compared to 4 and 3) and the HX3H mutant was totally incapable of complementing the HU phenotype (Fig. 3B, row 6 compared to 4 and 3). Taken together, our data suggest that the two iron-binding motifs in Apd1p are indispensable for cellular defense against HU challenge.
Apd1p is an iron-binding protein
From genetic assays, we observed that disruption of iron-binding motifs in APD1 perturbed its protective function against HU (Fig. 3). To formally address whether Apd1p is an iron-binding protein, we performed a series of biochemical experiments. We expressed His-Apd1p in E. coli and purified it to high purity as judged by SDS-PAGE (Fig. 4A). The apparent size of Apd1p was about 36 kDa which was close its predicted size (Fig. 4A). The faint band at 72 kDa might represent the dimeric form of Apd1p (Fig. 4A). To verify the presence of Fe-S cluster experimentally, we expressed and purified WT and mutant Apd1p proteins. Strikingly, Apd1p WT displayed a reddish brown color on non-denaturing PAGE gel (Fig. 4B, lane 1), indicating that Apd1p might be an iron-binding protein. Through site-directed mutagenesis, we disrupted the iron-binding motifs in Apd1p (Fig. 3A) and generated Apd1p-CX3C and Apd1p-HX3H mutants. These two mutant proteins expressed efficiently in bacteria, but the reddish brown color was not seen in either mutant (Fig. 4B, lanes 2 and 3 compared to 1), suggesting that these motifs are critical for iron incorporation into Apd1p protein.
Apd1p is an iron-containing protein.
(A-B) Expression and purification of recombinant Apd1p. E. coli-produced recombinant His6-Apd1p was purified with Ni-NTA-Superflow resin. Proteins were eluted and subjected to 15% SDS-PAGE (for Apd1p WT) or 15% non-denaturing PAGE (for Apd1p WT, CX3C and HX3H) for size determination and color visualization, respectively. The SDS-PAGE gel was stained with Coomassie blue. The arrow points to the major Apd1p band. The asterisk denotes the dye front. The cropped non-denaturing gels are shown in the figure and the full-length gels are presented in Supplementary Figure 2. The stained and unstained gels were run under the same experimental conditions. (C) UV-visible spectra of recombinant Apd1p proteins. Purified proteins (3 mg each) were subjected to photometric scanning at the indicated wavelength range. (D) Determination of iron concentration by ICP-MS. The concentration of Apd1p was determined by BCA micro-scale assay using BSA as a standard. To measure protein concentration of Apd1p (right panel), three calibration assays were performed with BSA to construct standard curves which represent the linear fits of experimental data. The concentration of Apd1p was then calculated to be 0.268 mM. To measure iron concentration (left panel), a set of iron standards ranging from 30 to 90 ppb was used to generate a calibration curve. Trace gallium at 2 ppb was the low calibration standard. The iron concentration in the Apd1p protein solution was obtained by performing a linear regression on the dataset and deriving an equation Y = A + BX, where A is the y intercept, B is the slope, Y is the normalized detection signal for iron and X is the iron concentration. The iron concentration in the original Apd1p solution was calculated to be 0.511 mM.
To corroborate that Apd1p is an iron-binding protein, the UV-visible absorption spectra of Apd1p WT and mutant proteins were obtained. Peaks at 450 nm and 570 nm were only seen in the spectrum of Apd1p WT, but not of either mutant (Fig. 4C). The absorption features were consistent with the presence of a [2Fe–2S] cluster. In further support of this, the relative amount of iron ions in Apd1p protein was determined by inductively-coupled plasma mass spectrometry (ICP-MS). The concentration of Apd1p protein was first measured by BCA micro-scale assay and iron ions were then analyzed by ICP-MS (Fig. 4D). Since 0.511 mM of iron was found in 0.268 mM of Apd1p protein solution, the Apd1p-to-Fe ratio is 1:1.9. Thus, Apd1p might function as a dimer and contain a [2Fe–2S] cluster.
HU sensitivity of apd1Δ cells is unlikely mediated through dNTP regulatory network
RNR is a major target of HU13. RNR generates DNA building blocks for replication and repair by catalyzing the conversion of NTP to dNTP through a free radical-based mechanism36,37. The active form of the RNR small subunit in budding yeast is a heterodimer of β and β′ subunits, encoded by the RNR2 and RNR4 genes, respectively38. β subunit houses a diferric tyrosyl radical cofactor (Fe2III-Tyr*) that is essential for initiation of nucleotide reduction in RNR large subunits. This initiation step controls the catalytic activity of the enzyme37. In a recent study of the in vivo formation of diferric tyrosyl radical in Rnr2p, it was demonstrated that an Fe-S protein, Dre2p, is required for optimal RNR activity and Tyr* generation23.
Since Apd1p is an Fe-S protein and loss of APD1 resulted in HU sensitivity, we asked whether Apd1p might also play a role in the electron-transfer of RNR in the context of dNTP production. To test this idea, we first determined whether apd1Δ cells might be sensitive to other nucleotide analogs 5-flourouracil (5-FU) and adenosine-2′-monophosphate (2-AMP), which serve as inhibitors of dNTP synthesis39. Different from the response to HU, apd1Δ cells were not sensitive to 5-FU and 2-AMP (Fig. 5A, row 2 compared to 1). To verify this result, we simultaneously deleted APD1 and either SML1 or RFX1. Sml1p is an RNR inhibitor40 and Rfx1p is an RNR transcriptional corepressor41. If the HU sensitivity in apd1Δ cells, as in the case of tsa1Δ cells8, is mediated through the dNTP regulatory network, it should be rescued when dNTP production is de-repressed in the absence of SML1 or RFX1. However, disruption of SML1 or RFX1 did not influence the HU sensitivity in apd1Δ cells (Fig. 5B, rows 5, 6, 11 and 12 compared to 2, 3, 8 and 9). Hence, the HU sensitivity in apd1Δ cells was unlikely due to dNTP dysregulation.
HU sensitivity of apd1Δ cells is not mediated through dNTP regulatory network.
(A) apd1Δ cells were not sensitive to nucleotide analogs. Ten-fold serial dilutions of strains BY4741 WT and BY4741 apd1Δ were spotted on YPD agar medium containing the indicated dose of 5FU or 2-AMP. Plates were incubated for 3–4 days at 30°C. (B) Deletion of RNR negative regulators in apd1Δ cells did not affect HU sensitivity. Ten-fold serial dilutions of strains BY4741 WT, BY4741 apd1Δ, BY4741 sml1Δ, BY4741 rfx1Δ, BY4741 apd1Δsml1Δ (clones I and II) and BY4741 apd1Δrfx1Δ (clones I and II) were spotted on YPD agar medium containing the indicated dose of HU. (C) Loss of APD1 sensitized cells to HU in both BY4741 and W303 genetic background. Ten-fold serial dilutions of strains W303 WT, W303 apd1Δ, BY4741 WT, BY4741 apd1Δ, BY4741 sod1Δ and BY4741 tsa1Δ were spotted on YPD agar medium, containing the indicated dose of HU. (D) Loss of APD1 did not affect the induction of RNRs in the presence of HU. W303 WT and W303 apd1Δ cells transformed with pRNR2/3-LacZ reporter were grown in SC-Ura liquid medium containing 50 mM of HU. Cells were collected when OD600 reached 0.5 and β-galactosidase (β-gal) assay was performed to evaluate RNR promoter activity. The pRNR2-LacZ activity recovered from control apd1Δ cells was arbitrarily set as 1. For quantitative RT–PCR analysis of RNR transcripts, logarithmically growing cells of the indicated strains in YPD were subjected to treatment with HU. Total RNA was extracted and 3 μg of RNA was used for cDNA synthesis. Quantitative PCR was performed to assess the levels of RNR transcripts. Relative RNA levels were normalized to the amount of ACT1 transcript. (E) Galactose-induced expression of Rnr1-3Myc-p. Western blotting was performed with mouse anti-Myc (Roche) and mouse anti-Pgk1p (Invitrogen) antibodies. (F) Overexpression of RNR1 in apd1Δ cells did not influence HU sensitivity. Ten-fold serial dilutions of the indicated strains carrying pGal-3Myc-RNR1 were spotted on SC-Ura medium containing either raffinose (−Gal) or galactose (+Gal) plus 200 mM of HU.
Besides, we also investigated whether RNR genes are induced in the absence of APD1. To this end, we examined the transcriptional induction of RNR2/3 in apd1Δ cells in the backgrounds of both W-303 and BY4741 strains. We first verified that loss of APD1 sensitized cells to HU (Fig. 5C, row 2 compared to 1 and row 4 compared to 3). In parallel, we also included positive control cells, like sod1Δ and tsa1Δ, which have been previously been shown8,42 to be sensitive to HU (Fig. 5C, rows 5–6 compared to 3). Next, transcriptional induction of RNR2 and RNR3 were analyzed by RNR2/3-LacZ reporter constructs43. Cells carrying respective reporter constructs were grown in liquid culture and treated with HU. Cells were not sensitive to HU at this low concentration. β-galactosidase assay was performed to measure transcriptional activity of RNR2 and RNR3 promoters. In line with results in Fig. 5B, loss of APD1 did not alter transcriptional induction of RNR genes (Fig. 5D, left panel). Consistent with this, induction of RNR2 transcripts was comparable between WT and apd1Δ cells in BY4741 background (Fig. 5D, right panel), indicating that loss of APD1 did not affect RNR transcription. Furthermore, we directly increased the production of dNTP by overexpressing a galactose-inducible RNR1 gene in apd1Δ cells. This construct has previously been shown to dramatically upregulate dNTP production and thereby contribute to exacerbated mutator phenotype44,45. It has also been used to study the relevance of dNTP production to reactive oxygen species-mediated mutagenesis8. The expression of Rnr1-3Myc-p was successfully induced with the addition of galactose (Fig. 5E, lanes 4–6 compared to 1–3). Under enforced Rnr1p expression, the HU sensitivity of apd1Δ cells remained unchanged (Fig. 5F, rows 5–6 compared to 4), suggesting that the overproduction of dNTP could not rescue the HU phenotype in apd1Δ cells. Notably, such sensitivity was unaffected by the KanMX gene deletion cassette, as the replacement of APD1 by HIS3 also resulted in identical phenotypes (Fig. 5B, rows 2–3 compared to 1 and rows 8–9 compared to 7; Fig. 5F, rows 5–6 compared to 4). Taken together, these results demonstrated that the suppression of HU sensitivity by APD1 was unlikely mediated through the dNTP regulatory network.
Suppression of HU sensitivity in apd1Δ cells by YAP1 and ATR1
Since modulation of dNTP production did not affect HU sensitivity in apd1Δ cells and our candidate gene approach to identify genetic partners of APD1 was not fruitful, we performed a genetic screen to search for high-copy-number suppressors of the HU phenotype. An expression library of all yeast genes was used in this screen and YAP1 was the only gene found to rescue the HU sensitivity in apd1Δ cells. A YAP1 plasmid was isolated from all 11 HU-sensitive clones obtained in the screen. To verify this, we constructed YAP1 expression plasmid and transformed it into apd1Δ cells. As shown in Fig. 6A, ectopic expression of YAP1 effectively complemented the HU sensitivity in apd1Δ cells (Fig. 6A, row 6 compared to 4). This was comparable to the re-expression of APD1 (Fig. 6A, row 6 compared to 5), indicating that YAP1 serves as a physiological suppressor of HU sensitivity in apd1Δ cells. Notably, yap1Δ cells were also sensitive to HU (Fig. 6A, row 2 compared to 3). Re-expression of YAP1 plasmid successfully rescued such phenotype (Fig. 6A, row 1 compared to 3), indicating the functionality of pYAP1 expression plasmid in these cells.
Suppression of HU sensitivity in apd1Δ cells by YAP1 and ATR1.
(A) Expression of YAP1 rescued the HU sensitivity in apd1Δ cells. Ten-fold serial dilutions of strains yap1Δ pRS415-YAP1, yap1Δ pRS415, WT pRS415, apd1Δ pRS415, apd1Δ pRS415-APD1 and apd1Δ pRS415-YAP1 were spotted on SC-L medium containing 240 mM of HU. Plates were incubated for 3–4 days at 30°C. (B) Expression of ATR1 suppressed the HU sensitivity in apd1Δ cells. Ten-fold serial dilutions of strains WT pRS415, apd1Δ pRS415, apd1Δ pRS415-APD1, apd1Δ pRS415-YAP1 and apd1Δ pRS415-ATR1 were spotted on SC-L medium containing 240 mM of HU. (C) Suppression of the HU sensitivity in apd1Δ cells by ATR1 was lost in the absence of YAP1. Ten-fold serial dilutions of strains apd1Δ pRS415-APD1, apd1Δ pRS415, apd1Δyap1Δ pRS415, apd1Δyap1Δ pRS415-APD1, apd1Δyap1Δ pRS415-YAP1 and apd1Δyap1Δ pRS415-ATR1 were spotted on SC-L medium containing 240 mM of HU.
One of the known YAP1 downstream targets is ATR1, a multi-drug resistance gene encoding a membrane transporter. Its expression has been shown to be transcriptionally upregulated by YAP133. ATR1 is required for cellular resistance to cytotoxic agents such as aminotriazole and boron.26,46 To explore whether ATR1 might also be involved in the suppression of HU sensitivity in apd1Δ cells, expression plasmid of ATR1 was constructed under the regulation of its own promoter. Complementation experiment was carried out. Expression of either ATR1 or YAP1 protected the apd1Δ cells from HU challenge to full extent as that of APD1 (Fig. 6B, rows 4–5 compared to 2–3). Thus, both ATR1 and YAP1 are fully capable of suppressing the deleterious effect of HU in APD1-null cells.
We next investigated the genetic interaction between ATR1 and YAP1 in the suppression of HU sensitivity in apd1Δ cells. To this end, we generated apd1Δ yap1Δ cells. Double deletion of APD1 and YAP1 exacerbated the HU sensitivity when compared to individual single mutants (Fig. 6C, row 3 compared to row 2 and to Fig. 6A, row 2), while re-expression of pAPD1 or pYAP1 reverted the cell growth phenotype back to its counterpart single mutant (Fig. 6C, row 3 compared to 4–5). This indicated the effectiveness and specificity of the genetic targeting. In this genetic background, re-expression of ATR1 plasmid, under the control of its own promoter, was ineffective in rescuing the HU phenotype (Fig. 6C, row 6 compared to 4–5). Thus, the protective effect of ATR1 was not seen in the absence of YAP1.
Discussion
We provided two key messages in this study. First, yeast Apd1p is an Fe-S protein. Second, loss of APD1 confers HU sensitivity. Our findings are consistent with the notion that dimeric Apd1p forms a [2Fe–2S] cluster which mediates its cellular function. As such, disruption of iron-binding motifs in Apd1p abolished its ability to protect cells against HU. Furthermore, the HU sensitivity in apd1Δ cells was not mediated through the dNTP regulatory network, but was suppressed by YAP1 or its target ATR1. Our findings revealed a new function of Apd1p in cellular defense against HU, which might be relevant to other thioredoxin-like ferredoxins.
Thioredoxin-like ferredoxins are a large family of Fe-S proteins with unknown biological function18. This study was started based on the hypothesis that Apd1p might be the electron donor for Tsa1p, a master peroxiredoxin in yeast9,10,16. Although we did not find any physical or genetic interaction between Apd1p and Tsa1p, we characterized Apd1p as an Fe-S protein critically involved in cellular defense against HU. On the other hand, we were unable to verify several results obtained for Apd1p in global analyses of drug susceptibility. Particularly, apd1Δ cells were not sensitive to H2O2. Based on our initial findings, we revised our model to hypothesize that Apd1p might operate through the dNTP regulatory network to exert its protective effect against HU. Although we could not positively define a role for dNTP production in the HU sensitivity of apd1Δ cells, we performed a genome-wide screen and identified YAP1 as a high-copy-number suppressor of the HU phenotype. The new information about Apd1p obtained in our study has provided the foundation for future investigations.
Our analysis of purified Apd1p protein by SDS-PAGE, color visualization, UV-visible spectrophotometric scanning and ICP-MS provided the first experimental evidence in support of the formation of a [2Fe–2S] cluster in Apd1p dimer. Although additional biophysical methods including electron paramagnetic resonance, nuclear magnetic resonance and Mössbauer spectroscopy could be used to characterize Apd1p further19,47, our biological assays have already indicated the role of redox and the essentiality of both iron-binding motifs in cellular function of Apd1p. Further investigations are required to shed more mechanistic light on exactly how the [2Fe–2S] cluster in Apd1p could mediate the protection from HU challenge.
Thioredoxins, glutaredoxins and peroxiredoxins serve as redox-sensitive molecular switches in cell signaling9,16, which are commonly mediated through the modulation of disulfide bond formation21,48. In this regard, our findings on Apd1p are generally consistent with the notion that it might function to modulate disulfide bond formation in its target. Our genetic analysis indicated the ability of Yap1p transcription factor to suppress the HU sensitivity in apd1Δ cells. Thus, Yap1p, but neither Tsa1p nor RNR as we originally hypothesized, might be a direct target of Apd1p. It will be of great interest to see how Apd1p might affect the transcriptional activity of Yap1p. Redox regulation of Yap1p by glutathione peroxidase Gpx3p, Tsa1p and Ybp1p has been documented49,50,51,52. Whether Apd1p might modulate Yap1p and related transcription factors in a similar manner warrants further study.
Our work reveals that the sensitivity of apd1Δ cells to HU can be suppressed by YAP1 or ATR1. Although Yap1p is known to play an important role in stress response31,32 and its expression is activated by HU34, we provided the first demonstration of the protective role of Yap1p and Atr1p against HU. Exactly how Yap1p modulates cellular response to HU remains murky. In one perspective, Atr1p is the target of Yap1p in this response since it is also capable of suppressing HU sensitivity in apd1Δ cells. However, the inability of ATR1 to suppress HU sensitivity in apd1Δ yap1Δ cells suggests a more complicated picture. The involvement of other Yap1p targets that cooperate with Atr1p is possible. One candidate is the Yap1p-regulated membrane transporter Flr1p, which is already known to mediate resistance to several cytotoxic drugs27,28,53. Indeed in some of these cases Flr1p and Atr1p are known to cooperate with each other.27,28 Alternatively, Yap1p might be absolutely required for expression or activity of Atr1p. In other words, Atr1p might not be expressed in the absence of YAP1. In our genetic analysis, YAP1 was found to rescue HU sensitivity in apd1Δ cells. In addition, apd1Δ yap1Δ cells were more sensitive to HU than apd1Δ or yap1Δ cells. These results raised the possibility that YAP1 and APD1 might operate in different pathways that confer resistance to HU. Nevertheless, additional biochemical and genetic analyses are required to clarify the roles of Yap1p and Atr1p in cellular defense against HU.
Methods
Strains and plasmids
Saccharomyces cerevisiae strains BY474154 and W303-1a and their isogenic strains (Table 1) were used. All knockout mutants were constructed by one-step gene deletion method55. Primers were listed in Table 2. Expression vector for APD1, YAP1 and ATR1 was based on pRS415 and the gene locus including promoter and open reading frame was PCR-amplified from yeast genomic DNA. Expression plasmids for APD1 mutants were constructed by site-directed mutagenesis as described8,56,57. The Myc tag sequence in APD1 and its mutants was introduced by PCR.
Plasmids pGal-RNR1, pRNR2-LacZ and pRNR3-LacZ were kindly provided by Dr. Stephen Elledge38,43.
RNA analysis
Total RNA was extracted by phenol/freeze RNA preparation method as described58. For RT-PCR, 3 μg of total RNA was used for cDNA synthesis. Quantitative RT-PCR was performed as described59. Relative mRNA levels were normalized to ACT1. PCR primers were listed in Table 2.
Protein analysis
Western blotting was performed essentially as described8,42,57. Yeast cells were harvested by centrifugation, followed by trichloroacetic acid extraction with the help of glass beads. β-galactosidase assay was carried out using a chemiluminescent method as described57.
Cell growth and cell cycle profiling
For growth curve analysis, logarithmically growing yeast cells, BY4741 wild type (WT) and apd1Δ, were inoculated into YPD medium to an OD600 of 0.1. Every indicated time point thereafter, cell density was monitored by OD600. Cell cycle profiling was performed as described previously60. First, yeast cells were grown to a density of 5 × 107 cells/ml in YPD medium. Cells were spun down and washed in PBS. Cells were then fixed in 1 ml of 70% ethanol for 1 hour at 4°C in dark. Subsequently, the fixed cells were treated with RNase A at 37°C for 1 hour. Cells were pelleted and resuspended in 1 ml of PBS containing 50 μg/ml propidium iodide. Samples were kept in dark at room temperature for 1 hour prior to flow cytometric analysis. DNA content of 20,000 cells from each sample was analyzed. To measure intracellular reactive oxygen species, cells were treated with 0.5 mM H2O2 and then with 10 μM DCF as described60.
Expression and purification of recombinant Apd1p
Bacterially expressed recombinant His-Apd1p was purified with the Ni-NTA-Superflow resin. Proteins were eluted and subjected to 15% SDS-PAGE (for Apd1p WT) or 15% non-denaturing PAGE (for Apd1p WT, CX3C and HX3H). After electrophoresis, the SDS-PAGE gel was stained with Coomassie blue to visualize the purified protein. The colored Apd1p band was directly seen on the non-denaturing gel.
UV-visible spectroscopy of recombinant Apd1p proteins
Each of the purified proteins was subjected to photometric scanning with a wavelength range of 260 to 750 nm and a scanning step size of 2 nm in 100 ms on a VarioskanTM Flash spectrometer.
Determination of iron concentration by ICP-MS
The concentration of Apd1p used for ICP-MS analysis was determined by bicinchoninic acid (BCA) micro-scale assay (Novagen) using bovine serum albumin (BSA) as a standard. Apd1p protein sample was diluted 50-fold prior to analysis. Calibration curve was obtained by measuring a set of iron standards (Fluka) ranging from 30 to 90 ppb (R = 0.9995). ICP-MS experiments were performed on an Agilent 7500a spectrometer (Agilent Technologies).
References
Entian, K. D. et al. Functional analysis of 150 deletion mutants in Saccharomyces cerevisiae by a systematic approach. Mol Gen Genet 262, 683–702 (1999).
Blackman, R. K. et al. Mitochondrial electron transport is the cellular target of the oncology drug elesclomol. PLoS One 7, e29798 (2012).
Postma, L., Lehrach, H. & Ralser, M. Surviving in the cold: yeast mutants with extended hibernating lifespan are oxidant sensitive. Aging 1, 957–960 (2009).
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Krogan, N. J. et al. Global landscape of protein complexes in the yeast Saccharomyces cerevisiae. Nature 440, 637–643 (2006).
Huang, M. E., Rio, A. G., Nicolas, A. & Kolodner, R. D. A genomewide screen in Saccharomyces cerevisiae for genes that suppress the accumulation of mutations. Proc Natl Acad Sci USA 100, 11529–11534 (2003).
Huang, M. E. & Kolodner, R. D. A biological network in Saccharomyces cerevisiae prevents the deleterious effects of endogenous oxidative DNA damage. Mol Cell 17, 709–720 (2005).
Tang, H. M. V., Siu, K. L., Wong, C. M. & Jin, D. Y. Loss of yeast peroxiredoxin Tsa1p induces genome instability through activation of the DNA damage checkpoint and elevation of dNTP levels. PLoS Genet 5, e1000697 (2009).
Rhee, S. G., Chae, H. Z. & Kim, K. Peroxiredoxins: a historical overview and speculative preview of novel mechanisms and emerging concepts in cell signaling. Free Radic Biol Med 38, 1543–1552 (2005).
Rhee, S. G., Woo, H. A., Kil, I. S. & Bae, S. H. Peroxiredoxin functions as a peroxidase and a regulator and sensor of local peroxides. J Biol Chem 287, 4403–4410 (2012).
Nyström, T., Yang, J. & Molin, M. Peroxiredoxins, gerontogenes linking aging to genome instability and cancer. Genes Dev 26, 2001–2008 (2012).
Yarbro, J. W. Mechanism of action of hydroxyurea. Semin Oncol 19, 1–10 (1992).
Hofer, A., Crona, M., Logan, D. T. & Sjöberg, B. M. DNA building blocks: keeping control of manufacture. Crit Rev Biochem Mol Biol 47, 50–63 (2012).
Madaan, K., Kaushik, D. & Verma, T. Hydroxyurea: a key player in cancer chemotherapy. Expert Rev Anticancer Ther 12, 19–29 (2012).
Meyer, Y., Buchanan, B. B., Vignols, F. & Reichheld, J. P. Thioredoxins and glutaredoxins: unifying elements in redox biology. Annu Rev Genet 43, 335–367 (2009).
Hanschmann, E. M., Godoy, J. R., Berndt, C., Hudemann, C. & Lillig, C. H. Thioredoxins, glutaredoxins and peroxiredoxins - molecular mechanisms and health significance: from cofactors to antioxidants to redox signaling. Antioxid Redox Signal 19, 1539–1605 (2013).
Chae, H. Z., Chung, S. J. & Rhee, S. G. Thioredoxin-dependent peroxide reductase from yeast. J Biol Chem 269, 27670–27678 (1994).
Meyer, J. Ferredoxins of the third kind. FEBS Lett 509, 1–5 (2001).
Xu, X. M. & Møller, S. G. Iron-sulfur clusters: biogenesis, molecular mechanisms and their functional significance. Antioxid Redox Signal 15, 271–307 (2011).
Pedone, E., Limauro, D., D'Ambrosio, K., De Simone, G. & Bartolucci, S. Multiple catalytically active thioredoxin folds: a winning strategy for many functions. Cell Mol Life Sci 67, 3797–3814 (2010).
Groitl, B. & Jakob, U. Thiol-based redox switches. Biochim Biophys Acta 1844, 1335–1343 (2014).
Wu, C. H., Jiang, W., Krebs, C. & Stubbe, J. YfaE, a ferredoxin involved in diferric-tyrosyl radical maintenance in Escherichia coli ribonucleotide reductase. Biochemistry 46, 11577–11588 (2007).
Zhang, Y. et al. Investigation of in vivo diferric tyrosyl radical formation in Saccharomyces cerevisiae Rnr2 protein: requirement of Rnr4 and contribution of Grx3/4 AND Dre2 proteins. J Biol Chem 286, 41499–41509 (2011).
Wysocki, R. et al. Transcriptional activation of metalloid tolerance genes in Saccharomyces cerevisiae requires the AP-1-like proteins Yap1p and Yap8p. Mol Biol Cell 15, 2049–2060 (2004).
Viau, C. et al. Sensitivity to Sn2+ of the yeast Saccharomyces cerevisiae depends on general energy metabolism, metal transport, anti-oxidative defences and DNA repair. Biometals 19, 705–714 (2006).
Kaya, A., Karakaya, H. C., Fomenko, D. E., Gladyshev, V. N. & Koc, A. Identification of a novel system for boron transport: Atr1 is a main boron exporter in yeast. Mol Cell Biol 29, 3665–3674 (2009).
Sundström, L., Larsson, S. & Jönsson, L. J. Identification of Saccharomyces cerevisiae genes involved in the resistance to phenolic fermentation inhibitors. Appl Biochem Biotechnol 161, 106–115 (2010).
Westman, J. O., Manikondu, R. B., Franzén, C. J. & Taherzadeh, M. J. Encapsulation-induced stress helps Saccharomyces cerevisiae resist convertible lignocellulose derived inhibitors. Int J Mol Sci 13, 11881–11894 (2012).
Kim, D. & Hahn, J. S. Roles of the Yap1 transcription factor and antioxidants in Saccharomyces cerevisiae's tolerance to furfural and 5-hydroxymethylfurfural, which function as thiol-reactive electrophiles generating oxidative stress. Appl Environ Microbiol 79, 5069–5077 (2013).
Pimentel, C. et al. Yap1 mediates tolerance to cobalt toxicity in the yeast Saccharomyces cerevisiae. Biochim Biophys Acta 1840, 1977–1986 (2014).
Lee, J. et al. Yap1 and Skn7 control two specialized oxidative stress response regulons in yeast. J Biol Chem 274, 16040–16046 (1999).
Toone, W. M., Morgan, B. A. & Jones, N. Redox control of AP-1-like factors in yeast and beyond. Oncogene 20, 2336–2346 (2001).
Coleman, S. T., Tseng, E. & Moye-Rowley, W. S. Saccharomyces cerevisiae basic region-leucine zipper protein regulatory networks converge at the ATR1 structural gene. J Biol Chem. 272, 23224–23230 (1997).
Dubacq, C. et al. Role of the iron mobilization and oxidative stress regulons in the genomic response of yeast to hydroxyurea. Mol Genet Genomics 275, 114–124 (2006).
Smith, S. et al. Mutator genes for suppression of gross chromosomal rearrangements identified by a genome-wide screening in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 101, 9039–9044 (2004).
Stubbe, J. & van Der Donk, W. A. Protein radicals in enzyme catalysis. Chem Rev 98, 705–762 (1998).
Nordlund, P. & Reichard, P. Ribonucleotide reductases. Annu Rev Biochem 75, 681–706 (2006).
Huang, M. & Elledge, S. J. Identification of RNR4, encoding a second essential small subunit of ribonucleotide reductase in Saccharomyces cerevisiae. Mol Cell Biol 17, 6105–6113 (1997).
Longley, D. B., Harkin, D. P. & Johnston, P. G. 5-fluorouracil: mechanisms of action and clinical strategies. Nat Rev Cancer 3, 330–338 (2003).
Chabes, A., Domkin, V. & Thelander, L. Yeast Sml1, a protein inhibitor of ribonucleotide reductase. J Biol Chem 274, 36679–36683 (1999).
Huang, M., Zhou, Z. & Elledge, S. J. The DNA replication and damage checkpoint pathways induce transcription by inhibition of the Crt1 repressor. Cell 94, 595–605 (1998).
Carter, C. D., Kitchen, L. E., Au, W. C., Babic, C. M. & Basrai, M. A. Loss of SOD1 and LYS7 sensitizes Saccharomyces cerevisiae to hydroxyurea and DNA damage agents and downregulates MEC1 pathway effectors. Mol Cell Biol 25, 10273–10285 (2005).
Zhou, Z. & Elledge, S. J. Isolation of crt mutants constitutive for transcription of the DNA damage inducible gene RNR3 in Saccharomyces cerevisiae. Genetics 131, 851–866 (1992).
Chabes, A. et al. Survival of DNA damage in yeast directly depends on increased dNTP levels allowed by relaxed feedback inhibition of ribonucleotide reductase. Cell 112, 391–401 (2003).
Chabes, A. & Stillman, B. Constitutively high dNTP concentration inhibits cell cycle progression and the DNA damage checkpoint in yeast Saccharomyces cerevisiae. Proc Natl Acad Sci USA 104, 1183–1188 (2007).
Kanazawa, S., Driscoll, M. & Struhl, K. ATR1, a Saccharomyces cerevisiae gene encoding a transmembrane protein required for aminotriazole resistance. Mol Cell Biol 8, 664–673 (1988).
Zhang, B. et al. Monothiol glutaredoxins can bind linear [Fe3S4]+ and [Fe4S4]2+ clusters in addition to [Fe2S2]2+ clusters: spectroscopic characterization and functional implications. J Am Chem Soc 135, 15153–15164 (2013).
König, J., Muthuramalingam, M. & Dietz, K. J. Mechanisms and dynamics in the thiol/disulfide redox regulatory network: transmitters, sensors and targets. Curr Opin Plant Biol 15, 261–268 (2012).
Delaunay, A., Pflieger, D., Barrault, M. B., Vinh, J. & Toledano, M. B. A thiol peroxidase is an H2O2 receptor and redox-transducer in gene activation. Cell 111, 471–481 (2002).
Veal, E. A., Ross, S. J., Malakasi, P., Peacock, E. & Morgan, B. A. Ybp1 is required for the hydrogen peroxide-induced oxidation of the Yap1 transcription factor. J Biol Chem 278, 30896–30904 (2003).
Okazaki, S., Tachibana, T., Naganuma, A., Mano, N. & Kuge, S. Multistep disulfide bond formation in Yap1 is required for sensing and transduction of H2O2 stress signal. Mol Cell 27, 675–688 (2007).
Tachibana, T. et al. A major peroxiredoxin-induced activation of Yap1 transcription factor is mediated by reduction-sensitive disulfide bonds and reveals a low level of transcriptional activation. J Biol Chem 284, 4464–4472 (2009).
Oskouian, B. & Saba, J. D. YAP1 confers resistance to the fatty acid synthase inhibitor cerulenin through the transporter Flr1p in Saccharomyces cerevisiae. Mol Gen Genet 261, 346–353 (1999).
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Longtine, M. S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998).
Wong, C. M., Siu, K. L. & Jin, D. Y. Peroxiredoxin-null yeast cells are hypersensitive to oxidative stress and genomically unstable. J Biol Chem 279, 23207–23213 (2004).
Kong, K. Y. E. et al. Cotranscriptional recruitment of yeast TRAMP complex to intronic sequences promotes optimal pre-mRNA splicing. Nucl Acids Res 42, 643–660 (2014).
Schmitt, M. E., Brown, T. A. & Trumpower, B. L. A rapid and simple method for preparation of RNA from Saccharomyces cerevisiae. Nucl Acids Res 18, 3091–3092 (1990).
Tang, H. M. V. et al. Requirement of CRTC1 coactivator for hepatitis B virus transcription. Nucl Acids Res 10.1093/nar/gku925 (2014).
Wong, C. M., Zhou, Y., Ng, R. W. M., Kung, H. F. & Jin, D. Y. Cooperation of yeast peroxiredoxins Tsa1p and Tsa2p in the cellular defense against oxidative and nitrosative stress. J Biol Chem 277, 5385–5394 (2002).
Acknowledgements
We thank Drs. Stephen Elledge and Mark Longtine for reagents and members of Jin laboratory for critical reading of manuscript. This work was supported by grants from the National Natural Science Foundation of China (Young Researcher Award 31200931 to H.-M.V.T.), S.K. Yee Medical Research Fund (2011 to D.-Y.J.) and Fogarty International Center of National Institutes of Health (R01TW008298 to C.-M.W.).
Author information
Authors and Affiliations
Contributions
H.M.V.T., H.Z.S., C.M.W. and D.Y.J. designed the experiments, analyzed data and wrote the manuscript. H.M.V.T., K.P., K.Y.E.K., L.H., L.C.C. and K.L.S. performed the experiments and analyzed data.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Tang, HM., Pan, K., Kong, KY. et al. Loss of APD1 in Yeast Confers Hydroxyurea Sensitivity Suppressed by Yap1p Transcription Factor. Sci Rep 5, 7897 (2015). https://doi.org/10.1038/srep07897
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07897
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.