Abstract
Multiplex real-time PCR with quantification of targets in a single fluorescence channel has been the demand in biotechnology industry. Here, we develop a novel analytical real-time PCR technique to detect multiple targets in a single fluorescence channel without melting curve analysis. In this technique, we show the intensity of the fluorescence signals of two discrete Tm targets is different at certain temperatures called detection temperatures, by which a high Tm target can be detected regardless of a low Tm target. We then identify the low Tm target by utilizing a change of the fluorescence signals between two different detection temperatures. Furthermore, it enables us to determine quantification of each target in a single channel, possibly facilitating convenient patient care for drug treatment in clinics.
Similar content being viewed by others
Introduction
The major advantage of real-time PCR is the development of homogeneous reactions, the amplification and detection of a target in real time1. As a result, real-time PCR has rapidly become the key tool for target and biomarker identification, expanding into applications as diverse as in vitro diagnostics (IVDs)2,3, food safety testing4,5 and pharmacogenomics6,7.
However, the biotechnology industry has been struggling to meet the increasing demand for assays that provide both high-level multiplexing and quantification of targets using existing real-time PCR technologies8,9. To date, the industry has made considerable progress toward achieving these goals. For example, multiplex real-time PCR has been possible by using either multiple fluorescence channels or melting curve analysis after amplification10,11. In particular, the latter has been the best way to detect multiple targets in a single fluorescence channel. However, these methods still has several downsides, including crosstalk between channels, longer turnaround time (TAT) due to end-point analysis after the PCR step and melting temperature (Tm) variations12,13. Moreover, it is still impossible to obtain an individual Ct (threshold cycle) value of each of multiple targets in a single fluorescence channel.
To overcome these challenges, we develop a novel analytical technique, MuDT (Multiple Detection Temperatures), which enables us to detect multiple targets in a single fluorescence channel without melting curve analysis. To demonstrate how MuDT works, we employ TOCE (Tagging Oligonucleotide Cleavage and Extension)14 technique to design oligonucleotides to detect the DNA target. TOCE utilizes indirect signal generation through the use of two novel components, Pitcher and Catcher. During each cycle, the Pitcher, a dual-purpose oligonucleotide that specifically binds to target DNA, is cleaved to release the unlabeled Extender that serves as a primer for an artificial template, a quenched-fluorescent molecule, the Catcher. Annealing and extension of the Extender on the Catcher generates Duplex Catcher, resulting in a fluorescence signal that is directly correlated to the quantity of the target DNA (Supplementary Fig. S1).
Overall, we verify that MuDT detects multiple targets in a single fluorescence channel. Notably, this method achieves Ct measurements for the targets, does not require melting curve analysis and avoids crosstalk between channels; thus, MuDT has the potential to deliver more comprehensive and actionable diagnostics, leading to improved patient care and reduced healthcare costs.
Results
Intensity of the fluorescence signal depends on detection temperature
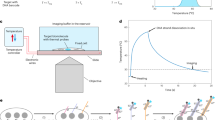
During real-time PCR reaction, the duplex-driven fluorescence signal (designated as a yellow circle) of the probes depends on the temperature because of their melting curves (Fig. 1). Therefore, different intensities of those signals at certain temperatures at each cycle are able to detect the targets with different Tm values (highlighted in gray). These temperatures are defined as “detection temperatures” at which the intensity of a target's fluorescence signal is measured.
Intensity of the fluorescence signals depends on detection temperature.
Detection temperature means a certain temperature where the unquenched fluorescence signal is measured at each cycle during real-time PCR reaction. If there are two targets representing different Tm profiles, the fluorescence signals between two targets would be different at certain detection temperatures (marked in gray). Therefore, these temperatures differentiate the high Tm target from the low Tm target, by which the high Tm target can be detected.
Accordingly, the intensity of the fluorescence signals from the targets with different Tm profiles can be ascertained by controlling the detection temperatures in MuDT technique, enabling targets to be distinguished from each other without melting curve analysis.
Identification of high Tm target by detection temperatures
By using TOCE technique, we targeted the genomic DNA of two pathogens, Chlamydia trachomatis (CT)15 and Neisseria gonorrhoeae (NG)15, with different Tm profiles, 75°C (high Tm) and 65°C (low Tm) respectively. We then confirmed that the designed components for targeting CT and NG yielded the unique and easily discernible Tm profiles (Fig. 2 and Supplementary Fig. S2). To demonstrate that detection temperatures control the intensity of fluorescence signals, we selected three different detection temperatures (60°C, 72°C and 95°C). After annealing and extension at 60°C at each cycle, additional steps for detection (5 s each at 60°C, 72°C and 95°C) were carried out and the outcomes were then compared. As expected, the fluorescence signals of both targets were vigorous at 60°C because this temperature is lower than their Tm values, insuring that fluorescence remains unquenched (Fig. 2a–b and Supplementary Fig. S2a–b). By contrast, no signal was detected for either target at 95°C due to fluorescence quenching at this temperature (Fig. 2a–b and Supplementary Fig. S2a–b). Importantly, we noticed that the two targets could be differentiated based on the dissimilar intensity of their unquenched fluorescence signals at a detection temperature of 72°C, at which the signal for NG was not detectable (Fig. 2a–b and Supplementary Fig. S2a–b).
Identification of the high Tm target by detection temperatures.
(a) A Multiple Detection Temperature (MuDT) reaction was assembled for the screening of Chlamydia trachomatis (CT) (high Tm) and the fluorescence signal was detected at three different detection temperatures (60°C, 72°C and 95°C) at each PCR cycle. The first three graphs represent the amplification plots for the target and the last graph contains the composite melting curves of the targets and three detection temperatures shown as dotted lines. (b and c) Procedures are the same as in (a), except that Neisseria gonorrhoeae (NG) (low Tm) alone (b) or both targets (c) were screened.
These results prompted us to examine the signals at the same detection temperatures when both targets are present in the reaction. Under this condition, we observed the CT signal was also detected at 60°C and 72°C, strongly suggesting that CT (high Tm target) can be detected regardless of the presence of NG (low Tm target) (Fig. 2c and Supplementary Fig. S2c). Therefore, we concluded that the appearance of the fluorescence signal at a detection temperature at which no signal is detected for the low Tm target indicates the presence of the high Tm target. In addition, a Ct value of the high Tm target can be determined at this detection temperature.
Detection of low Tm target by granting Ct value
We next asked how the NG target with low Tm is recognized. To this end, we devised a novel analytical method that uses ΔRFU (a change in Relative Fluorescence Units) of the targets between two different detection temperatures at each cycle during PCR reaction (Supplementary Fig. S3). So as to demonstrate this method, we first determined an arbitrary threshold that is defined as a minimum value over the ΔRFU of the high Tm target between two different detection temperatures, ensuring cutting off the ΔRFU value of the high Tm target. The determined threshold was then used to detect the low Tm target with its Ct value. For example, if both targets are present or the low Tm target is present alone, ΔRFU between the two detection temperatures would be significantly higher than the determined threshold; thus, such a result would indicate the presence of the low Tm target. Subsequently, the Ct value of the low Tm target can be obtained from the amplification plot of the ΔRFU of the low Tm target and the threshold eliminating the value of the high Tm target (designated by an arrow in Supplementary Fig. S3).
To test this idea, we plotted the ΔRFU of the CT target between the two detection temperatures (60°C and 72°C) used in Figure 2 (Fig. 3a and Supplementary Fig. S4). A threshold was then determined by eliminating the ΔRFU of the CT target. Indeed, the amplification plot was higher than the determined threshold when both targets were present or when NG was present alone (Fig. 3a and Supplementary Fig. S4). Using this information, we calculated a Ct value of the NG target from the cycle number at which the arbitrary threshold crossed the amplification plot (designated by an arrow in Fig. 3a and Supplementary Fig. S4).
Detection of the low Tm target by granting a Ct value.
(a) The arbitrary threshold derived from the ΔRFU of the high Tm target at two different detection temperatures was applied to calculate the Ct value of Neisseria gonorrhoeae (NG) (low Tm), designated by an arrow. NTC is no-target control. (b) Calculation of the Ct values with a serial dilution of NG gDNA (1 ng–100 fg). Left panel: NG alone; right panel: both Chlamydia trachomatis (CT) (high Tm) and NG. The averages (AVE) of the Ct values of three different experiments are shown. R2 indicates correlation coefficient, whereas the slope represents the efficiency of PCR. ΔRFU represents the value deducted the RFU at 72°C from that at 60°C. The table shown represents the concentration of each reaction.
To demonstrate that the obtained Ct value of the low Tm target (NG) correlates to its concentration, we plotted the ΔRFU values of the NG target with serially diluted NG genomic DNA (1 ng–100 fg). As a result, we could measure different Ct values that certainly correlate to the concentration of NG gDNA (Fig. 3b, Supplementary Fig. 5a and 5b). We then evaluated the correlation coefficient (R2) to determine how well the data fit the standard curve. To this end, we plotted the Ct values against log10[NG gDNA] to obtain the amplification efficiency curve. The curve was linear across a 105-fold range of input NG gDNA (R2 > 0.99), demonstrating that the Ct values of the low Tm target calculated with MuDT technique were directly correlated to the quantity of the target DNA, even when CT and NG were both present (Fig. 3b).
Application of MuDT technique to detection of sexually transmitted infections in clinical samples
To validate our result, we applied this technique to a commercially available Anyplex™ II STI-7 kit for detecting multiple pathogens in clinical specimens that cause sexually transmitted infections (STI)16. This kit is based on TOCE technique so that targets are detected by melting curve analysis. As shown in Table 1, it contains four channels each of which has two targets with different Tm values. To test whether MuDT can identify multiple pathogens in clinical samples by using this diagnostic product, we first selected two detection temperatures (72°C and 60°C). We then determined a respective arbitrary threshold from the ΔRFU of high Tm targets between these two detection temperatures to discover each low Tm target. As a result, the different intensities of the fluorescence signals at 72°C enabled us to detect the high Tm targets in each channel with their own Ct value. We also acquired an individual Ct value of the low Tm targets in each channel by using the determined thresholds (Table 1). These results confirm the capacity of MuDT in clinical diagnostics.
Discussion
In this study, we demonstrated that MuDT is not a new type of oligo-based real-time PCR technique. Rather, it is a novel method for analyzing real-time PCR data, by which multiple targets are obviously detected in a single channel without melting curve analysis. Therefore, MuDT can apply its novel analytical method to any other hybridization-based chemistry17 including molecular beacons18, FRET probes19 and any combination of these probes.
Moreover, MuDT technique opens the door for mingling with different fluorescence signal chemistries. Currently, hydrolysis- and hybridization-based chemistries are two major methods to generate fluorescence signal in real-time PCR. For instance, TaqMan probe uses the hydrolysis of the probe by 5′ to 3′ exonuclease activity of polymerase20, whereas TOCE is based on hybridization chemistry by which duplex formation of the Catcher generates fluorescence signal. We predicted that the use of a TaqMan probe in MuDT reaction enables us to design the targets with considerably high Tm value. Because the hydrolysis-based probe generates its fluorescence signal even at 95°C. However, the hybridized probe does not emit its signal at this temperature by denaturation of the probe (Fig. 2). Therefore, by using a TaqMan probe, it consequently ensures wider selection of detection temperatures, eventually detecting more targets in a single fluorescence channel. To demonstrate the applicability of TaqMan probe in MuDT, we tested a TaqMan probe as a target with high Tm. As a result, two targets with different fluorescence chemistries were also detected in a single channel (Supplementary Fig. S6). This result confirmed the possible scenario for multiplexing for detection of more than two targets in a single channel in MuDT reaction.
In addition, MuDT will upgrade any real-time PCR instrument for more than double capacity of multiplexing without increasing number of channels demanding expensive costs and to develop a universal single reagent (assay) without crosstalk between channels.
Clinically, co-infection of pathogens is dangerous to patients. Therefore, the necessary assay(s) should be multiplexing to detect all of them simultaneously and each target should be quantified as well. However, current real-time PCR chemistries are impossible to provide quantitative measurements for multiple targets in a single channel. Here, we probed that MuDT grants each Ct value to the targets in a single channel. Therefore, the multi-quantitation capability of this technique will enable clinicians to gain a complete package of diagnostic information including the diagnosis of disease, degree of disease progression and severity and monitoring of treatment.
In conclusion, we achieved the longstanding goal for more than 20 years in real-time PCR systems by developing a novel target-detection technique. With these successful results, we anticipate that MuDT will generate convenient multiplexing with quantification, better assay system and easier patient care.
Methods
Preparation of target DNA
Bacteria and genomic DNA [Chlamydia trachomatis (ATCC VR-1500) and Neisseria gonorrhoeae (ATCC 700825D)] were purchased from ATCC (VA, USA). DNAs extracted from swab and urine specimens were used for the detection of sexually transmitted infections. The QIAamp DNA Mini Kit (Qiagen, Germany) was used to purify genomic DNA.
Designing of Oligonucleotides
The oligonucleotide sequences used in this study are listed in Supplementary Table S1. Oligonucleotides were purchased from Integrated DNA Technologies Inc. (IA, USA) and Biosearch Technologies Inc. (CA, USA).
MuDT reaction and melting curve analysis for the high Tm target
Multiple Detection Temperature (MuDT) reactions were carried out in the final volume of 20 μl containing 1 ng of each genomic DNA (CT or NG, or both), 10 pmol of each primer set (NG-F/NG-R and CT-F/CT-R), 5 pmol of each Pitcher (NG-P and CT-P1) and 1 pmol of each Catcher (NG-C and CT-C) in buffer containing 2 units of Taq DNA polymerase, 200 μM of dNTPs and 2 mM of MgCl2. The whole PCR steps are as follows: denaturation for 15 min at 95°C and 50 cycles of denaturation (30 s at 95°C), annealing and extension (60 s at 60°C) and additional steps for detection (5 s each at 60°C, 72°C and 95°C) in a CFX96 thermocycler (Bio-Rad, USA). Fluorescence signals were detected at three different detection temperatures (additional steps: 60°C, 72°C and 95°C) at each cycle. Upon completion of the reaction, a melting curve was obtained by cooling the reaction mixture to 55°C, holding at 55°C for 30 s and then increasing the temperature from 55°C to 95°C stepwise (0.5°C/step) with a 5 s hold between each step. Fluorescence was measured continuously during the temperature rises to monitor dissociation of double-stranded DNAs. Melting peaks were derived from the melting curve data (CFX96 Manager).
MuDT reaction for the low Tm target
MuDT reactions were carried out in the final volume of 20 μl containing 1 ng of each genomic DNA (CT or NG, or both), 10 pmol of each primer set (NG-F/NG-R and CT-F/CT-R), 5 pmol of each Pitcher (NG-P and CT-P1) and 1 pmol of each Catcher (NG-C and CT-C) in buffer containing 2 units of Taq DNA polymerase, 200 μM of dNTPs and 2 mM of MgCl2. The whole PCR steps are as follows: denaturation for 15 min at 95°C and 50 cycles of denaturation (30 s at 95°C), annealing (60 s at 60°C) and extension (30 s at 72°C) in a CFX96 thermocycler (Bio-Rad, USA). Fluorescence signals were detected at annealing step (60°C) and extension step (72°C) at each cycle.
MuDT reaction for linearity of the low Tm target
MuDT reactions were performed in a final volume of 20 μl containing 1 ng of CT genomic DNA, serially diluted NG genomic DNA (1 ng–100 fg), 10 pmol of each primer set (NG-F/NG-R and CT-F/CT-R), 5 pmol of each Pitcher (NG-P and CT-P1) and 1 pmol of each Catcher (NG-C and CT-C) in buffer containing 2 units of Taq DNA polymerase, 200 μM of dNTPs and 2 mM of MgCl2. The whole PCR steps and detection of the fluorescence signals followed the same procedure as the MuDT reaction for detection of the low Tm target.
MuDT reaction and melting curve analysis for detection of sexually transmitted infections
Anyplex™ II STI-7 Detection (Catalog Number: SD7700Y, Seegene, Korea) was used for detection of sexually transmitted infections. Reactions were performed according to the protocol described by the product's manufacturer. Fluorescence signals were detected at annealing step (60°C) and extension step (72°C) at each cycle. Upon completion of the reaction, a melting curve was obtained by cooling the reaction mixture to 55°C, holding at 55°C for 30 s and then increasing the temperature from 55°C to 85°C stepwise (0.5°C/step) with a 5 s hold between each step. Fluorescence was measured continuously during the temperature rises to monitor dissociation of double-stranded DNAs. Melting peaks were derived from the melting curve data (CFX96 manager).
MuDT reaction using a TaqMan probe
MuDT reactions were carried out in the final volume of 20 μl containing 1 ng of each genomic DNA (CT or NG, or both), 10 pmol of each primer set (NG-F/NG-R and CT-F/CT-R), 5 pmol of Pitcher (NG-P), 1 pmol of Catcher (NG-C) and 1 pmol of TaqMan probe (CT-P2), in buffer containing 2 units of Taq DNA polymerase, 200 μM of dNTPs and 2 mM of MgCl2. The whole PCR steps and detection of the fluorescence signals followed the same procedure for the MuDT reaction for the low Tm target.
Calculation of a Ct of the low Tm target
During MuDT reactions, the fluorescence signal was observed at two different detection temperatures (60°C and 72°C) at each cycle. To obtain the Ct for the low Tm target, we first deducted the RFU at 72°C from the RFU at 60°C and subtracted the calculated value by baseline. The ΔRFU between the two detection temperatures (72°C and 60°C) was then plotted after selection of a suitable threshold and then the Ct value was defined as the fractional cycle at which ΔRFU reached the threshold level.
References
Higuchi, R., Fockler, C., Dollinger, G. & Watson, R. Kinetic PCR analysis: real-time monitoring of DNA amplification reactions. Biotechnology (NY) 11, 1026–1030 (1993).
Espy, M. J. et al. Real-time PCR in clinical microbiology: applications for routine laboratory testing. Clin Microbiol Rev 19, 165–256 (2006).
Gunson, R. N., Collins, T. C. & Carman, W. F. Practical experience of high throughput real time PCR in the routine diagnostic virology setting. J Clin Virol 35, 355–367 (2006).
Hoorfar, J. Rapid detection, characterization and enumeration of foodborne pathogens. APMIS Suppl 133, 1–24 (2011).
Maurin, M. Real-time PCR as a diagnostic tool for bacterial diseases. Expert. Rev. Mol.Diagn. 12, 731–754 (2012).
Postollec, F., Falentin, H., Pavan, S., Combrisson, J. & Sohier, D. Recent advances in quantitative PCR (qPCR)applications in food microbiology. Food Micobiol 28, 848–861 (2011).
Hiratsuka, M., Sasaki, T. & Mizugaki, M. Genetic testing for phamacogenetics and its clinical application in drug therapy. Clin. Chimica Acta 363, 177–186 (2006).
Zhong, Q. et al. Multiple digital PCR: breaking the one target per color barrier of quantitative PCR. Lab Chip 11, 2167 (2011).
Settanni, L. & Corsetti, A. The use of multiplex PCR to detect and differentiate food- and beverage-associated microorganisms: a review. J. Microbiol. Methods. Rev. 69, 1–22 (2007).
Fu, G., Miles, A. & Alphey, L. Multiplex detection and SNP genotyping in a single fluorescence channel. Plos One 7, e30340 10.1371/journal.pone.0030340 (2012).
Huang, Q. et al. Multiplex fluorescnece melting curve analysis for mutation detection with dual-labeled, self-quenched probes. Plos One 6, e19206 10.1371/journal.pone.0019296 (2011).
Whilet, D. M & Sloots, T. P. Melting curve analysis using hybridization probes: limitations in microbial molecular diagnostics. Pathol. 37, 254–265 (2005).
Whiley, D. M., Mackay, I. M., Syrmis, M. W., Witt, M. J. & Sloots, T. P. Detection and differentiation ofherpes simplex virus type 1 and 2 by a duplex LightCycler PCR that incorporates an internal control PCR reaction. J. Clin. Virol. 30, 32–38 (2004).
Lee, D. H. TOCE: Innovative Technology for High Multiplex Real-time PCR. Seegene bulletin 1, 5–10 (2012).
Low, A. J. et al. Neisseria gonorrhoeae and Chlamydia trachomatis infection in HIV-1-infected women taking antiretroviral therapy: a prospective cohort study from Birkina Faso. Sex Trasm. infect. 90, 100–103 (2014).
Choe, H. S. et al. Performance of AnyplexTM II multiplex real-time PCR for the diagnosis of seven sexually transmitted infections: comparison with currently available methods. Int. J. Infect. Dis. 17, e1134–1140 (2013).
Ranasinghe, R. T. & Brown, T. Fluorescnece based strategies for genetic analysis. Chem. Commun. 44, 5487–5502 (2005).
Tyagi, S. & Kramer, F. R. Molecular beacons: probes that fluoresce upon hybridizayion. Nature Biotec 14, 303–308 (1996).
Witter, C. T., Herrmann, M. G., Moss, A. A. & Rasmussen, R. P. Continuous fluorescence monitoring of rapid cycle DNA amplification. Bio Techniques 22, 130–138 (1997).
Holland, P. M., Abramson, R. D., Watson, R. & Gelfand, D. H. Detection of specific polymerase chain reaction product by utilizing the 5′ to 3′ exonuclease activity of Thermus aquaticus DNA polymerase. Proc. Natl. Acad. Sci. 88, 7276–7280 (1991).
Acknowledgements
We thank In-Taek Hwang and Dae-Hoon Lee for critical reading of the manuscript. This work was supported by Seegene, Inc.
Author information
Authors and Affiliations
Contributions
Y.L., D.K. and K.L. performed the experiments. J.C. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare competing financial interests: patent application that is related to the research in this paper.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Lee, YJ., Kim, D., Lee, K. et al. Single-channel multiplexing without melting curve analysis in real-time PCR. Sci Rep 4, 7439 (2014). https://doi.org/10.1038/srep07439
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07439
This article is cited by
-
Quantitative multiplex real-time polymerase chain reaction assay for the detection of Helicobacter pylori and clarithromycin resistance
BMC Microbiology (2023)
-
Seasonal epidemiological and clinical characteristics of pediatric patients with human parainfluenza virus infection by serotype: a retrospective study
Virology Journal (2022)
-
Significant Expansion of Real-Time PCR Multiplexing with Traditional Chemistries using Amplitude Modulation
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.