Abstract
The efficacy and mechanisms of therapeutic action are largely described by atomic bonds and interactions local to drug binding sites. Here we introduce global connectivity analysis as a high-throughput computational assay of therapeutic action – inspired by the Google page rank algorithm that unearths most “globally connected” websites from the information-dense world wide web (WWW). We execute short timescale (30 ps) molecular dynamics simulations with high sampling frequency (0.01 ps), to identify amino acid residue hubs whose global connectivity dynamics are characteristic of the ligand or mutation associated with the target protein. We find that unexpected allosteric hubs – up to 20Å from the ATP binding site, but within 5Å of the phosphorylation site – encode the Gibbs free energy of inhibition (ΔGinhibition) for select protein kinase-targeted cancer therapeutics. We further find that clinically relevant somatic cancer mutations implicated in both drug resistance and personalized drug sensitivity can be predicted in a high-throughput fashion. Our results establish global connectivity analysis as a potent assay of protein functional modulation. This sets the stage for unearthing disease-causal exome mutations and motivates forecast of clinical drug response on a patient-by-patient basis. We suggest incorporation of structure-guided genetic inference assays into pharmaceutical and healthcare Oncology workflows.
Similar content being viewed by others
Introduction
The first principles of protein-drug interactions focus on the geometric, chemical and physical properties of the orthosteric binding site – for instance the ATP-binding site for ATP-competitive protein kinase inhibitors1,2,3,4,5. Given allosteric regulation of protein function and emergence of drug resistance mutations outside of ligand-binding sites6,7,8, three complementary pioneering efforts have attempted to characterize molecular interactions from a structural standpoint (without bias of the ligand-binding site). These efforts include analysis of sequence alignments to identify likely pathways of allosteric communication9; studying coupling between distant sites via computing inter-residue structural contacts10,11,12; and molecular dynamics (MD) simulations for mapping coupled conformational changes13.
Past sequence-based methods have relied on thousands of evolutionary-related protein sequences to identify pathways of allosteric communication14. Some of the structure-based methods have truncated residue-residue interaction paths and used large distance thresholds to compensate for rotamer or other considerations15,16. MD simulations are low in throughput, owing to the need for long simulation timescales17. Diverse analytic methods such as root mean square deviation (RMSD) of defined motifs, clique and community patterns, correlation matrices, principal component analysis (PCA), energetic analysis (e.g. WHAM) and mutual information have been developed to decode the simulated MD trajectories18,19. There is a paucity of high-throughput, structure-guided computational assays that can be readily integrated into the emerging array of precision medicine technologies targeting oncology and other complex genetic diseases20,21.
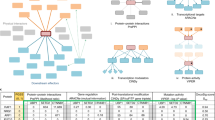
Here we introduce a high-throughput computational method to assay protein functional perturbation via short-timescale MD simulations and global connectivity analysis of the resultant trajectories (see Methods and Figure S1). These perturbations can involve small molecule ligand binding and/or amino acid mutations. In the case of a small molecule ligand of interest, the ligand bound protein crystal structure is considered. This protein-ligand complex is superposed onto an apo state (unbound) reference crystal structure. The ligand is then extracted onto the reference structure and the resulting reference-ligand structural complex is subjected to MD simulations (see Methods). In the case of amino acid mutations, the mutation of interest is computationally-introduced in the relevant position and the mutant protein is then subjected to MD simulations. The MD simulation itself is short-timescale (30 ps) and high sampling rate (0.01 ps). This is to intentionally capture only immediate atomic perturbations (see illustrative Movie S1). These include movements of side-chain and backbone atoms, but exclude pronounced conformational changes (such as significant loop movements) that are generally incorporated into current MD approaches – e.g. DFG-in to DFG-out transitions in kinases17,18,19. Next, we represent every conformation (without any bias from selection of few conformations) as a distinct Connectivity Graph – with residues as nodes, residue-residue contacts as edges and amino acid sequence considerations enabling edge weight assignment. For each graph (conformation) we compute, on a residue by residue basis, centrality measures such as Degree Centrality (Figure 1A) that captures local connectivity and Eigen Centrality (Figure 1B) that captures global connectivity (see Methods)22,23. This process is repeated with multiple random seeds that represent a range of initial atomic velocities to test the reproducibility of the computed results.
Connectivity of protein 3-D structures.
(A.) Degree centrality captures “local connectivity” for each node of a Graph. A caveat of degree centrality is that it considers nodes like “A” and “C” – that are involved in a cluster of densely connected nodes (each hence having a “high degree”) – to be less important than nodes like “D” that are exclusively connected to a large number of “low degree” nodes. (B.) Eigenvector centrality captures “global connectivity” of each node in the Graph – e.g. Node “A” is more important than node “D” – such nodes provide a “pathway” for allosteric communication between different parts of a protein structure such as nodes “B” and “C”. (C.) Each node is colored based on its global connectivity score on a linear scale (white = 0, black = 1). For inactivated (non-phosphorylated) TYK2 bound to ATP, local connectivity of an exemplary conformation has distributed “hubs” with limited discrimination of their relative importance. The Arg-1159 residue has been highlighted by a red box (D.) For the same conformation, global connectivity analysis results in a cluster of tightly coupled hubs. The Arg-1159 residue has been highlighted as a red box (E.) For the same TYK2-ATP conformation, global connectivity measures like Eigen centrality provides a superior ranking scheme compared to local connectivity measures like Degree centrality.
As highlighted here for the Janus kinase (JAK) enzyme family member tyrosine kinase 2 (TYK2), local connectivity emphasizes distributed hubs (Figure 1C) and global connectivity highlights a single cluster of tightly inter-coupled hubs (Figure 1D). The global connectivity measure yields a ranking scheme with better discrimination of amino acid residues compared to the local connectivity measure (Figure 1E, Figure S2). Global connectivity analysis is able to identify functionally important residues with high statistical significance for protein kinases such as TYK2 (p-value = 0.00036; see Methods). We noted a Fractal pattern of global connectivity dynamics for TYK2 bound to ATP – wherein sub-sampling at one tenth of the original sampling rate (i.e. 0.1 ps) retained the relevant information content. Ten different random seeds (representing a range of possible initial atomic velocities for the MD simulation) resulted in similar mean global connectivity measures for the amino acid residues constituting the TYK2-ATP structural complex (Figure S3). These appealing characteristics motivated further investigation of global connectivity analysis with protein kinases as a model system.
Results
ATP binding by a non-phoshorylated kinase increases global connectivity of residues along a path linking the orthosteric active site to the allosteric phosphorylation site
Non-phosphorylated (inactivated) TYK2 enzyme is considered here as an illustrative example. JAK kinase enzymes such as TYK2 (and JAK1, JAK2, JAK3) are popular drug targets for inflammation and oncology indications24,25,26. The reference crystal structure used for this analysis is PDB ID 4GVJ sans the ADP molecule. The ATP bound reference structure is compared with the Apo state structure. The residues Arg-1159, Asp-1023, Ser-1082, Glu-1071, Trp-1067, Ser-1086 and Leu-1024 have higher mean global connectivity in the former over the latter state (Figure S4 – ATP bound; Figure S5 – apo; Figure S6 – contrast). These residues form a contiguous set of residues distributed from a remote allosteric site (abutting the phosphorylation site on TYK2) to the ATP-binding site (and active site) on TYK2. Specifically, every residue other than Asp-1023 are physically distant from the ATP-binding site on TYK2 (Figure S7).
Change in global connectivity of select allosteric residues relate to the binding energy of a set of ATP-competitive orthosteric kinase inhibitors
The residues Arg-1159, Glu-1071 and Ser-1082 constitute an allosteric hub – i.e. high mean global connectivity; (Figure S8) with low variation across conformations (i.e. limited high frequency fluctuations; Figure S9). This allosteric hub is around 20 angstroms from the ATP-binding site, but within 7 angstroms of the Tyr-1054 phosphorylation site on inactivated TYK2 (Figure 2A). This allosteric hub is also highly conserved across Eukaryotic protein kinases (EPKs) and especially across the human kinome (Figure S10)27. The identified allosteric hub was examined for a series of Genentech inhibitors targeting non-phosphorylated TYK224. A correlation was observed between the total global connectivity of residues constituting the allosteric hub and the dissociation constant (Ki) for each compound (Figure S11). This implies that the global connectivity of the allosteric hub is related to the Gibbs Free Energy of Inhibition for this series of compounds (Figure 2B). The global connectivity of the residues constituting the allosteric hub is shown relative to other residues of TYK2 (Figure 2C; highlighted by * for Arg-1159; ∧ for Glu-1071; + for Ser-1082). The highly effective TYK2 inhibitors diminish the global connectivity of the hub residues more significantly.
Global connectivity of residues constituting the allosteric hub of inactive TYK2 captures the potency of TYK2 inhibitors.
(A.) The hub residues Arg-1159 (R1159 - pink), Glu-1071 (E1071 - orange) and Ser-1082 (S1082 - blue) are shown as spheres on inactivated (non-phosphorylated) TYK2 structure (gray cartoon). This hub is about 20 angstroms from ATP (sticks of green – carbon atoms) and around 7 angstroms from Tyr-1054 (Y1054 – yellow). (B.) The mean global connectivity of the hub residues (y-axis) is plotted against the Gibbs Free Energy of Inhibition (ΔGinhibition – x-axis) for a small set of inactive TYK2 inhibitors from Genentech, Pfizer, Merck, Abbott Laboratories and AVEO Pharmaceuticals (Table S1-S2). (C.) Heatmaps (top row) and spectra (bottom row) for TYK2 bound to ATP, cmp 41 (Ki = 240 nM), cmp 1 (Ki = 32 nM) and cmp 19 (Ki = 1.8 nM) – from left to right. The amino acid sequence of TYK2 is reflected on the x-axis of both the heatmap and spectra renderings and the columns corresponding to the hub residues are highlighted as follows – Arg-1159 (*); Glu-1071 (∧); and Asp-1082 (+). For the heatmaps, the y-axis is the conformation number from the MD simulation (ranging from 1 through 300) and the color of each entry is based on the hot rendering for global connectivity (black = none; red = medium; yellow = high; white = maximum). For the spectra, the y-axis is mean global connectivity (range 0 to 1).
Global connectivity analysis as a predictive tool of therapeutic efficacy
The VEGFR-selective inhibitors Tivozanib (Aveo Pharmaceuticals) and Axitinib (Pfizer) – with kinome partition indices ~1 and no cross reactivity to TYK2 – are considered as blinded negative controls28. The potent TYK2 inhibitors cmp6 (Merck; ic50 = 1 nm) and Staurosporine (Ki = 0.1 nm) are considered as blinded positive controls29. For these compounds, the mean global connectivity of the TYK2 allosteric hub – near Tyr-1054 (phosphorylation) site – is an effective indicator of the experimentally determined TYK2 inhibition potential (Figure 3A, Figure 3B). The heatmap and spectra renderings for these compounds shows the more potent TYK2 inhibitors Staurosporine and cmp6 deplete the allosteric hub residues' global connectivity more significantly than the TYK2-agnostic Tivozanib and Axitinib (Figure 3C).
Predictions of positive and negative controls for inactive TYK2 inhibition.
(A.) The test negative controls (VEGFR selective Tivozanib, Axitinib) and positive controls (Merck cmp6; ic50 = 1 nM, Staurosporine; Ki = 1 nM) are examined here along with Genentech's cmp1 (Ki = 32 nM) and cmp35 (Ki = 1.4 nM) for reference. (B.) For each ligand-bound TYK2 structure, the hub residues are highlighted as blue spheres and Tyr-1054 that gets phosphorylated by upstream kinases is shown as yellow spheres. (C.) Heatmaps (top row) and spectra (bottom row) for the apo state of TYK2 as well as TYK2 bound to Tivozanib Axitinib, cmp 6 and staurosporine – from left to right. The amino acid sequence of TYK2 is reflected on the x-axis of both the heatmap and spectra renderings and the columns corresponding to the hub residues are highlighted as follows – Arg-1159 (*); Glu-1071 (∧); and Asp-1082 (+). For the heatmaps, the y-axis is the MD conformation number and the color of each entry is based on the hot rendering for global connectivity (black = none; red = medium; yellow = high; white = maximum). For the spectra, the y-axis is the mean global connectivity.
Information theoretic perspective to the therapeutic inhibition of an enzyme drug target
We use a quantitative measure termed Dispersion (see Methods) to compare protein conformations in the presence of different compounds. Given the increase of global connectivity across non-hub residues for the more potent TYK2 inhibitors (Figure 2C, Figure 3C), we hypothesized that this phenomenon may factor into the mechanism of action for ATP-competitive TYK2 inhibitors. Consistent with this hypothesis, we find that the global connectivity of Arg-1159 and Glu-1071 are depleted and distributed instead to other residues on TYK2 – for the more potent inhibitors (Figure 4A). This is further consistent across multiple algorithms that assess global connectivity23. Quantitatively capturing the Dispersion measure of inactive TYK2 with various compounds shows higher dispersion for potent inhibitors compared to ATP (Figure 4B). This suggests an increased Entropy in the inhibited state of the kinase (over the Apo and ATP-bound states) from an information theoretic standpoint30 (Figure S12, Figure S13).
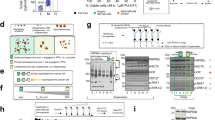
Dispersion as the mechanism of action of Type II kinase inhibitors.
(A.) Each amino acid residue is shown as a node (circle) colored based on its global connectivity with a linear scale (white = 0; black = 1). The ATP-bound TYK2 reference structure and the cmp23-bound TYK2 reference structure are compared herein (cmp23 is an effective TYK2 inhibitor). The Arg-1159 amino acid residue is highlighted as a red square. (B.) higher dispersion for potent inhibitors compared to ATP. (C.) R1159A point mutation reduces the global connectivity of the hub residues whereas randomly selected control mutations R941A have no effect on the hub residues. The Arg-1159 amino acid residue is highlighted as a red square. (D.) Inactivated (non-phosphorylated) BRAF kinase structure bound to ATP is shown. The Arg-704 and Glu-623 residues (blue spheres) are around 20A from the ATP site and within 5A of the Arg-671 methylation site. These residues constitute the hub for BRAF kinase. The Arg-704 residue is highlighted as a red square in the graph rendering.
Contributions of ultra-conserved structural elements towards carcinogenesis
The Arg-Glu salt bridge has been shown to be an integral feature of the Eukaryotic protein kinase fold and a “center hub of connectivity between structurally-conserved kinase motifs” using the phosphorylated Protein Kinase A (PKA) model system27. Mutation of either residue renders the mutants less effective substrates for PKA phosphorylation by upstream kinases, in addition to altering catalytic activity and enzyme regulation27. These experimental results suggest that the Arg-Glu salt bridge is important for the active state of kinases. In non-phosphorylated TYK2, R1159A and E1071A point mutations dampen the global connectivity of the hub compared to other randomly-selected control mutations (Figure 4C, Figure S14). We hypothesize that reducing the global connectivity of the phosphorylation-site (or methylation-site) – leads to destabilization of activation motifs and diminished upstream kinase activity. We suggest that this may be a broader mechanism of Type II kinase inhibitors (that target non-phosphorylated/inactivated kinases). In addition to TYK2, we find that the 1159-1071 equivalent Arg-Glu pair is the global connectivity hub for the serine-threonine oncoprotein kinase BRAF31 in the non-phosphorylated form (Figure 4D). More generically, we suggest that ultra-conserved structural elements (such as the R1159-E1071 pair of TYK2) within each oncoprotein family may be prime candidates for experimental and clinical studies to determine disease causality.
Given that both the illustrative tyrosine kinase (TYK2) and the serine-threonine kinase (BRAF) suggest the same hub in their inactive (non-phosphorylated) states, we consider a third kinase – ABL1 kinase involved in the oncogenic Bcr-ABL gene translocation that has been implicated as causal to leukemia. The goal is to investigate whether all inactive kinases have an identical hub (analogous to the Arg-1159 residue of TYK2). The alternative possibility is that each kinase may have a hub in the vicinity of its phosphorylation site (or other such activation trigger).
Mechanism of mutation induced clinical drug resistance from targeting an oncoprotein kinase
Analysis of Bcr-ABL kinase also provided us the opportunity to analyze the evolution of drug resistance. Here we highlight the T315I gatekeeper drug resistance mutation as an illustrative example. The spectra of wild-type and T315I (gatekeeper) mutant forms of ABL1 kinase bound to ATP as well as a series of inhibitors from Novartis, Pfizer, Ariad and Deciphera pharmaceuticals were analyzed (Figure 5A)32,33,34. This reveals an inactive ABL1 hub constituted of Arg-362, Ser-385 and Ile-360– distinct from the location of hubs for inactive BRAF and TYK2 kinases. As expected, the total global connectivity of the ABL1 hub residues is correlated with the experimentally-measured binding kinetics for inhibitors targeting both wild type and T315I mutant forms (Figure 5B). The Arg-362, Ser-385 and Ile-360 hub is around 15 angstroms from the ATP binding site on ABL1 kinase, but in direct contact with the Tyr-393 residue that is phosphorylated by upstream kinases – which trigger ABL1 kinase activation (Figure 5C). The increase in global connectivity of the hub residues in the ATP-bound T315I mutant state over the ATP-bound wild type state (Figure 5A), suggests the mutant state is more easily activated by upstream kinases. This hints at evolution of drug resistance being opportunistic for the growing tumor, i.e. drug-induced selection of the mutation enables increased oncogenic signaling.
Mechanism of kinase drug resistance.
(A.) The global connectivity spectra is plotted for each ATP-competitive Bcr-ABL kinase inhibitor against both the wild-type (top row) and T315I gatekeeper mutant (bottom row) forms of Bcr-ABL. The amino acid sequence of Bcr-ABL kinase is captured on the x-axis. The hub residues Arg-362 (*) and Ser-385 (∧) are highlighted. Ile-360 of the hub is not highlighted. (B.) Plot of the experimentally measured ic50 (nM) for WT and T315I mutant forms of Bcr-ABL kinase (x-axis) versus the computed net global connectivity of the hub residues (y-axis) is shown for ATP, Bosutinib (Pfizer), Imatinib (Novartis), Ponatinib (Ariad) and Rebastinib (Deciphera pharmaceuticals). (C.) The identified hub on Bcr-ABL kinase is Arg-362, Ser-385 and Ile-360 is highlighted as blue spheres. This hub is over 15 angstroms from the ATP binding site where an inhibitor molecule is shown (pink - carbon atoms). The hub is also over 15 angstroms from the gatekeeper residue (Thr-315 – orange spheres) that is frequently mutated into Ile-315 in several types of drug-resistant cancer. The hub residues are in direct physical contact with Tyr-393 that gets phosphorylated by upstream kinases leading to activation of Bcr-ABL.
Given the distinct hub of Bcr-ABL kinase, we suggest that each oncoprotein harbors a characteristic hub. The reduction of the global connectivity of hub residues may factor into the mechanisms governing therapeutic protein kinase inhibition.
Mechanistic insights into constitutive activity of somatically-mutated oncoprotein
Clinically-observed EGFR kinase mutations that sensitize patients differentially to the inactive (non-phosphorylated) EGFR kinase inhibitors Gefitinib and Erlotinib35,36,37 were analyzed. By comparing the ATP-bound structures of non-phosphorylated (inactive) EGFR kinase and phosphorylated (active) EGFR kinase, we find an increase in global connectivity of Arg-958, Glu-884, Gln-894 and Asp-896 in the active state (Figure 6A). The oncogenic EGFR L858R mutation is one of the most common non small cell lung cancer (NSCLC) somatic mutations, observed from sequencing of both solid tumor biopsies and circulating tumor cells (CTCs). This mutation renders the EGFR kinase constitutively active. The ATP-bound non-phosphorylated EGFR kinase harboring the oncogenic L858R mutation has a global connectivity spectra that correlates with the ATP-bound, phosphorylated EGFR kinase (Figure 6B). The global connectivity of the Arg-958, Glu-884, Gln-894 and Asp-896 amino acid residues are especially prominent.
Mechanisms of kinase inhibitor differential drug sensitization.
(A) non-phosphorylated EGFR + ATP (blue) compared with phosphorylated (active) EGFR + ATP (termed pATP – red); (B) The EGFR L858R mutation is seen to make the connectivity spectra of non-phosphorylated EGFR + ATP closely resemble that of active (phosphorylated) EGFR + ATP (termed pATP). (C) EGFR T790M mutant does not resemble pEGFR unlike the EGFR L858R mutant shown in B. (D–F) increased inhibition potency for Erlotinib against EGFR L858R mutant compared to EGFR WT protein and subsequent decrease in potency for Erlotinib against EGFR L858R + E884K double mutant. (G–I) increased inhibition potency for Gefitinib against EGFR L858R mutant compared to EGFR WT protein and subsequent further increase in potency for Gefitinib against EGFR L858R + E884K double mutant. (J) EGFR L861Q activated more than T790M but less than L858R.
Unlike the L858R mutation, non-phosphorylated EGFR kinase harboring the oncogenic T790M “gatekeeper” mutation is not seen to be a good correlate of the phosphorylated EGFR kinase from their spectra (Figure 6C). These results suggest global connectivity analysis as a useful method of predicting oncoprotein functional state for the somatic cancer mutations.
Structural insights into genetic sensitivity, drug resistance and clinical efficacy of targeted cancer therapy
The differential genetically-induced drug sensitivities for the FDA-approved targeted lung cancer drugs Erlotinib and Gefitinib was examined. Patients harboring the oncogenic EGFR kinase double mutation L858R + E884K have a clearly defined differential clinical response to these drugs36,38. The increased inhibition potency for Erlotinib against the oncogenic EGFR L858R mutant (over the EGFR WT protein) correlates with decreased global connectivity of the hub residues (Arg-958, Glu-884, Gln-894 and Asp-896) in the mutant (Figure 6D – wild type, Figure 6E – L858R mutant). The subsequent decrease in potency for Erlotinib against the oncogenic EGFR L858R + E884K double mutant correlates well with the increased global connectivity of the hub residues (Figure 6D – wild type, Figure 6F – L858R + E884K mutant). On the other hand, the increased inhibition potency for Gefitinib against EGFR L858R mutant (compared to EGFR WT protein) and subsequent further increase in potency for Gefitinib against the EGFR L858R + E884K double mutant correlates well with the increased global connectivity of the hub residues in the mutants (Figure 6G – wild type, Figure 6H – L858R, Figure 6I – L858R + E884K). The oncogenic EGFR L861Q mutation is known to constitutively activate the kinase more than the gatekeeper T790M mutation but less than the oncogenic L858R mutation, in agreement with the total global connectivity of the hub residues on EGFR kinase (Figure 6J)38.
These results suggest that global connectivity analysis may be a useful, high-throughput cancer diagnostic tool to augment the current array of DNA sequencing and sequence-based analytics tools. Mutations such as EGFR E884K are especially important to truly realize the promise of precision medicine. Such mutations that occur in less than 1% of lung cancer patients, emerge as an important determinant of therapeutic efficacy and choice of treatment for these patients. There are literally hundreds of thousands of somatic mutations such as EGFR kinase L858R and E884K that have been recorded from the first thousands of cancer patients whose biopsy genomes have been sequenced and deposited in the COSMIC database. Rare mutations (like EGFR E884K) can clearly impact therapeutic efficacy, but are largely ignored by the majority of current clinical biomarker and companion diagnostics methods39. This underlines the need to develop novel technologies that can identify those genomic alterations that are most likely driving each cancer subtype and acutely influencing drug response.
Structural basis for therapeutic inhibition of a phosphorylated (active) protein kinase
Given the generally improved selectivity of Type II kinase inhibitors over Type I inhibitors that target phosphorylated kinases40, we analyzed the hub residues of the phosphorylated (active) TYK2 state. Examination of the active TYK2 structure via global connectivity analysis suggests an orthosteric hub that includes residues of the DFG motif, the HRD motif and the ATP binding site (Figure 7A). A small set of Type I TYK2 inhibitors from Genentech, Pfizer, Merck and Aveo pharmaceuticals dampen the global connectivity of the orthosteric hub residues (Figure 7B). The total global connectivity of the hub residues also correlates with the experimentally determined Ki measures and the Gibbs free energy of inhibition (ΔGinhibition) measures for these compounds (Figure 7C; Figure S15). The heatmaps and spectral renderings for these compounds also support these observations (Figure 7D).
An orthosteric hub dictates potency of inhibitors targeting phosphorylated (activated) TYK2.
(A.) An exemplary conformation of the ATP-bound structure of pTYK2 highlighting the orthosteric hub of Phe-1042, Gly-1043, Lys-930 and His-1021 (blue spheres) is shown. An exemplary conformation of the Apo (unliganded) state of pTYK2 shows a general shift of the global connectivity towards the N-lobe, within the ATP-site proximal area (blue spheres). The potent inhibitor (Genentech cmp33 with pTYK2 Ki of 0.4 nM) bound pTYK2 structure that sees a complete shift of global connectivity away from the ATP-site (blue spheres). (B.) The high global connectivity of the ATP-bound pTYK2 structure in the orthosteric hub (red box) is shown (in the top graph), in sharp contrast with the fully depleted global connectivity of the hub (red box) for the Genentech cmp33 bound pTYK2 structure (in the bottom graph). (C.) Free energy of inhibition (ΔGinhibition) determined from the experimentally-determined dissociation constant (Ki) measures of a series of pTYK2 inhibitors from Pfizer, Genentech and AVEO pharmaceuticals (Table S3) is plotted on the x-axis (in KJ/mol) with the computed global connectivity of the orthosteric hub (on the y-axis) for each compound-pTYK2 complex. A near linear relationship is observed between these measures. (D.) Global connectivity heatmaps (top row) and spectra (bottom row) for pTYK2 with ATP and inhibitor molecules is shown, highlighting the hub residues His-1021 (+) and Phe-1042 (*). Gly-1043 of the pTYK2 hub is not highlighted for clarity.
These results highlight the predictive potential of global connectivity analysis across diverse kinases and distinct modes of inhibition. We suggest that the switch from the phosphorylation site proximal allosteric hub (in the inactivated state) to the active site proximal orthosteric hub (in the activated state) correlates with TYK2 enzyme activation in cells (Figure S16). Given that allosteric inhibitors are generally associated with desirable selective pharmacology41 but are challenging to develop, rationally engineering orthosteric inhibitors that dampen the global connectivity of allosteric hub residues may present a new approach to achieve selective Type I inhibitors (Figure S17).
Discussion
The method introduced in this study relies on capturing global connectivity modulation during a short timescale (~few tens of picoseconds) interaction between small molecule ligands (such as drug candidate compounds) and their target protein 3-D structure. It is utterly impossible to reveal the entire ensemble of conformational dynamics for a drug-protein interaction by a 30 picosecond MD simulation. While what is revealed in this study is probably related to the initial adaptation of the protein target to the bound ligand, it is very surprising that global connectivity analysis is able to capture “information” indicative of drug efficacy from such poorly sampled MD trajectories of protein kinases. As an alternative to simulating several hundreds of nanoseconds of MD simulation to even partially explore the conformational dynamics of a protein 3-D structure of interest, an alternate approach is to sample different initial conformations of the target protein bound to the ligand of interest using short-timescale MD simulations and global connectivity analysis. Such an approach may reveal a more comprehensive picture regarding how protein structure encodes orthosteric function and allosteric regulation.
Despite the caveat of MD simulations, the high-throughput nature of the approach presented in this study suggests the promise of mechanistically-driven virtual screening in the Cloud (Figure S18). Incorporating the effects of possible single point and combinations of “synthetic mutations” into analyses of the therapeutic potency for each compound takes us a step closer to “precision medicine”. Incorporating such analytics into the earliest stages of drug discovery may enable the structure-based selection of compounds that can be strong candidates for promotion into preclinical and clinical settings. Such an approach presents a strong complement to current emphasis on toxicology and intellectual property as the chief determinants of promoting compounds through a pharmaceutical pipeline. Selecting compounds with the least potential for clinical drug resistance probability offers the enticing possibility of increasing progression free survival for patients suffering from mutation-prone, complex diseases such as cancer.
Across diverse disciplines, connectivity analysis via centrality measurements have provided remarkable insights about how the “structure” of a Graph encodes “function” via its constituent nodes23,42. For instance, design of telecommunications switching networks with certain fault tolerance/resilience properties share several connectivity characteristics similar to the protein structures we analyze in this study. A nationwide telecom network may have several hubs (such as New York City, Dallas, Denver, Los Angeles etc.) that aggregate traffic from local sources (such as NJ, CT etc.). The fault tolerant performance of such a network (typically quantified as the “economic value” lost due to failure of one or more switching nodes) is usually determined by the hubs that aggregate several “dense local networks” (e.g. NY aggregating NJ, CT traffic) as opposed to hubs whose fan-out (degree) may be large but mostly sparse “local networks” (e.g. Dallas – several south/central states are aggregated in the Dallas node). Such a network when modeled as a graph will have NY as one of the highest ranking nodes compared to Dallas albeit the NY node in the graph having a possibly lower degree than the Dallas node. An outage in the NY node that is serving several other dense clusters (such as NJ, CT and also has important transatlantic connections) does far more economic damage than an outage in Dallas. Like the global connectivity analysis of the protein structures we cover in this paper, the Principal Eigenvector of the Telecom network graph will show a higher rank for NY as compared to Dallas. This analogy to telecommunications switching networks motivates how the global connectivity analysis introduced in this study may be superior to the current protein structure analysis benchmarks (e.g. local connectivity analysis such as select hydrogen bonds linking an enzyme catalytic site to the substrate molecule being considered as a key determinant of binding affinity and kinetics).
More broadly, in the field of structural molecular biology, a majority of past theoretical efforts to understand protein-ligand interactions – using measures such as dissociation constants (Ki, pKi), concentration benchmarks (ic50) and energies (e.g. Gibbsfree energy of binding – ΔGbinding) – have focused on local connectivity (e.g. few Hydrogen bonds or Van der Waals contacts) of the binding or interface region. In such cases, the importance of an amino acid residue has been obtained by accounting almost exclusively for residues that share a direct interaction or contact with it (Figure 1A). This is in sharp contrast to the all-pervading internet that routinely employs Global connectivity algorithms such as Larry Page's Google Page Rank22 and Jon Kleinberg's Hubs and Authorities43 to search and retrieve the most “important” sources from an information-dense World Wide Web (WWW).
Recently, there has been an emerging interest in exploring the applications of Google page rank like algorithms and elastic network computations to study biochemical networks44,45. There have also been studies that break down ligand-protein interaction energies into a residue-by-residue basis, to gain more insights into how interaction energies are distributed across a protein 3-D structure46,47. Future analysis of protein-ligand and protein-protein complexes via Global connectivity analysis has the potential to reveal intricacies of intramolecular communication as well as mechanisms underlying mutational pathogenesis.
Methods
Reference structures for the TYK2, BRAF, Bcr-ABL and EGFR protein kinases
Chemically-diverse ATP-competitive small molecules have been developed against both activated (phosphorylated) TYK2, as well as inactivated (non-phosphorylated) TYK2 (Tables S1-S3). The TYK2 structures with PDB IDs 4GVJ (non-phosphorylated) and 3LXN (phosphorylated) are selected as references with their ligands deleted. Protein superposition is used to extract the ligands of interest, as explained earlier. The resulting TYK2-ligand co-complex structures are considered for further analysis. The structures with PDB ID 3UE4, 3W32 and 2GS6 are used as reference structures for extracting Bcr-ABL, non-phosphorylated EGFR and phosphorylated EGFR kinase inhibitors respectively.
Preparation of the protein-ligand co-complex structure for molecular dynamics simulation
Each co-complex structure is edited to remove any crystallographic artifacts from the original crystal of the target protein. Specifically, missing side chains are modeled back in, unresolved loops are modeled back in and solvent molecules as well as ions are deleted. The edited co-complex is then typed with the CharmM force field. The fully prepared and typed protein-ligand co-complex 3-D structure is subjected to energy minimization with the Smart Minimizer algorithm of Discovery Studio 4.0.0.13259 (up to 4000 iterations or an RMS gradient of 0.0001 angstroms). The resulting optimized initial pose for the co-complex structure is used for further analysis. All computation was carried out over the Amazon Web Services EC2 Cloud platform.
Short-timescale high sampling rate MD simulations
To avoid biases from crystallographic resolution, rotamer-possibilities for side-chains and other related caveats of these “static” kinase-ligand 3-D structural models, MD simulations were performed on the co-complex structures. Each co-complex structure was solvated with the Generalized Born with Simple Switching (GBSW) model. The CharmM force field was used to type this system. The system was subjected to an unconstrained Molecular Dynamics (MD) cascade – starting with a heating phase of 4 picoseconds, time step of 2 femtoseconds, initial temperature of 50 Kelvin, target temperature of 300 Kelvin and non-bonded list radius of 14 angstroms. The next step of the MD cascade is the equilibration phase of 30 picoseconds duration, time step of 2 femtoseconds and target temperature of 300 Kelvin. Each system is noted to have equilibrated post this step, by ensuring that the net kinetic energy oscillates about a mean energy value for the period of observation. In the post-equilibration production phase of 30 picosecond duration, the following parameters are set – time step of 2 fs and target temperature of 300 Kelvin. A high sampling frequency of 0.01 ps per conformation is used to obtain a total of 3000 sampled conformations for the kinase-ligand co-complex during the 30 ps production phase. The dynamics cascade is carried out in Discovery Studio 4.0.0.13259 over the Amazon Web Services EC2 Cloud platform. Every one of the sampled conformations of the MD trajectory is considered. All ligands are modeled as uncharged/neutral (including ATP) without considering any ions (such as Mg2+).
“Digitizing the atomic contact space” to compute a Graph capturing each 3-D conformation
Next we “digitize” the atomic contact space for each computed protein conformation. For this purpose, we consider the set of oxygen, nitrogen, sulfur atoms, i.e. {O, N, S} – termed Atoms of Interest (AOI) – constituting each protein structural conformation. The reason for this choice is that only these three atoms showed maximal shift in net root mean square deviation (RMSD) during the course of the short time scale (30 ps) simulation, whereas Carbon (C) atoms did not have any significant movements. Including C atoms – that is, considering all heavy atoms – was further seen to be detrimental to observing significant shifts in global connectivity for different ligands and mutations. Thus, only the polar {O, N, S} atoms were considered for further analysis.
A set of Python-based programs were developed for these computations and automated using PiCloud. The Gephi package was used to visualize graphs as nodes and edges (http://gephi.org/), the networkx package used for graph computations (https://networkx.github.io/) and the matplotlib package was used for heatmap computations (http://matplotlib.org/). Each of the computed 3D protein structural conformations is represented as a Graph where the vertices are residues (amino acids) of the structure. For a protein of N amino acid residues, there will be N nodes in the graph. An edge (or “connection”) exists between any two residues (vertices) in this graph if there is at least one AOI in one residue within a Euclidean distance threshold of 4.5 angstroms from at least one AOI on the second residue. Pairs of residues harboring such contacts between AOI are termed to be “connected” to each other. Two distinct Graphs were constructed for each conformation: one to capture the discretized atomic interactions mentioned above (Connectivity Graph) and the other to capture the physical distance between the Alpha Carbon atoms of amino acids (Distance Graph). The two graphs have identical structure (same number of nodes and the edges that connect the nodes) but with different edge weights. The edge weight assignment for both these Graphs are described below.
For the Connectivity Graph, the “weight” of each residue-residue edge (or “connection”) is determined using a combination of location based edge weight assessment and sequence based edge weight assessment. For location based edge weight assessment, AOI-AOI contacts are weighted 16:4:1 depending on whether they are of the sidechain-sidechain, sidechain-backbone or backbone-backbone nature respectively. For sequence based edge weight assessment, contacts between AOI on two residues that are twelve or more residues apart in the sequence space are weighted 4 times more than other AOI contacts. The total edge weight between a pair of residues (nodes) in the protein (graph) is given by the sum of individual weights calculated between the “set of all considered AOI” on the residue pair. For each of the conformations obtained from the MD simulation of the target protein, one Connectivity Graph is determined as described above that captures the “digitized” information of that 3-D protein conformation. Hence, there are a total of 3000 such Connectivity Graphs that are computed – each corresponding to one of 3000 conformations that together constitute the entire MD trajectory of the target protein.
For the Distance Graph, the “weight” of each residue-residue edge (or “connection”) is just the physical distance (in Angstroms) of the Alpha Carbon atoms between the two residues.
Computing the Global Connectivity Temporal Function for each amino acid residues
A NxN adjacency matrix representation of the Connectivity Graph (Connectivity Matrix) is used for computing the global connectivity of each amino acid residue. The Connectivity Matrix is positive semi-definite and hence guaranteed to have an Eigenvector corresponding to the maximum positive Eigenvalue (Principal Eigenvector). Furthermore, all components (each component representing a certain amino acid residue of the protein conformation) of the Principal Eigenvector are positive and constitute the ranking of the amino acid residues. For example, in a TYK2 protein kinase, the global connectivity of Arg-1159 is given by the component corresponding to that residue in the Principal Eigenvector. The Principal Eigenvector is computed for each of the 3000 matrices/graphs obtained from the MD simulation, resulting in the Global Connectivity Temporal Function.
Computing mean global connectivity for each residue on the target protein kinase
The Fast Fourier Transform (FFT) of the Global Connectivity Temporal Function is computed for all N amino acid residues of the target protein. This results in N Fourier-transformed frequency domain functions. Each of these function's DC (zero frequency) values provide the mean (average) global connectivity for the N amino acid residues constituting the target kinase. The higher order harmonics are captured as follows: FFT1 (first harmomic), FFT2 (second harmonic), FFT3 (third harmonic), FFT4 (fourth harmonic), etc.
Identifying “hub” residues on target protein kinase with consistently high global connectivity
The residues with consistently high global connectivity throughout the MD trajectory are identified for the natural/endogenous ligand (ATP), provided the same residues do not have high global connectivity in the apo state (hub residues). The hub residues are obtained by subtracting the global connectivity temporal function of the ATP-bound state from the global connectivity temporal function of the apo state of the kinase.
Estimation of inhibitor potency for each ATP-competitive kinase inhibitor
The potency of inhibition for each drug candidate is estimated by comparing the mean measures of the identified “hub” residues for the natural ligand with the corresponding measures for the drug candidate. Those drug candidates that elicit relatively more reduced mean measures for the selected hub residues are found to be more potent inhibitors of the target kinase. Conversely, those drug candidates that elicit a relatively less pronounced reduction of mean measures for the selected hub residues are found to be less effective inhibitors.
Estimation of the Dispersion of global connectivity for each compound
The Floyd's algorithm is used on the Distance Graph to compute the shortest path between all the residue pairs of the protein conformation ( paths). This information is used along with the global connectivity obtained for each amino acid residue (node weight) to compute a path score for each such shortest path comprising a certain set of nodes in the Distance Graph. The path score is simply the sum of the weights of the nodes that constitute a shortest path. The physical intuition here is that the forces are transmitted across the shortest paths and all possible shortest paths are computed (in order to help us reason about the entire conformation in a holistic fashion). The Area under the curve of the plot formed by arranging path scores in decreasing order for all possible paths provide the measure of skewness or Dispersion (Figure S12 is an illustration of such a plot). Figure 4B is an example that shows the Area under the curve measure for multiple compounds. Note that the aggregate scalar Dispersion provides a quantitative measure for the visualization of Global Connectivity distribution (e.g. plotted as a graph in Figure 4A, with darker nodes having larger Global Connectivity). A higher value of Dispersion is equivalent to a graph visualization that has a larger number of dark(er) spots compared to a lower value of Dispersion. Both the scalar Dispersion measure and the global connectivity distribution capture the same information for a given protein structure.
paths). This information is used along with the global connectivity obtained for each amino acid residue (node weight) to compute a path score for each such shortest path comprising a certain set of nodes in the Distance Graph. The path score is simply the sum of the weights of the nodes that constitute a shortest path. The physical intuition here is that the forces are transmitted across the shortest paths and all possible shortest paths are computed (in order to help us reason about the entire conformation in a holistic fashion). The Area under the curve of the plot formed by arranging path scores in decreasing order for all possible paths provide the measure of skewness or Dispersion (Figure S12 is an illustration of such a plot). Figure 4B is an example that shows the Area under the curve measure for multiple compounds. Note that the aggregate scalar Dispersion provides a quantitative measure for the visualization of Global Connectivity distribution (e.g. plotted as a graph in Figure 4A, with darker nodes having larger Global Connectivity). A higher value of Dispersion is equivalent to a graph visualization that has a larger number of dark(er) spots compared to a lower value of Dispersion. Both the scalar Dispersion measure and the global connectivity distribution capture the same information for a given protein structure.
Statistical testing of ranking each residue's relative importance using global connectivity
Let us consider a protein structure of N amino acid residues where T is the number of top ranked amino acids considered as a suitable threshold for the effectiveness of prediction. Let us consider the following different cases:
Case #1 – Assuming only one site is functionally-relevant (e.g. ATP binding site)
Let us consider the Event where at least one among k amino acids of a functionally-important site (e.g., ATP binding site) are identified among the top T ranked amino acids:


(An effective ranking scheme, by definition, requires that T is much smaller than N)
Case #2 – Assuming two sites are both functionally-important (e.g. ATP-binding, active sites)
Let us consider the case where k1 and k2 are the number of amino acids in sites 1 and 2 respectively. The Event may be defined as finding at least one amino acid from each site (of ki amino acids) in the Top T ranked residues. Note that this is a stringent requirement that both sites are represented in the Top ranked residues. Here ki is the event that at least 1 of the amino acid residues in the ith site is in the Top ranked residues. As an example, P(k1) is the probability that at least one amino acid residue among k1 amino acid residues are found within the top T residues.

Using Equation 1 in Equation 2 yields:

Case #3 – Assuming three sites are all functionally-important sites (e.g. ATP-binding site; Active site, Phosphorylation site)
Let us consider the case where k1, k2 and k3 are the number of amino acids in sites 1, 2 and 3 respectively (with k1 + k2 + k3 < T). The Event may be defined as finding at least one amino acid from each site (of ki amino acids) in the Top T ranked residues. Here ki is the event that at least 1 of the amino acid residues in the ith site is in the Top ranked residues. From the well-known Inclusion-Exclusion principle:

Using equation (2) in equation (4) and rearranging the terms yields:






In a similar fashion, based on principle of inclusion-exclusion, one can compute the p-value for any number of functionally-important sites, etc.
References
Ringe, D. & Petsko, G. How Enzymes Work. Science 320, 1428–29 (2008).
Gilson, M. K. & Zhou, H.-X. Calculation of protein-ligand binding affinities. Ann. Rev. Biophys. Biomol. Struct. 36, 21–42 (2007).
Ciulli, A., Williams, G., Smith, A. G., Blundell, T. L. & Abell, C. Probing Hot Spots at Protein-Ligand Binding Sites: A Fragment-Based Approach Using Biophysical Methods. J. Med. Chem. 49, 4992–5000 (2006).
Singh, N. & Warshel, A. Absolute binding free energy calculations: On the accuracy of computational scoring of protein–ligand interactions. Prot: Struct, Func and Bioinfo. 78, 1705–1723 (2010).
Huang, D., Lafleur, K., Nevado, C. & Caflisch, A. Kinase selectivity potential for inhibitors targeting the ATP binding site: a network analysis. Bioinformatics 26, 198–204 (2010).
Szilágyi, A., Nussinov, R. & Csermely, P. Allo-network drugs: extension of the allosteric drug concept to protein- protein interaction and signaling networks. Curr Top Med Chem. 13, 64–77 (2013).
Cui, Q. & Karplus, M. Allostery and cooperativity revisited. Prot. Sci. 17, 1295–1307 (2008).
Changeux, J.-P. & Edelstein, S. J. Allosteric Mechanisms of Signal Transduction. Science 308, 1424–1428 (2005).
Süel, G. M., Lockless, S. W., Wall, M. A. & Ranganathan, R. Evolutionarily conserved networks of residues mediate allosteric communication in proteins. Nat. Struct. Mol. Biol. 10, 59–69 (2002).
Venkatakrishnan, A. J. et al. Molecular signatures of G-protein-coupled receptors. Nature 494, 185–194 (2013).
Weinkam, P., Pons, J. & Sali, A. Structure-based model of allostery predicts coupling between distant sites. Proc. Nat. Acad. Sci. 109, 4875–4880 (2012).
Cheng, T. M. K., Lu, Y.-E., Vendruscolo, M., Lio, P. & Blundell, T. L. Prediction by Graph Theoretic Measures of Structural Effects in Proteins Arising from Non-Synonymous Single Nucleotide Polymorphisms. PLoS Comput Biol. 4, e1000135 (2008).
Weiss, D. R. & Levitt, M. Can Morphing Methods Predict Intermediate Structures? J. Mol. Biol. 385, 665–674 (2009).
Morcos, F. et al. Direct-coupling analysis of residue coevolution captures native contacts across many protein families. Proc. Nat. Acad. Sci. 108, 1293–1301 (2011).
Vishveshwara, S., Brinda, K. V. & Kannan, N. Protein Structure: Insights from Graph Theory. J. Theor. Comput. Chem. 1, 87 (2002).
Soundararajan, V. et al. Networks link antigenic & receptor-binding sites of influenza hemagglutinin: Mechanistic insight into fitter strain propagation. Sci. Rep. 1, 200 (2011).
Shan, Y., Arkhipov, A., Kim, E. T., Pan, A. C. & Shaw, D. E. Transitions to catalytically inactive conformations in EGFR kinase. Transitions to catalytically inactive conformations in EGFR kinase. Proc. Nat. Acad. Sci. 110, 7270–7275 (2014).
Shukla, D., Meng, Y., Roux, B. & Pande, V. S. Activation pathway of Src kinase reveals intermediate states as targets for drug design. Nat. Comm. 5, 3397 (2014).
Ghosh, A., Vishveshwara, S. Variations in Clique and Community Patterns in Protein Structures during Allosteric Communication: Investigation of Dynamically Equilibrated Structures of Methionyl tRNA Synthetase Complexes. Biochem J. 47, 11398–11407 (2008).
Shrager, J. & Tenenbaum, J. M. Rapid learning for precision oncology. Nat Rev Clin Oncol. 11, 109–118 (2014).
Hu, R., Wang, X. & Zhan, X. Multi-parameter systematic strategies for predictive, preventive and personalised medicine in cancer. EPMA J. 4, 2 (2013).
Brin, S. & Page, L. The anatomy of a large-scale hypertextual Web search engine. Comp Net and ISDN Sys. 33, 107–17 (1998).
Langville, A. & Meyer, C. Google's Page Rank and Beyond: The Science of Search Engine Rankings (Princeton University Press, New Jersey, 2006).
Liang, J. et al. Lead identification of novel and selective TYK2 inhibitors. Euro J Med Chem. 67, 178–187 (2013).
Argiriadi, M. A. et al. Enabling structure-based drug design of Tyk2 through co-crystallization with a stabilizing aminoindazole inhibitor. BMC Struct Biol. 12, 22 (2012).
Chrencik, J. E. et al. Structural and Thermodynamic Characterization of the TYK2 and JAK3 Kinase Domains in Complex with CP-690550 and CMP-6. J. Mol. Biol. 400, 413–433 (2010).
Yang, J. et al. A Conserved Glu–Arg Salt Bridge Connects Coevolved Motifs That Define the Eukaryotic Protein Kinase Fold. J. Mol. Biol. 415, 666–679 (2012).
McTigue, M. et al. Molecular conformations, interactions and properties associated with drug efficiency and clinical performance among VEGFR Tyrosine Kinase inhibitors. Proc. Nat. Acad. Sci. 109, 18281–18289 (2012).
Tsui, V. et al. A new regulatory switch in a JAK protein kinase. Proteins 79, 393–401 (2011).
Shannon, C. E. A Mathematical Theory of Communication. Bell Sys Tech J. 27, 379–423 (1948).
Gould, A. E. et al. Design and Optimization of Potent and Orally Bioavailable Tetrahydronaphthalene Raf Inhibitors. J. Med. Chem. 54, 1836–1846 (2011).
Pritchard, J. P., Lauffenburger, D. A. & Hemann, M. T. Understanding resistance to combination chemotherapy. Drug Resist Updat. 15, 249–257 (2012).
Panjarian, S., Iacob, R. E., Chen, S., Engen, J. R. & Smithgall, T. E. Structure and Dynamic Regulation of Abl Kinases. J. Biol. Chem. 288, 5443–5450 (2013).
Tanneeru, K. & Guruprasad, L. Ponatinib Is a Pan-BCR-ABL Kinase Inhibitor: MD Simulations and SIE Study. PLoS ONE 8, e78556 (2013).
Guillermo Paez, J. et al. EGFR Mutations in Lung Cancer: Correlation with Clinical Response to Gefitinib Therapy. Science 304, 1497–1500 (2004).
Kobayashi, S. et al. EGFR Mutation and Resistance of Non–Small-Cell Lung Cancer to Gefitinib. N Engl J Med 352, 786–792 (2005).
Dowell, J. E. & Minna, J. D. Chasing Mutations in the Epidermal Growth Factor in Lung Cancer. N Engl J Med 352, 830–832 (2005).
Tang, Z. et al. Disruption of the EGFR E884-R958 ion pair conserved in the human kinome differentially alters signaling and inhibitor sensitivity. Oncogene 28, 518–33 (2009).
Eisenstein, M. Foundation medicine. Nat Biotech. 30, 14 (2012).
Liu, Y. & Gray, N. S. Rational design of inhibitors that bind to inactive kinase conformations. Nat. Chem. Biol. 2, 358–364 (2006).
Endicott, J. A., Noble, M. E. M. & Johnson, L. N. The Structural Basis for Control of Eukaryotic Protein Kinases. Ann. Rev. of Biochem. 81, 587–613 (2012).
Barabási, A.-L. Scale-Free Networks: A Decade and Beyond. Science 325, 412–413 (2009).
Kleinberg, J. Authoritative sources in a hyperlinked environment. J. of ACM 46, 604–632 (1999).
Hudelson, M., Mooney, B. L. & Clark, A. E. Determining polyhedral arrangements of atoms using PageRank. J. Math. Chem. 50, 2342–2350 (2012).
Yang, Z., Májek, P. & Bahar, I. Allosteric Transitions of Supramolecular Systems Explored by Network Models: Application to Chaperonin GroEL. PLoS Comp. Biol. 10.1371/journal.pcbi.1000360 (2009).
Tominaga, Y. & Jorgensen, W. L. General model for estimation of the inhibition of protein kinases using Monte Carlo simulations. J Med Chem. 47, 2534–49 (2004).
McGillick, B. E., Balius, T. E., Mukherjee, S. & Rizzo, R. C. Origins of resistance to the HIVgp41 viral entry inhibitor T20. Biochem. J. 49, 3575–92 (2010).
Acknowledgements
The authors thank Paul Fatheree, Jim Janc, Mathai Mammen (Theravance – San Francisco CA); Jared Toettcher (Princeton University – Princeton NJ); Ashwin Murali (Harvard Medical School – Boston MA); Justin Rettenmaier (University of California San Francisco – San Francisco CA); Ahmed Samatar (Merck – Boston MA); Mikhail Shapiro (California Institute of Technology – Pasadena CA); Mark Murcko (Schrödinger – Cambridge MA); Ken Butenhof (Accelrys/Biovia – San Diego CA); Kim Robasky (Quintiles – Rayleigh-Durham NC); Phillip Snyder, Alice Tsai, Mac Johnson and Mark Namchuk (Vertex Pharmaceuticals – Boston MA) for interesting discussions. The authors thank Ken Elkabany and Aaron Staley (PiCloud – San Francisco CA), Tina Yeh (Accelrys/Biovia – San Diego CA) and Salma Rafi (Schrödinger – Cambridge MA) for excellent technical support. The authors acknowledge financial support from the angel investors of Lumen Biomics.
Author information
Authors and Affiliations
Contributions
V.S. and M.A. envisioned the study, performed the experiments and analyzed the data. V.S. performed the structural studies and wrote the manuscript. M.A. formulated the theoretical framework, developed all the metrics, engineered all the computer programs and performed the relevant statistical analysis. V.S. and M.A. reviewed the data as well as the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information File
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Soundararajan, V., Aravamudan, M. Global connectivity of hub residues in Oncoprotein structures encodes genetic factors dictating personalized drug response to targeted Cancer therapy. Sci Rep 4, 7294 (2014). https://doi.org/10.1038/srep07294
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07294
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.