Abstract
Mitochondrial proteins are coded by nuclear (nDNA) and mitochondrial (mtDNA) genes, implying a complex cross-talk between the two genomes. Here we investigated the diversity displayed in 104 nuclear-coded mitochondrial proteins from 1,092 individuals from the 1000 Genomes dataset, in order to evaluate if these genes are under the effects of purifying selection and how that selection compares with their mitochondrial encoded counterparts. Only the very rare variants (frequency < 0.1%) in these nDNA genes are indistinguishable from a random set from all possible variants in terms of predicted pathogenicity score, but more frequent variants display distinct signs of purifying selection. Comparisons of selection strength indicate stronger selection in the mtDNA genes compared to this set of nDNA genes, accounted for by the high hydrophobicity of the proteins coded by the mtDNA. Most of the predicted pathogenic variants in the nDNA genes were restricted to a single continental population. The proportion of individuals having at least one potential pathogenic mutation in this gene set was significantly lower in Europeans than in Africans and Asians. This difference may reflect demographic asymmetries, since African and Asian populations experienced main expansions in middle Holocene, while in Europeans the main expansions occurred earlier in the post-glacial period.
Similar content being viewed by others
Introduction
The genomic era is enabling the production of massive datasets (such as The Cancer Genome Atlas1) created by large consortia focused on addressing the genetics behind complex diseases. The disease oriented projects have been complemented by consortia aiming to characterize the diversity of general human populations, such as HapMap2 and 1000 Genomes3. These extensive datasets are providing valuable evidence about the global frequency of single nucleotide variants or polymorphisms (SNVs or SNPs), including those variants that have been reported to cause disease as well as those merely predicted to be pathogenic. For instance, the analysis of the pilot phase of the 1000 Genomes Project estimated that on average each person carries approximately 250 to 300 loss-of-function SNVs (LOF) in annotated genes and 50 to 100 SNVs previously implicated in inherited disorders and that the rate of de novo germline base substitution mutations is approximately 10−8 per base pair per generation3. An update on the 1000 Genomes project4 revealed numbers that are even more astounding: individuals typically carry more than 2,500 non-synonymous variants at conserved positions, 20–40 variants identified as damaging at conserved sites and about 150 LOFs (stop-gains, frameshift indels in coding sequence and disruptions to essential splice sites). However, as most of these variants are common (>5%) or low-frequency (0.5–5%), their pathological potential can be questioned. If only the rare (<0.5%) variants are considered as potentially pathological, as those authors recommend, the numbers decrease: 130–400 non-synonymous variants per individual, 10–20 LOF variants, 2–5 damaging mutations and 1–2 variants identified previously from cancer genome sequencing. Another work focusing in 114 genes selected by an expert panel for their association with medically actionable genetic conditions possibly undiagnosed in adults showed that among the 1,000 exomes surveyed, a total of 585 instances of 239 unique variants were identified as disease causing in the Human Gene Mutation Database, of which 16 unique autosomal-dominant variants in 17 individuals were assessed to be pathogenic or likely pathogenic and one participant had two pathogenic variants for an autosomal-recessive disease5.
This considerable amount of potentially pathogenic variants in the general population is a consequence of the recent exponential expansion of the population size displayed by all current human populations6, leading to an excess of rare genetic variants, which are estimated to be quite young in the timescale of the modern human evolution. In fact, Fu et al.7, by using a simulation approach to generate series of coalescent trees for specific demographic models (having the limitation of a very high variance of the age estimate for each SNV), showed that approximately 73% of all protein-coding SNVs and approximately 86% of SNVs predicted to be deleterious arose in the past 5,000–10,000 years and that disease genes contained a significantly higher proportion of recently arisen deleterious SNVs than other genes. Another very important genomic signature driven by past demography is the fact that Europeans have an excess of deleterious variants in essential and Mendelian disease genes compared to Africans5,7,8 matching a weaker purifying selection due to the out-of-Africa dispersal around 60 thousand years ago9. Dorschner et al.5 quantified frequencies of ~3.4% for European descent and ~1.2% for African descent of the high-penetrance pathogenic SNVs in adults. Lohmueller et al.8, by performing exon resequencing, described that for SNPs segregating only in European-Americans (EAs) or in African-Americans (AAs), the proportion of non-synonymous SNPs was significantly higher in the EA sample (55.4%) than in the AA sample (47.0%; p < 2.3 × 10−37); as well as for SNPs that are inferred to be ‘probably damaging’ (15.9% in EA; 12.1% in AA; p < 3.3 × 10−11); and that in general, EAs have more homozygous genotypes for synonymous, non-synonymous and for the damaging allele at ‘probably damaging’ SNPs than AAs, while AAs have more heterozygous genotypes for all classes of SNPs. However, Boyko et al.10 found that the distributions of fitness effects of new mutations do not differ between AAs and EAs, although the alleles segregating in EAs were on average more deleterious than those segregating in African Americans consistent with Lohmueller et al.8. Abecasis et al.4 confirmed that low-frequency variants show substantial geographic differentiation and advanced that this is further increased by the action of purifying selection (whose strength is driven by evolutionary conservation and coding consequence) and that rare-variant load varies substantially across biological pathways.
In the context of complex diseases, mitochondrial disorders epitomize the complexity as they result from the interplay between the mitochondrial (mtDNA) and nuclear (nDNA) genomes, with each genome using different models of transmission and even genetic codes (in the human mtDNA genetic code UGA codes for tryptophan and not for termination; AUG codes for methionine and not for isoleucine; AGA and AGG do not code for arginine and were once thought to be termination codons11; AUA and AUG are both initiation codons)12. The maternally non-recombining human mtDNA is a double-stranded circular molecule of ~16.6-kilobase (kb), coding for 13 essential components of the respiratory chain (seven subunits of complex I - NADH–ubiquinone oxidoreductase; one of complex III subunit – ubiquinol cytochrome c reductase; three of the complex IV - cytochrome c oxidase; and two of complex V - ATP synthase), two ribosomal RNA genes and 22 transfer RNA genes. The majority of the mitochondrial respiratory chain polypeptides are coded by biparental inherited nuclear genes, as also happens for many proteins important for maintaining the mtDNA, such as the mtDNA polymerase γ (POLG1), the helicase-like protein Twinkle, the TP, TK, DGK and ANT1 protein products that are responsible for maintaining an appropriate balance of free nucleotides within the mitochondrion and TFAM, TFBM1 and TFBM2 which are needed for intra-mitochondrial transcription and translation13.
Disruption of either nDNA or mtDNA encoded genes can cause mitochondrial dysfunction and new insights into these disorders must necessarily address the basic forces which act upon the two highly differentiated genomes14. We previously characterized the effect of selection upon the mtDNA genome through conducting a phylogenetic evaluation of the observed non-synonymous mutations versus all possible single nucleotide variations and a quantitative analysis of the pathogenicity potential of these mutations15. Evidence pointed for the observable effect of purifying selection acting upon the mtDNA non-synonymous potentially-pathogenic mutations (MutPred pathogenicity score higher than 0.7; in a scale between 0, no pathogenicity, to 1, highly pathogenic16). This purifying selection is common to human and other mammals, such as cow, dog, pig and chimpanzee17. Interestingly, this selection is absent on cancer tissues whose characteristics render them free from many normal selective forces18. A special case of cancer, which is designated as the oncocytic tumor and is characterized by an abnormally high number of mitochondria and mtDNA, is significantly associated with highly-pathogenic non-synonymous mtDNA mutations, indicating a positive selection of this phenotype18.
In this work, we investigated the pathological potential of all non-synonymous SNPs that could occur from single nucleotide substitutions across 104 nuclear genes encoding mitochondrial proteins, including both oxidative phosphorylation proteins and regulatory proteins (of transcription, replication, stress protective, fission and fusion) (Supplemental Data Table S1). Although these genes only account for a subset of the estimated 1500–2000 genes encoding proteins targeted to mitochondria, they encode major proteins involved in mitochondrial pathways. We contrasted observed population diversity from the 1000 Genomes project with theoretical values for all possible non-synonymous mutations to ascertain the effect of purifying selection in these genes. We further compared signs of selection in these nDNA genes against mtDNA genes. We also investigated in these nuclear genes the proportion of individuals in this population database carrying potentially pathogenic SNPs and compared such values across different population groups. By focusing on the mitochondrial pathways, we shed light on the burden of mitochondrial diseases worldwide.
Results
In the 1000 Genomes database, we identified 706 non-synonymous variants in the 104 nuclear genes from 1,092 individuals, although 12 of these variants were not in Hardy-Weinberg equilibrium and therefore were removed from further analysis. A total of 360 non-synonymous variants were observed once; and 97 were observed twice, usually in two individuals from the same population (hence likely shared by recent common ancestry). By comparing the probability distributions of the pathogenicity scores in observed amino acid variants and in the 171,873 possible amino acid variants (Figure 1), we detected clear signs of purifying selection. The very rare variants (minor allele frequency; MAF < 0.1%) were not statistically different from the group of all possible variants (p = 0.34 by t-test), even though it was the largest group (478 variants). All other groups (156 rare with 0.1% < MAF < 1%; 27 intermediate with 1% < MAF < 5%; and 33 common with MAF > 5%) were significantly different from the set of all possible variants (p = 1.9 × 10−9; p = 0.00071; p = 7.8 × 10−12, respectively). This pattern is similar to the one observed for the mtDNA genes15.
Assessing Selection as a Function of the Pathogenicity Score Across Variant Frequency Classes.
Probability distributions of all possible and observed amino acid variant pathogenicity scores for (A) very rare variants (MAF < 0.1%); (B) rare variants (0.1% < MAF < 1%); (C) intermediate variants (1% < MAF < 5%); and (D) common variants (MAF > 5%). The solid line denotes the pathogenicity score distribution for all possible single nucleotide variants in the 104 genes assessed, for comparison. Pathogenicity scores calculated using MutPred, with high values representing greater pathogenicity.
Although both nDNA and mtDNA encoded mitochondrial genes are under the effect of purifying selection, the set of all possible variants for the nuclear genes has a mean pathogenicity score of 0.544, which is lower than the mean value displayed by the mtDNA variants, of 0.640. The difference is even more remarkable when comparing the distributions of the pathogenicity scores (Figure 2A). For all possible variants in the mtDNA encoded proteins, the distribution is strongly skewed to high predicted pathogenicity scores, while the distribution for the nuclear encoded genes is more moderate. These results have a great influence in measuring the effect of selection, which can be done by dividing the distribution of pathogenicity scores for all observed protein variants by the distribution of scores for all possible variants (Figure 2B). Selection functions for both mtDNA and nDNA genes were consistent with an exponential decay (nDNA: adjusted R2 = 0.748 and p-value = 1.28 × 10−5; mtDNA: adjusted R2 = 0.956; p-value = 4.812 × 10−6), although the exponential decay is especially strong for the mtDNA, indicating a stronger purifying selection.
Assessing Selection as a Function of the Pathogenicity Score in nDNA and mtDNA Encoded Proteins.
(A) Probability distributions of all possible and observed amino acid variant pathogenicity scores in nDNA and mtDNA encoded proteins. (B) The selection function for the amino acid variants in nDNA and mtDNA encoded proteins, defined by dividing the observed distributions of pathogenicity scores by the distribution of scores for all possible variants.
Our data indicate a differential potential for pathogenicity between the two genomes, with much higher potential pathogenicity in the mtDNA. One well-known property of proteins coded by vertebrate mtDNA is their high hydrophobicity compared to the proteins coded by the nDNA. We further investigated whether this chemical property could account for the different theoretical pathogenicity patterns of the proteins.
We began by ascertaining if the amino acids that have different codons between nDNA and mtDNA, due to the differing genetic codes, could be the main contributors to this issue (Figure S6), since this directly affects the range of amino acid changes that can be created by a single nucleotide change. The differences between the genetic codes would mean more opportunities to create tryptophan and methionine in mtDNA through single nucleotide variations, compared with more expected isoleucine and arginine in nDNA. Indeed, comparing with our nDNA gene set, mtDNA coded proteins show higher proportions of tryptophan and methionine and lower proportion of arginine, but also much higher percentage of isoleucine. This observation suggests that there is no simple correspondence between the difference in genetic codes and differences in hydrophobic amino acid content of proteins. Interestingly, pathogenicity values are higher in almost all amino acids for mtDNA coded proteins, meaning that the pathogenicity risk of a non-synonymous mutation in the mtDNA is higher than for a mutation in the same amino acid in the nDNA coded genes for mitochondrial proteins. This difference is present even in amino acids with the same genetic code in both nDNA and mtDNA, ruling that difference out as an explanation for the greater predicted pathogenicity of mtDNA variants (Figure S14.A, Supplemental Results Section 4).
The impact of hydrophobicity is clearer when we performed the analyses at the protein level (Figure 3A). All mtDNA coded proteins except for ATP8 have higher average hydrophobicity and higher average MutPred score than the proteins coded by nDNA (correlations are presented in Supplemental Results Section 1). These results hold true when using SIFT as a measure of pathogenicity (see Figure S10 Supplemental Results Section 2), suggesting that the solvent accessibility incorporated in MutPred algorithm does not critically influence the correlation between MutPred and hydrophobicity values. Curiously, ATP8 has been shown to be the most polymorphic mtDNA gene in both human (in the upper extreme when considering the amount of polymorphic sites or only the set of polymorphisms with a frequency of at least 0.1%14) and other species (presenting the higher average of radically changing amino acid properties per residue19). Further confirmation is evident when plotting the difference in mean pathogenicity score between nDNA and mtDNA per amino acid, which is strongly associated with the amino acid hydrophobicity (Figure 3B, p = 0.0013, R2 = 0.45 by linear regression). Between nDNA and mtDNA encoded proteins, hydrophobic amino acids show little difference in pathogenicity, but less hydrophobic ones display clear difference in it (Figure S14.B, Supplemental Results Section 4).
Assessing Hydrophobicity Influence at the Protein and Amino Acid Level, Using GRAVY Measure.
(A) Average pathogenicity score for each of the nDNA and mtDNA coded proteins vs average hydrophobicity of the protein. (B) Distribution of the difference in mean pathogenicity score between nDNA and mtDNA for each amino acid vs amino acid hydrophobicity. The line shows a linear fit (R2 = 0.45, p = 0.0013).
This positive correlation between overall pathogenicity and pathogenicity in proteins is consistent with previous findings of stability disruption in pathogenic variants20 as well as the enrichment of structural features, such as helix and sheet propensity and depletion of intrinsically disordered regions21,22 at sites of pathogenic variation. Indeed, we found that the average propensity for intrinsically disordered regions (assessed by VSL2B predictor23) was significantly lower for the set of 13 mtDNA-encoded proteins compared to the 104 nuclear DNA coded proteins (0.229 vs. 0.421; p = 2.5 × 10−7; t-test), with no significant differences in protein average length (294.2 vs. 253.1; p = 0.533; t-test) in the two groups. The higher propensity towards intrinsic disorder for nDNA proteins may be related to requirements for mitochondrial import or to their functional roles24,25. In the 117 proteins assessed in this study, the VSL2B values were highly correlated (R = −0.73, p < 0.0001) with the hydrophobicity values, preventing any useful multivariable model testing involving both VSL2B and hydrophobicity simultaneously. The Aliphatic index was also highly correlated with the hydrophobicity (R = 0.89, p < 0.0001). Considering this, we performed a multivariate linear regression analysis for the MutPred pathogenicity score with hydrophobicity and the Instability Index. In this multivariate model both hydrophobicity (p = 1.1 × 10−4) and the instability index (p = 4.6×10−6) are significantly associated with the pathogenicity score.
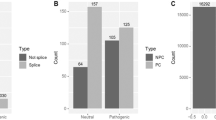
We then turned to the question of how many potentially pathogenic SNPs in these nuclear genes for critical mitochondrial proteins were observed in each individual and compared these numbers across different population groups (Figure 4). We restricted this analysis to non-admixed populations in the 1000 Genomes database by leaving out all Hispanic populations and the single African American population, while retaining the “Utah residents with Northern and Western European ancestry” as part of the European population. Such criterion finally selected 850 individuals. For this assessment of predicted pathogenicity we were conservative and further considered only the group of variants that satisfy all these criteria: (1) showing consensus predicted pathogenicity among three widely used tools (MutPred, Polyphen and SIFT; see Table S2); (2) being in HW equilibrium in all three continental population groups; and (3) having a frequency >0.5%. A total of 202 individuals (23.8% of the 850 individuals; Table S6) had at least one predicted pathogenic hit in the 104 nDNA genes coding mitochondrial proteins. There were 154 SNVs that were predicted pathogenic, for which the vast majority of subjects that were carriers had only one of these SNVs (183; 90.6%). A minor proportion of individuals had two of these SNVs (17; 8.4%) or three SNVs (2; 1.0%). Proportionally, a higher fraction of Africans display at least one predicted pathogenic SNV (32.4%), than Asians (28.3%) and Europeans (16.1%). The proportions of carriers and non-carriers of predicted pathogenic variants in the three population groups are highly significant different (p = 0.000009), due to the low proportion of carriers in Europeans (two by two population groups comparisons: Africans × Asians, p = 0.355; Africans × Europeans, p = 0.000013; Asians × Europeans, p = 0.00018). The vast majority (97.4%) of the predicted pathogenic SNVs was limited to a single population group and just two of these SNVs were shared between Africa and Asia and another two between Africa and Europe. Proportionally to the sample size, Africans had more non-synonymous SNPs than Europeans and Asians, but their average MutPred was 0.641 compared with 0.702 and 0.677 in the latter, respectively. Thus, these data seem to contradict the expectation based on the higher proportion of non-synonymous mutations in non-African populations in the whole genome. However, Abecasis et al.4 showed that the population bias in the non-synonymous variation proportion varies greatly between different pathways. For the oxidative phosphorylation pathway, which highly matches the nDNA genes studied here, the authors estimated a value of 0.595 for the excess of non-synonymous per kb for the rare SNPs (MAF < 0.5%), while typically the corresponding value across all proteins for rare SNPs is in the range 1–2. However, that value for rare SNPs in the oxidative phosphorylation pathway is at the low end of the range 0.5–1.5 found for overall common variants (the set already pruned by purifying selection). This indicates for the oxidative phosphorylation pathway a stronger purifying selection on the nuclear encoded genes than on nDNA genes in general.
Evaluating Pathogenic Mutations Across Population Groups.
(A) Number of overall predicted pathogenic mutations on our 104 genes across population groups (top number) and when divided by OXPHOS/non-OXPHOS function (bottom pair of numbers). (B) Frequency of carriers of predicted pathogenic mutations in the three population groups. Error bars are 95% confidence intervals and p-values are from Fisher's exact tests.
We also checked the OMIM and the COSMIC databases for these 104 genes. OMIM reports 155 non-synonymous SNPs, mostly displaying very high MutPred values (Figure 5). Of these, 19 were detected in the 1000 Genomes database, eight with MutPred score >0.7 (all in single individuals except one SNP in two African individuals from the same population) and the remaining 11 with MutPred score <0.7 (mostly present in several individuals, consistent with lower pathogenicity). The COSMIC dataset comprises 603 non-synonymous SNPs in these nDNA genes and its MutPred distribution curve resembles the curve for all possible non-synonymous variants, showing that the somatic mutations reported are indistinguishable from a random selection from the set of all possible amino acid SNPs and have therefore escaped the effects of purifying selection, as also occurs for somatic mtDNA mutations in cancer18. Only nine of the 603 non-synonymous SNPs in the COSMIC dataset were found in the 1000 Genomes database with MutPred values varying from 0.454 to 0.875. The non-synonymous mutations reported in OMIM, COSMIC and 1000 genomes for these 104 genes have a high bias to affect the residue arginine (Figure 6A), as also occurs overall in the genome26, but this bias is surprisingly strong in the OMIM dataset. The authors point out that as four out of the six codons for arginine contain CpG dinucleotides (in the non-wobble positions, so always non-synonymous), the 10–50 times higher mutation rate in those dinucleotides than in others leads to a higher mutability of arginine when compared with the other amino acids. In the OMIM dataset, a remarkably large proportion (41.5%) of these arginine mutations are to tryptophan (Figure 6B), while the corresponding proportion is only 11.5% in 1000 Genomes and 9.3% in the COSMIC datasets.
Pathogenic Non-synonymous Mutations in 1000 Genomes, OMIM and COSMIC.
Frequencies of amino acids in the 104 nDNA encoded proteins and of amino acids mutated in 1000 Genomes, OMIM and COSMIC datasets in which (A) represents the reference amino acid and (B) the variant amino acid. For comparison, the “frequency in protein” value is repeated in both panels.
Discussion
Our results show that the very rare variants (MAF < 0.1%) in the nDNA genes encoding mitochondrial proteins are consistent with a random set from all possible variants with no discernable selection having had time to occur upon them. All other classes of variant frequencies do display distinct signs of purifying selection eliminating variants with high pathogenicity scores, to such an extent that for common variants no more than 10% display MutPred pathogenicity values higher than 0.7. Hydrophobicity seems to account for the very high pathogenic values that variants can reach in mtDNA genes, when comparing to the nDNA coded genes. The high hydrophobicity of these proteins is one of the reasons proposed for the retention of the mitochondrial genome27, as it would be difficult to transport their protein products into the organelle if coded by the nDNA. Other authors believe the transfer of genes from mitochondria to the nucleus is still ongoing and, given enough time, the mtDNA will be lost28, while a third hypothesis states that certain genes have been preferentially maintained in mitochondria to locally adjust gene expression according to the redox or metabolic state of individual mitochondria29,30. Whatever the reason, the maintenance of the mtDNA requires a complex cross-talk between the two genomes. It is curious that despite the fact that the average protein pathogenicity does increase as the protein hydrophobicity increases, at the extreme high range of protein hydrophobicity (for values higher than 0.377, which encompasses all mtDNA encoded proteins except ATP8 and includes a few nDNA encoded proteins; Figure 3A and Supplemental Results Section 1) the relationship appears to change to a negative correlation. We should be careful in interpreting this evidence, as at that range of the graph there are very few data points.
Possibly the high hydrophobicity of proteins coded by mtDNA also rendered this genome more prone to very pathogenic mutations. Recently, Badgujar et al.31 showed that most mutations in the BRCA1 gene are located at the hydrophobic core and that the evaluation of its pathogenicity (by applying functional assays) depends on how they destabilize the hydrophobic environment of the protein structure. They even suggest that small molecules that could stabilize the hydrophobic environments of BRAC1, reducing the pathogenicity of the mutations, could be used as potential drugs for targeted cancer therapy. Also, for the hereditary human forms of encelophalopathies or prion diseases, four of its mutations occur in the hydrophobic core of the PrP prion protein and these have been shown to have a significant effect on the dynamics and stability of the protein, including the propensity to misfolding32. This hydrophobicity-driven faster selection force in mtDNA-coded compared with nDNA-coded mitochondrial proteins is even more significant when taking in consideration that selection has more power to determine variation in larger populations, while genetic drift has more influence in smaller populations33 and that the effective population size of a nuclear allele is four times that of a mitochondrial allele.
The African populations had more potential pathogenic mutations (when being conservative in this classification) in our gene set than did the European and Asian populations, although the average MutPred pathogenicity score was lower: 32.4% individuals with one pathogenic mutation and 0.641 average MutPred pathogenicity score in Africans; 28.3% and 0.677 in Asians; and 16.1% and 0.702 in Europeans. The issue of European and Asian populations having more non-synonymous mutations than Africans is still under debate. All authors agree that demography was very important, but they differ in the modeling parameters/complexity used to evaluate selection overall the genome, getting at contradictory results5,7,8,10. A major demographic difference between African and non-African populations was the older expansion experienced in Africans, leading to the out-of-Africa migration, setting the time zero for European and Asian populations – many of the models rely greatly on this event. But the class of rare and potentially pathogenic mutations consists primarily of new mutations, dating not more than 5,000–10,000 years ago. Worldwide mtDNA phylogenetic studies34,35,36,37,38,39 are showing that despite the occurrence of previous population expansions, all human populations experienced a dramatic effective population size expansion since the Holocene, 11,700 years ago, when current climatic conditions were established, leading to the emergence of agriculture and domestication, which further fuelled the population expansions. This is the time frame for the development of the rare pathogenic variants – so, all population groups should display comparable signs for the rare class of variants. Heterogeneity can result from differences in the population expansion strength across space and time within the Holocene. In Europe the main population expansions took place at the post-glacial period (the last glacial maximum was more aggressive in the northern hemisphere), with a lower impact of the Neolithic on European populations38,40,41. In contrast, in Africa and Asia recent population expansions merging the more recent Neolithic diffusions in the regions (in the last 5 millennia), seem to have been significant36,37,42,43,44. Further heterogeneity can be caused by the differential purifying selection strengths on proteins in different biochemical pathways – which is compatible with the observation that the rare-variant load varies substantially across biological pathways and population groups4. Thus our identified significant lower rate of predicted pathogenic SNP carriers in Europe is compatible with a strong purifying selection in the oxidative phosphorylation pathway (coded by nDNA) pruning more efficiently the main post-glacial European variant pool and less so the main middle-Holocene/Neolithic Asian and African variant pools. The evidence found here of a faster selection in the mtDNA genome than in the nuclear genome also fits nicely with our previous finding that there were no differences in the purifying selection strength between population groups for mtDNA coded OXPHOS proteins15, efficiently eliminating potentially pathogenic mtDNA mutations consistently in African, Asian and European populations.
Methods
The genetic diversity of the mitochondrial proteins coded by the nuclear and mitochondrial genomes
We selected 104 mitochondrial proteins coded by nuclear genes (Table S1), including both oxidative phosphorylation (OXPHOS) proteins and regulatory proteins (of transcription, replication, stress protective, fission and fusion). These nDNA encoded mitochondrial proteins were chosen for the importance of these pathways for mitochondrial diseases, hence the interest in the predicted pathogenicity of variants in these proteins. Both sets of proteins have to migrate to the mitochondrion, but only the OXPHOS proteins will be integrated together with the mtDNA coded ones in complexes inserted in the mitochondrial inner membrane. As we are mainly interested in non-synonymous mutations, the upstream and downstream flanking regions of the gene were ignored. The genetic diversity information for each gene coding these 104 proteins was extracted from the 1000 genomes ensembl browser (http://browser.1000genomes.org), through the available tools that generate VCF files. Currently, 1000 Genomes displays information for 1,092 individuals, of which 246 are from Africa (including African-Americans; 185 without that admixed population), 379 from Europe, 286 from Asia and 181 from America. The polymorphic positions were inputed into the “variant effect predictor” to check its location in exons/introns and status, in order to infer the list of observed non-synonymous positions (Tables S2 and S3). Genotypes and population frequencies of these polymorphisms were ascertained in the 1000 Genomes web site. The Hardy-Weinberg equilibrium (HWE) was checked by using the software Arlequin45, for the global population level for each SNP and in the cases were values were statistically significant (p < 0.001), the HWE was checked within each of the four continental populations and SNPs were removed if they had p < 0.001 in any of the continental populations.
We compared with information from 14,642 complete mtDNA genomes (available in the GenBank database, for the 1000 Genomes project and our own unpublished complete sequences; Table S4), representing worldwide populations. The OMIM and COSMIC datasets were also checked for mutations in these genes (Tables S7 and S8).
Pathogenicity inference
A PERL script was used to infer all possible non-synonymous variants from all possible SNPs in the 104 nDNA genes, based on the GRCh37 human reference for one of the protein coding transcripts (usually the one coding the longer protein sequencing) available for each gene in the 1000 Genomes ensembl browser (the selected transcript identifiers are reported in Table S1). The list of all possible non-synonymous variants was then assessed by the MutPred algorithm46 to infer pathogenicity scores. MutPred generates probability-like scores for amino acid substitutions to estimates their pathogenicity. We used the same version of MutPred (1.2) as before17 in calculating the pathogenicity scores for variants in the nuclear genes. The MutPred scores for the observed non-synonymous mutations were checked in this list. Other pathogenicity scores, such as PolyPhen47 and SIFT48 were inferred in the tool “variant effect predictor” of 1000 Genomes ensembl browser, for the observed polymorphisms, in order to correlate the prediction between the three classifiers (Supplemental Results Section 2). Furthermore, SIFT predictor was estimated for all possible non-synonymous variants in the nDNA and mtDNA coded proteins, by using SIFT online tools (http://sift.jcvi.org/).
Protein physicochemical properties
Several physicochemical properties (Table S5), such as hydrophobicity or hydropathy, deduced from the protein sequences were determined by using the ProtoParam tool available on the ExPASy Server49. We were mainly interested in having a measure of hydrophobicity or hydropathy of the protein and this tool provides the grand average of hydropathy (GRAVY) as the sum of hydropathy values of all the amino acids divided by the number of residues in the sequence, based on the Kyte and Doolittle scale50, which takes into account values from water-to-vapor energy transfer and from internal-external distribution of amino acid residues. We also estimated the: Instability Index, which provides an estimate of the stability of the protein in a test tube, based on the observation that the occurrence of certain dipeptides is significantly different in the unstable proteins compared with those in the stable ones; and Aliphatic Index defined as the relative volume occupied by aliphatic side chains (alanine, valine, isoleucine and leucine), which may be regarded as a positive factor for the increase of thermostability of globular proteins.
We also used the MitoProt II web site51 to estimate other hydrophobicity measures based on several hydrophobicity scales. The estimated measures are: H17, the maximal local hydrophobicity of a segment; MesoH (Mesohydrophobicity), the average regional hydrophobicity over an extended region of the sequence; MuHD, the maximal Eisenberg's hydrophobic moment with 6 angles of 75″, 95″, 100″ and 105″, with a scanning window of 18 residues; Hmax, the maximal hydrophobicity of 7 neighboring residues on the maximal alpha segment, with alpha having the values of 75, 95, 100 and 105 degrees. The scales used are: the Kyte and Doolittle (KD); the Goldman, Engelman and Steitz scale (GES)52, which quantifies the free energy of water-oil transfer for residues in an a-helical structure; the Gunnar von Heijne scale 1 (GVH1)53, a statistical scale obtained from the amino acid frequencies in the central part of a transmembrane segment with respect to the non-membranous stretches; and the Eisenberg's consensus scale (ECS)54, designed to mitigate the effects of outlaying values in any one scale produced by the peculiarities of the method and is simply an average of four scales. We tested the effect of some of these diverse measures in the results (Supplemental Results Section 3).
We used the VSL2B for protein disorder prediction, which is applicable to proteins with more than 25 amino acids and can accurately identify short disordered regions that are often misclassified by other disorder predictors.
Statistical analyses
All comparisons of mean values were calculated in Origin 7 (www.originlab.com), by using two-tailed t-tests assuming unequal variances. The observed non-synonymous SNP lists for nDNA and mtDNA coded proteins consisted in single instances of each variant, as it is difficult to infer if a nDNA variant present in several individuals is shared by ancestry or by state. Although it is easier to infer recurrence in the mtDNA, we followed the same rule, for comparison sake. Nonetheless, we evaluated the influence of the nDNA SNPs' frequency in the MutPred pathogenicity inference by performing comparisons between very rare variants (MAF < 0.1%), rare (0.1% < MAF < 1%), intermediate (1% < MAF < 5%) and common (MAF > 5%).
A selection function for the amino acid variants was calculated by dividing the probability distributions for the set of all observed non-synonymous mutations and all possible non-synonymous mutations (created by a single nucleotide change from the reference sequence) into ten equal intervals of the MutPred pathogenicity scores (0 to 0.1, etc.) and taking the ratio of the two probabilities within each bin. The nonlinear curve fit of the selection function to the pathogenicity score and the adjusted R2 were calculated in Origin 7.
The multivariate general linear regression analysis for MutPred values with hydrophobicity and Instability Index was performed in R.
References
Hudson, T. J. et al. International network of cancer genome projects. Nature 464, 993–998, 10.1038/nature08987 (2010).
Project, T. I. H. The International HapMap Project. Nature 426, 789–796, 10.1038/nature02168 (2003).
Abecasis, G. R. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073, 10.1038/nature09534 (2010).
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65, 10.1038/nature11632 (2012).
Dorschner, M. O. et al. Actionable, Pathogenic Incidental Findings in 1,000 Participants' Exomes. Am J Hum Genet 93, 631–640, 10.1016/j.ajhg.2013.08.006 (2013).
Coventry, A. et al. Deep resequencing reveals excess rare recent variants consistent with explosive population growth. Nat Commun 1, 131, 10.1038/ncomms1130 (2010).
Fu, W. et al. Analysis of 6,515 exomes reveals the recent origin of most human protein-coding variants. Nature 493, 216–220, 10.1038/nature11690 (2013).
Lohmueller, K. E. et al. Proportionally more deleterious genetic variation in European than in African populations. Nature 451, 994–997, 10.1038/nature06611 (2008).
Soares, P. et al. The Expansion of mtDNA Haplogroup L3 within and out of Africa. Mol Biol Evol 29, 915–927, 10.1093/molbev/msr245 (2012).
Boyko, A. R. et al. Assessing the evolutionary impact of amino acid mutations in the human genome. PLoS Genet 4, e1000083, 10.1371/journal.pgen.1000083 (2008).
Temperley, R., Richter, R., Dennerlein, S., Lightowlers, R. N. & Chrzanowska-Lightowlers, Z. M. Hungry Codons Promote Frameshifting in Human Mitochondrial Ribosomes. Science 327, 301–301, 10.1126/science.1180674 (2010).
Anderson, S. et al. Sequence and organization of the human mitochondrial genome. Nature 290, 457–465 (1981).
Chinnery, P. F. in Human Mitochondrial DNA and the Evolution of Homo sapiens Vol. 18 Nucleic Acids and Molecular Biology (ed H.-J.; Macaulay Bandelt, V.; Richards M.) Ch. 1, 3–15 (Berlin Heidelberg, Springer-Verlag, 2006).
Pereira, L. et al. The diversity present in 5140 human mitochondrial genomes. Am J Hum Genet 84, 628–640, 10.1016/j.ajhg.2009.04.013 (2009).
Pereira, L., Soares, P., Radivojac, P., Li, B. & Samuels, D. C. Comparing Phylogeny and the Predicted Pathogenicity of Protein Variations Reveals Equal Purifying Selection across the Global Human mtDNA Diversity. Am J Hum Genet 88, 433–439, 10.1016/j.ajhg.2011.03.006 (2011).
Li, B. et al. Automated inference of molecular mechanisms of disease from amino acid substitutions. Bioinformatics 25, 2744–2750, 10.1093/bioinformatics/btp528 (2009).
Soares, P. et al. Evaluating purifying selection in the mitochondrial DNA of various mammalian species. PLoS One 8, e58993, 10.1371/journal.pone.0058993 (2013).
Pereira, L., Soares, P., Maximo, V. & Samuels, D. C. Somatic mitochondrial DNA mutations in cancer escape purifying selection and high pathogenicity mutations lead to the oncocytic phenotype: pathogenicity analysis of reported somatic mtDNA mutations in tumors. BMC Cancer 12, 53, 10.1186/1471-2407-12-53 (2012).
da Fonseca, R. R., Johnson, W. E., O'Brien, S. J., Ramos, M. J. & Antunes, A. The adaptive evolution of the mammalian mitochondrial genome. BMC Genomics 9, 119, 10.1186/1471-2164-9-119 (2008).
Wang, Z. & Moult, J. SNPs, Protein Structure and Disease. Hum Mutat 17, 263–270 (2001).
Mort, M. et al. In Silico Functional Profiling of Human Disease-Associated and Polymorphic Amino Acid Substitutions. Hum Mutat 31, 335–346 (2010).
Vacic, V. et al. Disease-Associated Mutations Disrupt Functionally Important regions of Intrinsic Protein Disorder. PLoS Comput Biol 8, e1002709 (2012).
Peng, K., Radivojac, P., Vucetic, S., Dunker, A. K. & Obradovic, Z. Length-Dependent Prediction of Protein Intrinsic Disorder. BMC Bioinformatics 7, 208 (2006).
Schmidt, O., Pfanner, N. & Meisinger, C. Mitochondrial Protein Import: From Proteomics to Functional Mechanisms. Nat Rev Mol Cell Bio 11, 655–667 (2010).
Homma, K., Fukuchi, S., Nishikawa, K., Sakamoto, S. & Sugawara, H. Intrinsically Disordered Regions Have Specific Functions in Mitochondrial and Nuclear Proteins. Mol Biosyst 8, 247–255 (2012).
de Beer, T. A. et al. Amino acid changes in disease-associated variants differ radically from variants observed in the 1000 genomes project dataset. PLoS Comput Biol 9, e1003382, 10.1371/journal.pcbi.1003382 (2013).
Claros, M. G. et al. Limitations to in vivo import of hydrophobic proteins into yeast mitochondria. The case of a cytoplasmically synthesized apocytochrome b. Eur J Biochem 228, 762–771 (1995).
Palmer, J. D. Organelle genomes: going, going, gone! Science. 275, 790–791 (1997).
Allen, J. F. Why chloroplasts and mitochondria contain genomes. Comp Funct Genomics 4, 31–36, 10.1002/cfg.245 (2003).
Race, H. L., Herrmann, R. G. & Martin, W. Why have organelles retained genomes? Trends Genet 15, 364–370 (1999).
Badgujar, D. C., Sawant, U., Mahadik, H., Gadewal, N. & Varma, A. K. Pathogenicity of mutations discovered in BRCA1 BRCT domains is characterized by destabilizing the hydrophobic interactions. J. Cancer Sci. Ther 4, 386–393 (2012).
van der Kamp, M. W. & Daggett, V. Pathogenic mutations in the hydrophobic core of the human prion protein can promote structural instability and misfolding. J Mol Biol 404, 732–748, 10.1016/j.jmb.2010.09.060 (2010).
Lanfear, R., Kokko, H. & Eyre-Walker, A. Population size and the rate of evolution. Trends Ecol Evol 29, 33–41, 10.1016/j.tree.2013.09.009 (2014).
Fernandes, V. et al. The Arabian cradle: mitochondrial relicts of the first steps along the southern route out of Africa. Am J Hum Genet 90, 347–355, 10.1016/j.ajhg.2011.12.010 (2012).
Pereira, L. et al. Population expansion in the North African late Pleistocene signalled by mitochondrial DNA haplogroup U6. BMC Evol Biol 10, 390, 10.1186/1471-2148-10-390 (2010).
Rito, T. et al. The first modern human dispersals across Africa. PLoS One 8, e80031, 10.1371/journal.pone.0080031 (2013).
Soares, P. et al. Climate change and postglacial human dispersals in southeast Asia. Mol Biol Evol 25, 1209–1218, 10.1093/molbev/msn068 (2008).
Soares, P. et al. The archaeogenetics of Europe. Curr Biol 20, R174–183, 10.1016/j.cub.2009.11.054 (2010).
Atkinson, Q. D., Gray, R. D. & Drummond, A. J. mtDNA variation predicts population size in humans and reveals a major Southern Asian chapter in human prehistory. Mol Biol Evol 25, 468–474, 10.1093/molbev/msm277 (2008).
Pala, M. et al. Mitochondrial DNA signals of late glacial recolonization of Europe from near eastern refugia. Am J Hum Genet 90, 915–924, 10.1016/j.ajhg.2012.04.003 (2012).
Fu, Q., Rudan, P., Paabo, S. & Krause, J. Complete mitochondrial genomes reveal neolithic expansion into Europe. PLoS One 7, e32473, 10.1371/journal.pone.0032473 (2012).
Gignoux, C. R., Henn, B. M. & Mountain, J. L. Rapid, global demographic expansions after the origins of agriculture. Proc Natl Acad Sci U S A 108, 6044–6049, 10.1073/pnas.0914274108 (2011).
Zheng, H. X. et al. Major population expansion of East Asians began before neolithic time: evidence of mtDNA genomes. PLoS One 6, e25835, 10.1371/journal.pone.0025835 (2011).
Zheng, H. X., Yan, S., Qin, Z. D. & Jin, L. MtDNA analysis of global populations support that major population expansions began before Neolithic Time. Sci Rep 2, 745, 10.1038/srep00745 (2012).
Excoffier, L., Laval, G. & Schneider, S. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 1, 47–50 (2005).
Zhidkov, I., Livneh, E. A., Rubin, E. & Mishmar, D. MtDNA mutation pattern in tumors and human evolution are shaped by similar selective constraints. Genome Res 19, 576–580, 10.1101/gr.086462.108 (2009).
Adzhubei, I., Jordan, D. M. & Sunyaev, S. R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr Protoc Hum Genet Chapter 7, Unit7 20, 10.1002/0471142905.hg0720s76 (2013).
Ng, P. C. & Henikoff, S. Predicting deleterious amino acid substitutions. Genome Res 11, 863–874, 10.1101/gr.176601 (2001).
Gasteiger, E. et al. in The Proteomics Protocols Handbook (ed J.M. Walker) 571–607 (Berlin Heidelberg, Springer-Verlag, 2005).
Kyte, J. & Doolittle, R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol 157, 105–132 (1982).
Claros, M. G. & Vincens, P. Computational method to predict mitochondrially imported proteins and their targeting sequences. Eur J Biochem 241, 779–786 (1996).
Engelman, D. M., Steitz, T. A. & Goldman, A. Identifying nonpolar transbilayer helices in amino acid sequences of membrane proteins. Annu Rev Biophys Biophys Chem 15, 321–353, 10.1146/annurev.bb.15.060186.001541 (1986).
von Heijne, G. Membrane protein structure prediction. Hydrophobicity analysis and the positive-inside rule. J Mol Biol 225, 487–494 (1992).
Eisenberg, D. Three-dimensional structure of membrane and surface proteins. Annu Rev Biochem 53, 595–623, 10.1146/annurev.bi.53.070184.003115 (1984).
Acknowledgements
FCT, the Portuguese Foundation for Science and Technology, supported this work through the research project PTDC/CS-ANT/113832/2009 and the personal grant to P.S. (SFRH/BPD/64233/2009). P.T. has a PhD grant from the People Programme (Marie Curie Actions) of the European Union's Seventh Framework Programme FP7/2007–2013/ under REA grant agreement no. 317184. The Instituto de Patologia e Imunologia Molecular da Universidade do Porto is an Associate Laboratory of the Portuguese Ministry of Science, Technology and Higher Education and is partially supported by FCT. L.P. had a mobility grant from The Luso-American Development Foundation.
Author information
Authors and Affiliations
Contributions
L.P. and D.C.S. designed the study and wrote the manuscript. L.P., P.S., P.T., T.R., A.W., B.L., P.R. and D.C.S. performed the analyses. All authors reviewed and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplemental Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Pereira, L., Soares, P., Triska, P. et al. Global human frequencies of predicted nuclear pathogenic variants and the role played by protein hydrophobicity in pathogenicity potential. Sci Rep 4, 7155 (2014). https://doi.org/10.1038/srep07155
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07155
This article is cited by
-
Phylogenetic and population-based approaches to mitogenome variation do not support association with male infertility
Journal of Human Genetics (2017)
-
60,000 years of interactions between Central and Eastern Africa documented by major African mitochondrial haplogroup L2
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.