Abstract
For several years, knowledge on the microbiome associated with marine invertebrates was impaired by the challenges associated with the characterization of bacterial communities. With the advent of culture independent molecular tools it is possible to gain new insights on the diversity and richness of microorganisms associated with marine invertebrates. In the present study, we evaluated if different preservation and processing methodologies (prior to DNA extraction) can affect the bacterial diversity retrieved from snakelocks anemone Anemonia viridis. Denaturing gradient gel electrophoresis (DGGE) community fingerprints were used as proxy to determine the bacterial diversity retrieved (H′). Statistical analyses indicated that preservation significantly affects H′. The best approach to preserve and process A. viridis biomass for bacterial community fingerprint analysis was flash freezing in liquid nitrogen (preservation) followed by the use of a mechanical homogenizer (process), as it consistently yielded higher H′. Alternatively, biomass samples can be processed fresh followed by cell lyses using a mechanical homogenizer or mortar & pestle. The suitability of employing these two alternative procedures was further reinforced by the quantification of the 16S rRNA gene; no significant differences were recorded when comparing these two approaches and the use of liquid nitrogen followed by processing with a mechanical homogenizer.
Similar content being viewed by others
Introduction
Phylum Cnidaria is a large, diverse and ecologically important group of relatively simple organisms that is widely distributed in marine environments1. Research on marine cnidarians experienced a significant advance over the last decades with the growing awareness on the vulnerability of certain key ecosystems (e.g. coral reefs) driven by direct or indirect anthropogenic actions2,3,4. Additionally, with the intensification of bioprospecting of marine invertebrates for drug discovery, as well as other biotechnological applications, researchers have started to target cnidarians in their quest for marine bioactive compounds5,6. Alongside with this new trend, there are growing evidences that microbes associated with marine invertebrates may be the true producers of such bioactive compounds or, at least, be partially involved in the process of biosynthesis of some of these molecules7. Several of these compounds are secondary metabolites produced by symbiotic microorganisms in chemical mediation and/or defense of interaction among marine microorganisms8. The microbiome of certain marine invertebrates may represent a remarkable proportion of the holobiont biomass, with anthozoan cnidarians being no exception and hosting abundant and diverse communities of bacteria9. Certain species able to secrete mucus may reach microbial concentrations up to 1000-fold higher than those observed in seawater10. While microbial communities associated with tropical reef building corals are already starting to be unraveled, those colonizing other groups of anthozoans are still largely unknown11. For several years, this gap of knowledge has been mainly due to the challenges associated with the characterization of bacterial communities using culture dependent approaches12. Only a small fraction of microbial symbionts can be cultured outside its cnidarian host using conventional culture media. The advent of culture independent molecular technologies [e.g. Denaturing Gradient Gel Electrophoresis (DGGE) and high–throughput DNA sequencing], made possible to overcome these bottlenecks and reveal the diversity and richness of microorganisms associated with marine invertebrates in general9,13,14. Anthozoan cnidarians are no exception to this breakthrough11,15,16. Regardless of the potential associated with the use of high–throughput DNA sequencing to profile the microbiome of marine invertebrates, the final results achieved are still largely dependent on the quality and quantity of DNA extracted from collected samples17. DNA quality and quantity is known to vary with the procedures employed for preserving and processing samples, as well as on the reliability of the DNA extraction method17,18.
Sea anemones, namely those hosting endosymbiotic photosynthetic dinoflagellates, are recognized to be important sentinel species19. These organisms may help researchers to monitor potential environmental shifts in temperate coastal waters triggered by global climate changes. Extreme bleaching events of Anemonia in the Mediterranean under abnormally warm water conditions20 are a good example on the suitability of these anthozoans as sentinel species. In light of the hologenome theory10, these anemones should be considered as holobionts21, a complex symbiosis between the cnidarian animal, its photosynthetic microalgae (e.g., zooxanthellae) and its complex community of associated microorganisms that play a key role on the overall health of the cnidarian host. Therefore, it is important to monitor potential shifts in the microorganisms associated with these sea anemones to understand how environmental disturbances may shape anemone individuals and populations.
Despite the existence of reports on the suitability of processing techniques to preserve samples and extract microbial DNA from marine invertebrates (e.g.17,22), only a few studies are currently available on sea anemones23,24. Given the current state of the art on this topic and the complexity/specificity of this biological matrix, we consider that it is relevant to standardize a protocol that can allow researchers to extract good-quality DNA in order to perform a reliable analysis of the bacterial communities associated with sea anemones. In line with this goal, here we used the snakelocks anemone Anemonia viridis (Forskål, 1775) as a model species to evaluate how different preservation and processing approaches could affect the quality of extracted DNA and the molecular profiles of bacterial communities retrieved from these organisms.
Results
The two-way ANOVA revealed that there was no significant (P = 0.496) interaction between the processing and preserving procedures that were tested and that the categorical factor processing did not significantly (P = 0.347) affected the values of Shannon's index of diversity (H′) calculated from the bacterial fingerprints generated from DGGE. However, the categorical factor preserving significantly (P = 0.022) affected H′ values. Experimental treatments employing freezing at −80°C differed significantly from those processing fresh samples or samples flash frozen with liquid nitrogen (P = 0.017 and P = 0.025, respectively). The highest average H′ value (±s.d.) was displayed by LN_H (H′ = 2.72 ± 0.17), while the lowest was that of F-80_H (H′ = 1.68 ± 0.79) (Figure 1).
Shannon's index of diversity (H′) calculated from DGGE community profiles of Bacteria detected on snakelocks anemone Anemonia viridis from each experimental treatment.
Values presented are means (+s.d.) of five independent replicates. Fr – Fresh (blue); NH – non-homogenized (full colored); H – maceration with homogenizer (pinstripe right); MP - maceration with mortar & pestle (pinstripe left); LN - freezing with liquid nitrogen followed by preservation at −80°C (green); F-80 – freezing and preservation at −80°C (red). Different letters represent significant differences (Tukey's test, P < 0.05).
The bacterial fingerprints recorded in the DGGE of the three experimental treatments promoting the highest H′ (in descending order, LN_H, Fr_H and Fr_MP) are illustrated in Figure 2.
Denaturing gradient gel electrophoresis (DGGE) based analysis of bacterial community composition in the snakelocks anemone Anemonia viridis (cropped image, full-length gel is presented in Supplementary Fig. S1).
The DGGE gel presented compares community fingerprints of 16S rRNA gene fragments amplified from DNA for the three experimental procedures displaying the highest H′ values calculated from DGGE community profiles of Bacteria detected in samples of snakelocks anemone Anemonia viridis: fresh samples processed with homogenizer (Fr_H); samples frozen with liquid nitrogen followed by processing with homogenizer (LN_H); and fresh samples processed with mortar & pestle (Fr_MP). Equal numbers represent samples originating from the same anemone.
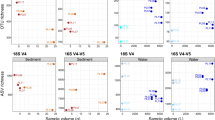
The first two axis of the PCO explained 65.1% of the variability recorded in the bacterial fingerprints of the three experimental treatments yielding the highest H′ (Figure 3). Samples from treatment LN_H are clearly clustered apart from those where samples were processed fresh (Fr).
PCO of the three experimental procedures displaying the highest H′ values calculated from DGGE community profiles of Bacteria detected in samples of snakelocks anemone Anemonia viridis: fresh samples processed with homogenizer (Fr_H); samples frozen with liquid nitrogen followed by processing with homogenizer (LN_H); and fresh samples processed with mortar & pestle (Fr_MP).
Real-time PCR quantification of 16S rRNA gene did not reveal the existence of any significant differences between the three procedures yielding the highest H′ (P = 0.008).
Discussion
The present study reveals that the use of community fingerprinting approaches such as PCR-DGGE is a robust technique to assess and/or optimize processing and preservation methodologies of biological samples destined for microbial communities analysis using molecular tools25,26. While it is true that PCR-DGGE only detects the more abundant taxa present in the sample being analyzed it also provides an excellent high-throughput tool for comparative community structure analysis, it allows researchers to determine and compare the relative abundance of different bacterial populations and therefore compare procedures, without the need to use more expensive and labor intensive techniques25,27,28. Indeed, by using this approach it was possible to verify that there are no significant interactions between preservation and processing procedures employed for samples of A. viridis meant to be used in bacterial diversity analysis using molecular techniques. Preservation was recorded to significantly affect H′ of bacterial communities retrieved from sea anemones, as already recorded for sponges17. It is now recognized by researchers that the preservation technique employed for marine invertebrate samples is a key point for molecular analysis of microbial communities29,30. In the present study it was possible to show that flash freezing and homogenizing (LN_H) collected samples consistently yielded the highest bacterial diversity from snakelocks anemones (Figure 1). It was also possible to verify that sea anemones tissue can also be processed fresh (e.g. Fr_H and Fr_MP) with satisfactory results if researchers have the constraint of not being able to flash freeze samples with liquid nitrogen.
According to the 16S rRNA gene quantification results from Real-time PCR, any of the three procedures was considered a suitable option to obtain bacterial DNA for molecular studies of bacterial communities from sea anemones.
The best results achieved in our study through the flash freezing of collected samples are in line with the fact that at such extremely low temperatures no DNA degradation occurs through enzymatic activity18,31,32,33; in this way the bacterial diversity retained is close to that present at sampling time.
Liquid nitrogen can be difficult to obtain and transport in remote locations and keeping samples frozen while in transit can at times be a challenging task. However, our results support the fact that flash freezing is indeed the most efficient approach when aiming to preserve biological samples from invertebrates for molecular analysis of their microbial communities29,30,33. Successfully retrieving microbial communities associated with these marine animals can be of paramount importance for biotechnological34 and/or ecological24 purposes. The extraction method can affect the diversity of microorganisms retrieved from sea anemones17. Nonetheless, as the extraction method employed in the present study displays a good compromise between the quantity and quality of extracted DNA, processing costs and processing time per sample, we recommend researchers to use our methodology. In the future, the use of standardized procedures for processing and preserving collected samples of sea anemones will allow researchers to perform reliable comparisons by ensuring homogeneity between studies. Moreover, it also makes possible the use of less expensive approaches (e.g. DGGE) to compare shifts in the relative abundance of the microbiome associated with these marine invertebrates.
Methods
Sample collection, preservation and processing
Five snakelocks anemones A. viridis were collected at low tide, in the intertidal region of Praia da Aguda (41°02′51.06″N; 8°39′14.20″W), Arcozelo, Portugal, in November 2011 and individually stocked in sterile plastic bags for immediate transportation to the laboratory.
Each of the five sea anemones collected was fragmented into 9 similar sized pieces using sterile scalpels blades along their radial axis; each piece included similar amounts of anemones body and tentacles, as well as a similar wet weight. A factorial experimental design employing three levels of preservation (samples used fresh, frozen at −80°C and flash frozen with liquid nitrogen) and three levels of processing (non-homogenized samples, samples homogenized with mortar and pestle and samples homogenized with a mechanical tissue homogenizer) was tested prior to DNA extraction. Briefly, this factorial design allowed us to evaluate 9 different experimental treatments, each with five independent replicates: Fr_NH (fresh samples non homogenized, where fresh samples were used directly for extraction of nucleic acids without any further processing or preservation); Fr_H (fresh samples were processed with the Omni Tissue Homogenizer (Omni International, Kennesaw, Georgia, USA) and used for DNA extraction without any further treatment); Fr_MP (fresh samples were processed with the mortar & pestle and used for DNA extraction without any further treatment); LN_H and LN_MP (samples were first preserved by flash freezing in liquid nitrogen and kept at −80°C and then homogenized with the Omni Tissue Homogenizer or mortar & pestle, respectively, prior to DNA extraction); F-80_H and F-80_MP (samples were first frozen and kept at −80°C and then homogenized with the Omni Tissue Homogenizer or mortar & pestle, respectively, prior to DNA extraction); LN_NH (samples were flash frozen in liquid nitrogen and kept at −80°C and used for DNA extraction without any further processing); and F-80_NH (samples were frozen and kept at −80°C and used for DNA extraction without any further processing) (see Figure 4 for a schematic representation of the experimental design).
Schematic representation of the experimental design employed to evaluate the effect of different processing and preservation approaches on bacterial diversity retrieved after performing bacterial DNA extraction and amplification of snakelocks anemone Anemonia viridis.
Fr – Fresh; NH – non-homogenized; H – maceration with homogenizer; MP - maceration with mortar & pestle; F-80 – freezing and preservation at −80°C; LN - freezing with liquid nitrogen followed by preservation at −80°C.
Extraction of nucleic acid
Nucleic acids were extracted from 0.5 g of sea anemone samples from each experimental treatment described above. All samples were homogenized using FastPrep® (Qbiogene Inc., USA) bead-beating system in combination with a mixture of beads (0.10 g Zirconia beads (0.1 mm) + 0.20 g glass beads (0.25–0.5 mm) + 0.20 g glass beads (0.75–1.0 mm) + 2 glass beads (2.85–3.45 mm)) (ROTH, DE) and Buffer SLX Mlus from E.Z.N.A.™ Soil DNA Kit (Omega Bio-Tek Inc., USA). Extraction was performed according to the instructions provided by the manufacturer. DNA was determined using Qubit™ dsDNA HS Assay Kits for Qubit® 2.0 Fluorometer (Invitrogen, Life Technologies Corporation) (see Supplementary Table ST1).
Bacterial community diversity
Bacterial community composition was evaluated by performing a DGGE based on DNA (16S rRNA gene). The bacterial fingerprints yielded by the DGGE were used as a proxy to evaluate the diversity of the bacterial community retrieved from sea anemones handled according to each of the preservation and processing combinations described above. NESTED PCR was used for a more efficient amplification of 16S rRNA gene fragments of bacterial genomic DNA extracted from sea anemone. In the first PCR, the universal bacterial primers F-27 (5′-AGAGTTTGATC(A/C)TGGCTCAG-3′) and R-1492 (5′-TACGG(C/T)TACCTTGTTACGACTT-3′) were used to amplify c. 1500 bp of the 16S rRNA gene35,36. The PCR reaction mixtures (25 μL) consisted of DNA template (1 μL), DreamTaq PCR Master Mix (2×) (12.5 μL) (Fermentas, Thermo Fisher Scientific Inc., USA), bovine serum albumin (BSA, 2.0 mg/mL) and forward (0.1 μM) and reverse primers (0.1 μM). After 5 min of denaturation at 94°C, 25 thermal cycles of 45 s at 94°C, 45 s at 56°C and 1.3 min at 72°C, the PCR was finished by an extension step at 72°C for 10 min. The amplicons obtained from the first PCR were then used as templates for a second PCR with bacterial DGGE primers F984-GC (5′- CGCCCGGGGCGCGCCCCGGGCGGGGCGGGGGCACGGGGGGAACGCGAAGAACCTTAC-3′) and R1378 (5′- CGGTGTGTACAAGGCCCGGGAACG -3′) (c. 473 bp)36. For these PCR reactions it was used the same quantity of DNA template, DreamTaq PCR Master Mix (2×), forward and reverse primers and 4% acetamide. Cycling conditions were of 4 min at 94°C for denaturation, 25 thermal cycles of 1 min at 95°C, 1 min at 53°C and 2 min at 72°C, with an extension step at 72°C for 7 min to finish the PCR reactions. All PCR amplification products were examined by electrophoresis on 1.0% agarose gels containing GelRed (Biotium Inc., USA). DGGE analysis was carried out on a DCode universal mutation detection system (Bio-Rad). The GC-clamped amplicons were applied to a double-gradient polyacrylamide gel (40–58% of denaturants) containing 6–10% acrylamide. The run was performed in 1× Tris-acetate–EDTA buffer at 58°C at a constant voltage of 160 V for 16 h. DGGE gels were silver-stained according to37. The solutions used were 10% (vol/vol) ethanol plus 0.5% acetic acid for fixation, 0.1% (wt/vol) silver nitrate for staining, freshly prepared developing solution containing 0.01% (wt/vol) sodium borohydride, 0.15% formaldehyde, 1.5% (wt/vol) NaOH, and, finally, 0.75% (wt/vol) sodium carbonate solution to stop the development. Gels were documented with a Molecular Imager chemiDoc XRS+ digitalize system (Bio-Rad). A total of five DGGEs were performed and analyzed: four DGGEs to cover all the samples of the nine experimental procedures and one DGGE including the procedures yielding the highest values of Shannon's index of diversity (H′) (see below on Statistical analysis).
DNA quantification using Real-time PCR
As Real-time PCR is an expensive laboratory procedure, it was solely used to determine the copy numbers of the 16S rRNA gene from the three preservation and processing combinations yielding the highest H′ values. The assays were performed with a StepOne real time PCR system (Applied Biosystems). The samples were quantified by determining the threshold cycle value and by comparing it to a standard curve to determine the copy number. Bacterial primers 968F (5′-AACGCGAAGAACCTTAC-3′) and 1401R (5′-CGGTGTGTACAAGACCC-3′) were used to amplify 16S rRNA gene fragments (c. 433 bp) from template DNA38. The real-time PCR master mix (20 μL) contained template DNA (1 μL), 2× SYBR Green Master Mix (Apllied Biosystems) and forward (0.1 μM) and reverse (0.1 μM) primers. The amplifications were carried out as following: initial denaturation (10 min at 95°C) was followed by 40 cycles of 15 s at 94°C, 30 s at 53°C and 30 s at 72°C and completed by fluorescence data acquisition at 80°C to dissociate the primers dimers. Product specificity was confirmed by melting point analysis (55°C to 95°C with a plate read every 0.5°C).
A fragment of the 16S rRNA gene amplified with primers U27 and 1492R (ca. 1450 bp)35, was used as a standard for the calibration curve. After amplification the standard was purified using the Geneclean II kit (MP Biomedical, France) and quantified with the Quant-iT dsDNA high sensitivity assay kit (Invitrogen, USA) and the Qubit fluorometer (Invitrogen, USA). The gene copy number in the initial standard curve was calculated considering the DNA content, the length of the fragment and the average weight of a base pair (650 Da). A standard curve was constructed by producing a ten times dilution series from 108 to 101 target gene copies per μL.
Sample copy numbers were log transformed and normalize to DNA input.
Statistical analysis
Bacterial fingerprints of each denaturing gradient gel were normalized using the GelCompar 4.0 software (Applied Maths, Belgium), as described by Smalla et al. (2001). Shannon's index of diversity (H′), was determined as H′ = −∑ pi ln pi, where pi is often the proportion of individuals belonging to the each species in the dataset of interest. The existence of significant differences in Shannon's index of diversity (H′) values calculated from each experimental treatment was investigated by using a two-way ANOVA (with processing and preserving procedures being used as the categorical factors and the test also accounting for interactions between both factors). The assumptions of normality and homogeneity of variance were verified by using the Shapiro-Wilks and Levene's test, respectively. Post hoc Tukey HSD test was used whenever significant differences at p < 0.05 were recorded. The two-way ANOVA was performed using the software STATISTICA® 7 (StatSoft Inc., USA).
A Principal Coordinates Analysis (PCO) was performed representing differences among experimental treatments along the first two axes. The raw data matrix was log (x + 1) transformed prior to the statistical analysis in order to place more emphasis on compositional differences among samples rather than on quantitative differences39. A similarity/difference matrix was latter constructed using the Euclidean distance. This multivariate statistical test was performed using Primer 6.1 with PERMANOVA add-on (Primer-E Ltd, UK).
Significant differences among treatments selected for DNA quantification using Real-time PCR were tested using a one-way ANOVA using also the software STATISTICA® 7 (StatSoft Inc., USA). The assumptions of normality and homogeneity of variance were verified by using the Shapiro-Wilks and Bartlett's test, respectively.
References
Daly, M. et al. The phylum Cnidaria: A review of phylogenetic patterns and diversity 300 years after Linnaeus. Zootaxa 127–182 (2007).
Carpenter, K. E. et al. One-third of reef-building corals face elevated extinction risk from climate change and local impacts. Science 321, 560–563 (2008).
Hoegh-Guldberg, O. et al. Coral reefs under rapid climate change and ocean acidification. Science 318, 1737–1742 (2007).
Hughes, T. P. et al. Climate change, human impacts and the resilience of coral reefs. Science 301, 929–933 (2003).
Leal, M. C., Madeira, C., Brandão, C. A., Puga, J. & Calado, R. Bioprospecting of marine invertebrates for new natural products - A zoogeographical and chemical perspective. Molecules 17, 9842–9854 (2012).
Rocha, J., Peixe, L., Gomes, N. C. M. & Calado, R. Cnidarians as a source of new marine bioactive compounds - An overview of the last decade and future steps for bioprospecting. Mar. Drugs 9, 1860–1886 (2011).
Shnit-Orland, M. & Kushmaro, A. Coral mucus-associated bacteria: a possible first line of defense. FEMS Microbiol. Ecol. 67, 371–380 (2009).
Paul, V. & Puglisi, M. Chemical mediation of interactions among marine organisms. Nat. Prod. Rep. 21, 189–209 (2004).
Di Camillo, C. G. et al. Biodiversity of prokaryotic communities associated with the ectoderm of Ectopleura crocea (Cnidaria, Hydrozoa). PLoS ONE 7, e39926 (2012).
Rosenberg, E., Koren, O., Reshef, L., Efrony, R. & Zilber-Rosenberg, I. The role of microorganisms in coral health, disease and evolution. Nat. Rev. Microbiol. 5, 355–362 (2007).
La Riviere, M., Roumagnac, M., Garrabou, J. & Bally, M. Transient shifts in bacterial communities associated with the temperate gorgonian Paramuricea clavata in the northwestern Mediterranean Sea. PLoS ONE 8, e57385 (2013).
Fuhrman, J. A. & Campbell, L. Marine ecology - Microbial microdiversity. Nature 393, 410–411 (1998).
Cleary, D. F. et al. Habitat- and host-related variation in sponge bacterial symbiont communities in Indonesian waters. FEMS Microbiol. Ecol. 85, 465–482 (2013).
White, J. R. et al. Pyrosequencing of bacterial symbionts within Axinella corrugata sponges: diversity and seasonal variability. PLoS ONE 7, e38204 (2012).
Bourne, D. G. et al. Coral reef invertebrate microbiomes correlate with the presence of photosymbionts. The ISME Journal 7, 1452–1458 (2013).
Lee, O. O. et al. Spatial and species variations in bacterial communities associated with corals from the Red Sea as revealed by pyrosequencing. Appl. Environ. Microbiol. 78, 7173–7184 (2012).
Simister, R. L., Schmitt, S. & Taylor, M. W. Evaluating methods for the preservation and extraction of DNA and RNA for analysis of microbial communities in marine sponges. J. Exp. Mar. Biol. Ecol. 397, 38–43 (2011).
Nagy, Z. T. A hands-on overview of tissue preservation methods for molecular genetic analyses. Org. Divers. Evol. 10, 91–105 (2010).
Winston, G. W. & Heffernan, L. M. Development and characterization of Sea Anemones as Bioindicators of Offshore Research Exploitation and Environmental Impact. U.S. Dept. of the Interior, Minerals Management Service, Gulf of Mexico OCS Region, New Orleans, Louisiana. OCS Study MMS. 99–0037. (1999).
Leutenegger, A. et al. Analysis of fluorescent and non-fluorescent sea anemones from the Mediterranean Sea during a bleaching event. J. Exp. Mar. Biol. Ecol. 353, 221–234 (2007).
Margulis, L. & Fester, R. Symbiosis as Source of Evolutionary Innovation: Speciation and Morphogenesis. (MIT Press, 1991).
Ferrara, G. B. et al. The assessment of DNA from marine organisms via a modified salting-out protocol. Cell. Mol. Biol. Lett. 11, 155–160 (2006).
Pinto, S. M., Fernandes-Matioli, F. M. C. & Schlenz, E. DNA extraction from sea anemone (Cnidaria: Actiniaria) tissues for molecular analyses. Genet. Mol. Biol. 23, 601–604 (2000).
Meron, D., Buia, M. C., Fine, M. & Banin, E. Changes in microbial communities associated with the sea anemone Anemonia viridis in a natural pH gradient. Microb. Ecol. 65, 269–276 (2013).
Cleary, D. F. R., Smalla, K., Mendonca-Hagler, L. C. S. & Gomes, N. C. M. Assessment of variation in bacterial composition among microhabitats in a mangrove environment using DGGE fingerprints and barcoded pyrosequencing. PLoS ONE 7, e29380 (2012).
Lai, X. et al. Denaturing gradient gel electrophoresis (DGGE) analysis of bacterial community composition in deep-sea sediments of the South China Sea. World J. Microbiol. Biotechnol. 22, 1337–1345 (2006).
Neilson, J. W., Jordan, F. L. & Maier, R. M. Analysis of artifacts suggests DGGE should not be used for quantitative diversity analysis. J. Microbiol. Methods 92, 256–263 (2013).
Liu, T. Mutational screening of hMLH1 and hMSH2 that confer inherited colorectal cancer susceptibility using denature gradient gel electrophoresis (DGGE). Methods Mol. Biol. 653, 193–205 (2010).
Dawson, M. N., Raskoff, K. A. & Jacobs, D. K. Field preservation of marine invertebrate tissue for DNA analyses. Mol. Mar. Biol. Biotechnol. 7, 145–152 (1998).
Gray, M. A., Pratte, Z. A. & Kellogg, C. A. Comparison of DNA preservation methods for environmental bacterial community samples. FEMS Microbiol. Ecol. 83, 468–477 (2013).
Abad, M. J., Bedoya, L. M. & Bermejo, P. Natural marine anti-inflammatory products. Mini-Rev. Med. Chem. 8, 740–754 (2008).
Post, R. J., Flook, P. K. & Millest, A. L. Methods for the preservation of insects for DNA studies. Biochem. Syst. Ecol. 21, 85–92 (1993).
Reiss, R. A., Schwert, D. P. & Ashworth, A. C. Field preservation of Coleoptera for molecular-genetic analyses. Environ. Entomol. 24, 716–719 (1995).
Leal, M. C., Calado, R., Sheridan, C., Alimonti, A. & Osinga, R. Coral aquaculture to support drug discovery. Trends in Biotechnol. 31, 555–561 (2013).
Weisburg, W. G., Barns, S. M., Pelletier, D. A. & Lane, D. J. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 173, 697–703 (1991).
Heuer, H., Krsek, M., Baker, P., Smalla, K. & Wellington, E. M. H. Analysis of actinomycete communities by specific amplification of genes encoding 16S rRNA and gel-electrophoretic separation in denaturing gradients. Appl. Environ. Microbiol. 63, 3233–3241 (1997).
Riesner, D. et al. Temperature-gradient gel electrophoresis of nucleic acids: analysis of conformational transitions, sequence variations and protein-nucleic acid interactions. Electrophoresis 10, 377–389 (1989).
Nübel, U. E, B., Felske, a., Snaidr, J., Wieshuber, a., Amann, R. I. et al. Sequence heterogeneities of genes encoding 16S rRNAs in Paenibacillus polymyxa detected by temperature gradient gel electrophoresis. J. Bacteriol. 178, 5636–5643 (1996).
PERMANOVA+ for PRIMER: Guide to software and statistical methods. (Primer-E Ltd, Plymouth, United Kingdom, 2008).
Acknowledgements
Joana Rocha is supported by the Portuguese Science Foundation (FCT) through a PhD Grant (SFRH/BD/33476/2008) from PhD Programme in Marine and Environmental Sciences. This work was supported by European Funds through COMPETE and by National Funds through the Portuguese Science Foundation (FCT) within project PEst-C/MAR/LA0017/2013. The authors are grateful to Ana Pires and Rita Polónia for technical support in DGGE.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: J.R., N.C.M.G. and R.C. Performed the experiments: J.R. Analyzed the data: J.R., N.C.M.G. and R.C. Contributed reagents/materials/analysis tools: R.C. Wrote the paper: J.R., F.J.R.C.C., L.P., N.C.M.G. and R.C.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Rocha, J., Coelho, F., Peixe, L. et al. Optimization of preservation and processing of sea anemones for microbial community analysis using molecular tools. Sci Rep 4, 6986 (2014). https://doi.org/10.1038/srep06986
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06986
This article is cited by
-
The antibacterial and antibiofilm activity of sea anemone (Stichodactyla haddoni) against antibiotic-resistant bacteria and characterization of bioactive metabolites
International Aquatic Research (2019)
-
The stable microbiome of inter and sub-tidal anemone species under increasing pCO2
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.