Abstract
Human leukocyte antigen (HLA)-DR4/HLA-DRB1*04 has been reported to be a risk factor for Vogt-Koyanagi-Harada disease (VKH) with various strength of association. Its sub-alleles were also found to be associated with VKH. However the results were inconsistent. In this study, we systematically searched the related literature, pooled the odds ratios (ORs) and 95% confidence interval (CI) of association of HLA-DR4/HLA-DRB1*04 or its sub-alleles with VKH from individual studies and explored the potential source of heterogeneity. A total of 1853 VKH patients and 4164 controls from 21 articles were included in this meta-analysis. The pooled OR of association of HLA-DR4/HLA-DRB1*04 and VKH was 8.42 (95% CI: 5.69–12.45). There were significant heterogeneity (I2 = 71%). Subgroup analysis indicated that ethnicity was the source of heterogeneity (all I2 = 0, ORs ranged from 2.09–13.69 in subgroups). The sub-alleles, HLA-DRB1*0404 (OR = 2.57), 0405 (OR = 10.31) and 0410 (OR = 6.52) increased the risk of VKH; 0401 (OR = 0.21) protected VKH; while other sub-alleles were not associated with VKH. Our meta-analysis confirmed the association between VKH and HLA-DR4/DRB1*04, found the strength of association is different in different ethnic groups and identified HLA-DRB1*0404, 0405 and 0410 as risk sub-alleles while 0401 as protective sub-allele.
Similar content being viewed by others
Introduction
Vogt-Koyanagi-Harada disease (VKH) is a systematic autoimmune disorder that affects tissues containing melanin such as the eyes, inner ears, meninges and skin1. The ocular manifestations are characterized by chronic bilateral, diffuse, granulomatous uveitis, which may lead to blindness. Several risk factors have been identified for VKH, including dark skin pigmentation, females, aged between 20 to 50 years and genetic factors2,3,4.
Human leukocyte antigen (HLA) system is the locus of genes that encode for major histocompatibility complex (MHC), which is a set of cell surface molecules mediating interaction of leukocyte5. Therefore, HLA plays an important role in immune system function as well as in the pathogenesis of autoimmune diseases, including VKH6. Almost 40 years ago, the association of HLA-BW22J and VKH was reported7. However, it could not be replicated in subsequent studies8. Later on, more articles have been published on the association of different types of HLA and VKH. Among them, the HLA-DR4 serotype, its corresponding allele HLA-DRB1*04 were mostly investigated9,10. Their results, however, are inconsistent, especially on the strengths of association with reported risk increases variably. The sample size of most studies is small. Recently, there is a review summarizing the genetic studies of VKH11. However, it is a narrative review and did not quantitatively synthesize the results from individual studies.
In this study, we performed a systematic review and meta-analysis to investigate the association between VKH withHLA-DR4/HLA-DRB1*04 and its sub-alleles, combine the small sample size studies and explore the sources of the inconsistency.
Methods
Search strategy
Literature search was performed on MEDLINE, Science citation index, SCOPUS databases using PubMed, Web of Science and SCOPUS search engines up to June 1, 2014. The following medical subject headings and keywords were used for search strategy: “human leukocyte antigen” OR “HLA” OR “major histocompatibility complex” OR “MHC” AND (“VKH” OR “Vogt Koyanagi Harada”). No language or year of publication restrictions was imposed.
Study selection
The retrieved records from literature search were reviewed by two reviewers independently (T.K.S, W.J.L). Any disagreement was solved by discussion and consensus together with a third author (H.Y.C). Included studies met the following pre-specified criteria: 1) Association study of HLA-DR4/HLA-DRB1*04 or its sub-alleles with VKH; 2) The number or percentage of HLA-DR4 serotype and/or HLA-DRB1*04 allele/sub-alleles must be provided in VKH cases and controls; 3) The type of article is an original research study, not a review, case report, or editorial comment. The studies that did not provide sufficient information even after contacting the corresponding author were excluded. Considering some studies may contain overlapping cases and/or controls, we paid close attention to the authors, study subject's geographic location and numbers of subjects. For duplicated studies, we selected the publication that contained the most number of VKH cases.
Data extraction
Data extraction was carried out by two reviewers independently (T.K.S, W.J.L). Any disagreement was solved by discussion and consensus together with a third author (H.Y.C). The following data from each included study were collected: author, year of publication, study design, ethnicity, the mean age and gender of cases and controls, diagnostic criteria used for VKH, counts or frequencies of HLA-DR4/HLA-DRB1*04 and its sub-alleles in cases and controls.
Risk of bias assessment
The quality evaluation was also carried out by two reviewers (T.K.S, W.J.L) independently. Further independent review and resolution by a third reviewer (C.H.Y.) was sought if the two reviewers disagreed. The risk of bias assessment considered 6domainsas suggested in the HuGENet handbook12: bias in ascertainment of cases, bias in ascertainment of controls, bias in genotyping controls, bias in population stratification, confounding bias, multiple test and selective outcome reports. Each item was classified by “yes/no” to risk of bias or as “unclear” if there was no sufficient information to assess.
Statistical analysis
Statistical analysis was performed using Review Manager (version 5.2.6.0; Copenhagen: The Nordic Cochrane Centre, The Cochrane Collaboration, 2012) and STATA (version 419.12.0.866, STATA Corp LP, College Station, Texas).For individual study, we calculated the odds ratio (OR) and 95% confidence interval (CI), pooled these data to compare HLA-DR4/HLA-DRB1*04 frequencies between VKH patients and controls. Between-study heterogeneity was assed using the Q statistics and quantified using the I2 statistic (I2 = 0–25%, low heterogeneity; I2 = 25%–50%, moderate heterogeneity; I2 = 50%–75% large heterogeneity; I2 = 75%–100%, extreme heterogeneity)13. Pooled ORs and 95% CIs were computed using fix-effects models when I2 < 25%, or random-effect models when I2 > = 25%. We performed ethnicity-based sub-group analysis to investigate the strength of association in different ethnicity. If ethnicity cannot explain heterogeneity, univariate random-effects meta-regression was used to investigate the potential sources of heterogeneity, such as, typing technique, publication language and publication year. Funnel plot and Egger's test were used to assess publication bias, sensitivity analyses were conducted by Exclusion sensitivity plot analysis. P value less than 0.05 were considered significantly except for Q test and Egger's test, where 0.1 was considered as significant level.
Results
General characteristic of included studies
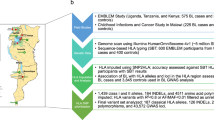
Studies selection process was showed in Figure 1. Among 404 records retrieved from databases, 169 were excluded in initial screening, 197 were excluded because they were unrelated studies, reviews, case reports or animal studies. After reviewing the full text, 11 studies were excluded because they had overlapping information in cases and/or controls with another publications14,15,16,17,18,19,20,21,22,23,24 and 4 were excluded because they investigated other variants but not HLA-DR4/HLA-DRB1*0425,26,27,28, two were excluded because there was no control29,30. Eventually, 21 studies6,9,10,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48 were retained that contributed data to association of VKH and HLA-DR4/HLA-DRB1*04. There were 19 articles in English, 2 in Chinese. The characteristics of included studies are presented in Table 1.
Study bias assessment
Possible bias of the included studies is showed in Table 2. Overall the quality of included studies was good. Two (10%) studies10,49 had bias of ascertainment in cases, 2 (10%) studies42,45 were unclear in bias in population stratification and one (5%) study10 had bias in ascertainment of controls. No study had bias in genotyping controls, confounding bias or selective outcome report.
Association of HLA-DR4/DRB1*04with VKH
In 19 case-control studies investigating the association of HLA-DR4/HLA-DRB1*04 with VKH, the ORs of individual studies ranged from 1.57 to 125.00. The heterogeneity across studies was high with I2 = 71%, p < 0.00001. Sub-group analysis based on ethnicity showed the I2 = 0 in all four subgroups, with OR = 13.69 (95%CI: 10.58, 17.69), 8.67 (95%CI: 2.75, 27.31), 4.79 (95%CI: 3.24, 7.09) and 2.09 (95%CI: 1.25, 3.48) in Eastern Asians, Italians, Hispanics and Indians respectively. There was significant difference among the subgroups (I2 = 71%, p < 0.00001, Figure 2A). The pooled OR of all studies was 8.42 (95% CI: 5.69, 12.45). The funnel plot did not show any obvious evidence of asymmetry (Figure 2B) and the p value of Egger's test was 0.457. Therefore, publication bias was not evident in this meta-analysis. Exclusion sensitivity plot showed that pooled OR was not influenced by omitting any study (Figure 2C).
Meta-analysis of the association of HLA-DR4/HLA-DRB1*04 with Vogt-Koyanagi-Harada (VKH) disease.
(A): Forest plot showing the odds ratios (ORs) of VKH carrying HLA-DR4/HLA-DRB1*04 in individual studies, sub-groups based on ethnicity and the pooled results. (B): Funnel plots for positive rate of HLA-DR4/HLA-DRB1*04 between VKH cases and controls. (C). Exclusion sensitivity plot showing the results of pooled ORs after omitting each study.
Association of HLA- DRB1*04 sub-alleles with VKH
The association of HLA-DRB1*0401, 0402, 0403, 0404, 0405, 0406, 0407, 0408, 0410, 0411, 0417, 0437 with VKH was investigated (Table 3 and Figure 3). Among them, no statistically significant association was found between VKH with HLA-DRB1*0402, 0403, 0406, 0407, 0410, 0411, 0417, or 0437. HLA-DRB1*0401 investigated in 6 studies. The individual OR was statistically significant in only one study, but the pooled OR was 0.21 (95% CI: 0.07–0.65, I2 = 0%, Figure 3A). HLA-DRB1*0404 was investigated in 6 articles and only two of them reported significant association with VKH. The pooled OR was 2.57 (95% CI: 1.54–4.32, I2 = 0%, Figure 3D). HLA-DRB1*0410 was investigated in 8 studies. Statistically significant association of HLA-DRB1*0410 with VKH was reported in two of them and the pooled OR was 6.52 (95% CI: 3.23–13.18, I2 = 0%, Figure 3I). HLA-DRB1*0405 was investigated in 12 studies and the pooled OR was 10.31 (95% CI: 5.56–19.11, I2 = 77%, Figure 3E). Meta-regression found that publication year, ethnicity or publication language cannot explain the between studies heterogeneity (all p > 0.05).
Discussion
The present meta-analysis, including 1853 VKH patients and 4164 controls from 21 articles, investigated the association of HLA-DR4/HLA-DRB1*04 and its sub-alleles with VKH. Our results indicate that HLA-DR4/HLA-DRB1*04 carriers have an increased risk of VKH with OR 8.42. The strength of this association is highest in Eastern Asian and lowest in Indians. Some of HLA-DRB1*04's sub-alleles HLA-DRB1*0404, 0405 and 0410 increased the risk of VKH; 0401 reduced risk of VKH; while 0402, 0403, 0406, 0407, 0410, 0411, 0417, or 0437 was not associated with VKH.
The association of HLA-DR4/HLA-DRB1*04 with VKH was reported in various ethnic populations. Our meta-analysis confirms these previous reports on the association of HLA-DR4/HLA-DRB1*04 with VKH. VKH appears to occur commonly among communities with dark pigment such as Native American, Arabian, Eastern Asian and Indian but not in blacks of sub-Saharan Africans or Caucasians50. It is known that the strength of HLA-disease associations can vary among different racial groups51. Also, different racial groups can have distinct HLA associations with a common, clinically identified disease. With the power of meta-analysis, here we pooled all the published results of the association between VKH with HLA-DR4/HLA-DRB1*04 and demonstrated that ethnicity was the source of heterogeneity. The OR was 13.69 (95%CI: 10.58, 17.69), 8.67 (95%CI: 2.75, 27.31), 4.79CI (95%CI: 3.24, 7.09) and 2.09 (95%CI: 1.25, 3.48) in Eastern Asian, Italian, Hispanic and Indian respectively.
In this meta-analysis, we not only confirmed the association of HLA-DR4/HLA-DRB1*04 with VKH, but also identified the sub-alleles, HLA-DRB1*0404, 0405, 0410 as the risk alleles of VKH, while HLA-DRB1*0401 as the protective allele for VKH. HLA genes and proteins are known to be highly polymorphic, i.e. they exist in many different forms among humans and play an essential role in recognition of immune system52. HLA-DR is a MHC class II cell surface receptor, which displays peptides antigens produced from the HLA-DRB1 gene to the immune system. If the immune system recognizes the peptides as foreign (such as viral or bacterial peptides), it triggers a response to attack the invading viruses or bacteria. It was reported that VKH patients are sensitized to melanocyte epitopes, while patients with HLA-DRB1*0405 recognize a broader melanocyte-derived peptide repertoire53. The functional correlation of HLA-DRB1*0404, 0410 or 0401 with melanocyte epitopes has not been reported. Possibly HLA-DRB1*0404 and 0410 may have broader melanocyte-derived peptide repertoire as 0405, while HLA-DRB1*0401 may have narrower melanocyte epitopes.
Among the sub-alleles of HLA-DRB1*04, HLA-DRB1*0405 was the most investigated allele. Statistically significant association of HLA-DRB1*0405 with VKH was reported in most original studies except in Levinson's article42. Our meta-analysis confirmed the positive association of HLA-DRB1*0405 with VKH, although there was some heterogeneity that cannot be explained. Statistical significant association of VKH with HLA-DRB1*0404, 0410 or 0401 was reported in only a few studies but not others. The inconsistency of results from different publications may be due to a small sample size in individual studies. With the power of meta-analysis, the sample size was pooled and increased and we were able to resolve the inconsistency among publications and identify HLA-DRB1*0404 and 0410 as risk alleles while 0401 as protective allele.
This meta-analysis suggests that in clinical practice, genotyping of HLA-DRB1*0404, 0410 and 0401 is recommended for VKH patients in addition to HLA-DRB1*0405. The genotyping of HLA-DRB1*0402, 0403, 0406, 0407, 0410, 0411, 0417, or 0437 is not necessary. Further studies are needed to investigate the functional implication of HLA-DRB1*0404, 0405, 0410 and 0401, which may provide further insight into the pathogenesis of VKH.
Our study has some limitations. First, we did not identify the source of heterogeneity in the association of HLA-DRB1*0405 with VKH after exploring ethnicity, publication year and publication language. Some studies did not give detailed data such as onset/study age, gender percentage. Therefore we could not estimate them further in meta-regression. Second, there may be some original studies not retrieved by the current literature search. We did not search the Japanese database. Although some Japanese medical journals are indexed in PubMed and Embase, we still cannot eliminate if some articles in Japanese or other language were missed. Third, the diagnostic criteria of VKH adopted in individual studies are different, which may contribute to the heterogeneity of association.
In conclusion, this meta-analysis demonstrates a strong association between HLA-DR4/HLA-DRB1*04 and VKH. The strength of association was variable in different ethnicities. The sub-alleles, HLA-DRB1*0404, 0405, 0410 were risk factors of VKH, while HLA-DRB1*0401 was the protective factor.
References
Moorthy, R. S., Inomata, H. & Rao, N. A. Vogt-Koyanagi-Harada syndrome. Surv Ophthalmol 39, 265–292 (1995).
Yang, P. et al. Clinical characteristics of Vogt-Koyanagi-Harada syndrome in Chinese patients. Ophthalmology 114, 606–614 (2007).
Sukavatcharin, S., Tsai, J. H. & Rao, N. A. Vogt-Koyanagi-Harada disease in Hispanic patients. Int Ophthalmol 27, 143–148 (2007).
Damico, F. M., Kiss, S. & Young, L. H. Vogt-Koyanagi-Harada disease. Semin Ophthalmol 20, 183–190 (2005).
Caillat-Zucman, S. Molecular mechanisms of HLA association with autoimmune diseases. Tissue Antigens 73, 1–8 (2009).
Davis, J. L. et al. HLA associations and ancestry in Vogt-Koyanagi-Harada disease and sympathetic ophthalmia. Ophthalmology 97, 1137–1142 (1990).
Tagawa, Y., Sugiura, S., Yakura, H., Wakisaka, A. & Aizawa, M. Letter: HLA and Vogt-Koyanagh-Harada syndrome. N Engl J Med 295, 173 (1976).
Douglass, S. N. & Douglass, J. M. Letter: Hla and vogt-koyanagi-harada syndrome. N Engl J Med 295, 788 (1976).
Shindo, Y., Ohno, S., Yamamoto, T., Nakamura, S. & Inoko, H. Complete association of the HLA-DRB1*04 and -DQB1*04 alleles with Vogt-Koyanagi-Harada's disease. Hum Immunol 39, 169–176 (1994).
Islam, S. M. et al. HLA class II genes in Vogt-Koyanagi-Harada disease. Invest Ophthalmol Vis Sci 35, 3890–3896 (1994).
Ng, J. Y., Luk, F. O., Lai, T. Y. & Pang, C. P. Influence of molecular genetics in Vogt-Koyanagi-Harada disease. J Ophthalmic Inflamm Infect 4, 20 (2014).
Little, J. & Higgins, J. P. T. (editors). The HuGENet™ HuGE Review Handbook, version 1.0. http://www.hugenet.ca (2006).
Higgins, J. P., Thompson, S. G., Deeks, J. J. & Altman, D. G. Measuring inconsistency in meta-analyses. BMJ 327, 557–560 (2003).
Shindo, Y., Inoko, H., Yamamoto, T. & Ohno, S. HLA-DRB1 typing of Vogt-Koyanagi-Harada's disease by PCR-RFLP and the strong association with DRB1*0405 and DRB1*0410. Br J Ophthalmol 78, 223–226 (1994).
Islam, S. M. et al. Influence of HLA-DRB1 gene variation on the clinical course of Vogt-Koyanagi-Harada disease. Invest Ophthalmol Vis Sci 35, 752–756 (1994).
Alaez, C. et al. Strong association of HLA class II sequences in Mexicans with Vogt-Koyanagi-Harada's disease. Hum Immunol 60, 875–882 (1999).
Arellanes-Garcia, L. et al. Strong association of Vogt Koyanagi Harada disease with DRB1-04 residues and DQB1*0302 in Mexican mestizo patients. Invest Ophthalmol Vis Sci 38, S526 (1997).
Flores-A, H. et al. Vogt-Koyanagi-Harada (VKH) in Mexicans is strongly associated with DRB1*0405, with no contribution of HLA-Class I loci. Tissue Antigens 75, 625–626 (2010).
Li, K. et al. Polymorphisms of FCRL3 in a Chinese population with Vogt-Koyanagi-Harada (VKH) syndrome. Mol Vis 15, 955–961 (2009).
Meng, Q. et al. PDCD1 genes may protect against extraocular manifestations in Chinese Han patients with Vogt-Koyanagi-Harada syndrome. Mol Vis 15, 386–392 (2009).
Mora, P., Bautista, N., Arellanes, L. & Gorodezky, C. HLA associations and clinical features in Mexican Mestizos with VOGT Koyanagi Harada desease (VKH). Invest Ophthalmol Vis Sci 37, S363 (1996).
Shindo, Y., Ohno, S., Nakamura, S., Onoe, K. & Inoko, H. A significant association of HLA-DPB1(*)0501 with Vogt-Koyanagi-Harada's disease results from a linkage disequilibrium with the primarily associated allele, DRB1(*)0405. Tissue Antigens 47, 344–345 (1996).
Yamamoto, T., Sasaki, T. & Ogawa, K. Correlation between HLA antigens and clinical manifestations in Harada disease. Rinsho Ganka 45, 1657–1661 (1991).
Yamamoto, J. H. et al. HLA-DR4 in VOGT-Koyanagi-Harada's disease in Brazilian patients. Invest Ophthalmol Vis Sci 37, S363 (1996).
Sheereen, A. et al. A study of KIR genes and HLA-C in Vogt-Koyanagi-Harada disease in Saudi Arabia. Mol Vis 17, 3523–3528 (2011).
Min, H. Y. et al. Polymorphism of HLA-DQB1 alleles in Chinese Han patients with Vogt-Koyonagi-Harada syndrome. Zhonghua yan ke za zhi 43, 355–360 (2007).
Martinez, J. A. et al. Vogt-Koyanagi-Harada syndrome in patients with Cherokee Indian ancestry. Am J Ophthalmol 114, 615–620 (1992).
Lin, C. P., Hou, P. K., Hung, P. T. & Chen, M. S. Disease pattern of uveitis in Taiwan. Invest Ophthalmol Vis Sci 37, S1038 (1996).
Abad, S. et al. Characteristics of Vogt-Koyanagi-Harada disease in a French cohort: ethnicity, systemic manifestations and HLA genotype data. Ocul Immunol Inflamm 16, 3–8 (2008).
Pezzi, P. P., Paroli, M. P., Priori, R., Da Dalt, S. & Corradi, R. Vogt-Koyanagi-Harada syndrome and keratoconjunctivitis sicca. Am J Ophthalmol 137, 769–770 (2004).
Alaez, C. et al. Major histocompatibility complex and strong human leukocyte antigen-DRB1 and gender association with Vogt-Koyanagi-Harada syndrome in Mexican Mestizos. Hum Immunol 72, 1198–1203 (2011).
Zhao, M., Jiang, Y. & Abrahams, I. W. Association of HLA antigens with Vogt-Koyanagi-Harada syndrome in a Han Chinese population. Arch Ophthalmol 109, 368–370 (1991).
Numaga, J., Matsuki, K., Tokunaga, K., Juji, T. & Mochizuki, M. Analysis of human leukocyte antigen HLA-DR beta amino acid sequence in Vogt-Koyanagi-Harada syndrome. Invest Ophthalmol Vis Sci 32, 1958–1961 (1991).
Zhang, X. Y., Wang, X. M. & Hu, T. S. Profiling human leukocyte antigens in Vogt-Koyanagi-Harada syndrome. Am J Ophthalmol 113, 567–572 (1992).
Weisz, J. M. et al. Association between Vogt-Koyanagi-Harada syndrome and HLA-DR1 and -DR4 in Hispanic patients living in southern California. Ophthalmology 102, 1012–1015 (1995).
Pivetti-Pezzi, P., Accorinti, M., Colabelli-Gisoldi, R. A. & Pirraglia, M. P. Vogt-Koyanagi-Harada disease and HLA type in Italian patients. Am J Ophthalmol 122, 889–891 (1996).
Xiao, T., Jiang, Y. & You, X. The association of HLA-DR4 gene subtypes with Vogt-Koyanagi-Harada syndrome. Zhonghua Yan Ke Za Zhi 33, 268–271 (1997).
Arellanes-Garcia, L. et al. HLA-DR is strongly associated with Vogt-Koyanagi-Harada disease in Mexican Mestizo patients. Ocul Immunol Inflamm 6, 93–100 (1998).
Goldberg, A. C. et al. HLA-DRB1*0405 is the predominant allele in Brazilian patients with Vogt-Koyanagi-Harada disease. Hum Immunol 59, 183–188 (1998).
Kim, M. H. et al. Association of HLA with Vogt-Koyanagi-Harada syndrome in Koreans. Am J Ophthalmol 129, 173–177 (2000).
Zhang, M., Qiu, C. & Hu, T. Association of HLA-DRB genes with Vogt-Koyanagi-Harada syndrome in a Chinese Han population. Zhongguo Yi Xue Ke Xue Yuan Xue Bao 22, 36–40 (2000).
Levinson, R. D. et al. HLA-DRB1 and -DQB1 alleles in mestizo patients with Vogt-Koyanagi-Harada's disease in Southern California. Hum Immunol 65, 1477–1482 (2004).
Horie, Y. et al. Tyrosinase gene family and Vogt-Koyanagi-Harada disease in Japanese patients. Mol Vis 12, 1601–1605 (2006).
Hou, S. P. et al. Small ubiquitin-like modifier 4 (SUMO4) polymorphisms and Vogt-Koyanagi-Harada (VKH) syndrome in the Chinese Han population. Mol Vis 14, 2597–2603 (2008).
Iqniebi, A. et al. HLA-DRB1 among patients with Vogt-Koyanagi-Harada disease in Saudi Arabia. Mol Vis 15, 1876–1880 (2009).
Tiercy, J. M., Rathinam, S. R., Gex-Fabry, M. & Baglivo, E. A shared HLA-DRB1 epitope in the DR beta first domain is associated with Vogt-Koyanagi-Harada syndrome in Indian patients. Mol Vis 16, 353–358 (2010).
Gupta, A., Kamal, S., Gupta, V., Bambery, P. & Kaura, B. HLA typing in Vogt-Koyanagi-Harada syndrome in North Indian patients. Ocul Immunol Inflamm 15, 89–97 (2007).
Nomura, S. et al. Clinical significance of HLA-DRB1*0410 in Japanese patients with idiopathic thrombocytopenic purpura. Blood 91, 3616–3622 (1998).
Hou, S. et al. Small ubiquitin-like modifier 4 (SUMO4) polymorphisms and Vogt-Koyanagi-Harada (VKH) syndrome in the Chinese Han population. Mol Vis 14, 2597–2603 (2008).
Read, R. W. et al. Revised diagnostic criteria for Vogt-Koyanagi-Harada disease: report of an international committee on nomenclature. Am J Ophthalmol 131, 647–652 (2001).
de Menthon, M., Lavalley, M. P., Maldini, C., Guillevin, L. & Mahr, A. HLA-B51/B5 and the risk of Behcet's disease: a systematic review and meta-analysis of case-control genetic association studies. Arthritis Rheum 61, 1287–1296 (2009).
Apanius, V., Penn, D., Slev, P. R., Ruff, L. R. & Potts, W. K. The nature of selection on the major histocompatibility complex. Crit Rev Immunol 17, 179–224 (1997).
Damico, F. M. et al. T-cell recognition and cytokine profile induced by melanocyte epitopes in patients with HLA-DRB1*0405-positive and -negative Vogt-Koyanagi-Harada uveitis. Invest Ophthalmol Vis Sci 46, 2465–2471 (2005).
Acknowledgements
This study was supported by the National Nature Science Foundation of China (81170853) and Guangdong Science and Technology Project (2011B031300013).
Author information
Authors and Affiliations
Contributions
H.C. designed the study. T.S., W.L. and L.Z. conducted the study. T.S. and H.C. analyzed the data. T.S. wrote the main manuscript text. J.C. and H.C. revised the manuscript. All authors reviewed and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Shi, T., Lv, W., Zhang, L. et al. Association of HLA-DR4/HLA-DRB1*04 with Vogt-Koyanagi-Harada disease: A Systematic Review and Meta-analysis. Sci Rep 4, 6887 (2014). https://doi.org/10.1038/srep06887
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06887
This article is cited by
-
Successive onset of Vogt-Koyanagi-Harada syndrome in father and son
BMC Ophthalmology (2023)
-
Unusual neurologic manifestations of Vogt-Koyanagi-Harada disease: a systematic literature review
BMC Neurology (2022)
-
Case with metastatic cutaneous malignant melanoma that developed Vogt–Koyanagi–Harada-like uveitis following pembrolizumab treatment
Documenta Ophthalmologica (2021)
-
Proliferative retinopathy as a feature of Vogt Koyanagi Harada Disease: a report of two cases
BMC Ophthalmology (2020)
-
Vogt-Koyanagi-Harada disease: a retrospective and multicentric study of 41 patients
BMC Ophthalmology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.