Abstract
Combinations of fluorescent proteins (FPs) are routinely used for multi-parameter in vivo imaging experiments to visualize tagged proteins or cell populations of interest. Studies involving FPs are often limited by spectral overlap, toxicity, relative quantum efficiency and the potential for immunological rejection upon transfer into a non-tolerant recipient. Here we evaluate the immunologic visibility of several commonly used FPs by the murine immune system and identify a spectrally compatible, immunologically tolerated combination of FPs well suited for in vivo two-photon imaging.
Similar content being viewed by others
Introduction
Two-photon laser scanning microscopy (TPLSM)1 has vertically enhanced our understanding of biological systems by allowing real-time deep tissue imaging2,3. Two-photon microscopes are usually equipped with titanium-doped sapphire (Ti:sapphire) lasers that generate pulsed near-infrared light3,4. Using a Ti:sapphire laser, excitation of a fluorophore is achieved when two photons are absorbed simultaneously5,6. This ensures that the probability of excitation is highest in the focal plane, thus minimizing undesirable excitation above and below this plane. Moreover, use of near-infrared light allows deeper penetration into tissues than can be achieved with conventional single photon lasers. In fact, imaging depths of greater than 1 mm have been reported in the literature7. Given these advantages and others, TPLSM has become an invaluable microscopy technique for imaging living systems and its use has expanded greatly in recent years2,3.
Because TPLSM allows design of 4-dimensional (4D) multi-parameter in vivo imaging experiments, it is routine to utilize panels of fluorescent probes, proteins, etc. to visualize multiple cell populations and/or structures of interest. This requires optimization to ensure compatibility among the selected fluorophores. Use of fluorescent proteins became a mainstay in the imaging community following the discovery of green fluorescent protein (GFP), which was isolated from the jellyfish Aequorea Victoria8,9,10. The palette of available fluorescent proteins (FPs) has expanded considerably since then to include derivatives of GFP such as cyan fluorescent protein (CFP) and yellow fluorescent protein (YFP)11. FPs have also been isolated from other species such as DsRed from the coral Discosoma12. DsRed is a tetrameric red fluorescent protein that was later used to generate derivatives like monomeric red fluorescent protein (mRFP), monomeric orange fluorescent protein (mOrange) and mCherry13. There are now more than 50 different FPs available with spectral properties ranging from blue to far red11,14. However, it is important to consider that not all FP combinations are ideal and some FPs can be toxic or even immunologically rejected upon transfer into a non-tolerant host15,16. We therefore set out in this study to evaluate the immunologic visibility of several commonly used FPs and to identify a combination of FPs that are bright, non-toxic, spectrally separable and immunologically tolerated following adoptive transfer.
At present, it is standard practice in two-photon imaging experiments to adoptively transfer FP-tagged cells from a FP-positive donor into a genetically identical FP-negative recipient. Because FPs are non-self, adoptive transfers have the potential to result in a FP-specific immune response16. To evaluate the immunological “rejectability” of three commonly used FPs (i.e. CFP, GFP and DsRed), we set up an adoptive transfer paradigm to monitor FP rejection over time. This was accomplished by crossing mice that express CFP, GFP, or DsRedT3 under the β-actin promoter with CD8+ T cell receptor (TCR) transgenic mice that recognize the glycoprotein (GP), amino acids 33–41, of lymphocytic choriomeningitis virus (LCMV) presented in H-2Db (referred to as P14 mice)17. The resultant F1 crosses yielded P14 T cells expressing CFP, GFP, or DsRed under the β-actin promoter. These cells were isolated from the spleen and 5,000 of each population were adoptively transferred into C57BL/6J (B6) recipients. One day later the mice were challenged intraperitoneally with lymphocytic choriomeningitis virus (LCMV) Armstrong strain, which results in an acute viral infection that is cleared in 8–10 days18. Following infection CFP, GFP and DsRed P14 cells divided extensively and could be monitored flow cytometrically at weekly intervals in the blood over time (Fig. 1A,B). All cell populations were observed at comparable frequencies in the blood at day 7 post-infection (Fig. 1A). LCMV-specific CD8+ T cell expansion is ordinarily followed by contraction, and, ultimately, the stable maintenance of memory T cells19. CFP and GFP P14 cells underwent contraction and were stably maintained in circulation 3 months following infection (Fig. 1A,B). In contrast, DsRed P14 cells rapidly contracted between 7 and 14 days post-infection and were no longer present in circulation by day 21. This is consistent with DsRed being recognized as a foreign protein and rejected by the murine immune system16.
Evaluation of the immune reaction to commonly used fluorescent proteins.
(A, B) Naïve B6 mice (n = 5 per group) were seeded i.v. with 5,000 fluorescent protein (FP)-expressing P14 CD8+ T cells and infected one day later i.p. with 2 × 105 PFU LCMV Arm. The percentage of FP-expressing CD8+ T cells were monitored in the blood at the indicated time points. Representative flow cytometric plots gated on CD45+ CD8+ T cells are shown in panel A. Pink boxes denote the percentage of FP-expressing CD8+ T cells. Note that DsRed+ P14 cells are completely eliminated by day 28. In panel B, data are plotted over time as the percentage of FP+ P14 cells (mean ± SD) remaining relative to day 7. The colored lines represent fitted curves depicting P14 decay over time. (C) The upper panel shows a schematic of the NSE-DsRedE2 construct used to generate B6 NSE-DsRedE2 transgenic mice. The lower bar graph shows that absolute quantity of DsRed mRNA in 1 μg of total RNA harvested from the denoted tissues. Data are represented as mean ± SD (n = 4 mice). (D) 5,000 DsRed+ P14 cells were adoptively transferred into naïve B6 or NSE-DsRed mice (n = 5 mice per group) and infected one day later with LCMV Arm. The fate of GFP+ P14 cells in B6 mice served as a control for this experiment. Normalized percentages (mean ± SD) are plotted over time in a manner identical to that described in panel B. DsRed+ P14 cells are maintained in NSE-DsRed but not B6 mice. (E) ELISAs were performed to detect anti-DsRed antibodies. Representative ELISA data are plotted for serially diluted sera obtained at day 15 post-infection from B6 and NSE-DsRed mice seeded with DsRed+ P14 cells (n = 5 mice per group). Anti-DsRed and anti-GFP antibodies served as positive and negative controls, respectively. (F) Curves analogous to those shown in panel E were generated at the denoted time points. Fluorescence emission from the third dilution is plotted versus time. Data are represented as mean ± SD (n = 5 mice per group). B6 mice seeded with DsRed+ P14 cells generate anti-DsRed antibodies, whereas NSE-DsRed mice do not. All data are representative of two independent experiments.
To definitively address this hypothesis and determine whether tolerance to DsRed could be established, we performed adoptive transfer studies in transgenic mice expressing DsRed (DsRed Express 2) under the neuron specific enolase promoter20 (NSE-DsRedE2) (Fig. 1C, D). Quantitative PCR revealed that DsRed was expressed at detectable levels in the brain and kidneys of NSE-DsRedE2 mice, potentially allowing for induction of peripheral tolerance to DsRed due to its expression as a self-protein. We next adoptively transferred DsRed P14 cells into NSE-DsRedE2 mice and compared their expansion and contraction kinetics following LCMV infection to cells transferred into wild type B6 controls. Importantly, DsRed P14 cells were stably maintained in NSE-DsRedE2 mice (comparable to GFP P14 cells) but not wild type B6 mice (Fig. 1D). To confirm that NSE-DsRedE2 mice were indeed immunologically tolerant to DsRed, we quantified serum antibody titers to DsRed over time (Fig. 1E, F). A marked increase in serum anti-DsRed antibody titers were observed in B6 mice between 7 and 14 days post-infection that paralleled the kinetics of DsRed P14 cell rejection in these mice (Fig. 1B, D). No detectable anti-DsRed antibody response was observed in NSE-DsRed mice, indicating that immunological tolerance to DsRed had been established.
Given the issues associated with DsRed and the degree of spectral overlap between CFP, GFP and YFP, we set out to develop a palette of FPs that were spectrally separable and would not be rejected in a B6 host. To this end, we generated two new transgenic reporter mice expressing mOrange or monomeric teal fluorescent protein (mTFP1) under the CMV early enhancer/chicken β-actin (CAG) promoter (Fig. 2A, B). mOrange is a monomeric derivative of DsRed that can be excited with longer wavelengths of two-photon light (~1000 nm) and emits between 549 and 565 nm12, making it spectrally separable from common GFP-derivatives like CFP and YFP. mTFP1, on the other hand, was previously engineered based on tetrameric CFP from Clavularia coral21. We selected mTFP1 because of its photostability, high quantum yield (~2-fold higher than enhanced GFP) and emission peak at 492 nm, which provided a superior replacement for CFP.
Generation of transgenic fluorescent protein reporter mice.
(A, B) The upper panels show schematics of the constructs used to generate transgenic mice expressing mTFP1 or mOrange under control of the CAG promoter. Flow cytometric dot plots in the lower panels (gated on CD45+ CD8+ cells) show FP expression in CD8+ T cells from three representative founder lines with low (left), intermediate (middle) and high (right) expression. Pink boxes and associated numbers denote the percentage of FP-expressing CD8+ T cells. Only founders with homogenous expression of mTFP1 or mOrange in CD45+ hematopoietic cells (right) were selected for backcrossing. (C, D) 5,000 mTFP1+ or mOrange+ P14 cells were adoptively transferred into naïve B6 mice (n = 5 mice per group) and infected one day later with LCMV Arm. Thy1.1+ mRFP+ P14 cells were also evaluated in this experiment as another monomeric derivative of DsRed. Panel C shows representative flow cytometric plots at day 7 and 28 post-infection. All plots are gated on CD45+ CD8+ T cells. Pink boxes and associated numbers denote the percentage of FP-expressing CD8+ T cells. Normalized data (mean ± SD) are plotted versus time in panel D. The dotted live denotes the threshold of detection. Note that all FP+ P14 cells are stably maintained in circulation following LCMV infection. Data shown in this figure are representative of two independent experiments.
Potential CAG-mOrange and CAG-mTFP1 founder lines were screened flow cytometrically for uniform FP expression in the hematopoietic system (Fig. 2A, B). Flow cytometric analyses revealed a mosaic of expression patterns, with the percentage of FP-positive hematopoietic cells ranging from 17% to greater than 95% (Fig. 2A, B). Mice with homogenous FP expression in >95% of circulating CD45+ hematopoietic cells were selected as founders for backcrossing. After establishing the founder lines, CAG-mOrange and CAG-mTFP1 mice were crossed with P14 mice to generate a traceable population of virus-specific CD8+ T cells to monitor in LCMV-infected adoptive transfer recipients. Following LCMV infection, mTFP1+ and mOrange+ P14 cells expanded comparably in B6 recipients and were maintained stably in circulation following the expected contraction phase (Fig. 2C, D). These data indicate that mTFP1 and mOrange were not recognized as being foreign in a B6 host. Because mOrange is a monomeric derivative of tetrameric DsRed, we postulated that the reduction in antigenic complexity contributed in part to the immunological ignorance of mOrange following adoptive transfer. Consistent with this theory, evaluation of a second monomeric DsRed derivative (i.e., mRFP1) revealed no evidence of rejection in B6 mice following LCMV infection (Fig. 2C, D).
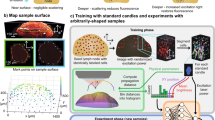
Having generated two new transferrable FP reporter lines, we lastly evaluated the ease with which the FPs expressed in these lines could be detected and spectrally separated following two-photon illumination in a living tissue. This was accomplished by conducting in vivo multi-parameter TPLSM experiments with the two-photon laser tuned to 920 nm. CAG-mOrange and CAG-mTFP1 mice were crossed with P14 mice to generate populations of traceable LCMV-specific CD8+ T cells. Naïve CD11c-YFP mice that express YFP in dendritic cells22 were seeded intravenously with naïve mTFP1 and mOrange P14 cells (5,000 of each population) and infected one day later with LCMV. For comparison, we seeded another group of CD11c-YFP mice with CFP and GFP P14 cells. CFP, GFP and YFP are used for multi-parameter two-photon imaging experiments, but the color combination is spectrally sub-optimal. Separation of the emission spectra using dichroic mirrors at 458, 495 and 525 nm revealed a considerable degree of overlap between the FPs (Fig. 3A, upper panel). This overlap becomes even more apparent in an unprocessed four-color TPLSM data set captured in the intact lymph node of a LCMV-infected CD11c-YFP mouse seeded with CFP and GFP P14 cells (Fig. 3A, middle panel; Movie 1). Images corresponding to the four channels of the NDD4 detector array show the original fluorescence emissions for second harmonic (channel 1), CFP (channel 2), GFP (channel 3) and YFP (channel 4). Note the considerable degree of fluorescence overlap between the channels, particularly in the merged image. Application of a spectral unmixing algorithm to the data set resulted in better separation between the channels, but some overlap remained (Fig. 3A, lower panel; Movie 1). This is evidenced by the failure to generate four pure channels (white, blue, green, red) in the resultant merged image (Fig. 3A, lower panel).
Optimization of spectral reassignment and cellular tracking for in vivo two-photon imaging.
(A, B) Graphs show the fluorescent protein emission spectra for the denoted FPs as well as second harmonic signal corresponding to collagen. The gray bars depict the dichroic mirrors used in the NDD4 external detector to separate the light into four distinct channels. The wavelengths of the dichroic mirrors are denoted above the gray bars. Naïve CD11c-YFP mice were seeded with 5,000 CFP+ P14 cells/GFP+ P14 cells (panel A) or 5,000 mTFP1+ P14 cells/mOrange+ P14 cells (panel B) and infected one day later with LCMV Arm i.p. 3D time lapses were acquired in the lymph node at day 7 post-infection. The upper panels show representative unmanipulated maximal projections generated at a single time point that depict the lymph node capsule defined by second harmonic signal (white, channel 1) as well as a subcapsular region containing T cells and antigen presenting cells. Images depict the light collected by the four detectors (channels 1 through 4) in the NDD4 external detector box. The “merge” image represents the combination of the four channels (Ch). The lower panels show the same maximal projections following spectral unmixing using Leica Application Suite AF software. The boxes beneath the images provide the wavelengths of light captured by each channel of the NDD4 detector array. See corresponding Movies 1 and 2. Images shown in this figure are representative of two independent experiments.
By comparison, the emission spectra for mTFP1, YFP and mOrange showed a much better degree of separation than the CFP/GFP/YFP color combination (Fig. 3B, upper panel) and using optimally placed dichroic mirrors at 458, 510 and 561 nm, the resultant original TPLSM data set revealed a lower degree of overlap between the channels. For example, pure blue and red coloration was observed in the original merged image (Fig. 3B, middle panel; Movie 2). Spectral unmixing further improved the quality of this data set and generated perfect separation of the data into four distinct channels (white, blue, green, red) (Fig. 3C, lower panel; Movie 2). These data demonstrate that mTFP1, YFP and mOrange are a well suited color combination for multi-parameter two-photon imaging experiments.
In conclusion, FPs have become an essential component of many in vivo two-photon imaging experiments and FP-tagged cells are often transferred into recipients despite the potential for immunological rejection of the foreign FP. In this study, we set up an adoptive transfer paradigm to evaluate the rejectability of several commonly used FPs. Surprisingly, we observed that only 1 of the 6 FPs tested (DsRed) was rejected upon adoptive transfer into a B6 recipient, consistent with a previous study showing that a cytotoxic lymphocyte response can be generated against this protein16. We further demonstrated that a humoral response is also mounted against DsRed and that rejection of this FP can be prevented upon establishment immunological tolerance. Interestingly, the dominant DsRedT3 epitope (SSLQDGCFI) previously reported to be recognized by CD8+ T cells in B6 recipients is mutated by one amino acid in mOrange and mRFP1 (SSLQDGEFI)16. It is conceivable that the substitution of a polar hydrophilic amino acid like cysteine (C) in DsRedT3 to a negatively charged glutamic acid (E) in mOrange/mRFP1 could impair CD8 T cell receptor recognition. Alternatively, we postulate that rejection of DsRed in non-tolerant mice is linked to its tetrameric structure and/or ability to aggregate23,24,25, as monomeric derivatives of DsRed were not viewed as foreign upon adoptive transfer. Tetrameric or aggregate DsRed may promote humoral immunity by eliciting multi-valent stimulation of B cell receptors following release from dead or dying cells26. DsRed aggregation may also facilitate uptake by antigen presenting cells and presentation to T cells.
To obtain a spectrally compatible color palette for multi-parameter two-photon imaging experiments, we generated two new transgenic FP reporter mice: CAG-mTFP1 and CAG-mOrange. mTFP1 is very photostable and has one of the highest quantum yields of any FP21. mOrange is also photostable13,27 and has the added advantage of being photoconvertible to a far-red protein28. Both proteins were immunologically tolerated by the B6 immune system and performed optimally in a TPLSM experiment when imaged with YFP. This contrasted with the CFP/GFP/YFP combination which retained some spectral overlap even after unmixing. Another advantage of using mTFP1/YFP/mOrange is that all three FPs (plus collagen) were excited with a single wavelength of two-photon light (920 nm), which has advantages over other FP combinations that require dual lasers set to different wavelengths29,30. Collectively, our studies should help advance future TPLSM experimentation by a providing a compatible, immunologically-tolerated FP combination for adoptive transfer studies.
Methods
Mice
C57BL/6J (B6/J), B6.129(ICR)-Tg(CAG-ECFP)CK6Nagy/J (actin-CFP), C57BL/6-Tg(CAG-EGFP)1Osb/J (actin-GFP), B6.Cg-Tg(CAG-DsRed*MST)1Nagy/J (actin-DsRed), B6.Cg-Tg(CAG-mRFP1)1F1Hadj/J (actin-mRFP1), B6 CD11c-YFP mice, B6 Thy1.1+DbGP33–41 TCR-tg (Thy1.1 P14) and B6 DbGP33–41 TCR-tg (P14) mice17 were bred and maintained under specific pathogen free conditions at the National Institute of Health (NIH). B6, actin-CFP, actin-GFP, actin-DsRedT3 and actin-mRFP1 were originally purchased from The Jackson Laboratory. CD11c-YFP mice22, generously provided by M. Nussenzweig (Rockefeller University, New York, NY), were backcrossed for three additional generations to obtain mice on a pure B6/J background. Fluorescent protein-expressing P14 cells were generated by setting up F1 crosses between P14 (or Thy1.1 P14) mice and each of the aforementioned actin transgenic reporter mice.
Transgenic mouse generation
All transgenic mice were generated by the National Institute of Mental Health (NIMH) Transgenic Core Facility. To generate mice expressing DsRed Express2 under the neuron specific enolase promoter (NSE-DsRedE2) a cDNA construct was made by replacing the cytomegalovirus (CMV) promoter in the pCMV DsRed-Express2 vector (Clontech) with the murine NSE promoter. The NSE promoter sequence (2060 bp) was PCR amplified using primers (fwd: 5′ATATATATTAATGTTGTTAAACCTTCGATTCCG3′; rev: 5′TATATGCTAGCTCGAGGACTGCAGACTCAG3′) containing the AseI and NheI restriction enzyme sites. This promoter was cloned into the pCMV-DsRedE2 vector following removal of the CMV promoter. A rabbit beta-globin polyadenylation signal sequence (462 bp) was PCR amplified from the pCAG-GFP vector (Addgene) and inserted downstream of DsRed-Express2 cassette at NotI restriction enzyme site. Transgenic mice expressing monomeric teal fluorescent protein (mTFP1) under the chicken β-actin promoter were generated by first PCR amplifying the entire 708 bp mTFP1 coding region using primers (fwd: 5′ATATATGAATTCGCCACCATGGTGAGCAAGGGCGAGG3′; rev: 5′ATATATCTC GAGTTACTTGTACAGCTCGTCCATG3′) containing EcoRI and XhoI restriction sites. This cDNA was cloned into the same sites following removal of the GFP sequence from the pCAG-GFP vector (Addgene). The resultant plasmid was digested with Sal I/HindIII and a 3102 bp fragment containing the CMV early enhancer/chicken β-actin (CAG) promoter, mTFP1 and PolyA sequence was prepared for microinjection into the pronuclei of fertilized mouse eggs. Mice expressing monomeric orange fluorescent protein (mOrange) under the CAG promoter were generated in a similar manner. The entire 711 bp coding sequence for mOrange was PCR amplified from the pmOrange Vector (Clontech) using the aforementioned primers and cloned into pCAG-GFP following removal of GFP with EcoRI and XhoI. A 2900 bp fragment consisting of the CAG promoter, mOrange and a PolyA sequence was cut with Sal I/HindIII and prepared for microinjection. To generate all transgenic mice, linearized constructs were injected into C57BL/6N eggs. Following selection of transgene positive founder lines, all mice were backcrossed onto the C57BL/6J background for 10 generations. Mice were confirmed to be on a pure C57BL/6J background using single-nucleotide polymorphism (SNP) analysis (Charles River).
Virus
LCMV Armstrong (Arm) clone 53b was propagated on BHK-21 cells and viral titers were established by performing plaque assays on Vero cells. Mice were infected intraperitoneally with 2 × 105 PFU of virus.
Mononuclear cell isolation
Splenocytes were mechanically disrupted through a 100-μm nylon mesh, treated for 5 min with red blood cell lysis buffer (0.14 M NH4Cl and 0.017 M Tris-HCl, pH 7.2), washed twice and resuspended in PBS containing 1% FBS. Peripheral blood mononuclear cells (PBMCs) were obtained by mixing 200 μl of blood with 10 ml of red blood cell lysis buffer on ice for 1 h. Afterward, PBMCs were centrifuged, washed and resuspended in PBS containing 1% FBS.
CD8+ T cell isolation and adoptive transfer
CD8+ T cells were purified from spleens of naïve P14 mice by negative selection using an EasySep Mouse CD8+ T cell enrichment kit (STEMCELL Technology) per the manufacturer's instructions. The purity of each preparation was flow cytometrically determined to be greater than 98%. One day prior to LCMV infection, naïve C57BL/6J mice were seeded intravenously with 5,000 fluorescent protein (CFP, mTFP1, GFP, mOrange, DsRed, or mRFP1) tagged CD8+ P14 T cells.
Flow cytometry
PBMCs were blocked with 3.3 μg/ml anti-mouse CD16/CD32 (Fc Block; BD) in PBS containing 1% FBS and 0.1% sodium azide for 10 min on ice. Cells were then surface stained with cocktails of the following antibodies: Thy1.1 FITC (53-6.7; BD Biosciences), CD8 Pacific Blue (5H10; Life Technologies), CD8a APC (53-6.7; BD Biosciences) and CD45.2 Alexa700 (104; Biolegend) for 20 min on ice. All cells were acquired using a digital flow cytometer (Digital LSR II; BD) and data were analyzed with FlowJo software (Tree Star).
Detection of anti-DsRed antibodies
Serum anti-DsRed antibodies were detected using the Amplex ELISA Development Kit for Mouse IgG with Amplex UltraRed reagent (Life Technologies). Corning 96-well ELISA plates were coated with 10 μg/ml recombinant DsRed (Clontech) and incubated overnight at 4°C. The next day plates were washed 3 times with PBS containing 0.1% Tween-20 and blocked overnight at 4°C with PBS containing 1% bovine serum albumin. All serum samples were then diluted serially (3-fold dilutions) from 1:10 to 1:1,771,470 and incubated for 1 h at room temperature (RT) on the DsRed coated plates. An anti-DsRed antibody (Clontech) was used as a positive control for this assay and an anti-GFP antibody (Clontech) was used as a negative control. Afterward, plates were washed 3× with PBS/Tween-20 and incubated for 30 min at RT with 50 ng/ml of goat anti-mouse IgG horseradish peroxidase, washed 3× with PBS/Tween-20 and incubated for 30 minutes with Amplex Red at RT in a dark chamber. Fluorescence was quantified using a Varioskan Flash fluorometer (530 nm excitation; 590 nm emission) (ThermoScientific).
Real-time PCR
All tissues were homogenized and subsequently treated with TRIzol before performing a column-based RNA purification using the PureLink RNA mini kit (Life Technologies). The extracted total RNA was then treated with amplification grade DNAse I (Life Technologies) to remove any contaminating genomic DNA. 250 ng of total RNA was used for cDNA synthesis using iScript™ cDNA Synthesis Kit (Bio-Rad). To determine the absolute copy number of DsRed mRNA in various tissue samples, 12.5 ng cDNA was used for quantitative real-time polymerase chain reactions (Q-PCR) with DsRed specific primers (fwd: 5′GGCACCTTCATCTACCACG3′ and rev: 5′ CTTGTGGATCTCGCCCTTC3′). Known quantities of linearized pCMV-DsRed plasmid (Clontech) served as a reference for standard curve generation. Q-PCRs were performed using SYBR Green PCR master mix (Applied Biosystems) and a CFX96 Real-Time PCR machine (Bio-Rad). Q-PCR reactions were run with an initial denaturation temperature of 95°C for 10 min, which was followed by 40 cycles of a three-step amplification (95°C for 30 s, 60°C for 20 s and 72°C for 20 s). All reactions were performed in duplicate and RNA samples without reverse transcriptase served as a negative control.
Two-photon microscopy
Two-photon imaging was performed as described previously31. Briefly, lymph nodes were glued to a plastic cover slip. The coverslip was then placed into a flow chamber perfused with 37°C high-glucose DMEM bubbled with a mixture of 95% O2 and 5% CO2. 3D time lapses were captured using a Leica SP5 two-photon microscope equipped with an 8,000-Hz resonant scanner, a 20× water-dipping objective (1.0 NA), a quad NDD4 external detector array and a Mai Tai HP DeepSee Laser (Spectra-Physics) tuned to 920 nm. Stacks consisting of 15 to 30 planes (2 μm step size) were acquired at 30 sec intervals beginning 50 μm below the cut surface. Signal contrast was enhanced by averaging 12 video frames per plane in resonance scanning mode. Fluorescence emission was passed through customized dichroic mirrors (Semrock) in the NDD4 external detector. Collagen structure in the lymph node was visualized by collecting second harmonic signal corresponding to emission wavelengths less than 458 nm. For simultaneous detection of second harmonic, CFP, GFP and YFP, the following dichroic mirrors were used: 458nm-LP, 495nm-LP, 525nm-LP. For detection of second harmonic, mTFP1, YFP and mOrange the 495 nm mirror was replaced with 510nm-LP and 525 nm with 562nm-LP (see Fig. 3A,B). Following acquisition, all images were spectrally unmixed using the Leica Application Suite AF software before final processing was performed in Imaris 7.3 Software (Bitplane).
Graphs
All graphing and curve fitting was performed using GraphPad Prism 6.0 (GraphPad Software, Inc.). The fitted curves shown in Figures 1B, 1D and 2D were generated using the equation for a one phase decay: Y = (Y0 − Plateau)*exp(−K*X) + Plateau. The curves in Figure 1E were generated using the equation for a sigmoidal dose-responses: y = Bottom + (Top − Bottom)/(1 + 10((LogEC50−X)*HillSlope)).
Animal ethics
All experiments involving mice were approved by the National Institutes of Health Animal Care and Use Committee and conducted in accordance with the guidelines set forth by the Association for Assessment and Accreditation of Laboratory Animal Care (AAALAC).
References
Denk, W., Strickler, J. H. & Webb, W. W. Two-photon laser scanning fluorescence microscopy. Science 248, 73–76 (1990).
Zinselmeyer, B. H. et al. Chapter 16. Two-photon microscopy and multidimensional analysis of cell dynamics. Methods Enzymol 461, 349–378 (2009).
Herz, J., Zinselmeyer, B. H. & McGavern, D. B. Two-photon imaging of microbial immunity in living tissues. Microsc Microanal 18, 730–741 (2012).
Moulton, P. F. Spectroscopic and laser characteristics of Ti:Al2O3 . J Opt Soc Am B 3, 125–133 (1986).
Goeppert-Mayer, M. Über Elementarakte mit zwei Quantensprüngen. Ann Phys 9, 273–295 (1931).
Kaiser, W. & Garrett, C. G. B. Two-photon excitation in CaF2:Eu2+. Phys Rev Lett 9, 453 (1961).
Kobat, D., Horton, N. G. & Xu, C. In vivo two-photon microscopy to 1.6-mm depth in mouse cortex. J Biomed Opt 16, 106014 (2011).
Shimomura, O., Johnson, F. H. & Saiga, Y. Extraction, purification and properties of aequorin, a bioluminescent protein from the luminous hydromedusan, Aequorea. J Cell Comp Physiol 59, 223–239 (1962).
Prasher, D. C., Eckenrode, V. K., Ward, W. W., Prendergast, F. G. & Cormier, M. J. Primary structure of the Aequorea victoria green-fluorescent protein. Gene 111, 229–233 (1992).
Heim, R., Cubitt, A. B. & Tsien, R. Y. Improved green fluorescence. Nature 373, 663–664 (1995).
Shaner, N. C., Steinbach, P. A. & Tsien, R. Y. A guide to choosing fluorescent proteins. Nat Methods 2, 905–909 (2005).
Matz, M. V. et al. Fluorescent proteins from nonbioluminescent Anthozoa species. Nat Biotechnol 17, 969–973 (1999).
Shaner, N. C. et al. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat Biotechnol 22, 1567–1572 (2004).
Drobizhev, M., Makarov, N. S., Tillo, S. E., Hughes, T. E. & Rebane, A. Two-photon absorption properties of fluorescent proteins. Nat Methods 8, 393–399 (2011).
Hadjantonakis, A. K., Macmaster, S. & Nagy, A. Embryonic stem cells and mice expressing different GFP variants for multiple non-invasive reporter usage within a single animal. BMC Biotechnol 2, 11 (2002).
Davey, G. M. et al. Identification of a MHC I-restricted epitope of DsRed in C57BL/6 mice. Mol Immunol 53, 450–452 (2013).
Pircher, H., Burki, K., Lang, R., Hengartner, H. & Zinkernagel, R. M. Tolerance induction in double specific T-cell receptor transgenic mice varies with antigen. Nature 342, 559–561 (1989).
Ahmed, R., Salmi, A., Butler, L. D., Chiller, J. M. & Oldstone, M. B. Selection of genetic variants of lymphocytic choriomeningitis virus in spleens of persistently infected mice. Role in suppression of cytotoxic T lymphocyte response and viral persistence. J Exp Med 160, 521–540 (1984).
Murali-Krishna, K. et al. Counting antigen-specific CD8 T cells: a reevaluation of bystander activation during viral infection. Immunity 8, 177–187 (1998).
Kwon, C. H. et al. Neuron-specific enolase-cre mouse line with cre activity in specific neuronal populations. Genesis 44, 130–135 (2006).
Ai, H. W., Henderson, J. N., Remington, S. J. & Campbell, R. E. Directed evolution of a monomeric, bright and photostable version of Clavularia cyan fluorescent protein: structural characterization and applications in fluorescence imaging. Biochem J 400, 531–540 (2006).
Lindquist, R. L. et al. Visualizing dendritic cell networks in vivo. Nature Immunol 5, 1243–1250 (2004).
Yarbrough, D., Wachter, R. M., Kallio, K., Matz, M. V. & Remington, S. J. Refined crystal structure of DsRed, a red fluorescent protein from coral, at 2.0-A resolution. Proc Natl Acad Sci U S A 98, 462–467 (2001).
Jakobs, S., Subramaniam, V., Schonle, A., Jovin, T. M. & Hell, S. W. EFGP and DsRed expressing cultures of Escherichia coli imaged by confocal, two-photon and fluorescence lifetime microscopy. FEBS Lett 479, 131–135 (2000).
Yanushevich, Y. G. et al. A strategy for the generation of non-aggregating mutants of Anthozoa fluorescent proteins. FEBS Lett 511, 11–14 (2002).
Rosenberg, A. S. Effects of protein aggregates: an immunologic perspective. AAPS J 8, E501–507 (2006).
Shaner, N. C. et al. Improving the photostability of bright monomeric orange and red fluorescent proteins. Nat Methods 5, 545–551 (2008).
Kremers, G. J., Hazelwood, K. L., Murphy, C. S., Davidson, M. W. & Piston, D. W. Photoconversion in orange and red fluorescent proteins. Nat Methods 6, 355–358 (2009).
Kawano, H., Kogure, T., Abe, Y., Mizuno, H. & Miyawaki, A. Two-photon dual-color imaging using fluorescent proteins. Nat Methods 5, 373–374 (2008).
Mahou, P. et al. Multicolor two-photon tissue imaging by wavelength mixing. Nat Methods 9, 815–818 (2012).
Zinselmeyer, B. H. et al. PD-1 promotes immune exhaustion by inducing antiviral T cell motility paralysis. J Exp Med 210, 757–774 (2013).
Acknowledgements
This work was supported by National Institutes of Health (NIH) intramural program. D. Nayak was supported by a NIH Intramural Competitive Fellowship. We would like to thank Dr. James Pickel and the NIMH Transgenic Core Facility for their assistance in generating the new transgenic fluorescent protein reporter mice described in this manuscript.
Author information
Authors and Affiliations
Contributions
S.G. and D.B.M. designed and completed the experiments. D.N. generated the transgenic mice. D.N. and D.B.M. wrote the manuscript. B.H.Z. suggested using OFP and mTFP1 for two-photon imaging, helped with data analysis and prepared the figures/movies. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Movie Legends
Supplementary Information
Movie 1
Supplementary Information
Movie 2
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Gossa, S., Nayak, D., Zinselmeyer, B. et al. Development of an Immunologically Tolerated Combination of Fluorescent Proteins for In vivo Two-photon Imaging. Sci Rep 4, 6664 (2014). https://doi.org/10.1038/srep06664
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06664
This article is cited by
-
Infection drives meningeal engraftment by inflammatory monocytes that impairs CNS immunity
Nature Immunology (2019)
-
Neural precursor cells form integrated brain-like tissue when implanted into rat cerebrospinal fluid
Communications Biology (2018)
-
T-bet-dependent NKp46+ innate lymphoid cells regulate the onset of TH17-induced neuroinflammation
Nature Immunology (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.