Abstract
The research in microbial communities would potentially impact a vast number of applications in “bio”-related disciplines. Large-scale analyses became a clear trend in microbial community studies, thus it is increasingly important to perform efficient and in-depth data mining for insightful biological principles from large number of samples. However, as microbial communities are from different sources and of different structures, comparison and data-mining from large number of samples become quite difficult. In this work, we have proposed a data model to represent large-scale comparison of microbial community samples, namely the “Multi-Dimensional View” data model (the MDV model) that should at least include 3 aspects: samples profile (S), taxa profile (T) and meta-data profile (V). We have also proposed a method for rapid data analysis based on the MDV model and applied it on the case studies with samples from various environmental conditions. Results have shown that though sampling environments usually define key variables, the analysis could detect bio-makers and even subtle variables based on large number of samples, which might be used to discover novel principles that drive the development of communities. The efficiency and effectiveness of data analysis method based on the MDV model have been validated by the results.
Similar content being viewed by others
Introduction
Microbes are ubiquitous on our planet and it is well-known that the total number of microbial cells on Earth is huge1,2. These organisms usually live in communities and each of these communities has a different taxonomical structure. As such, microbial communities would serve as the largest reservoir of genes and genetic functions for a vast number of applications in “bio”-related disciplines, including biomedicine, bioenergy, bioremediation and biodefense3. Since over 90% of strains in a microbial community could not be isolated or cultivated4, metagenomic methods have been used to analyze a microbial community as a whole. Such an approach has enabled exploring relationships among microbes, their communities and habitats at the most fundamental genomic level. Furthermore, environments have profoundly and delicately shaped the microbial community structures, thus making microbial communities from different conditions or time-points different, as well as making it possible for communities from similar types of environment to be significantly different5.
With the advancement of microbial community analysis, it is now possible to conduct sample collection, DNA extraction and taxonomical structure analysis by an efficient pipeline6,7 for large number of samples. These efforts, together with the advanced methods for rapid sample comparison8,9 have enabled the monitoring of microbial communities in time-course and under different conditions. For example, microbial community analyses have been conducted for monitoring of human microbial communities5,10,11,12, environmental samples of ocean microbial communities13 and soil microbial communities14.
As large-scale metagenomic analyses become a clear trend in microbial community analysis, data-mining methods should keep pace. Based on large volume of microbial community samples, it is becoming more and more important to perform in-depth data-mining for valuable biological information on a large scale. Currently many tools such as Mothur15, QIIME16 and MEGAN17 provide metagenomic analysis methods for microbial communities, which mostly focus on samples alone and ignore the connections to the environmental factors. And some of these tools also face difficuties in throughput and data-volume when handreds of samples are to be compared and integrated for mining. The basic data-mining requirements are to unveil the correlations between communities and key factors (taxa, environmental factors, etc.), as well as the effect of these factors on the changes of these communities. For advanced data-mining method development, we believe they should have at least two properties: firstly the method should be capable of handling large-scale datasets and secondly the analysis results should be profound enough to show the underlining relationships among microbial community structures, their environments and the ever-changing organisms within samples.
Though microbial community data are from different sources and of different structures, a large-scale comparison of them could be presented based on a uniformed data model, namely the “Multi-Dimensional View” (MDV) data model that should at least include 3 aspects (Figure 1, for details refer to “Methods” section): samples profile (S), taxa profile (T) and meta-data (environmental conditions including sampling time, condition, etc.) profile (V). In other words, MDV = {S, T, V}. Among these, “meta-data” profile includes all environmental and temporal variables for microbial communities, such as host/habitat for human microbiota, temperature, pH value, etc. This 3-aspect view (Figure 1 (A)) is a simplified model that could include more views such as different batch of experiments and so on to become the extended MDV model (Figure 1 (B)).
The data model for comparison of a number of microbial community samples.
(A) The 3-aspect view for the comparison data model. (B) Meta-data could be extended to include multiple environmental and temporal variables including habitat, pH value, etc. Among these meta-data variables, some are highly related to human habitat samples, while others are highly related to environmental samples.
Based on this MDV model, the digging of biological relationships from communities could be summarized as the data-mining from the MDV = {S, T, V} space and the above-mentioned two key aspects for data-mining method development become very natural and clear: the deep data-mining would essentially echo the effective clustering of those basic elements in the MDV model and efficiency requirements echo the needs for fast process of such clustering. Thus the effective and efficient clustering of basic elements in the MDV model would be the core for the success of large-scale microbial community data-mining.
In this work, we focused on inferring the correlation between the taxa profile (T) and meta-data (V) by data-mining method in the MDV model, i.e., comparison of samples with different meta-data. We have proposed a method for the rapid data comparison and correlation analysis among microbial community samples based on the MDV model, which is supported by High-Performance Computation for rapid process. This method has also been applied on 3 sets of samples from different conditions including human-associated habitats, soil and marine water, each of which has a large number of samples. These datasets are of different complexity and comes with different meta-data, therefore they are suitable for assessment of data model and data analysis methods. The comparison and correlation analysis results based on these datasets have showed excellent performance of our method for in-depth data-mining from massive number of microbial community samples.
Results
Microbial community samples
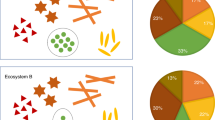
We have evaluated the efficiency of sample comparison and correlation analysis method in MDV spaces based on 3 microbial community datasets. The 3 sets of microbial community samples were gathered from different environments, each having a large number of samples (Figure 2). Dataset A contains 258 human-associated microbial community samples from 3 different habitats of 6 individuals, which were produced by Caporaso, et al., PNAS 201118 and Caporaso, et al., Genome Biology, 201119 (refer to Table S1 in supporting information File S1 for details); Dataset B contains 40 microbial samples from marine surface water sampled at 3 different time-points, which were produced by Caporaso, et al., PNAS 201118 (refer to Table S2 in supporting information File S1 for details); Dataset C contains 42 soil microbial community samples of 3 different locations, produced by the same work as Dataset B (refer to Table S3 in supporting information File S1 for details). These 3 datasets thus represented broad-based microbial communities that also have important biological applications. All of these microbial community samples' sequencing data were produced by Illumina GAIIx from 16S rRNA genes.
The 3 microbial community datasets used in this study, represented in 3D views according to the MDV data model.
Each dataset correspond to a MDV model with different {S, T, V} space. The MDV cubes were generated using SVG (Scalable Vector Graphics) and photos were captured by one of the authors (Xiaoquan Su) in-house.
Results on human-associated habitat microbial community samples
The commensal microorganisms living in our gut20,21, skin22,23 and various other places have key roles in our physiology24, including our immune responses and metabolism, as well as in various human diseases25. Since hosts and sampling times would significantly affect the structure of human-associated habitat microbial communities, the combination of large amount of samples together with their meta-data would serve as a good benchmark for testing analysis methods.
In this case study, we have obtained 258 human-associated habitats microbial community samples from 3 different habitats (gut samples from feces, skin samples from palms and oral samples from tongue) of 6 individuals (Table 1). In the MDV model, |S| = 258 and V = {Host, Habitat}. Among the 6 hosts, 2 (Female 5 and Male 6) were from the same family, which were obtained from Caporaso, et al., Genome Biology, 201119, while others were from different families (Female 1, Male 2, Male 3 and Male 4) with samples' sequences produced by different primers, which were obtained from Caporaso, et al., PNAS 201118.
We have first generated pair-wise similarity matrices with all 258 samples based on their taxonomical structure among samples ((S, T) space of the MDV model) from different families (Figure 3 (A)) and the same family (Figure 3 (B)), respectively. Then we used hierarchical-based clustering methods based on similarly matrices to examine the relationship among different human microbiota (for details refer to “Methods” section). Results (Figure 3) have shown that samples from the same habitat were clustered together and samples from skin and oral environment shared more common structures, yet community structures for samples within gut were significantly different. This clustering pattern by habitats indicated that among the various meta-data (V space of the MDV model, including family background (possibly related to diet26), host and habitats), habitat played a more important role in shaping the community structures for these samples. Further probing of the bio-marker taxa in (T, V) space of the MDV model (for details refer to “Methods” section) that caused such pattern has shown that Bacteroidaceae and Clostridiaceae (dominating gut microbial communities), Prevotellaceae and Pasteurellaceae (dominating oral microbial communities) and Corynebacterineae (dominating skin microbial communities) were the most prominent taxa (Table 2) that could distinguish samples from different habitats.
Similarity matrix of Human-associated habitat microbial community samples.
(A) Hosts were from different families. (B) Hosts were from the same family. Each tile represents a similarity value between two samples from a color gradient between red and green: red color indicates higher similarity value and green color indicates lower similarity value, with red/green shades in between indicating intermediate values.
We noticed that among the hosts in different families, most samples from the same host could be clustered together for each habitat (Figure 3 (A)). Only few samples labeled with “Male_4_Gut” were divided into two groups probably due to the reason that sequences produced by different primers were from the different regions of 16S rRNA gene). Additionally, among family members (Female 5 and Male 6), samples of the same habitat could not be distinguished by host (Figure 3 (B)). The most abundant taxa in samples from Female 5 and Male 6 include Bacteroidaceae (P-value = 0.346), Prevotellaceae (P-value = 0.777), Pasteurellaceae (P-value = 0.809) and Streptococcus (P-value = 0.741) which showed high similarity in relative abundances due to the strong effect from small-scale environment of the same family26, thus making the differentiation difficult.
Furthermore, we conducted the PCoA (Principal Coordinates Analysis) analysis based on sample similarity matrix from the same family to examine the correlation of the microbial community patterns to hosts and habitats. It was obvious in the PCoA results (Figure 4) that samples could be differentiated by habitats, but samples from the same habitats but different family members were mixed together because they shared similar community structure patterns.
Results on microbial community samples from marine water
Marine microbial communities play a very important role in the regulation of carbon and nitrogen circulation of the globe27 and they contain important genes for a wide application area such as bioenergy, bioremediation, etc28. However, marine samples are very diverse in their structure as well as function, making knowledge discovery from them quite challenging.
In this work, we applied our method to analyze 40 microbial samples produced by Caporaso, et al., PNAS 201118 from marine surface water of Newport Beach Pier, CA, US collected at different time-points (seasons)18. These samples were collected from 3 different time-points (seasons) at the same location. In the MDV model, |S| = 40 and V = {Time, Temperature}. We used hierarchical-based method to evaluate the relationships among all marine water communities and density-based clustering methods MCODE29 (for details refer to “Methods” section) to examine the major differentiation factors during time-course based on the pair-wise similarity matrix.
Results from Figure 5(A) and Figure 5 (B) indicated that all samples could be divided into three groups by the meta-data of sampling time-point (V space in the MDV model). Since these marine water samples were collected from a similar site (a near-coast site) and water-depth (surface) yet at 3 different time-points (seasons) with different water temperature, the microbial community structures showed high correlation with V = water temperature in the MDV model in Figure 5 (B), which has also been proven in other works30. Detailed analyses on bio-markers in (T, V) space of the MDV model have shown that the relatively abundant and most dynamic taxa for these samples include Flavobacteriaceae (P-value = 0.00095), Prochlorococcus (P-value = 0.00056) and Rhodobacteraceae (P-value = 0.00056) (Figure 5(C)), all of which were sensitive to water temperature as well. Additionally, from Figure 5 (A) we observed that though each cluster of samples had high intra-cluster similarity, samples from time-point 2 were not similar enough with any of the samples from time-point 1 and time-point 3, indicating that meta-data for samples from time-point 2 might be drastically different. Our analyses on the above 3 most dynamic taxa have also shown that compared to samples from time-point 1 and time-point 3, samples from time-point 2 always have different taxa abundances with regard to Flavobacteriaceae, Prochlorococcus and Rhodobacteraceae (Figure 5(C)).
Clustering and bio-marker analysis results of marine samples.
(A) Hierarchical-based clustering results to discover the relationships among samples, in which the more similar the two samples the deeper dark red color. (B) Density-based clustering result to examine the major differentiation factors, in which nodes represent samples and edges between nodes indicated that their similarities were above the threshold of 85%. (C) The relative abundances distribution for all marine water samples for 3 most dynamic taxa in marine samples.
Results on microbial community samples from soil
Soil microbial communities belong to a type representing the most important communities on land for regulation of the carbon and nitrogen circulation on earth31,32 and they were directly related to agriculture researches30. Soil microbial communities also represented the most complex, diverse and dynamic communities on earth33.
We have used 42 soil microbial community samples of 3 different places each with different pH values from the work of Caporaso, et al., PNAS 201118 to demonstrate the performance of our method. For the soil samples, both 3′ reads and 5′ reads which were the sequencing results of 16S rRNA genes in two complementary directions by different primers were generated and analyzed together (Table 3). In the MDV model, |S| = 42 and V = {Type, Location, pH, Primer}. We then processed the samples with hierarchical-based clustering method (for details refer to “Methods” section) based on their similarity matrix to discover the corresponding environmental patterns.
From the results (Figure 6) we observed that all samples could be divided into 3 groups, mainly by the pH values of the sampling environments. We also noticed that in each group, samples sequenced by 3′ primer and 5′ primer could be distinguished from the clustering results due to the technical specification of sequencing that sequences produced by 3′ primer and 5′ primer were from different regions of 16S rRNA genes. We also verified our results using the Fast UniFrac34 algorithm and obtained similar results (refer to Figure S1 in Supporting information File S1 for details).
We further investigated the correlation between the community structures of soil samples and their environment factors by PCoA (Principal Coordinates Analysis) in (T, V) space of the MDV model. Results in Figure 7 (A) elucidated the high correlation of the community structure to the pH values: both 3′ reads samples and 5′ reads samples were ordered from alkalinity soil to acid soil (from pH 8.3 to pH 4.9) and sample from the acid and semiacid environment were more similar (samples from pH 4.9 soil and pH 6.1 soil), which has been proved by Fierer et al., PNAS 200635.
Then we performed the bio-marker analysis to discover the abundant key taxa that strongly correlated with Vi = pH value. As soil microbial communities were much more complex with a huge number (>1,000) of species in each sample, a taxon with more than 5% relative proportion in the community was already very abundant. The abundance variation of taxa Sphingomonadaceae (Pearson correlation coefficient R = 0.9537, abundances 0.6%–15%), Rubrobacterineae (R = 0.9696, abundances 0.9%–5.5%) and Micromonosporineae (R = 0.9296, abundances 0.5%–5.3%) had strong positive correlation with pH values, as well as Burkholderiaceae (R = −0.9832, abundances 0.3%–3.4%) were highly negative correlated to pH values, which would be the reason behind the strong correlation of community structure with pH values (Figure 7 (B)). In addition, there was no significant correlation (|R| < 0.7) for pH values and other abundant taxa. This further confirmed that the pH values might affect soil microbial communities significantly through the changes of these abundant taxa35.
Efficiency analysis
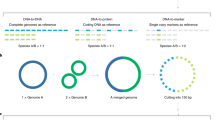
We have also evaluated the running time of data-mining analysis including similarity matrix construction, clustering and correlation analysis, based on the 3 sets of microbial communities. Benefited by the GPU based High Performance Computing (HPC)9 in the most time-consuming process of similarity matrix construction (Figure 8, pie charts), the overall computing speed of GPU achieved more than 60 times speed-up compared to computing speed of CPU, with 16 cores (Figure 8, bar charts). This HPC strategy has made possible data-mining on 258 samples (dataset A) to be completed within only 2 minutes, out of which nearly 30% of time was spent on clustering and correlation analyses.
Discussion and Conclusion
As large amount of metagenomic data could be accumulated quickly from various microbial community profiling projects using NGS, it is becoming more and more important to perform in-depth analysis of microbial communities, as well as data-mining for valuable yet hidden biological principles that controls the dynamic changes of microbial community samples. The basic questions based on such a large amount of samples would be the comparison and correlation analysis which include the understanding of relationships among communities, key factors (taxa, environmental factor, etc.) for such relationships, as well as the effect of environmental and/or temporal factors on community dynamics.
One apparent yet critical problem for data-mining from large number of microbial communities is the heterogeneity of samples (different sources, different meta-data, different structure, etc.). In this work, we have proposed a data model to represent large-scale comparison of these samples, namely the “multi-dimensional view” data model (MDV = {S, T, V}) that consisted of 3 basic aspects: sample profile, taxa profile and meta-data profile. The effective and efficient analysis among different elements in the MDV model is the core for the success of large-scale microbial community data-mining. We have also proposed a method for the rapid data comparison and correlation analysis among microbial community samples based on the MDV model, which is supported by High-Performance Computation for rapid process. The comparison and correlation analysis results based on datasets from various sampling conditions showed excellent performance for in-depth data-mining from massive number of microbial community samples.
The MDV model is not only restricted by sample clustering, but could also be used for taxa clustering as well. Based on taxa clustering (in T space), important biomarkers for distinguishing samples could be discovered36,37. Clustering from another angle of meta-data (in V space) would also help to distinguish important environmental or temporal factors that would affect the dynamics of microbial community samples. These future works based on the MDV model would serve well for more data-mining and in-depth understanding of the underlining principle controlling the functions and evolution of various microbial communities, which would also have great potential in applications.
Methods
The MDV data model
The “Multi-Dimensional View” (MDV) data model includes 3 aspects (Figure 1): sample profile (S), taxa profile (T) and meta-data profile (V), which could be integrated by formula 1:

In this 3-dimensional view (3D view), sample profiles S = (s1, s2, ..sn) contains the ID and basic information about the samples; taxa profiles T = (t1, t2, …, tm) contains community structure information about the taxa, their relative abundances in different samples and their phylogenetic relationship (represented by fphylogeney in Formula 1); meta-data profiles V = (v1, v2, …, vq) contains the meta-data (sampling time, environment condition, etc.) of all samples. In this work, we focus on analysing the relationships among samples with different meta-data. This is equivalent to inferring the correlation between the taxa profile (T) and meta-data (V) by data-mining in the MDV model, which could also be describe by Formula 2:

The data-mining method
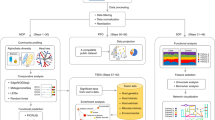
The rapid data-mining procedure includes community structure analysis, similarity-matrix construction, sample clustering and correlation with meta-data based on the MDV model. The overall scheme is illustrated in Figure 9:
Microbial community structure analysis
The community structure profiles of all samples are parsed out from their 16S rRNA gene sequences by high efficient metagenomic analysis tool Parallel-META36 (version 2.0). Parallel-META maps the 16S rRNA sequences of each sample by MegaBLAST37 to the reference database to identify the taxonomical classification and phylogenetic relationship of each species. In this work we use the GreenGenes38 core-set (release date: May 2009) as the reference database and 1E-30 as the expectation value for MegaBLAST based database mapping.
Similarity matrix construction
The similarity matrix reflects the similarity of samples in S = (s1, s2, ……, sn) space based on their taxonomical structure data T = fphylogeny(t1, t2, ……, tm). The similarity score between two microbial community samples evaluates as a quantitative similarity (always a float value between 0% and 100%) calculated by Meta-Storms8,9 algorithm based on the community structure analysis results. The similarity matrix of N samples that consisted by N*N pairs represents pair-wise similarity, in which each pair indicated the similarity score of one sample pair. Based on the permutation test results in our previous work8, a similarity score of 85% or higher indicates significant similarity between 2 samples.
Clustering methods
Clustering methods includes hierarchical-based method and density-based method from MDV = {S, T, V} space. The hierarchical-based clustering elucidates the relationships among the microbial community samples and sample groups, while the density-based clustering focuses on discovering sample groups with significant difference defined by a given threshold. The density-based clustering is also used for validity check for the results of hierarchical-based clustering.
-
a
The hierarchical-based clustering method is implemented by “HClust” function of CRAN R39 and results are visualized by MetaSee software40 and “gplots” package (Gregory R., et al., gplots: Various R programming tools for plotting data. http://CRAN.R-project.org/package=gplots) of CRAN R. In the hierarchical-based clustering, distances among different clusters were evaluated using the “average linkage” (http://stat.ethz.ch/R-manual/R-devel/library/stats/html/hclust.html) method.
-
b
The density-based clustering method is implemented by MCODE29 and results are visualized in Cytoscape software41. Based on permutation tests8, similarity score of 85% or higher indicates the significant similarity between 2 samples. In the density-based clustering analysis we select 85% as the threshold for significant difference.
Correlation and bio-marker selection methods
The correlation analysis attempts to discover relationships between taxa profiles T = fphylogeney(t1, t2, ……, tm) space and V = (v1, v2, ……, vq) space based on the clustering results to deduce the fcorrelation (T, V) in Formula 2. The Principal Coordinates Analysis (PCoA) are used to elucidate the correlation between community structures and meta-data based on the similarity matrix, which is implemented by “vegan” package (ari Oksanen, et al., vegan: Community Ecology Package. http://CRAN.R-project.org/package=vegan) of CRAN R. Then we also select the bio-markers which are considered as abundant taxa that have high correlation with the meta-data and clustering results. For the numerical meta-data (such as pH value, temperature, etc.), we calculate the Pearson correlation coefficient (R) between abundance values of specified taxa and meta-data and select the taxa with R value equal to or larger than 0.9 which indicate the significant correlation between abundance values and meta-data. For the discrete meta-data (such as human-associated habitat, location, etc.), we perform the Wilcoxon and Kruskal rank-sum test and select the taxa with P-value smaller or equal to 0.01, which indicate the significant difference of abundance values among different meta-data.
High-performance computing
The MDV data model has been considered for parallel processing of sample comparison. The similarity among microbial community samples are evaluated by the similarity scores in (T, V) space of the MDV model. The similarity score between each sample pair is calculated by Meta-Storms8 algorithm with time complexity of Nlog(N) (N is the number of species existing in one sample). However, as the amount of samples increases, the overall time complexity of M 2* Nlog(N) (M is the number of samples) based on pair-wise comparison always leads to an unacceptable running time.
In this work, we have performed the calculation of the similarity matrix for massive number of samples using GPU-Meta-Storms9 based on NVIDIA Tesla M2075 GPU hardware (448 stream processors, 6 GB onboard memory). To calculate the similarity matrix of N samples, N * N threads are launched in GPU with many-core architecture to let each similarity score in the matrix be processed by one independent thread in parallel (Figure 10). To fully utilize the GPU-based computation power, we have also designed optimization strategies including global memory alignment, register recalling allocation and shared memory utilization in I/O (Input/Output) operations to improve the overall performance by GPU computing.
References
Whitman, W. B., Coleman, D. C. & Wiebe, W. J. Prokaryotes: The unseen majority. P Natl Acad Sci USA 95, 6578–6583 (1998).
Proctor, G. N. Mathematics of microbial plasmid instability and subsequent differential growth of plasmid-free and plasmid-containing cells, relevant to the analysis of experimental colony number data. Plasmid 32, 101–130 (1994).
National Research Council. The New Science of Metagenomics: Revealing the Secrets of Our Microbial Planet, (The National Academies Press, Washington, DC, 2007).
Jurkowski, A., Reid, A. H. & Labov, J. B. Metagenomics: a call for bringing a new science into the classroom (while it's still new). CBE Life Sci Educ 6, 260–265 (2007).
Muegge, B. D. et al. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332, 970–974 (2011).
Kuczynski, J. et al. Experimental and analytical tools for studying the human microbiome. Nat Rev Genet 13, 47–58 (2012).
Knight, R. et al. Unlocking the potential of metagenomics through replicated experimental design. Nature Biotechnology 30, 513–520 (2012).
Su, X., Xu, J. & Ning, K. Meta-Storms: Efficient Search for Similar Microbial Communities Based on a Novel Indexing Scheme and Similarity Score for Metagenomic Data. Bioinformatics (2012).
Su, X., Wang, X., Jing, G. & Ning, K. GPU-Meta-Storms: Computing the structure similarities among massive amount of microbial community samples using GPU. Bioinformatics (2013).
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009).
Turnbaugh, P. J. & Gordon, J. I. The core gut microbiome, energy balance and obesity. J Physiol 587, 4153–4158 (2009).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Yooseph, S. et al. The Sorcerer II Global Ocean Sampling expedition: expanding the universe of protein families. PLoS Biol 5, e16 (2007).
Fierer, N. et al. Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc Natl Acad Sci U S A 109, 21390–21395 (2012).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75, 7537–7541 (2009).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7, 335–336 (2010).
Huson, D. H., Mitra, S., Ruscheweyh, H. J., Weber, N. & Schuster, S. C. Integrative analysis of environmental sequences using MEGAN4. Genome Res 21, 1552–1560 (2011).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. P Natl Acad Sci USA 108, 4516–4522 (2011).
Caporaso, J. G. et al. Moving pictures of the human microbiome. Genome Biol 12, R50 (2011).
Kau, A. L., Ahern, P. P., Griffin, N. W., Goodman, A. L. & Gordon, J. I. Human nutrition, the gut microbiome and the immune system. Nature 474, 327–336 (2011).
Brown, C. T. et al. Gut microbiome metagenomics analysis suggests a functional model for the development of autoimmunity for type 1 diabetes. PLoS One 6, e25792 (2011).
Capone, K. A., Dowd, S. E., Stamatas, G. N. & Nikolovski, J. Diversity of the human skin microbiome early in life. J Invest Dermatol 131, 2026–2032 (2011).
Kong, H. H. & Segre, J. A. Skin Microbiome: Looking Back to Move Forward. J Invest Dermatol (2011).
Solt, I., Kim, M. J. & Offer, C. The human microbiome. Harefuah 150, 484–488 (2011).
Boerner, B. P. & Sarvetnick, N. E. Type 1 diabetes: role of intestinal microbiome in humans and mice. Ann N Y Acad Sci 1243, 103–118 (2011).
Song, S. J. et al. Cohabiting family members share microbiota with one another and with their dogs. Elife 2, e00458 (2013).
Jacquez, G. M. A k nearest neighbour test for space-time interaction. Stat Med 15, 1935–1949 (1996).
Zhang, T., Ding, J. L. & Wang, F. Simulation of image multi-spectrum using field measured endmember spectrum. Guang Pu Xue Yu Guang Pu Fen Xi 30, 2889–2893 (2010).
Bader, G. D. & Hogue, C. W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics 4, 2 (2003).
Inderjit & van der Putten, W. H. Impacts of soil microbial communities on exotic plant invasions. Trends Ecol Evol 25, 512–519 (2010).
Fierer, N. et al. Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J 6, 1007–1017 (2012).
Charlop-Powers, Z., Owen, J. G., Reddy, B. V., Ternei, M. A. & Brady, S. F. Chemical-biogeographic survey of secondary metabolism in soil. Proc Natl Acad Sci U S A 111, 3757–3762 (2014).
Wagg, C., Bender, S. F., Widmer, F. & van der Heijden, M. G. Soil biodiversity and soil community composition determine ecosystem multifunctionality. Proc Natl Acad Sci U S A (2014).
Hamady, M., Lozupone, C. & Knight, R. Fast UniFrac: facilitating high-throughput phylogenetic analyses of microbial communities including analysis of pyrosequencing and PhyloChip data. ISME J 4, 17–27 (2010).
Fierer, N. & Jackson, R. B. The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci U S A 103, 626–631 (2006).
Su, X., Xu, J. & Ning, K. Parallel-META: efficient metagenomic data analysis based on high-performance computation. BMC Systems Biology 6, S16 (2012).
McGinnis, S. & Madden, T. L. BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Research 32, W20–W25 (2004).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72, 5069–5072 (2006).
Dessau, R. B. & Pipper, C. B. “R”--project for statistical computing. Ugeskr Laeger 170, 328–330 (2008).
Song, B., Su, X., Xu, J. & Ning, K. MetaSee: An Interactive and Extendable Visualization Toolbox for Metagenomic Sample Analysis and Comparison. PLoS One 7, e48998 (2012).
Pedamallu, C. S. & Posfai, J. Open source tool for prediction of genome wide protein-protein interaction network based on ortholog information. Source Code Biol Med 5, 8 (2010).
Acknowledgements
This work is supported by Chinese Academy of Sciences' e-Science grant (grant number INFO-115-D01-Z006), Ministry of Science and Technology's high-tech (863) grant (grant number 2012AA02A707 and 2014AA021502), National Natural Science Foundation of China grant (grant number 61103167, 31271410 and 61303161) and NSFC-DFG's Sino-Germany collaboration grant (grant number GZ878). We thank NVIDIA for providing us with the high-performance computing hardware through QIBEBT-NVIDIA CUDA Research Center.
Author information
Authors and Affiliations
Contributions
K.N. and X.S. conceived of and proposed the idea. K.N. and X.S. designed the study. X.S., J.H. and S.H. developed the algorithm. X.S. performed the data analysis. X.S. and K.N. contributed to editing and proof-reading the manuscript. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
File S1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Su, X., Hu, J., Huang, S. et al. Rapid comparison and correlation analysis among massive number of microbial community samples based on MDV data model. Sci Rep 4, 6393 (2014). https://doi.org/10.1038/srep06393
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06393
This article is cited by
-
Effect of biological activated carbon filter depth and backwashing process on transformation of biofilm community
Frontiers of Environmental Science & Engineering (2019)
-
Less primary fistula failure in hypertensive patients
Journal of Human Hypertension (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.