Abstract
Capture of circulating tumor cells (CTCs) from peripheral blood of cancer patients has major implications for metastatic detection and therapy analyses. Here we demonstrated a microfluidic device for high efficiency and high purity capture of CTCs. The key novelty of this approach lies on the integration of a microfilter with conical-shaped holes and a micro-injector with cross-flow components for size dependent capture of tumor cells without significant retention of non-tumor cells. Under conditions of constant flow rate, tumor cells spiked into phosphate buffered saline could be recovered and then cultured for further analyses. When tumor cells were spiked in blood of healthy donors, they could also be recovered at high efficiency and high clearance efficiency of white blood cells. When the same device was used for clinical validation, CTCs could be detected in blood samples of cancer patients but not in that of healthy donors. Finally, the capture efficiency of tumor cells is cell-type dependent but the hole size of the filter should be more closely correlated to the nuclei size of the tumor cells. Together with the advantage of easy operation, low-cost and high potential of integration, this approach offers unprecedented opportunities for metastatic detection and cancer treatment monitoring.
Similar content being viewed by others
Introduction
Circulating tumor cells (CTCs) in peripheral blood of cancer patients are reliable biomarkers for metastatic detection and treatment monitoring. The challenge is to capture them with a high efficiency and high purity, despite their extreme rarity and high phenotype heterogeneity1,2,3,4,5,6,7,8,9,10. Currently, two approaches are in competition, depending on whether or not the capture is affinity-based. The affinity-based capture relies on immunochemical interaction between magnetic beads11,12,13,14,15,16 or patterned structures17,18,19,20 and tumor cells that express special surface markers such as EpCAM, an epithelial cell adhesion molecule over expressed by majority of tumor cells. These methods are efficient for capturing CTCs of epithelial phenotypes but not applicable to those with down-regulated or lost epithelial markers. Indeed, it is known that tumor cells of epithelial origin may undergo epithelial to mesenchymal transition21,22. In contrast, the non-affinity methods, including centrifuging deflection23,24,25,26,27, dielectrophoretic separation28,29,30 and size-based filtration12,31,32,33,34,35,36,37,38,39,40,41,42,43,44, are able to isolate both epithelial and mesenchymal phenotypes, which are more appropriate for analyses of tumor heterogeneity, tumor drug resistance, etc. However, these approaches are technologically biased and it is often difficult to reach a trade-off between capture efficiency, purity and cell viability which are all important criteria for both fundamental and clinical studies. For example, the size-based filtration has been developed for more than several decades but the most of reported results were obtained by using track-etched polycarbonate filters12,31,32,33,34,35. The main problem of this approach is contamination: the track-etched cylindrical holes of the filter have generally high retention ratio of non-tumor cells if the size of the holes is sufficiently small to reach high tumor cell capture efficiency. Furthermore, the randomly distributed holes are prone to significant coalescence of two or more holes allowing tumor cells to escape from the filter. More recent studies have shown that a variety of microfabrication techniques could be used to manufacture filters of different hole-types and densities36,37,38,39,40,41,42,43,44,45, but none of them were able to meet the critical requirements for both clinical application and fundamental studies.
Ideally, the size-based filtration of CTCs has to be efficient for non-tumor cell clearance with high flow-rate but low transfilter pressure, which implies the use of two-dimensional filters with high density and conical shaped holes. In addition, the filters should be mechanically and thermally stable, optically transparent and biologically compatible for downstream genotype and phenotype analyses. In this work, such filters were produced by micro-aspiration assisted molding of UV curable polyethylene glycol diacrylate (PEGDA). To prevent clogging and reserve a high integration potential, the fabricated filters were inserted into microfluidic devices with cross-flow injectors. We show both experimentally and theoretically the advantages of such a platform for high efficiency and high purity capture of CTCs.
Results
Cancer cell capture
We fabricated conical microhole filters which could be easily integrated into a two-level microfluidic device (Fig. 1). Both filter and microfluidic device were made of biocompatible materials (PEGDA and polydimethylsiloxane or PDMS, respectively) with rather standard fabrication processes (Supplementary Fig. S1). PDMS is now widely used for fast prototyping of microfluidic devices whereas PEGDA can be easily functionalized to tune the protein or cell adhesion properties of the filter surfaces. The sample was loaded into the microfluidic device from the inlet of the top chamber (inlet 1) and collected from the outlet of the bottom chamber (outlet 2), passing through the microfilter sandwiched between the two structured PDMS layers (Supplementary Fig. S2).
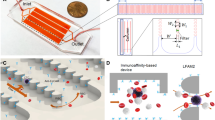
Microfluidic device with integrated conical-hole filter for capture of circulating tumor cells.
(a) Schematic of the experimental setup and CTC filtering. A filter made of PEGDA with conical holes is sandwiched between two patterned PDMS layers and the sample was loaded from inlet 1 and collected from outlet 2, passing through the filter with a syringe pump for constant pulling speed. (b) Photograph of a microfluidic device operating with a test blood sample. (c) Photograph of a 9 mm diameter filter aside a 1 cent euro coin. (d) SEM image of a conical holes array. Scale bar: (d) 40 µm.
We fabricated 30 µm thick PEGDA filter with 30 µm pitch conical holes at constant angle of 13° (Supplementary Fig. S3). For comparison, filters with conical holes of three different entrance diameters (5.5, 6.5 and 8.0 µm) were produced over an area of disk of two different diameters (6 and 9 mm). For demonstration, HT-29 cancer cells were spiked in a diluted blood sample of a healthy person at concentration of 103 cells/ml and the filtration experiments were performed with a filter of 6.5 µm conical holes at a constant flow rate, controlled with a syringe pump at a constant pulling speed (0.2 ml/min). As results, we recovered the most of the spiked HT-29 cells on the filter with a limited number of normal blood cells (Fig. 2) and fluorescence images were recorded after immunostaining (green for cytokeratin or CK positive, red for CD45 positive, blue for nuclei). Since CK is a common tumor marker, by counting the total number of cells injected into the device and the number of CK positive cells on the filter, we obtained a capture efficiency of ~95% (see Supplementary Information for the definition of capture efficiency). More systematically, the capture efficiency could be evaluated with cancer cells spiked in PBS solutions at different concentration (see below), which lead to comparable values. In order to assess the capture purity, we defined clearance efficiency for white blood cells (WBCs) of the filter (see Supplementary Information). Since CD45 is a common marker of WBCs, by counting the number of CD45 positive cells on the filter and the total number of through holes of the filter, we obtained a WBC clearance efficiency of 96%. This has also been confirmed by counting CD45 positive cells on the filer with blood samples of cancer patients and healthy donors (see below).
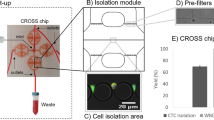
HT-29 cells and WBCs on the surface of a filter.
(a–c) Fluorescence images of HT-29 cells and WBCs stained with anti-CK-AlexaFluor 488 (green) and anti-CD45-PE (red), respectively. Nuclei were stained in blue with DAPI. Here HT-29 cells were spiked in a blood sample and the filtration was done with a filter of 6.5 µm hole diameter. (d) Merged image of (a), (b) and (c). Scale bar: (a–d) 30 µm.
Validation with clinical samples
Blood samples donated by 15 cancer patients with metastasis and 6 healthy donors were used to test the potential clinical relevance of the device. The volume of each blood sample was 2 ml. Without red blood cells lysing, we diluted all samples in PBS at a volume ratio of 1:2 before filtration. The filtration experiments were performed with a filter of 6.5 µm conical holes at a flow rate of 0.2 ml/min. After filtration and immunostaining, fluorescence images of the cells captured on the filters were recorded (green for CK positive, red for CD45 positive, blue for nuclei of all cells). Figure 3 shows one example of the fluorescence images obtained with the blood sample of a lung cancer patient. Again, both CK positive and CD45 positive cells were captured, validating the feasibility of CTC capture by using our device. Here, the most of observed CD45 positive cells are in the hole, which may be slightly deformed. These fluorescence images show a clear correlation between DAPI and CK positive/CD45 negative or CD45 positive/CK negative, which allowed us to attribute the CK positive and CD45 negative cells to CTCs. Statistically, CTCs could be detected in the blood samples of a large portion of cancer patients but not in that of healthy donors (Table 1). Among 15 blood samples of the cancer patients, only 5 did not show up the CTC signature, which may be due to the limited volume of our blood samples. As expected, the identification of captured CTCs in our device was much facilitated because only a few hundreds WBCs were found on the surface of the filter (Fig. S4). In addition to the cancer cells and WBCs, some undefined CK and CD45 negative cells may also show up, but their presence did not affect the detection of CTCs. More systematic analyses will be conducted for a more detailed clinical validation of the device.
Flow rate and cell density dependencies
The flow rate dependency of the cell capture efficiency was studied by using HT-29 cells spiked into PBS at 100 cells/ml concentration. In the range of flow rate under consideration (0.2–2.0 ml/min with filters of 6.5 µm holes diameter and 6 mm diameter filter area), we observed increased capture efficiency when decreasing the flow rate (Fig. 4a). A further increase of the capture efficiency (>95%) should be possible with a smaller flow rate. For the same capture efficiency (95%), the flow rate was increased by a factor of 2.25 when the 6 mm diameter filter was replaced by the 9 mm diameter filter, which corresponds to a decrease of the filtration time from 18 min to 8 min for a 6 ml sample. Similarly, we evaluated the cell density dependency of the capture efficiency (Fig. 4b) by enumerating cell numbers on the 6 mm diameter filter after immunostaining. As expected, the capture efficiency of our device is almost the same for different cell densities due to the fact that the number of captured cell (10–100) is much less than the number of holes of the filter (104 ~ 105) and there should be no change of the transfilter pressure under given experimental conditions. Thus, the capture efficiency we evaluated with 100 cells/ml spiked in blood should be reasonable for the estimation of CTC capture efficiency of our devices.
Evaluation of cell capture performance.
(a) Capture efficiency as a function of flow rate for 100 HT-29 cells spiked in 1 ml PBS. (b) Capture efficiency of 10, 20, 50, 100 HT-29 cells spiked in 1 ml PBS at a flow rate of 0.2 or 0.5 ml/min, respectively. (c) Cell-type dependency of capture with filter of different hole sizes at a flow rate of 0.5 ml/min. (d) Cell and nuclei sizes measured with a suspension of cells stained by Hoechest33342. The error bars represent the standard derivation (s.d.) of three measurements.
Cell type and hole size dependencies
To assess the reliability of our device for different tumor cell types, the capture efficiency has been evaluated for three cancer cell lines (MCF-7, HT-29 and U87), each spiked in PBS at a 100 cells/ml concentration and injected into the device at a flow rate of 0.2 or 0.5 ml/min. Not surprisingly, remarkable differences of the capture efficiency could be observed due to the size difference of these tumor cells (Fig. 4c). Here, three types of filters of different hole diameters (5.5, 6.5, 8.0 μm) were used, which also allowed us to evaluate the hole size dependency of the capture efficiency. Clearly, the smaller the hole size, the higher the capture efficiency. Using the filters of 5.5 µm diameter holes, a capture efficiency of 98%, 96% and 85% could be achieved for MCF-7, HT-29 and U87, respectively. Increasing the hole size from 5.5 µm to 6.5 µm has negligible effect on the capture of MCF-7 but the filter of 8.0 µm diameter holes resulted in a much decreased capture efficiency, suggesting an optimal size (~6.5 µm) of conical holes for MCF-7 capture. The filters of 6.5 µm diameter holes could also be accepted for HT-29 capture but filters of 5.5 µm in diameter or smaller is preferable for U87 capture. To understand the observed cell type dependencies of micro-hole capture, the size of each cell type and their nuclei were determined (Fig. 4d) by analyzing the bright field images of suspended cells and the fluorescence of the cell nuclei stained with 0.5 µg/ml Hoechst 33342 in a suspension of cells in PBS. As expected, MCF-7 has the largest size of cell body and nuclei, giving rise to the best capture efficiency for the given hole size of the filter (6.5 µm). U87 and HT-29 have comparable cell sizes, but U87 has much smaller nuclei, which led to capture efficiency significantly lower than that of HT-29 using the same filter of 6.5 µm hole diameters. Since the nuclei are much more difficult to deform than the cytoplasm, the entrance diameter of conical holes for filtration should be correlated to the size of corresponding nucleus rather than the size of the cell.
Cell viability assessment
HT29 cells spiked in PBS were used to evaluate the cell viability after filtration with 6.5 µm conical-hole filters. About 90% of the tumor cells could be captured when conducting the assay using 104 spiked cells at a flow rate of 0.5 ml/min. Calcein AM and EthD-1 were used to stain live and dead cells respectively, indicating that more than 95% of captured HT-29 cells were alive. Cell culture was tested with the same filter after filtration (Fig. 5a and 5b). After 3 days, live cells localized on the surface of the filter formed cell clusters due to growth (Fig. 5c). As can be seen, most of the clusters have a cell number less than 6, due probably to the single cell proliferation initially captured on the filter at different places. Considering a doubling time of ~23 h, not all captured cells are capable to expand. Some of cells need time for adaptation or repairing, some others could be lost during culture medium exchange, etc. Among the larger clusters, half have more than 8 cells per cluster, which is due probably to the capture of more than one cell in close proximity to each other.
Cell culture and viability test.
(a) Bright field image of cells on the PEGDA filter after capture and culture for 3 days. (b) Immunostaining fluorescence of HT-29 cells after Calcein AM/EthD-1staining. (c) Statistic data of cell number per cluster at day 4. Scale bar: (a) and (b) 50 μm. The error bars represent the s.d. of three times of counting with a number of randomly selected areas.
Discussion
We put forward the novel design of the filter with high density conical holes to achieve a useful compromise between the capture efficiency, purity and viability of CTCs. In general, the capture efficiency of CTCs increases with the decrease of the hole-size but a too much small hole size leads to poor capture purity because of the accumulation of undesired blood cells on the filter. The optimal hole-size preventing the loss of precious targeting cells should be comparable or smaller than the nuclei size of the CTCs, which can also be smaller or overlap with the size of erythrocytes. However, considering the fact that the shapes of erythrocytes can be extensively changed under the influence of mechanical forces in a flow or in a restraint area of circulation, erythrocytes may easily penetrate into the filter hole under the influence of applied transfilter pressure. The question is how erythrocytes can easily escape from the hole. The red blood cells (RBCs) are non-nucleated cells in form of disks of 6.2 ~ 8.2 µm diameters and 0.8 ~ 2.5 µm thickness. Although abundant, they can pass quickly through microholes of comparable sizes. WBCs have comparable or larger sizes. Once in the holes, their retention depends mainly on: (i) the transfilter pressure which cannot be too high for insuring viability of captured CTCs; (ii) the interaction with hole's wall; and, (iii) the shape of the hole. To illustrate the advantage of using conical holes, we consider a model of soft particles squeezed in two types of holes (cylindrical and conical) with the same size at the entrance. In simplified geometries shown in Figure 6a and 6b, one can relate the built-up pressure to the surface tension of the particle according to Laplace's law,

where γ is the surface tension, R1 and R2 are respectively the radius of the curvature of the tailing and leading edges of the particle. Thus, Δp = 0 for cylindrical holes and Δp > 0 for conical holes because of the difference between the two curvatures. In the case of conical holes and with a surface tension47 of ~3.0 × 10−5 N/m and a radius of curvature of tailing and leading edge R1 = 3.25 µm and R2 = 2R1, respectively, Δp ~ 9.2 Pa. Although this built-up pressure is rather small, it may efficiently stimulate WBCs to escape the conical holes, suggesting a better clearance of the conical-hole filter than that of the cylindrical ones. For comparison, our model was tested by performing capture of tumor cells under the same experimental conditions but using the two types of filtration holes. HT-29 cells were spiked into blood samples and then loaded into the devices at a flow rate of 0.2 ml/min. After capture and immunostaining, we deduced a capture efficiency of 96% (98%) and a WBC clearance efficiency of 96% (69%) with a conical (cylindrical) hole filter (Fig. 6c). As expected, the capture efficiencies of both filters were pretty high but much more WBCs were found on the filter of cylindrical holes showing a decreased capture purity (Supplementary Fig. S5). The retention of WBCs and other blood cells may have other side effects such aggregation due to the release of clogging factors from the deformed or lysed cells, making the cylindrical-hole clearance more difficult and thus increasing the transfilter pressure which is undesirable for a constant-flow filtration.
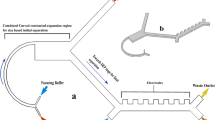
Model and comparison of cylindrical- and conical-hole filters.
(a, b) Schematic of cell retention in a cylindrical and conical hole: Cells squeezed in two filter types can have different built up pressure due to the difference in surface tension of their leading and tailing edges. (c) Cancer capture efficiency and WBC clearance efficiency with tumor cells spiked in donor blood at a flow rate of 0.2 ml/min. The error bars represent the S.D. of three measurements.
The cell viability is one of the most critical issues in current research of CTC capture since living cells can be expanded and used for downstream phenotypic and genotypic analyses. In principle, a low transfilter pressure should be applied during filtration to avoid the stress induced cellular damage. In our experiments, the transfilter pressure was controlled by the flow rate with a syringe pump. Consider a non-Newtonian fluid through a cylindrical hole at a constant flow rate, the transfilter pressure can be calculated by33

where a is the hole radius and L is the thickness of the hole, η is the viscosity and Q the flow rate of the liquid. For given a, η and Q, ΔP increases with L. In the case of conical holes, the calculation is more complex. In the limit of very large aperture angles, the thickness of the hole can be considered as infinitely small so that a minimal transfilter pressure is obtained. For the filter composed by an array of holes, the transfilter pressure should be inversely proportional to the number of holes. In our case, the number of holes in a 6 (9) mm diameter filter is 3.6 × 104 (8.2 × 104), which is huge comparing to the number of CTCs that can be captured. With a flow rate of 0.2 ml/min and a blood sample of viscosity of 3~4 × 10−2 Pa. s, we obtained a transfilter pressure of 213 ~ 284 (95 ~ 126) Pa for a 30 µm thick cylindrical-hole filter of 6 (9) mm diameter. In the case of conical holes with very large angles, the thickness of the filter can be considered to be very small (L ~ 0). Accordingly, Eq. (2) leads to a transfilter pressure of 24 ~ 32 (10 ~ 14) Pa, which means that the transfilter pressure of conical holes is one order of magnitude lower than that of 30 µm thick cylindrical-holes. Such a decrease of transfilter pressure can lead to a significant increase of the cell viability46,47,48,49, proving again the advantage of conical-hole filters. Moreover, such a small transfilter pressure will remain the same during filtration if the clearance of the filter is efficient. Note also that the small transfilter pressure resulting from the flow rate we applied should be significantly smaller than the stiffness of the most of tumor cells, which guarantees high cell viability during and after filtration. As expected, the most of captured tumor cells were immobilized in the hole area but on the surface of the filter due to the small size of the hole and the low transfilter pressure (Supplementary Fig. S6).
Finally, numeric simulation has been performed to compare the flow dynamics with the two types of filters using simplified geometry and boundary conditions (Supplementary Fig. S7). The results show that for the same transfilter pressure the flow rate across the conical holes is significantly larger than that across the cylindrical ones. In other words, under conditions of constant flow rate, the transfilter pressure of the conical holes should be significantly smaller than that of cylindrical ones, in agreement with the above analyses. These results also clearly show a flow focusing effect, suggesting the necessity of introducing a cross-flow component to flush gently away the cells accidently deposed on the surface of dead areas at equidistance of neighboring holes to avoid clogging of blood cells even in the absence of RBCs lysis. In our devices, this has been achieved by designing a number of radial channels connected between the inlet channel and the upper chamber (Supplementary Fig. S2). Effectively, this unique design allowed us to gain in WBC clearance efficiency. Nevertheless, this cross flow component should also be small enough to avoid additional flow stress. For a flow rate of 0.2 ml/min and a filter of 6.5 µm holes and 9 µm diameter, the maximum value of induced shear stress is ~8 Pa for a cell size of 15 µm diameter, which is reasonably small to avoid cell damage but sufficient to flush away the non-adherent cells. Note also that once a tumor cell is captured by a hole the flow can bypass this cell, reducing further the pressure and/or shear stress induced cell damage.
In conclusion, we developed a microfluidic device with a conical-shaped microhole array and a micro-injector with cross-flow components. This device was first validated by filtration of tumor cells spiked in blood and then used for detection of CTCs in blood samples of cancer patients. Furthermore, three tumor cell lines were tested with microhole arrays of different hole-sizes, showing a clear cell-type dependency of the capture efficiency and suggesting a (cell) nuclei size dependent design of microholes. By using tumor cells spiked in both blood of donors and PBS solution, we estimated a capture efficiency of ~95%. By using blood samples of both healthy donors and cancer patient, we estimated a WBC clearance efficiency of ~96%, which is significantly higher than that with a filter of cylindrical holes. Finally, tumor cells after filtration could be cultured on the same filter for several days, showing a cell viability of ~95%, which is generally required for more clear assessments of the captured CTCs. Together with the simplicity of the method and the possibility to integrate other microfluidic functions into the same device, more systematic studies are warranted to elucidate the underlying cellular and molecular mechanism of CTCs.
Methods
Fabrication of conical-hole microfilters
PEGDA filters were fabricated by UV assisted molding with a PDMS mold (Supplementary Fig. S1). First, a chromium mask was fabricated by UV photolithography to define microhole arrays of 5.5, 6.5, 8.0 µm diameters with 30 µm pitch sizes, using a pattern generator (μPG 101, Heidelberg Instr.). Then, the mask was spin coated with a 30 µm thick resist (AZ40XT, MicroChem) and backside exposed to a UV light at incident angle of 20°at a low rotation speed. After development, conical microholes were obtained, showing a half conical angle of ~13°. Afterward, a mixture of PDMS pre-polymer and cross-linker (GE RTV 615) at ratio of 10:1 was poured on the patterned resist layer. After curing at 80°C for 2 h, the PDMS layer was peeled off, resulting in a negative-tone replica (conical pillars) of the microhole array. Then, the patterned PDMS layer was placed on a glass slide and degased in a desiccator for 15 min. Finally, the cavity between PDMS layer and glass slide was filled with a photosensitive mixture of PEGDA and 2-hydroxy-2-methylpropiophenone at ratio of 1 v/v%., followed by a UV exposure. Similarly, 100 µm thick PEGDA rings of 13 mm out diameter and 6 or 9 mm inner diameter were prepared. They were then bonded to the filters (small diameter hole side) to enhance their mechanic stability.
Microfluidic device integration
Two-layer PDMS devices were fabricated by conventional soft-lithography. The bottom PDMS layer (Supplementary Fig. S2a) of the device was obtained with a mold of 100 µm thick photoresist layer (SU8 3050, MicroChem). The upper PDMS layer (Supplementary Fig. S2b) was prepared with a mold of PMMA produced by a computer numerical control (CNC) machining tool (Mini-Mill GX, Minitech Machinery Co.). Here one inlet and one outlet were designed in both layers to facilitate the post-filtration experiments. After separation of the upper PDMS layer, holes of inlet and outlet were punched. After separation of the bottom layer, a through hole of 9 mm diameter was punched. Then, the surfaces of both layers were treated with plasma for a few second before embedding the PEGDA filter. Finally, holes of the inlet and outlet of the bottom layer were punched and the whole assembly was thermally bonded to a glass slide.
CTC isolation
Three cell lines (MCF-7, HT-29 and U87) were prepared in 10% FBS and 1% penicillin/streptomycin with their respective media, McCoy's 5A Modified Medium and Dulbecco's Modified Eagle Medium (DMEM) at 37°Cwith 5% CO2 supplementation for 3–4 days. Prior to filtration tests, cells grown to confluence were detached by Trypsin and diluted into PBS or PBS diluted healthy human blood (VPBS:Vblood = 1:1) to the desired density. Human blood was collected from healthy donors in collection tube with EDTA to prevent coagulation and used within 24 h. A metallic needle was mounted on the bottom of an Eppendorf tube and then inserted in the inlet of the upper layer of the device to facility injection. Before sample introduction, the device was rinsed with PBS. Then, the sample was introduced from the Eppendorf tube by negative pressure generated with a syringe pump connected the outlet of the bottom layer. By varying the speed of the syringe pump, the flow rate of the sample across the filter could be controlled. After filtration of blood samples, the device was rinsed with 300 µl PBS once from inlet 1 to outlet 2 and once from inlet 2 to outlet 2 for more clear observation.
Immunostaining and microscopic imaging
The captured cells were fixed and treated inside the device. 4% paraformaldehyde (PFA) diluted in PBS at 10 μl/min for 15 min was injected from inlet 1 to outlet 2, followed by PBS rinsing at 50 μl/min for 6 min. Similarly, 0.2% Triton X-100 in PBS was injected into the device at 10 μl/min for 10 min for permeabilization and 300 μl 1% BSA in PBS was introduced at 10 μl/min for 30 min to block out non-specific bindings. Afterwards, cells were stained with dye solutions (passing from inlet 1 to outlet 1). First, 600 μl immunostaining solution, containing 5 μg/ml Anti-Pan-cytokeratin (AE1/AE3) Alexa Fluor 488- (1:100 diluted in PBS; eBioscience SAS) and Anti-Human CD45 PE (1:20 diluted in PBS, eBioscience SAS, France), was injected at 10 μl/min for 1 h to stain tumor cells and WBCs, respectively. Then, the nucleus were stained by 4′,6-diamidino-2-phenylindole (DAPI,100 nM, Life Technologies) at 10 μl/min for 10 min. Finally, the device was rinsed by 300 µl PBS for 6 min and fluorescence images were obtained with an inverted optical microscope (Zeiss, Axiovert 200) equipped with a digital CCD camera (Evolution QEI).
Cell culture after capture
A filter of 6.5 µm hole-diameter treated with Maxgel ECM mixture (1:25 diluted in complete McCoy's 5A medium, Sigma-Aldrich) at 37°C for 1 h was used for culture test. After filtration of HT-29 cells spiked in PBS at a flow rate of 0.2 ml/min, the upper PDMS layer could be peeled off and the filter was took out gently and placed into a petri dish. 70 μl culture medium was dropped on the filter in the center of the ring and then incubated for 1 h for cell attachment. Then, 1.5 ml medium was added gently around the ring for culture at 37°C in a 5% CO2 incubator.
Cell viability assay
Cell viability was studied by live/dead assay. Briefly, 2 μM of Calcein AM and 2 μM EthD-1 were added on the filter after filtration for live and dead cell staining, respectively. After 30 min incubation at 37°C and 5% CO2, cells were analyzed with a fluorescence microscope, as described above. Cell viability was calculated by live cells number divided by total cells number.
Scanning electron microscopy observation
Samples were fixed in PBS containing 4% formaldehyde for 30 minutes. Then, they were rinsed twice with PBS buffer and immersed in 30% ethanol (in DI) for 30 minutes. Afterward, the samples were dehydrated in a graded series of ethanol with concentrations of 50%, 70%, 80%, 90%, 95% and 100%, respectively, each for 10 min and dried with a nitrogen gas flow. Before observation, a 2 nm thick gold layer was deposited on the samples by sputtering. The observation was performed with a scanning electron microscope (Hitachi S-800) operated at 10 kV.
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–74 (2011).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Nguyen, D. X. et al. Metastasis: from dissemination to organ-specific colonization. Nat. Rev. Cancer 9, 274–84 (2009).
Klein, C. A. Parallel progression of primary tumours and metastases. Nat. Rev. Cancer 9, 302–312 (2009).
Hyun, K. A. & Jung, H. I. Advances and critical concerns with the microfluidic enrichments of circulating tumor cells. Lab Chip 14, 45–56 (2014).
Arya, S. K., Lim, B. & Rahman, A. R. Enrichment, detection and clinical significance of circulating tumor cells. Lab Chip 13, 1995–2027 (2013).
Sun, Y. F. et al. Circulating tumor cells: advances in detection methods, biological issues and clinical relevance. J. Cancer Res. Clin. Oncol. 137, 1151–73 (2011).
Paterlini-Brechot, P. & Benali, N. L. Circulating tumor cells (CTC) detection: clinical impact and future directions. Cancer Lett. 253,180–204 (2007).
Erica, D. et al. Rare Cell Capture in Microfluidic Devices. Chem. Eng. Sci. 66, 1508–1522 (2011).
Parkinson, D. R. et al. Considerations in the development of circulating tumor cell technology for clinical use. J. Transl. Med. 10, 138 (2012).
Cristofanilli, M. et al. Circulating tumor cells, disease progression and survival in metastatic breast cancer. N. Engl. J. Med. 351, 781–91 (2004).
Farace, F. et al. A direct comparison of cellsearch and iset for circulating tumour cell detection in patients with metastatic carcinomas. Br. J. Cancer 105, 847–853 (2011).
Myung, J. H., Gajjar, K. A., Saric, J., Eddington, D. T. & Hong, S. Dendrimer-Mediated Multivalent Binding for the Enhanced Capture of Tumor Cells. Angew. Chem. Int. Ed. Engl. 50, 11769–72 (2011)
Hoshino, K. et al. Microchip-based immunomagnetic detection of circulating tumor cells. Lab Chip 11, 3449–57 (2011).
Liu, Y. J. et al. A micropillar-integrated smart microfluidic device for specific capture and sorting of cells. Electrophoresis 28, 4713–22 (2007).
Saliba, A. E. et al. Microfluidic sorting and multimodal typing of cancer cells in self-assembled magnetic arrays. Proc. Natl. Acad. Sci. USA 107, 14524 (2010).
Nagrath, S. et al. Isolation of rare circulating tumour cells in cancer patients by microchip technology. Nature 450, 1235–9 (2007).
Stott, S. L. et al. Isolation of circulating tumor cells using a microvortex-generating herringbone-chip. Proc. Natl. Acad. Sci. USA 107, 18392–18398 (2010).
Wang, S. et al. Highly efficient capture of circulating tumor cells by using nanostructured silicon substrates with integrated chaotic micromixers. Angew. Chem. Int. Ed. Engl. 50, 3084–8 (2011).
Chen, Y. et al. Rare cell isolation and analysis in microfluidics. Lab Chip 14, 626–45 (2014).
Thiery, J. P. Epithelial-mesenchymal transitions in tumour progression. Nat. Rev. Cancer 2, 442–54 (2002).
Yu, M. et al. Circulating breast tumor cells exhibit dynamic changes in epithelial and mesenchymal composition. Science 339, 580–584 (2013).
Lv, P., Tang, Z., Liang, X., Guo, M. & Han, R. P. S. Spatially gradated segregation and recovery of circulating tumor cells from peripheral blood of cancer patients. Biomicrofluidics 7, 034109 (2013).
Hou, H. W. et al. Isolation and retrieval of circulating tumor cells using centrifugal forces. Sci. Rep. 3, 1259 (2013).
Sun, J. et al. Double spiral microchannel for label-free tumor cell separation and enrichment. Lab Chip 12, 3952–3960 (2012).
Sun, J. et al. Size-based hydrodynamic rare tumor cell separation in curved microfluidic channels. Biomicrofluidics 7, 011802 (2012).
Karabacak, N. M. et al. Microfluidic, marker-free isolation of circulating tumor cells from blood samples. Nat. Protoc. 9, 694–710 (2014).
Gupta, V. et al. ApoStream™, a new dielectrophoretic device for antibody independent isolation and recovery of viable cancer cells from blood. Biomicrofluidics 6, 024133 (2012).
Gascoyne, P. R. & Shim, S. Isolation of circulating tumor cells by dielectrophoresis. Cancers (Basel) 6, 545–79 (2014).
Shim, S. et al. Dielectrophoresis has broad applicability to marker-free isolation of tumor cells from blood by microfluidic systems. Biomicrofluidics 7, 011808 (2013).
Seal, S. H. A Sieve for the isolation of cancer cells and other large cells from the blood. Cancer 17, 637–642 (1964).
Khato, J., Sato, H., Suzuki, M. & Sato, H. Filtrability and flow characteristics of leukemic and non-leukemic tumor cell suspension through polycarbonate filters in relation to hematogenous spread of cancer. J. Exp. Med. 128, 273–84 (1979).
Holdich, R. et al. Pore design and engineering for filters and membranes. Phil. Trans. R. Soc. A 364, 161–174 (2006).
Vona, G. et al. Isolation by size of epithelial tumor cells - a new method for the immunomorphological and molecular characterization of circulating tumor cells. Am. J. Pathol. 156, 57–63 (2000).
Desitter, I. et al. A new device for rapid isolation by size and characterization of rare circulating tumor cells. Anticancer Res. 31, 427–41 (2011).
Zheng, S. et al. Membrane microfilter device for selective capture, electrolysis and genomic analysis of human circulating tumor cells. J. Chromatogr. A1162, 154–161 (2007).
Tan, S. J., Yobas, L., Lee, G. Y., Ong, C. N. & Lim, C. T. Microdevice for the isolation and enumeration of cancer cells from blood. Biomed. Microdev. 11, 883–892 (2009).
Hosokawa, M. et al. Size-selective microcavity array for rapid and efficient detection of circulating tumor cells. Anal. Chem. 82, 6629–6635 (2010).
Zheng, S. et al. 3D microfilter device for viable circulating tumor cell (CTC) enrichment from blood. Biomed. Microdev. 13, 203–213 (2011).
Hur, S. C., Mach, A. J. & Di Carlo, D. High-throughput size based rare cell enrichment using microscale vortices. Biomicrofluidics 5, 022206 (2011).
Lim, L. S. et al. Microsieve labchip device for rapid enumeration and fluorescence in situ hybridization of circulating tumor cells. Lab Chip 12, 4388–4396 (2012).
Lin, H. K. et al. Portable filter-based microdevice for detection and characterization of circulating tumor cells. Clin. Cancer Res. 16, 5011–5018 (2012).
Coumans, F. A. W. et al. Filter characteristics influencing circulating tumor cell enrichment from whole blood. PLoS ONE 8, e61770 (2013).
Hosokawa, M. et al. Size-based isolation of circulating tumor cells in lung cancer patients using a microcavity array system. PLoS ONE 8, e67466 (2013).
Adams, D. L., Amstutz, P., Tang, C. M. et al. The systematic study of circulating tumor cell isolation using lithographic microfilters. RSC Adv. 4, 4334–4342 (2014).
Lim, C. T., Zhou, E. H. & Quek, S. T. Mechanical models for living cells--a review. J. Biomech. 39,195–216 (2006).
Kuo, J. S. et al. Deformability considerations in filtration of biological cells. Lab Chip 10, 837–42 (2010).
Harouaka, R. A., Nisic, M. & Zheng, S. Y. Circulating tumor cell enrichment based on physical properties. J. Lab. Autom. 18, 455–68 (2013).
Harouaka, R. A. et al. Flexible micro spring array device for high-throughput enrichment of viable circulating tumor cells. Clin. Chem. 60, 323–333 (2014).
Acknowledgements
This work was supported by the European Commission under contract no. IRSES-GA-2009-247641 (Microcare) and Agence de Recherche Nationale under contract no ANR-13-NANO-0011-01. Y.T. is grateful to the Chinese Scholar Council for grant.
Author information
Authors and Affiliations
Contributions
Project planning and design: Y.C., J.S., Y.C. and L.W.; Experiments: Y.T., J.S. and S.L.; Numerical simulation: L.W.; manuscript: Y.C. Y.T., J.S.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
SUPPLEMENTARY INFO
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Tang, Y., Shi, J., Li, S. et al. Microfluidic device with integrated microfilter of conical-shaped holes for high efficiency and high purity capture of circulating tumor cells. Sci Rep 4, 6052 (2014). https://doi.org/10.1038/srep06052
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep06052
This article is cited by
-
Separation of fine particles and liquid using a physical trapping method via a stepped structure
Microsystem Technologies (2024)
-
From the teapot effect to tap-triggered self-wetting: a 3D self-driving sieve for whole blood filtration
Microsystems & Nanoengineering (2023)
-
Comparative application of microfluidic systems in circulating tumor cells and extracellular vesicles isolation; a review
Biomedical Microdevices (2023)
-
3D spiral channels combined with flexible micro-sieve for high-throughput rare tumor cell enrichment and assay from clinical pleural effusion samples
Bio-Design and Manufacturing (2022)
-
Pillar based microfluidic approach to sorting of microparticle mixtures at various particle ratios
Microfluidics and Nanofluidics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.