Abstract
Spatiotemporal mapping of the position and orientation of nano-machinery inside complex and dynamic cellular environments is essential for the detailed understanding of many bio-physical processes. For the genuine observation of such biomolecular dynamics with high signal to noise ratio and reduced disturbance from the labeling probes, reduction in the size of nano-bio labels and simplification of techniques for their observation are important. Here we achieve this using polarized dark field scattering micro-spectroscopy (PDFSMS), in its simplest form so that it is deployable in several experiments. We not only locate tiny gold nanorods (GNRs) of size 30 (length) × 10 nm (diameter) inside HEK293 cells but also demonstrate mapping of their in-situ polarization patterns using a novel method. Real time observations of rotating GNR with DFSMS and PDFSMS are used to resolve in-plane and out-of-plane rotational modes of GNR. We have shown that PDFSMS itself can provide complete information about the state of GNR. A step ahead, we demonstrate the application of PDFSMS to track three dimensional rotational dynamics of transferrin-conjugated GNRs inside live HEK293 cells. These first-time observations of the three dimensional intracellular rotational dynamics of tiny GNRs using PDFSMS present a new landmark in single particle scattering spectroscopy.
Similar content being viewed by others
Introduction
Development of techniques to study nano-bio interactions at a single particle level is of great importance to understand the complexity of biological functions1,2,3. In these studies, it is important to monitor spatial as well as temporal behavior of systems of interest. Although there has been extensive use of gold nanoparticles (GNPs) in biomedical applications4 and studies at nano-bio interfaces5, when it comes to monitoring orientational and rotational dynamics of molecules, gold nanorods (GNRs) are better nano-bio labels6 due to their anisotropic thermal7 and optical properties8,9. Different experimental methods have used these properties of GNRs to image and track single molecule orientation in vitro and in vivo6,10. Different techniques involve defocused or dual wavelength dark field imaging, two photon luminescence imaging or dark field imaging through bifringent crystal etc11,12,13,14. Differential interference contrast (DIC) imaging technique has been developed for single particle orientation and rotational tracking (SPORT) of GNR-labeled motor proteins and intracellular cargos in live cells10,15,16,17,18,19,20,21,22.
While there have been a few observations of single GNRs by optical techniques, the dimensions of the particles were above 60–75 nm (L) × 20–40 nm (D) for in vivo observations15,16,17,18,19. However, full potential of such single particle studies is realized when size of the probe is smaller than the system under observation. Smaller probes have lesser influence on cellular processes and allow observation of genuine dynamics. For example, observations of the rotational dynamics involved in functions of FoF1-ATP synthase6 and F1-ATPase23 were done using probes such as 77 (L) × 39 (D) nm GNRs and 1,000–2,600 (L) × 10 (D) nm actin filaments, respectively while size of these proteins is in the range of ~10 nm24,25. Even smaller probes would enable better understanding of the dynamics. Rotational dynamics of plasmonic particles below 40 nm have never been observed in cellular systems, although orientations have been discerned from the samples deposited on a substrate11,12,26. In the present work, we present polarized dark field scattering micro-spectroscopy (PDFSMS) of small GNRs of 30 (L) × 10 (D) nm (labeled GNR30X10 later) in complex cellular environments. Use of ultrasmall GNRs and VNIR light source instead of a laser source reduces the possibility of heating and minimizes the effect of probe itself on the system under observation13,14,27,28,29. Unlike previous studies, we have not used any confocal set-up or complex arrangement for polarized illumination or collection11,12,13,14,27,28,29. With a regular dark field set-up equipped with linear polarizer (analyzer), we not only determine orientation of GNRs but also map their in-situ polarization patterns (referred as polar maps). We also discuss the effect of cellular environment on the changing polarizability and scattering spectrum of GNR. Real time observations of rotating GNRs with DFSMS and PDFSMS have also shown that in-plane and out-of-plane rotational modes of GNR can be resolved using simple dark field microscopy. The observations on three dimensional movements of GNRs of the kind reported here has not been done so far. We have provided novel methods to map in-situ polar patterns of GNRs and resolve their rotational modes. Hence PDFSMS alone can provide complete information about the state of GNR such as three dimensional motion, orientation, rotational modes and whether it is an aggregate or single GNR. To the best of our knowledge, these are first time observations of three dimensional ultrasmall nanoparticle dynamics with simpler instrumentation.
Results
Set-up and principle of mapping the in-situ polar maps of GNRs
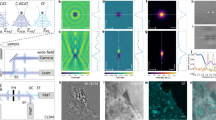
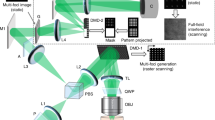
The key to achieve successful mapping of polarization patterns is to utilize the refraction of light rays when they pass through an analyzer with minute angular tilt. As shown in Fig. 1a, a regular dark field set-up was used for these measurements. Magnified view of the set-up in Fig. 1b explains the principle of in-situ polar mapping. Due to the intrinsic polarizability of GNR, scattered light has a major component polarized along the longitudinal axis of GNR. When this light passes through an analyzer, a minute angle in the mounting of the analyzer gives rise to refraction which causes the image to be shifted by a few pixels in the direction of the analyzer axis. This results in the mapping of a filled circle for scattering sources with no polarizability, as in the case of spherical GNP (Fig. 2b and supporting information (SI) Video S1). But for GNRs, circular mapping is modulated by variations in the scattering intensity (Fig. 2a and Video S2). Size distribution for the same is shown in (SI Fig. S1). Fig. 2b shows dark field images (b1, b2) of a single spherical GNP and an anisotropic GNP captured at different angles of the analyzer. As all GNPs are not perfectly spherical in a typical synthesis, such collection of particles can always be seen in a sample. Unlike GNR30X10, in this case, the aspect ratio is less with larger diameter as suggested by the scattering spectrum (Fig. S2). On rotation of the analyzer, paths tracked by both these particles shown in Fig. 2b (b3) follow a circle. Corresponding polar maps, calculated by SI Equation 1, are shown in Fig. 2b (b4). Comparison of polar plots calculated by SI Equation 2 and polar maps prove that the particle orientation is reproduced sincerely (Fig. S2). Here polar plots provide angular distribution of scattering intensity integrated over particle spot (in image) area. But polar maps provide visualization of how these changes occur in space. Polar mapping is sensitive to even smaller deviations in the shape of the nanostructure from an isotropic sphere. Hence it works also for GNPs which are not exactly spherical (Fig. S3, S4). For such GNPs, value of polarization anisotropy (P, calculated by SI Equation 3) was ~0.4 whereas for GNRs it was ~1.0. A transmission electron microscopic (TEM) image of GNR30x10 used in this study is shown in Fig. 1c, along with a high resolution image of one particle.
In-situ polarization pattern mapping was achieved by displacement of the image caused by a tilted analyzer.
(a) Schematic illustration of the experimental set-up used for in-situ polar mapping of GNRs. (b) Principle of polar mapping is explained in this ray diagram. Briefly, this is possible due to minute angular tilt (δϕ) in the mounting of the analyzer which causes refraction of the polarized light scattered by GNR. Figures are not to scale. (c) A transmission electron microscopic (TEM) image of the GNR30x10 is shown along with a high resolution image of the tip of one GNR (inset).
Imaging, spectroscopy and polar mapping of single GNRs.
(a) Dark field images of a single GNR30X10 captured at different angles of the analyzer. Corresponding variations in the scattering intensity are shown in the form of a polar plot. (b) Dark field images of a single spherical GNP and a single anisotropic GNP at different angles of analyzer are shown in b1 and b2. Only difference between the images is the analyzer orientation, shown by the double headed arrow, ‘↕’ for 0 degree (b1) and ‘↔’ for 90 degrees (b2). Path tracked by both these particles are shown in the image b3 and corresponding polar maps determined by SI Equation 1 are shown in the image b4. (c) Scattering spectra of a single GNR and an aggregate of GNRs formed upon interaction with bovine serum albumin (BSA) are shown along with the spectrum of halogen lamp used for illuminating the sample.
Figure 2c shows the lamp spectrum with maximum in the region where LSPR (longitudinal surface plasmon resonance) of GNR30X10 occurs (check Fig. S5 for DDA simulated spectrum). This helps in reducing the noisy background from the cellular components which scatter mostly in the blue-green region (Fig. S6). This facilitates thorough hyperspectral analysis of the changes in the scattering spectra of GNR which are discussed in greater detail in the next section.
Spectroscopic identification of single GNPs, GNRs and GNR aggregates in vitro and in vivo. It is well known that SPR (surface plasmon resonance) of nanostructures is very sensitive to the changes in the surrounding environment8. Hence before proceeding to in vivo mapping measurements, we have established a method to identify distinct scattering features of small GNRs in different environments. GNP40 are included in this comparison because GNR30X10 aggregates and GNP40 look almost similar in their dark field images but can be distinguished only on the basis of their spectral features as explained later. Figure 3 shows the consolidated data obtained after statistical analysis of the single particle SPR peak positions of GNP40, GNR30X10 and GNR30X10 aggregates in vitro and in vivo. Examples of spectra corresponding to each SPR peak distribution are given in Fig. S7 and Gaussian curve fits for their peak positions are given in Fig. S8. Figures 3a & 3b show optical dark field images of GNP40 and GNR30X10. Although the volume of GNR30X10 is much smaller than that of GNP40 (~1/15 of GNP40), GNP40 has peak distribution from 500–600 nm but GNR30X10 has a distribution of SPR peaks from 600–800 nm (Fig. 3e). Gold nanorods exhibit this property due to aspect ratio dependence of their SPR. It is expected that cetyltrimethylammonium bromide (CTAB)-stabilized particles (although cleaned) aggregate on cell membrane either due to receptor clustering30 or due to interaction with proteins in cell culture medium31. Hence to monitor changes in the scattering signal of GNR30X10 after aggregation or clustering, we have done analysis of GNR30X10 samples incubated with BSA (GNR@BSA) and compared their SPR peak distribution with the scattering signals obtained from GNR30X10 treated HEK293 cells. TEM images and UV-Vis absorption spectra of GNR@BSA samples are shown in Fig. S9 and S10, respectively. Figure 3e shows the SPR distribution of GNR@BSA. The various SPR peaks are labeled as shown in Fig. 3e. We see that those scattering spectra of GNR@BSA have only one SPR peak (labeled, GNR@BSA:S1P1); their peak distribution is centered at ~740 nm with small fraction of peaks at ~530 nm (see their relative area). However, many scattering spectra of GNR@BSA have two SPR peaks; first one centered at ~530 nm (labeled, GNR@BSA:S2P1) and the other one centered at ~730 nm (labeled, GNR@BSA:S2P2). To confirm the presence of two SPRs, we have simulated the scattering spectrum of an aggregate of six GNR30X10 as shown in Fig. 3f & 3g. We see that although GNR30X10 exhibits only one prominent SPR, an aggregate exhibits two strong SPRs. The possible reason behind this is the plasmonic coupling between multiple GNR30X10 which enhances the TSPR (transverse surface plasmon resonance) of GNR30X10 and closely packed GNRs in an aggregate tend to behave as a single particle equivalent to a GNR of reduced aspect ratio. This would give equally prominent TSPR and LSPR. As a result of this plasmonic coupling, a red shift in LSPR of GNR@BSA can also be observed. GNP40:S1P1 is red shifted as compared to GNR@BSA:S2P1 (Fig. 3e); possibly due to smaller diameter of GNR30X10. Ensemble UV-Vis absorption spectra also show that SPR for GNP40 occurs at 532 nm, whereas TSPR of GNR30X10 occurs at 514 nm (Fig. S11). It suggests the possibility that major component of GNR@BSA:S1P1 arises due to chains or chain-shaped aggregates of GNR30X10. It is known from previous studies that zwitterionic molecules such as small peptides present in cellular environment may give rise to chain-shaped aggregates of GNRs32. Observation of two prominent peaks was also supported by the results obtained by analyzing hyperspectral image of HEK293 cells treated with GNRs (GNR@HEK293). It was observed that the distribution of GNR@HEK293:S2P1 and GNR@HEK293:S2P2 matches with the distribution of GNR@BSA:S2P1 and GNR@BSA:S2P2, except that there is a small increase in the baseline for GNR@HEK293 scattering spectra in the region around 600 nm (Fig. S6). Increase in the baseline can be due the background scattering from surrounding cellular components. It should be noted that, although appearance of two prominent SPR peaks in experimental results matches the simulated data, this observation cannot be generalized. This is because GNR aggregation is hard to control in complex biological environments and many different geometries or combinations can exist inside cells. Hence on the basis of these observations, we can not only locate aggregates of small GNRs in complex cellular environment but also differentiate between an intact GNR and an aggregate based on the number and position of SPR peaks. It should be emphasized that although GNP40 and GNR@BSA look almost similar in optical dark field images, they can be distinguished clearly using their spectra. In the next section we compare the polar mapping and effect of various environments on the polarizability of GNR30X10.
Spectroscopic identification of single GNPs, GNRs and GNR aggregates in vitro and in vivo.
Dark field images of various samples at 100× magnification. (a) GNPs of 40 nm diameter (GNP40). (b) GNR30X10 (c) GNR30X10 aggregates obtained by treating it with BSA. (d) GNR30X10 aggregates in HEK293 cells. (e) Consolidated data obtained from the Gaussian curve fitting of the statistical distributions of single particle SPR peaks of the samples (a-d). In labels, ‘S’ stands for the total number of SPR peaks in the scattering spectrum and ‘P’ stands for a serial number of SPR peak when it is counted from lower wavelength to higher wavelength. Area of each of the Gaussian peak represents the number of pixels in the hyperspectral image whose scattering maximum falls within its base width. (f) Surface images of the model dipole arrays generated for DDA simulations of samples (a-d). Please note the relative size of these nanostructures. (g) Corresponding DDA simulated scattering cross sections of nanostructures on unpolarized white light excitation. Color coding is given such that simulated spectra can be compared with the experimental results obtained for the samples (a-c).
Effect of various environments on the polarizability of GNR30X10
To check the applicability of polar mapping in different environments, multiple observations were made. Figure 4a and 4b show the polar plots and corresponding maps for a single GNP40 and a single GNR30X10, respectively. For GNP40 with small polarization anisotropy, the mapped pattern is just a filled circle but for GNR30X10, it exhibits a filled ‘figure 8’ pattern. In Fig. 4b, 4c and 4d we can see how the scattering spectra, polar plots and polar maps of GNR30X10 change sequentially from the free state, during interaction with HEK293 cell membrane and after entering the cell. Scattering spectrum for GNR30X10 on HEK293 cell surface (Fig. 4c) suggests that the GNR is at an initial stage of clustering or aggregation. It should be noted that, although due to diffraction limit of optical microscopy, an aggregate appears like a single particle; it can be very well distinguished by its spectroscopic features. It can be seen that polar plot and map exhibited by GNR30X10 aggregate inside the cell is similar to an aggregate of GNR30X10 treated with BSA (Fig. 4e & 4e). Figure S12 provides an example of such a mapping done over the whole HEK293 cell, treated with GNR30X10. Please note the changes in P for all these samples, it justifies that unless the nanostructure under observation is a perfect sphere, despite the small value of P, maps are efficient in determining in-situ orientation. Hence with the criterion discussed above, PDFSMS alone can provide complete information about the state of small GNRs in cellular environments.
Effect of various environments on the polarizability of GNR30X10.
Each column in this figure shows (top to bottom) a polar plots, polar maps and single particle scattering spectrum for, (a) GNP40, (b) GNR30X10, (c) GNR30X10 on the HEK293 cell surface, (d) GNR30X10 inside HEK293 cell and (e) GNR30X10 treated with BSA. Polar plots indicate the nature of GNP or GNR being examined. Inset images in each of the polar map images show dark field images of the particles when analyzer was (left) parallel (↔) and (right) perpendicular (↕) to the longitudinal axis defined by the orientation of the analyzer where maximum scattering occurs.
Real time rotational dynamics of GNR30X10 using PDFSMS
For in vitro observations of rotational dynamics, we have monitored freely moving GNRs in a viscous solution of PEG (GNR@PEG). Rotational dynamics of GNRs can occur at the time scale of microseconds to seconds. In this study, due to limitations of the temporal resolution of the camera, we could perform time lapse measurements only at the temporal resolution of 100 ms. This allows us to observe longer time scale rotational dynamics of GNRs. Figures 5a and 5b show the time-lapse images of a freely moving single GNR30X10 in such a solution captured with and without analyzer, respectively. In our previous observations with GNR@BSA, we have observed that SPR peak position of GNR changes after interaction with BSA; but forGNR@PEG, SPR position remains constant (Fig. S13). This suggests the existence of single isolated GNRs instead of aggregates. For GNRs with deionized water as surrounding medium (GNR@DI), TSPR in single GNR scattering spectrum is nearly zero. But in the case of GNR@PEG, possible adsorption of PEG on the GNR surface gives rise to increased SPR width and non-zero TSPR component (Fig. S13). Due to this, when orientation of GNR@PEG is perpendicular to the analyzer axis, its scattering intensity is not zero. For particles like this, non-zero TSPR gives an additional advantage of discrimination between orientation change and Z axis movement (see below). Change in color - mainly variations in the red channel scattering intensity of GNR30X10 - can be used to determine its orientation. Whereas width of the 2D Gaussian fitted to GNR spot (WG) in the RGB (red-green-blue) image can be used to determine its movement along the Z axis. From the temporal changes in WG (Fig. 5a (blue curve) and 5b (green curve)), we see that WG decreases to less than ~60% of the WG value for in focus GNR, only if its movement occurs along the Z axis. Generally on defocusing, scattering based image of GNR becomes diffuse, ring shaped and reduced signal to noise ratio leads to reduction in WG. Such images are shown for GNR@DI in Video S3. As per these observations, our system can resolve 1 micron movement along Z axis. Since color is important for the determination of in focus GNR30X10 orientation, images were color saturated instead of improving brightness and contrast required for the observation of defocused GNR30X10 patterns. For orientation measurements, intensity variations that occur due to Z axis movement of the scatterer can be eliminated by subtracting the green channel intensity from the red channel intensity. Details of this baseline correction are explained in SI Discussion 1 with an example of a rotating GNR30X10 in HEK293 cell, in comparison with a vesicle, the latter can well be considered as an isotropically scattering particle. Figure 5b shows that changes in color and scattering intensity of GNR can also be observed up to some extent even without analyzer in the optical path. The reason behind this is explained in the next section, with the help of DDA simulations.
Real time rotational dynamics of GNR30X10 using PDFSMS.
(a) Time-lapse (100 ms) images of a single GNR30X10 in viscous solution of PEG captured through the analyzer. To fit in smaller space, images are stacked such that time (seconds) for each image can be calculated by adding tX and tY time coordinates and can be correlated with the graph shown below. Corresponding variations in the scattering intensity (magenta curve) and 2D Gaussian width (blue curve) of the particle spot are shown in the graph below. Arrow indicates a time point when GNR moves along Z axis, goes completely out of focus and reverts back to the focal plane. (b) Images of a single GNR30X10 from the same sample captured without the analyzer are shown with corresponding variations in the scattering intensity (red curve) and spot width (green curve). Arrow has same meaning as in (a). (R-G) indicates that variations are shown for the difference between red and green channel scattering intensity. Next figures show DDA simulated scattering cross sections of a single GNR30X10 performing different rotational modes. Arrows with letter P show allowed polarizations while illumination of GNR in Z direction. Descriptions of rotational modes are: (c) out-of-plane rotation mode with unpolarized illumination, (d) in-plane rotation mode with unpolarized illumination and (e) in-plane rotation mode with polarized illumination.
Observations of a rotating GNR with and without analyzer
Figures 5(c,d & e) show the results of DDA simulations of GNR30X10 performing in/out of plane rotation. Upon unpolarized white light excitation, scattering cross section of GNR performing out-of-plane rotation will vary, as shown in Fig. 5c. Hence orientation of GNR performing such rotation can be sensed even without polarized collection as we have said before. But for a GNR performing in-plane rotation, polarized illumination is necessary to determine its orientation (Fig. 5d & 5e); conversely, upon unpolarized illumination, this rotational mode can be observed by polarized collection. This also suggests the possibility that rotational modes of GNR illuminated with unpolarized light can be resolved by simultaneous observation with and without analyzer. In the absence of a simultaneous DFSMS and PSFSMS imaging setup, separate measurements were performed on GNRs rotating inside a viscous PEG solution using DFSMS and PSFSMS separately. As shown in Fig. 5a, when observed with PDFSMS, changes in the color of GNR can be seen for both in-plane and out-of-plane rotations. But when GNR was seen with DFSMS, changes in the color of GNR can be seen only for out-of-plane rotation. Here in this study we have focused only on the observations that can be done with a simple setup of PDFSMS. Next section discusses the feasibility of such observations of real time rotational dynamics of GNR30X10 inside HEK293 cell and development of a methodology for the same.
3D Three dimensional spatiotemporal mapping of GNR@Tf inside HEK293 cell
In case of GNR@PEG, when GNR moves along the Z axis, we observed a decrease in WG. However, for GNR@Tf@HEK293 (transferrin-conjugated GNR inside HEK293 cell), when GNR@Tf goes drastically out of focus, an increase in WG was observed. This is attributed to the noise in the environment inside cell; because of which when GNR goes completely out of focus, noisy surrounding results in a fit with the 2D Gaussian of larger width. Also, in such observations, GNR movement occurs in three dimensions in space; because of which, camera gain and focus settings need to be changed and optimized time to time. All these changes are mentioned in detail at each place wherever it has been made. Figure 6a shows the scattering spectrum of GNR@Tf when it was attached on the cell membrane. It still shows a sharp SPR at ~600 nm, suggesting the existence of single GNR30X10. Figure 6c shows the position of GNR inside the cell; it shows that GNR under observation has tracked its path in a constricted environment between the nucleus and the cell membrane. Figure S14 shows an atlas of all the images of GNR corresponding to the movement of GNR inside cell. Observed events can be explained in detail as follows. During the first one minute, apart from translational motion, no considerable change in the orientation and WG of GNR was observed (red color track in Fig. 6f). This initial data collected at slightly lower gain of camera is shown in Fig. S15. After that, the gain was optimized for proper observation of GNR and kept constant throughout the measurements (indicated by green arrow in Fig. 6f). Then fusion between GNR and another vesicle was observed. Video S4 shows this binding event; where it can be seen that when GNR moves or gets attracted towards the vesicle, its scattering intensity increases. This is reflected in the increase in WG with corresponding increase in the scattering intensity (Fig. 6d & 6e). Observation of this fusion event also explains the possible reason behind red shift in the scattering spectrum of GNR (Fig. 6b). Then after minor focus adjustment, GNR was monitored for ~1 min; during which it kept moving towards the interior of the cell. Then it started fluctuating along the Z direction and went slightly out of focus which was reflected in the decrease of WG but still it was possible to monitor it (hence no focus adjustment was necessary at this point). But after ~20 seconds, when it reached in the crowded and constricted interior of the cell, its movement along the Z direction increased and it went quite out of focus. Then the focus was adjusted to an optimized position to monitor the GNR. In this highly interactive environment, it resulted in enormous fluctuations in the value of WG and orientational changes for the next 2 min; which we have tentatively interpreted as a process in which TfR dissociates from GNR. However, after this process, when GNR started coming out of the interior, its fluctuations in the Z direction decreased considerably. Then it kept moving towards cell membrane with steady values of WG, intervened by large variations at some places. These large variations can be interpreted as due to collisions with other vesicles and attempts of GNR to come outside the cell. Here due to large scattering from vesicles, a minor fluctuation in the path tracking occurs. By correlating temporal variations in the scattering intensity (R-G) and WG (shown by blue line in Fig. 6d & 6e), it can be inferred that GNR movements along Z direction occur by out-of-plane rotations. Studies have shown that for Tf or GNR@Tf an average time required for penetration of the cell membrane is ~7 min10,33. This implies that movement of GNR inside cell starts 7 min after it begins interacting with cell surface proteins10,33. Our observations over a time span of ~10 min starts with the movement of GNR@Tf from cell membrane towards interior of the cell and then back towards the cell membrane at the end of the observation (Video S5). This is ~half of the transit time (21 min) that is reported for transferrin receptors33, which suggests that our observations start much after the entry of GNR inside the cell and may involve dissociation of GNR from TfR so that GNR follows a different path to return towards the cell membrane.
3D Three dimensional spatiotemporal mapping of GNR@Tf inside HEK293 cell.
(a) Scattering spectrum of a single GNR@Tf attached on HEK293 cell membrane. (b) Scattering spectrum of the same GNR@Tf after entering inside the cell. Insets show corresponding hyperspectral images. (c) Image of a HEK293 cell showing the position of the monitored GNR@Tf inside the cell. The GNR is marked with a square. Inset shows an enlarged view of the GNR. Light scattered in the Z direction was collected through analyzer, whose orientation is shown by yellow double arrow. (d) Temporal variations in the scattering intensity (R-G) of GNR@Tf. Time scale corresponds to the axis of the graph shown below. Pink vertical bars show the region where microscope focus was adjusted on the particle after it went out of the focal plane. (e) Temporal variations in the 2D Gaussian width of GNR spot. Labeled events are discussed in the manuscript text. (f) Path of GNR@Tf inside the HEK293 cell. Green arrow shows the time point from where temporal data of the GNR is shown. Color of the trace corresponds to the time scale of graphs (d,e). Please note that the background image is just to give a rough idea of the position of GNR inside the cell.
Discussion
To conclude, we have used PDFSMS in its simplest form to develop a methodology for three dimensional spatiotemporal orientational mapping of tiny GNR30X10. We have shown that minute angular tilt in the analyzer can be used as an advantage to map in-situ polarization patterns (called as polar maps) and hence orientation of GNRs. Using statistical analysis of single particle SPR peaks and DDA simulations, we have given a criterion to distinguish GNR aggregates from GNPs and single GNRs. It is shown that aggregation of GNR30X10 induced by proteins in cellular environment gives rise to two prominent SPR peaks. This criterion was used to compare the spectroscopic features of samples such as GNP and GNR on glass substrate, GNR on cell membrane and an aggregate of GNR on glass substrate and inside the cell; the data were correlated with the polar plots and maps to understand the effect of environment on the polarizability of these particles. We have also demonstrated PDFSMS for real time observations of the rotational dynamics of GNR30X10 in cellular environment. From the observations of freely moving GNR in viscous solution of PEG, it was inferred that movement of GNR along Z axis can be tracked by decrease in WG, the width obtained by 2D Gaussian fitting of GNR spot in dark field image. In contrast, in a noisier cellular environment, WG increases whenever GNR movement occurs along the Z axis. Observation made on tracking of GNR@Tf inside HEK293 cells suggest that motion along Z axis occurs with out-of-plane rotations. To summarize, we have shown that simply DFSMS with an analyzer can be potentially used for understanding the overall state of an anisotropic nanostructure in complex environments.
As a future direction, our data suggests the possibilities that Intracellular path of Tf and GNR@Tf might be different and this may also involve dissociation of Tf itself from GNR. We plan to investigate behavior of GNRs using simultaneous polarized and unpolarized DFSMS to resolve the rotational modes of GNR. Combination of this technique with real time signal controlled microscope stage will be useful for further improvements in the three dimensional rotation and orientation tracking of GNR-labeled intracellular components.
Methods
Polarized dark field scattering micro-spectroscopy
HSI measurements were performed using Cytoviva hyperspectral imaging system equipped with Cytoviva™ high resolution dark field condenser (oil immersion). UPLFLN 100XOI2 oil immersion objective (Olympus) with NA of 1.3–0.6 and working distance of 0.2 mm was used for magnification. Analyzer was kept in the optical path after the objective. A L1090 - Halogen Lamp (400–1,000 nm) with Aluminum Reflector (International Light Technologies Inc., USA) was used for illumination of the sample. Optical images were collected using Dagexcel-M Color Camera – Cooled (CCD, 2,048 × 2,048). Spectra were measured using the Specim V10E spectrometer (400–1,000 nm) at a spectral resolution of ±1.3 nm. To tackle with the lower scattering cross section of ultrasmall nanoparticles, measurements were performed at full power illumination of source (150 W) and enhanced camera gain. For live cell experiments CV-30 CytoViva™ Environment Chamber provided by Warner Instruments was used. DMEM and GNR solutions were pumped using NE-300 Just Infusion™ Syringe Pump.
Cell culture and maintenance
HEK293 cells were cultured in Dulbecco's Modified Eagle Medium (DMEM) containing 10% foetal bovine serum (FBS) and antibiotics (100 units of penicillin and 10 µg streptomycin/mL). For steady state observations with fixed cells, following protocol was used. HEK293 cells were cultured on poly-L-lysine-coated 0.145 mm thick Nexterion® clean room cleaned glass coverslips (SCHOTT) in six well plates. Once cells reached ~80% confluency, they were washed twice with 1X phosphate buffered saline (PBS) and 2 mL DMEM containing 100 µL GNR was added. These cells were incubated for 10 min inside the incubator at 37°C. Then cells were washed thrice with 1XPBS and incubated for 8 min with 4% paraformaldehyde solution in 1XPBS. Then cells were washed thrice with PBS and incubated with ethanol gradient (25%, 50%, 75%, 100%) 10 min each. Then cells were dried and mounted with DPX mountant on 1 mm thick ultrasonically cleaned glass slide covered with 0.145 mm thick Nexterion® clean room cleaned glass coverslips (SCHOTT) for HSI measurements.
For live cell experiments, HEK293 cells were cultured on poly-L-lysine-coated 40 mm round glass slides (Warner instruments) in 60 mm petri dishes. After reaching 80% confluency, cells were washed once with 1XPBS and fresh DMEM. Then the slide was mounted on CV-30 set-up immediately and after initial pumping, flow of DMEM was maintained at 0.05 mL/hr. A three way valve was used to switch between DMEM and (DMEM + GNR) solutions.
Image analysis
Image processing, particle tracking and RGB analysis was done using NIH ImageJ34 which included particle tracker plugins developed by MOSAIC group (ETH, Zurich)35 and Fabrice Cordeli (Institut Curie, Orsay, France). Details of image processing algorithms are given in SI Method 1. MATLAB scripts were developed for polar mapping, tracking the integrated RGB intensity of moving particle images and to extract particle images separately from large area images. For 2D Gaussian fitting of particle images, a script developed by Gero Nootz (MATLAB CENTRAL File ID: #37087) was used. Spectral image analysis was done using ENVI and homebuilt software LSPRPEAK1.001 (Fig. S16).
DDA simulations
DDA simulations were done using DDSCAT7.3 package developed by Draine and Flatau36,37,38. Refractive index data provided by Johnson and Christy et al. was used for gold39. Particle or aggregate models were generated by DDSCAT7.3 code36,37,38 and images were generated by Paraview software40. In order to simulate spectra with unpolarized excitation, scattering cross sections were averaged for different polarizations of incident electromagnetic wave varying from 0–360° at an interval of 10°. To match with the experimental data, simulated spectra were corrected using lamp spectrum.
References
Stender, A. S. et al. Single cell optical imaging and spectroscopy. Chem. Rev. 113, 2469–2527 (2013).
Mattoussi, H., Palui, G. & Na, H. B. Luminescent quantum dots as platforms for probing in vitro and in vivo biological processes. Adv. Drug Delivery Rev. 64, 138–166 (2012).
Kawashima, N. et al. Reversible dimerization of egfr revealed by single-molecule fluorescence imaging using quantum dots. Chem. Eur. J. 16, 1186–1192 (2010).
Dreaden, E. C., Alkilany, A. M., Huang, X., Murphy, C. J. & El-Sayed, M. A. The golden age: Gold nanoparticles for biomedicine. Chem. Soc. Rev. 41, 2740–2779 (2012).
Alkilany, A. M., Lohse, S. E. & Murphy, C. J. The gold standard: Gold nanoparticle libraries to understand the nano-bio interface. Acc. Chem. Res. 46, 650–661 (2013).
Ishmukhametov, R., Hornung, T., Spetzler, D. & Frasch, W. D. Direct observation of stepped proteolipid ring rotation in e. Coli fof1-atp synthase. EMBO J. 29, 3911–3923 (2010).
Ma, H., Bendix, P. M. & Oddershede, L. B. Large-scale orientation dependent heating from a single irradiated gold nanorod. Nano Lett. 12, 3954–3960 (2012).
Khlebtsov, B. N. & Khlebtsov, N. G. Multipole plasmons in metal nanorods: Scaling properties and dependence on particle size, shape, orientation and dielectric environment. J. Phys. Chem. C 111, 11516–11527 (2007).
Lereu, A. L. et al. Spectroscopy and imaging of arrays of nanorods toward nanopolarimetry. Nanotechnology 23, 045701 (2012).
Gu, Y., Sun, W., Wang, G. & Fang, N. Single particle orientation and rotation tracking discloses distinctive rotational dynamics of drug delivery vectors on live cell membranes. J. Am. Chem. Soc. 133, 5720–5723 (2011).
Li, T. et al. Three-dimensional orientation sensors by defocused imaging of gold nanorods through an ordinary wide-field microscope. ACS Nano 6, 1268–1277 (2012).
Failla, A. V., Qian, H., Qian, H., Hartschuh, A. & Meixner, A. J. Orientational imaging of subwavelength au particles with higher order laser modes. Nano Lett. 6, 1374–1378 (2006).
Xiao, L., Qiao, Y.-X., He, Y. & Yeung, E. S. Three dimensional orientational imaging of nanoparticles with darkfield microscopy. Anal. Chem. 82, 5268–5274 (2010).
Xiao, L., Wei, L., Liu, C., He, Y. & Yeung, E. S. Unsynchronized translational and rotational diffusion of nanocargo on a living cell membrane. Angew. Chem. Int. Ed. 51, 4181–4184 (2012).
Ha, J.-W., Sun, W., Wang, G.-F. & Fang, N. Differential interference contrast polarization anisotropy for tracking rotational dynamics of gold nanorods. Chem. Commun. 47, 7743–7745 (2011).
Sun, W., Gu, Y., Wang, G. & Fang, N. Dual-modality single particle orientation and rotational tracking of intracellular transport of nanocargos. Anal. Chem. 84, 1134–1138 (2012).
Ha, J. W., Sun, W., Stender, A. S. & Fang, N. Dual-wavelength detection of rotational diffusion of single anisotropic nanocarriers on live cell membranes. J. Phys. Chem. C 116, 2766–2771 (2012).
Wang, G., Sun, W., Luo, Y. & Fang, N. Resolving rotational motions of nano-objects in engineered environments and live cells with gold nanorods and differential interference contrast microscopy. J. Am. Chem. Soc. 132, 16417–16422 (2010).
Gu, Y. et al. Revealing rotational modes of functionalized gold nanorods on live cell membranes. Small 9, 785–792 (2013).
Marchuk, K., Ha, J. W. & Fang, N. Three-dimensional high-resolution rotational tracking with superlocalization reveals conformations of surface-bound anisotropic nanoparticles. Nano Lett. 13, 1245–1250 (2013).
Xiao, L., Ha, J. W., Wei, L., Wang, G. & Fang, N. Determining the full three-dimensional orientation of single anisotropic nanoparticles by differential interference contrast microscopy. Angew. Chem. Int. Ed. 51, 7734–7738 (2012).
Gu, Y. et al. Rotational dynamics of cargos at pauses during axonal transport. Nat. Commun. 3, 2037 (2012).
Noji, H., Yasuda, R., Yoshida, M. & Kinosita Jr, K. Direct observation of the rotation of f1-atpase. Nature 386, 299–302 (1997).
Palanisami, A. & Okamoto, T. Torque-induced slip of the rotary motor f1-atpase. Nano Lett. 10, 4146–4149 (2000).
Junge, W. Atp synthase and other motor proteins. Proc. Natl. Acad. Sci. U. S. A. 96, 4735–4737 (1999).
Yasuda, R., Noji, H., Yoshida, M., Kinosita, K. & Itoh, H. Resolution of distinct rotational substeps by submillisecond kinetic analysis of f1-atpase. Nature 410, 898–904 (2001).
Xiao, L., Qiao, Y., He, Y. & Yeung, E. S. Imaging translational and rotational diffusion of single anisotropic nanoparticles with planar illumination microscopy. J. Am. Chem. Soc. 133, 10638–10645 (2011).
Ha, J. W., Marchuk, K. & Fang, N. Focused orientation and position imaging (fopi) of single anisotropic plasmonic nanoparticles by total internal reflection scattering microscopy. Nano Lett. 12, 4282–4288 (2012).
Biswas, S., Nepal, D., Park, K. & Vaia, R. A. Orientation sensing with color using plasmonic gold nanorods and assemblies. J. Phys. Chem. Lett. 3, 2568–2574 (2012).
Bartczak, D., Sanchez-Elsner, T., Louafi, F., Millar, T. M. & Kanaras, A. G. Receptor-mediated interactions between colloidal gold nanoparticles and human umbilical vein endothelial cells. Small 7, 388–394 (2011).
Rosman, C. et al. A new approach to assess gold nanoparticle uptake by mammalian cells: Combining optical dark-field and transmission electron microscopy. Small 8, 3683–3690 (2012).
Sudeep, P. K., Joseph, S. T. S. & Thomas, K. G. Selective detection of cysteine and glutathione using gold nanorods. J. Am. Chem. Soc. 127, 6516–6517 (2005).
Bleil, J. D. & Bretscher, M. S. Transferrin receptor and its recycling in hela cells. EMBO J. 1, 351–355 (1982).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH image to imagej: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Sbalzarini, I. F. & Koumoutsakos, P. Feature point tracking and trajectory analysis for video imaging in cell biology. J. Struct. Biol. 151, 182–195 (2005).
Draine, B. T. & Flatau, P. J. Discrete-dipole approximation for scattering calculations. JOSA A 11, 1491–1499 (1994).
Draine, B. T. & Flatau, P. J. Discrete-dipole approximation for periodic targets: Theory and tests. JOSA A 25, 2693–2703 (2008).
Flatau, P. J. & Draine, B. T. Fast near field calculations in the discrete dipole approximation for regular rectilinear grids. Opt. Express 20, 1247–1252 (2012).
Johnson, P. B. & Christy, R. W. Optical constants of the noble metals. Phys. Rev. B 6, 4370–4379 (1972).
Ahrens, J., Geveci, B. & Law, C. [Paraview: An end-user tool for large data visualization]. Visualization Handbook [Hansen, C. D. & Johnson, C. R. (ed.)] [717–]. (732Elsevier Inc., Burlington, 2005).
Acknowledgements
We thank the Department of Science and Technology, India for constantly supporting our research on nanomaterials. K.C. and T.P. thank James Beach, Director, Technology Department, CytoViva, Inc., USA for discussions. K.C. thanks Prashant Kumar, Department of Biotechnology, Indian Institute of Technology Madras, India for discussions on designing the cell culture experiments.
Author information
Authors and Affiliations
Contributions
K.C. and T.P. designed the experiments. K.C. performed experiments. K.C. and T.P. analyzed the data and wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supporting Information
Supplementary Information
Supporting Video S1
Supplementary Information
Supporting Video S2
Supplementary Information
Supporting Video S3
Supplementary Information
Supporting Video S4
Supplementary Information
Supporting Video S5
Supplementary Information
Supporting Video S6
Supplementary Information
Supporting Video S7
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Chaudhari, K., Pradeep, T. Spatiotemporal mapping of three dimensional rotational dynamics of single ultrasmall gold nanorods. Sci Rep 4, 5948 (2014). https://doi.org/10.1038/srep05948
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05948
This article is cited by
-
IL-6-induced response of human osteoblasts from patients with rheumatoid arthritis after inhibition of the signaling pathway
Clinical and Experimental Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.