Abstract
Dust storms include particulate matter that is transported over land and sea with biota that could impact downwind ecosystems. In addition to the physico-chemical compositions, organismal diversities of dust from two storm events in southern Israel, December 2012 (Ev12) and January 2013 (Ev13), were determined by pyro-sequencing using primers universal to 16S and 18S rRNA genes and compared. The bio-assemblages in the collected dust samples were affiliated with scores of different taxa. Distinct patterns of richness and diversity of the two events were influenced by the origins of the air masses: Ev13 was rich with reads affiliated to Betaproteobacteria and Embryophyta, consistent with a European origin. Ev12, originated in north-Africa, contained significantly more of the Actinobacteria and fungi, without conifers. The abundance of bacterial and eukaryotic reads demonstrates dissemination of biological material in dust that may impose health hazards of pathogens and allergens and influence vegetation migration throughout the world.
Similar content being viewed by others
Introduction
Dust storms, considered major contributors to global aerosols1, transport desert soils to the atmosphere and substantially impact the global environment. Estimates of global dust emissions from soils to the atmosphere vary between 1 and 3 pg per year2. Climatic variations affect the land surfaces of aeolian (wind-driven) systems3 and droughts have significantly contributed to increased dust emissions4,5. The contribution of anthropogenic soils, e.g. agricultural fields and industrial yards, are of increasing concern6. Up to 50% of the total atmospheric dust originates from disturbed soils7, which may contain different particle compositions from that of natural dust. The windblown dust can travel tens of thousands of kilometers before being deposited1, depending on the particle characteristics (size, chemistry) and the air-mass properties (e.g., velocity, density, height)8. The size and chemical compositions vary in space and time and may determine the potential impact on air quality and human health9,10.
Airborne dust particles contain different microorganisms that are exceedingly mobile in space and time11,12. For example, the number of culturable microorganisms in the US Virgin Islands increased during African dust-storm events by about 8 fold, from 0.013 L−1 of air under clear atmospheric conditions to 0.10513,14. Dust storms originating in the Saharan desert heavily affect the south-eastern Mediterranean basin, mostly during the winter and spring15; qualitative and quantitative analyses of dust-associated fungal communities revealed distinct pattern of distribution in the atmosphere of Haifa (Israel)16,17.
Airborne dust contains a variety of chemicals and microbial agents such as bacteria, fungi and viruses where some of them are pathogenic and pose a risk to the ecosystem and human health as the clouds traverse regions11,12,18. The composition of microorganisms is still not well-defined and taxonomic studies of organisms' diversity in the outdoor air have just started to emerge. In recent years, conventional molecular approaches have widely been used to study the diversity and community composition of prokaryotes and eukaryotes in air and outdoor dusts17,19,20,21,22,23,24, but the methods exploited are incomplete. Clone libraries, Restriction Fragment Length Polymorphism, Denaturative Gradient Gel Electrophoresis, Ribosomal Intergenic Spacer Analysis and PCR-single Strand Conformation Polymorphism are usually restricted to less than 500 sequences or patterns. These approaches, albeit providing general information on the structure of explored communities, are therefore not sufficient for meaningful comparisons.
In this study, the high throughput sequencing method (pyrosequencing) was used to explore, the structure of the prokaryotic and eukaryotic communities in airborne samples of dust collected in an urban environment in the Negev (southern Israel) following two distinct desert storm events.
Results and Discussion
The arena
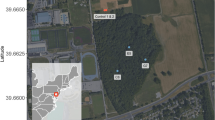
Samples collected in an urban environment in the Negev (southern Israel) were used as a case study. The Negev is located at the margin of the natural dust sources and is frequently subjected to such storms with common duration of several hours to one-day25,26. Recent studies relied mainly on satellite images (MODIS) and the HYSPLIT model of air mass backward trajectories to indicate the geographic source of the observed bio samples1,27,28,29,30. Satellite images of the studied region during the dust storm events (each lasted less than a day), the air mass trajectories and wind directions at the level of the measurement point (consistent with the HYSPLIT trajectories) are displayed in Fig. 1. Both storms were associated with a synoptic system of the (Cyprus) cold low-pressure that typically moves eastward over the Mediterranean Sea27. The trajectory of the December air mass (Ev12) extended over North Africa. Due to a deeper cold low in January 2013 (996 mb) than in December 2012 (1004 mb) the origin of the January air mass (Ev13) was attributed to South Europe (Fig. 1).
Satellite images (MODIS) of the studied region during the dust storms along with air mass transport at different heights above ground level (AGL) derived from Backward Trajectories model NOAA/ARL HYSPLIT-4 (credit to: www.noaa.gov) and equivalent wind directions at 10 m above surface level from the HCMR POSEIDON System (credit to: www.poseidon.hcmr.gr).
Red arrowheads indicate the sampling site. Right hand side panels: daily recorded averages of major meteorological variables and pollutants. AT – air temperature; RH – relative humidity; WS – wind speed; PM – particulate matter; ST – settled dust.
Physical and chemical properties
As is generally accepted, stronger winds and lower temperatures associated with lower pressure were observed here too (Fig. 1): the levels of major air gas pollutants were not affected by the dust events and remained relatively low as in non-dusty days in the studied region, but those of PM10 (particulate matter ≤ 10 μm in diameter) in both storms were about 20-fold higher than the background value in the area at non-dusty days (42 μg m−3), as also observed in other strong dust storms in the Negev during the last decade10. High levels of PM during dust storm events raise air pollution above the standard values of air quality10,31. In arid environments, hourly PM concentrations during dust storms can reach even 10 mg m−332.
The particle size distributions of the dust were typically bi-modal for both samples with a cutoff at about 25 μm (Fig. 2). The January sample (Ev13) included higher content of finer particles (45.1%) than that (34.0%) of December's (Ev12) and smaller sizes of the coarser population (peaks at 50 and 60 μm respectively). The possible sources of the finer particles, which can be carried longer distances, are North-African and South-European soils (see mass trajectories and compatible wind directions 10 m above the surface in Fig. 1), whereas the possible proxy source is the Negev soils located close to Be'er Sheva. In both Ev12 and Ev13 samples, there were relatively high (> 22%) frequencies of the class weight of respirable particles (< 10 μm) (Fig. 2) that affect human health9.
Typical elements were found in both dust samples (Table S1), the most common of which were Si, as in mineral soils, silt and sand fractions (> 50 μm)33 and Ca originating in sedimentary environments (such as dry lakes), calcareous soils and rocks in the arid land of the Mediterranean basin. The higher percentage of Si in Ev12 (Table S1) may indicate a longer terrestrial transport of the dust (Fig. 1). No significant differences in the contents of Al, Fe, K, Mg and Na were found between the dust samples, but some differences were found in the contents of minor elements: Co was lacking in Ev12; Cu, Os, Ni and Ga were not observed in Ev13.
Richness and diversity of organisms in storm dust
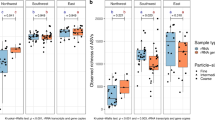
The four libraries represent true diversity reasonably well: the average Good's coverage of the reads in both samples (at 97% cut-off) was 94.5% for eukaryotes and 86% for bacteria (Table 1). On the other hand, the relatively low number of observed Operational Taxonomic Units (OTUs; Table 1 and Fig. S1), reaching 46% and 58% saturation (Sobs/SChao; Table 1) for eukaryotes and bacteria respectively, suggests an under-sampling of the dust; diversity may however be biased by tendency of Chao index to overestimate species richness34. The bacterial diversity in both samples was significantly higher than the eukaryotic and the total richness of the dust sample Ev12 was higher than of Ev13 (Table 1).
Venn diagrams (Fig. S2) demonstrate a higher number of bacterial and eukaryotic OTUs that were unique in both dust samples, implying different origins of the biota. The bacterial samples shared 200 OTUs (Fig. S2A), representing the majority of the reads, 54% and 62% for Ev12Bac and Ev13Bac respectively. Most of the unique 1,014 OTUs of Ev12Bac (Table S2) are singletons (i.e., OTU containing a single sequence), implying a relatively low coverage (83%) and high values of richness and diversity of this sample. PCR bias associated with initial amplicon generation may impose distortions in the observed community structure, particularly over-estimating rare taxa. Detection limit of the latter may be affected by the approach employed in quality control as well, but pyrosequencing itself appears not to impose significant bias of overall community structure estimates35. Eukaryotic samples shared only 34 OTUs (Fig. S2B, Tables S3 and S4), which also represented the majority of the sequences, 66% and 47% for Ev12Euk and Ev13Euk. Higher diversity and richness was also observed in Ev12Euk than in Ev13Euk (Table 1).
Prokaryotic sequences (Fig. 3A)
Reads of 16S rRNA genes that were retrieved from both dust storm samples belong to the following phyla: Acidobacteria, Actinobacteria, Bacteroidetes, Deinococcus-Thermus, Firmicutes, Proteobacteria, Cyanobacteria, in addition to unclassified bacteria. Reads of Gemmatimonadetes and Chloroflexi were found only in Ev12Bac and Verrucomicrobia, only in Ev13Bac. On a class level, the distributions were similar, with the most dominant Alphaproteobacteria (44.0% and 46.7% for Ev12Bac and Ev13Bac respectively), Actinobacteria (23.9% and 17.5%) and Betaproteobacteria (4.0% and 15.5%).
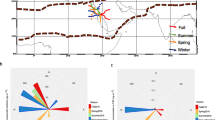
Distributions of rRNA genes reads, retrieved from the dust samples Ev12 and Ev13.
(a) Bacterial 16S reads at the class level. Affiliation of classes to different phyla are: 1–4 Acidobacteria; 5 – Actinobacteria; 6–7 – Armatimonadetes; 8–10 – Bacteroidetes; 11 – Chloroflexi; 12 – Deinococcus-Thermus; 13–15 – Firmecutes; 16 – Gemmatimonadetes; 17–21 – Proteobacteria; 22 – Verrucomicrobia; 23 – unclassified. (b) Eukaryotic 18S reads at the kingdom-class levels. Affiliation of taxonomic categories to highest taxonomic ranks are: 1–16 – Fungi kingdom (1–7 – Ascomycota phylum; 8–14 – Basidiomycota phylum; 15 – unclassified Dikarya superphylum; 16 – Basal fungal lineages phylum); 17–19 – Metazoa kingdom (17–18 – Arthropoda phylum; 19 – unclassified Metazoa); 20–21 – Alveolata superphylum (Ciliophora phylum); 22–28 – Viridiplantae phylum; 29 – unclassified eukaryota phylum.
Actinobacteria
This one of the dominant bacterial phylum/class is made up of gram-positive bacteria that are widely distributed in both terrestrial and aquatic ecosystems. Some of the Actinobacteria are spore forming which range from motile zoospores to specialized propagules which resist desiccation and mild heat. Most of the sequences are affiliated within the order Actinomycetales (95.7% and 99.3% for Ev12Bac and Ev13Bac respectively). The Geodermatophilaceae (22.9% and 15.2%), Micrococcaceae (22.9% and 42.3%) and unclassified-actinomycetales (27.6% and 23.5%) are the dominated families. The Actinobacteria reads belong to 3 major genera, Kocuria, Arthrobacter and Blastococcus, the species of which are found in various environments such as plants, soils, sediments and rhizomes. Among all bacterial reads, 4.7% in Ev12Bac and 20.5% in Ev13Bac displayed variable similarity (84–100%) to the Kocuria species, some with 99% homology to K. rosea (KC689298, FJ745378). Several groups are using metagenomic framework to study airborne microbial communities36,37. During a severe smog event over Beijing, for example, Cao et al.36 found that the most abundant bacterial phylum, order and species were respectively Actinobacteria, Actinomycetales and Geodermatophilus obscurus, Modestobacter marinus, Blastococcus saxobsidens, Kocuria rhizophila and Micrococcus luteus.
Kocuria spp. belonging to Micrococcaceae are considered as non-pathogenic commensals that colonize the oropharynx, skin and mucosa, but K. rosea is an opportunistic pathogen in immune-compromised patients38 and K. kristinae is associated with acute cholecystitis39. In Ev12Bac and Ev13Bac, 11.4% and 14% respectively are closely related to Arthrobacter spp. (e.g., HF585203, JX164047, JN680244, AJ576068, AB522428), which are widely distributed in nature, particularly soil. Their overall pathogenic potential is rather low, though some Arthrobacter stains were isolated from clinical specimens40. Most of the 10.9% and 7.6% of the reads from both samples respectively, affiliated with Blastococcus spp., are identical to B. saxobsidens (EU977834) and B. aggregatus (FR865889), found on rock surfaces in the Mediterranean basin41. Very few sequences (11 and 7 respectively) were identical to those of the genus Microbacterium found in soil and sewage, some of which have been recognized as pathogens in humans, causing e.g., endophthalmitis42.
Alphaproteobacteria
This class is a group of gram-negative bacteria that comprise most phototrophic genera, symbionts of plants and animals and a group of pathogens. Orders Rhodobacterales (28.9% and 59.5%) and Rhizobiales (44.6% and 17.1%), are among the largest revealed in both Ev12Bac and Ev13Bac respectively; the former are dominant and ubiquitous primary surface colonizers in temperate coastal waters of the world43 and the latter include nitrogen fixing symbionts and pathogenic to animals and plants44. Alphaproteobacteria in both samples showed similar distribution on a family and genus levels and was dominated by the following genera: Paracoccus (5.3% and 15.8%), Rubellimicrobium (15.5% and 24.5%) and Skermanella (6.7% and 9.3%). Some of the ubiquitous spp. of the genus Brevundimonas are considered to be opportunistic pathogens in immune-compromised hosts45, e.g., B. vesicularis and B. diminuta are frequently isolated species in human infections. Only 16 and 14 reads here were identical to B. vesicularis (KC494336, GU430201, HM755555).
Betaproteobacteria
This is the 3rd most dominant class, with 89% and 83% of the reads for Ev12Bac and Ev13Bac respectively, belonging to the order Burkholderiales that includes several pathogenic genera e.g., Burkholderia and Bordetella, but none of these reads belongs to them. Among the reads of this order, 43.6% and 62.7% are identical to Massilia yuzhufengensis (JQ409016) isolated from ice drilled in Yuzhufeng Glacier, Tibetan Plateau, China46.
Unclassified bacteria
A large portion of these reads in Ev12Bac and Ev13Bac (19.1% and 8.6% respectively) were revealed by the RDP database; when analyzed against NCBI GenBank, 56.5% and 11.9% of these were identical to the cyanobacterial species Oscillatoria nigroviridis PCC 7112 (CP003614).
Viability of pathogenic microorganisms
Viability is a meaningful parameter in developing predictive models for disease dispersal29. An average of about 70% viable bacteria has recently been estimated by fluorescent staining in anticyclone air28, but the majority (>99%) of microorganisms from the environment resist cultivation in the laboratory36,47, hence culture dependent methods cannot be used to assess viability of the pathogens in dust samples.
Eukaryotic sequences (Fig. 3B)
Dothideomycetes
This is the largest and most diverse class of ascomycetes that includes several plant pathogens (e.g., Phaeosphaeria nodorum, Venturia inaequalis). Ev12Euk and Ev13Euk contained 13.5% and 3.15% such reads respectively, most of them (8.9% and 1.5%) are highly similar (99–100%) to Cladosporium spp. (e.g. JX273066, JN974018, JN546118), some of which cause infections of the skin and toenails, sinusitis and pulmonary infections. Their airborne spores are significant allergens48 and in large amounts they can severely affect asthmatics49. Dominant sequences retrieved from indoor dust of urban area in central Finland were identical to C. cladosporioides and C. herbarum23. The rest of the Dothideomycetes reads (4.3% and 1.3%) were identical to Alternaria spp. (e.g., KC584600, KC584596, KC584599), major plant pathogens and common humans allergens48, causing hay fever or hypersensitivity reactions that sometimes lead to asthma49. Many health disorders are caused by these fungi, which grow on skin, mucous membranes, the eyeballs and the respiratory tract50. Alternaria and Epicoccum spp. (producing multicellular dictyospores > 10 μm) are abundant allergenic fungi, mostly in the larger particle-size ranges (da > 44.7 μm), whereas Cladosporium is an abundant allergenic fungal genus, distributed evenly across all the particle-sizes ranges51. Viable microbial populations, including presumptive plant pathogens Alternaria infectoria and Chaetomium globosum, have recently been detected in Asian air samples even after traveling 10 days across the Pacific Ocean in the free troposphere, information that has significant implications for epidemiology29.
Tremellomycetes
This fugal class (Agaricomycotina, Basidiomycota) is a dimorphic, nutritionally heterogeneous group comprising of saprotrophs, animal parasites, severe human pathogens and fungicolous species52. Ev12Euk and Ev13Euk contain 12% and 3.3% of such reads that are highly similar to Cryptococcus spp. and Filobasidium spp.; about 60% of them are identical to C. albidus (HQ231895) that occasionally causes moderate-to-severe diseases, specifically meningitis, in patients with compromised immunity53.
Mitosporic Ascomycota
These fungi comprise a heterogeneous group and represent more than half of Ascomycota lacking a sexual state; many pathogenic fungi in plants and mammals, including humans, belong to this group54. Ev12Euk and Ev13Euk contain 3.9% and 0.5% reads respectively belonging to this group.
Embryophyta
This is the most familiar subkingdom of green plants (Viridiplantae), informally called land plants, excluding green algae55 and was represented by 10.4% and 79.2% in Ev12Euk and Ev13Euk respectively. This high percent of plant reads in the latter is consistent with southern Europe as the origin of the air mass in January 2013. Among them, 3.6% and 2.7% in Ev12Euk and Ev13Euk respectively displayed 98–100% homology to leafy trees from the Morus spp. (GU476477 and L24398), Moringa oleifera (U42786), Plocosperma buxifolium (HQ384684), Metteniusa tessmanniana (AM421127) and Olea europaea (L49289). Additional 3% of Embryophyta reads in each Ev12Euk and Ev13Euk are identical to Bryum pseudotriquetrum (KC291525), Blindia acuta (AF023681) and Pottia truncata (X95935). Among the Embryophyta reads, 2.3% and 40.4% in Ev12Euk and Ev13Euk respectively are highly similar (98–99%) to Cratylia sp. (JX158808) and chickpea Cicer arietinum (AHII01138308) that belong to Fabaceae commonly known as legume, the 3rd-largest land plant family and economically important56. Dust events that transport pollen long-distances introduce vegetation changes and are prone to errors in studies (paleoecology) that interpret past local vegetation based on presence of pollen11.
Sequences affiliated with class Pinopsida (mostly conifers) were the 2nd largest group (25.4%) of the Embryophyta in Ev13Euk, none of them were found in Ev12Euk. This group included reads highly similar (99–100%) to e.g., Cupressus gigantea (EF053166), Tetraclinis articulata (EU161293), Juniperus morrisonicola (EF673744), Chamaecyparis pisifera (EF053165), Picea morrisonicola (AB026939), Pinus luchuensis (D38246). Finally, flowers and grass also appeared only in Ev13Euk as 2.8% of the Embryophyta reads that are similar (97–100%) to Arabidopsis thaliana (X16077), Arctium lappa (JF703098), Tagetes sp. Nickrent 3061 (U42501), Sinapis alba (X17062), Lolium multiflorum (AY846367), Festuca rubra (AF168844) and other species. Air masses originating in southern Europe do not usually harbor particles but include primary biological aerosol consisting of viruses, bacteria, fungal spores and plant pollen. The January dust storm here may have collected the particles in the Sinai desert on its way to the Negev.
Alveolata
This superphylum is a monophyletic group of primarily single-celled eukaryotes that have adopted extremely diverse modes of nutrition such as predation, photoautotrophy and intracellular parasitism. Most alveolates fall into 3 main subgroups: ciliates, dinoflagellates and apicomplexans. Ciliates are one of the most important groups of protists common to water tarns and soils and have many ecto- and endosymbiotic members, as well as some obligate and opportunistic parasites. Both dust samples (Ev12Euk and Ev13Euk) encompass 1.7% and 0.1% respectively of such reads; most reads from the former are identical to Halteria spp. with typical size of 20–50 μm and predominantly found in fresh water habitats.
Arthropoda
Members of this phylum are exoskeleton-containing invertebrates with a segmented body and jointed appendages and include insects, arachnids, crustaceans and myriapods. Ev12Euk contained 2.6% reads, most of which are affiliated with unclassified Metazoa/Animalia (with similarity of ≤93% in the BLAST database) and some belong to infraclass Neoptera—a group that includes most of the winged insects. Ev13Euk included only 0.26% such sequences.
Unclassified eukarya
According to SILVA database classification, reads from Ev12Euk and Ev13Euk include respectively 47.5% and 1.8% unclassified eukaryotes; they were therefore further analyzed using the BlastN program: about half (23%) of the Ev12Euk reads were identical to Rhizophlyctis rosea (NG017175), a soil cellulose-decomposing zoospores-producing fungus57. Additional 4.5% were identical to other ubiquitous zoospore-producing soil-dwelling fungal species belonging to Gaertneriomyces, Spizellomyces, Powellomyces and other genera (e.g., JN020240, GU568157, JN940943, FJ827646, AY349038, FJ827660, HQ901755, HQ901759). Additional 7.3% of Ev12Euk unclassified reads were identical to sequences of nematodes such as Aphelenchus avenae (AY284640 and AB731165), which is mycophagous and capable of withstanding droughts58. No nematodes were observed in Ev13Euk. The remaining (12.7%) of unclassified eukaryotic reads from Ev12Euk appeared as singletons with low similarity to known sequences. Unclassified Ev13Euk reads include only 69 sequences, 9 of which showed high similarity (97%) to an uncultured ciliate clone QD09 (HQ909037).
Concluding remarks
The results demonstrate that the diversity of organisms in airborne dust is higher than previously reported and may rival that of the other (terrestrial or aquatic) environments.11,12 Dust of Ev13, of south European origin, was rich with reads affiliated to Betaproteobacteria and to Embryophyta (land plants) in general, particularly conifers. On the other hand, dust of Ev12 (of north African origin) contained significantly more Actinobacteria, fungi, unclassified bacteria and eukaryotic sequences. Reads affiliated to allergenic and pathogenic species existed in both airborne dust samples. Intercontinental aerobiology studies will aid in developing predictive models for disease dispersal29. Despite its limitations47, culturing data may be valuable by knowing what species remain viable after long distance atmospheric transport30.
This study will likely affect the methodology to analyze climatic factors, soil sources, levels of particulate matter and air biology associated with dust storms and the assessment of the possible impacts on the ecosystems and hazards to public health.
Methods
Dust sample collection and physico-chemical measurements
The samples of outdoor dust were collected in Be'er Sheva (Israel) on a building roof within the campus of Ben-Gurion University of the Negev, during storm events that occurred in the “dust season” (October - May): on December 20, 2012 (Ev12) and on January 7, 2013 (Ev13). Both events were typically strong storms in the studied area. The spatial extent of each event was recorded by satellite images (MODIS Level 1 and VIIRS Level 1) (Fig. 1). The air mass transport in the region was detected through Backward Trajectories model (NOAA/ARL HYSPLIT-4)59 prior and during the events. The wind directions at 10 m level were retrieved by the POSEIDON System. The following major pollutants and meteorological variables were recorded: NO2 (ppb), NO (ppb), CO (ppm), SO2 (ppb), O3 (ppb), wind speed (m s−1), air temperature (°C) and relative humidity (%). Atmospheric concentrations (µg m−3) of PM10 and PM2.5 and total settled dust (g m−2) were recorded simultaneously during the storms in a monitoring system nearby the dust samplers.
The sterilized settling-dust collectors consist of a rectangular plastic tray (40 × 25 × 10 cm) filled with layers of glass/quartz marbles (10 mm in diameter). Atmospheric dust particles that cross the tray aperture are trapped in the marble matrix due to the matrix cohesion and roughness along with reduced air velocity near the surface of the tray. At the end of each event the dust samples were moved into sterilized glass vials for size distribution and elemental analyses. Size distributions of particles over the range of 0.08 to 2000 μm were obtained by a high-resolution laser diffractometer (ANALYSETTE 22 MicroTec Plus). Each sample was dispersed by sonication (at 38 kHz) in a Na-hexametaphosphate solution (0.5%), then transferred to a fluid module of the instrument (containing deionized water) and subjected to 3 consecutive 1 min runs at a medium pump speed of 6 L min−1. The data were processed using the Mie scattering model (RI = 1.56, AC = 0.01) with an error < 5.0%. The MasControl software was employed to determine the following relevant parameters: mean size, median, modes in multiple modal distributions, sorting values, size fraction weights.
Elemental analyses were performed by X-Ray Fluorescence (XRF) method using XRF Spectrometer, Panalytical Co., model Axios [WDXRF (wavelength dispersive), 1 kW]. Omnian software was used for quantitative analysis.
DNA extraction, PCR amplification and pyrosequencing
Total DNA was extracted from 0.5 g of the 2 dust samples (Ev12 and Ev13) using a Power Soil Isolation Kit (MoBio Laboratories, Carlsbad, CA) with the bead-beating protocol supplied by the manufacturer and quality and concentration were assessed using a NanoDrop ND 1000 spectrophotometer (NanoDrop Technologies, Wilmington, DE).
Low concentration (~5 ng μl−1) of the total genomic DNA of each sample was amplified using a Biometra TGradient thermocycler (Biometra, Gottingen, Germany) and 2 pairs of primers: the universal bacterial primers 8F, AGAGTTTGATYMTGGCTCAG and 907R, CCGTCAATTCMTTTGAGTTT to amplify a ~900 bp region of the 16S rRNA genes and the universal eukaryotic primers NSF370/18, AGGGYTCGAYYCCGGAGA and NSR1787/18, CYGCAGGTTCACCTACRG targeting 18S rRNA genes amplifying the ~1400 bp. The reaction mixture included 12.5 μl ReddyMix (ABgene, Surrey, UK), 1 µl of 10 μM concentrations of each primer (forward and reverse), 1 µl of 25 mM bovine serum albumin, 1–3 µl of the sample genomic DNA (10–40 ng µl−1) and water for a total volume to 25 µl. An initial denaturation hot start of 4 min at 95°C was followed by 30 cycles of the following incubation pattern: 94°C for 30 s, 54°C for 30 s and 72°C for 1–1.5 min.
The amplicons were submitted to Molecular Research laboratory (MR DNA, Shallowater, TX) for PCR optimization and pyrosequencing analyses utilized Roche 454 (454 Life Sciences, Branford, CT). The company's primer 27Fmod AGRGTTTGATCMTGGCTCAG was used to obtain bacterial reads of the hypervariable regions V1-V2 from the dust samples (Ev12 and Ev13), designated Ev12Bac and Ev13Bac and the primer Euk528F CCGCGGTAATTCCAGCTC was used for the V4 eukaryotic reads designated Ev12Euk and Ev13Euk.
Analysis of organismic diversity
In total, 15,019 bacterial and 15,424 eukaryotic sequence reads were recovered and subjected to MOTHUR software version 1.9.160, leaving 4020 bacterial reads for each sample, Ev12Bac and Ev13Bac and 3779 eukaryotic reads for each Ev12Euk and Ev13Euk, for further considerations.
The sequences were trimmed using MOTHUR software for removal of primers, barcodes, ambiguous nucleotides, long homo-polymers, reads below a minimum quality score of 25 and sequences shorter than 150 bases. Sequences were aligned, checked for chimeras, filtered and classified using the Ribosomal Database Project II website (RDP Release 10) for bacteria (Table S2) and SILVA reference files for eukarya (Tables S3 and S4). Reads classified as Chloroplast, Mitochondria or “unknown” were also removed. The sub.sample function in MOTHUR, used to randomly select equally-sized reads from each library, further reduced data sets for subsequent analyses. The pyrosequencing population data were also analyzed by multiple sequence alignments using the dist.seqs function60.
MOTHUR operational taxonomic units (OTUs) analyses were conducted (by a 0.03 distance level) to present α-diversity: Chao1 richness estimators, the inverse Simpson diversity index and rarefaction curves at 0.03 distances, which were plotted on a line chart using Microsoft Excel. PC-ORD 6 software (MJM Software Design, OR) was used to calculate beta-diversity.
Raw sequencing data was deposited in the MG-RAST (metagenomics.anl.gov) archive under accession numbers: 4534886.3 for Ev12Bac; 4534887.3 for Ev12Euk; 4534888.3 for Ev13Bac; 4534889.3 for Ev13Euk.
References
Yu, H. et al. Aerosols from overseas rival domestic emissions over North America. Science 337, 566–569 (2012).
Goudie, A. S. Dust storms: recent developments. J. Environ. Manag. 90, 89–94 (2009).
Katra, I. & Lancaster, N. Surface-sediment dynamics in a dust source from spaceborne multispectral thermal infrared data. Remote Sens. Environ. 112, 3212–3221 (2008).
Wang, X., Zhou, Z. & Dong, Z. Control of dust emissions by geomorphic conditions, wind environments and land use in northern China: An examination based on dust storm frequency from 1960 to 2003. Geomorphology 81, 292–308 (2006).
Jilbert, T. et al. Climate-controlled multidecadal variability in North African dust transport to the Mediterranean. Geology 38, 19–22 (2010).
Mulitza, S. et al. Increase in African dust flux at the onset of commercial agriculture in the Sahel region. Nature 466, 226–228 (2010).
Tegen, I., Werner, M., Harrison, S. P. & Kohfeld, K. E. Relative importance of climate and land use in determining present and futre global soil dust emission. Geoph Res Lett 31, L05105 (2004).
Kok, J. F. Does the size distribution of mineral dust aerosols depend on the wind speed at emission? Atmos. Chem. Phys. 11, 10149–10156 (2011).
Vodonos, A. et al. The impact of desert dust exposure on hospitalization due to the exacerbation of chronic obstructive pulmonary disease. Air Qual. Atmos. Health 10.1007/s11869-014-0253-z (2014).
Krasnov, H., Katra, I., Koutrakis, P. & Friger, M. D. Contribution of dust storms to PM10 levels in an urban arid environment. J. Air Waste Manag. Assoc. 64, 89–94 (2014).
Kellogg, C. A. & Griffin, D. W. Aerobiology and the global transport of desert dust. Trends Ecol. Evol. 21, 638–644 (2006).
Polymenakou, P. N. Atmosphere: A source of pathogenic or beneficial microbes? Atmosphere 3, 87–102 (2012).
Griffin, D. W., Garrison, V., Herman, J. R. & Shinn, E. A. African dust in the Caribbean atmosphere: microbiology and public health. Aerobiologia 17, 203–213 (2001).
Griffin, D. W. et al. Atmospheric microbiology in the northern Caribbean during African dust events. Aerobiologia 19, 143–157 (2003).
Israelevich, P. L., Levin, Z., Joseph, J. H. & Ganor, E. Desert aerosol transport in the Mediterranean region as inferred from the TOMS aerosol index. J. Geophs. Res. 107, (D21) 4572 (2002).
Schlesinger, P., Mamane, Y. & Grishkan, I. Transport of microorganisms to Israel by Saharan dust clouds. Aerobiologia 22, 259–273 (2006).
Grishkan, I., Schlesinger, P. & Mamane, Y. Influence of dust storms on concentration and content of fungi in the atmosphere of Haifa, Israel. Aerobiologia 28, 557–564 (2012).
Griffin, D. W. Atmospheric movement of microorganisms in clouds of desert dust and implications for human health. Clin. Microbiol. Rev. 20, 459–477 (2007).
Janke, T. et al. Analysis of the fungal flora in environmental dust samples by PCR–SSCP method. Curr. Microbiol. 67, 156–169 (2013).
Jeon, E. M. et al. Impact of Asian dust events on airborne bacterial community assessed by molecular analyses. Atmos. Environ. 45, 4313–4321 (2011).
Maron, P. A. et al. Assessing genetic structure and diversity of airborne bacterial communities by DNA fingerprinting and 16S rDNA clone library. Atmos. Environ. 39, 3687–3695 (2005).
Yamaguchi, N., Ichijo, T., Sakotani, A., Baba, T. & Nasu, M. Global dispersion of bacterial cells on Asian dust. Sci. Rep. 2, 525 (2012).
Pitkäranta, M. et al. Analysis of fungal flora in indoor dust by ribosomal DNA sequence analysis, quantitative PCR and culture. Appl. Environ. Microbiol. 74, 233–244 (2008).
Polymenakou, P. N., Mandalakis, M., Stephanou, E. G. & Tselepides, A. Particle size distribution of airborne microorganisms and pathogens during an intense African dust event in the eastern Mediterranean. Environ. Health. Persp. 116, 292–296 (2008).
Dayan, U., Heffter, J., Miller, J. & Gutman, G. Dust intrusion events into the mediterranean basin. J. Appl. Meteorol. 30, 1185–1199 (1991).
Ganor, E., Osetinsky, I., Stupp, A. & Alpert, P. Increasing trend of African dust, over 49 years, in the eastern Mediterranean. J. Geophys. Res. 115, D07201 (2010).
Creamean, J. M. et al. Dust and biological aerosols from the Sahara and Asia influence precipitation in the western US. Science 339, 1572–1578 (2013).
Murata, K. & Zhang, D. Transport of bacterial cells toward the Pacific in northern hemisphere westerly winds. Atmos. Environ. 87, 138–145 (2014).
Smith, D. J. et al. Free tropospheric transport of microorganisms from Asia to North America. Microb. Ecol. 64, 973–985 (2012).
Smith, D. J. et al. Intercontinental dispersal of bacteria and archaea by transpacific winds. Appl. Environ. Microbiol. 79, 1134–1139 (2013).
Mitsakou, C. et al. Saharan dust levels in Greece and received inhalation doses. Atmos. Chem. Phys. 8, 7181–7192 (2008).
Ganor, E., Stupp, A. & Alpert, P. A method to determine the effect of mineral dust aerosols on air quality. Atmos. Environ. 43, 5463–5468 (2009).
Kalderon-Asael, B., Erel, Y., Sandler, A. & Dayan, U. Mineralogical and chemical characterization of suspended atmospheric particles over the east mediterranean based on synoptic-scale circulation patterns. Atmos. Environ. 43, 3963–3970 (2009).
Toti, D. S., Coyle, F. A. & Miller, J. A. A structured inventory of Appalachian grass bald and heath bald spider assemblages and a test of species richness estimator performance. J. Arachnol, 28, 329–345 (2000).
Lee, C. K. et al. Groundtruthing next-gen sequencing for microbial ecology–biases and errors in community structure estimates from PCR amplicon pyrosequencing. PLoS One, 7, e44224 (2012).
Cao, C. et al. Inhalable microorganisms in Beijing's PM2.5 and PM10 pollutants during a severe smog event. Environ. Sci. Technol. 48, 1499–1505 (2014).
Yooseph, S. et al. A metagenomic framework for the study of airborne microbial communities. PLoS One 8, e81862 (2013).
Altuntas, F. et al. Catheter-related bacteremia due to Kocuria rosea in a patient undergoing peripheral blood stem cell transplantation. BMC Infect. Dis. 4, 62 (2004).
Ma, E. S. et al. Kocuria kristinae infection associated with acute cholecystitis. BMC Infect. Dis. 5, 60 (2005).
Funke, G. et al. Isolation of Arthrobacter spp. from clinical specimens and description of Arthrobacter cumminsii sp. nov. and Arthrobacter woluwensis sp. nov. J. Clin. Microbiol. 34, 2356–2363 (1996).
Urzi, C., Salamone, P., Schumann, P., Rohde, M. & Stackebrandt, E. Blastococcus saxobsidens sp. nov. and emended descriptions of the genus Blastococcus Ahrens and Moll 1970 and Blastococcus aggregatus Ahrens and Moll 1970. Int. J. Syst. Evol. Microbiol. 54, 253–259 (2004).
Funke, G., Haase, G., Schnitzler, N., Schrage, N. & Reinert, R. R. Endophthalmitis due to Microbacterium species: case report and review of Microbacterium infections. Clin. Infect. Dis. 24, 713–716 (1997).
Dang, H. Y., Li, T. G., Chen, M. N. & Huang, G. Q. Cross-ocean distribution of Rhodobacterales bacteria as primary surface colonizers in temperate coastal marine waters. Appl. Environ. Microbiol. 74, 52–60 (2008).
Carvalho, F. M., Souza, R. C., Barcellos, F. G., Hungria, M. & Vasconcelos, A. T. R. Genomic and evolutionary comparisons of diazotrophic and pathogenic bacteria of the order Rhizobiales. BMC Microbiol. 10, 37 (2010).
Chi, C. Y., Fung, C. P., Wong, W. W. & Liu, C. Y. Brevundimonas bacteremia: two case reports and literature review. Scand. J. Infect. Dis. 36, 59–61 (2004).
Shen, L. et al. Massilia yuzhufengensis sp. nov., isolated from an ice core. Int. J. Syst. Evol. Microbiol. 63, 1285–1290 (2013).
Kaeberlein, T., Lewis, K. & Epstein, S. S. Isolating “uncultivable” microorganisms in pure culture in a simulated natural environment. Science 296, 1127–1129 (2002).
Simon-Nobbe, B., Denk, U., Poll, V., Rid, R. & Breitenbach, M. The spectrum of fungal allergy. Int. Arch. Allergy Immunol. 145, 58–86 (2008).
Denning, D. W., O'Driscoll, B. R., Hogaboam, C. M., Bowyer, P. & Niven, R. M. The link between fungi and severe asthma: a summary of the evidence. Eur. Respir. J. 27, 615–626 (2006).
Singh, B. & Denning, D. W. Allergic bronchopulmonary mycosis due to Alternaria: case report and review. Med. Mycol. Case Rep. 1, 20–23 (2012).
Yamamoto, N. et al. Particle-size distributions and seasonal diversity of allergenic and pathogenic fungi in outdoor air. ISME J. 6, 1801–1811 (2012).
Cooper, C. R. [Yeasts pathogenic to humans]. The Yeasts: A Taxonomic Study [Kurtzman, C. P., Fell, J. W. & Boekhout, T. (eds)] [1339–1372] (Elsevier, London, 2011).
Endo, J. O., Klein, S. Z., Pirozzi, M., Pirozzi, C. & Hull, C. M. Generalized Cryptococcus albidus in an immunosuppressed patient with palmopustular psoriasis. Cutis 88, 129–132 (2011).
Taylor, J. W., Jacobson, D. J. & Fisher, M. C. The evolution of asexual fungi: Reproduction, speciation and classification. Annu. Rev. Phytopathol. 37, 197–246 (1999).
Niklas, K. J. & Kutschera, U. The evolution of the land plant life cycle. New Phytol. 185, 27–41 (2010).
Doyle, J. J. Phylogeny of the legume family: an approach to understanding the origins of nodulation. Annu. Rev. Ecol. Syst. 25, 325–349 (1994).
Chambers, T. C. & Willoughby, L. G. The fine structure of Rhizophlyctis rosea, a soil phycomycete. J. Roy. Micro. Soc. 83, 355–364 (1964).
Browne, J. A. et al. Dehydration-specific induction of hydrophilic protein genes in the anhydrobiotic nematode Aphelenchus avenae. Eukaryot. Cell 3, 966–975 (2004).
Draxler, R. R. & Rolph, G. D. HYSPLIT (HYbrid Single-Particle Lagrangian Integrated Trajectory), NOAA Air Resources Laboratory, http://ready.arl.noaa.gov/HYSPLIT.php(2014) (date of access: 14/01/2014).
Schloss, P. D. et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Acknowledgements
This investigation was partially supported by grants from the Israeli Environment and Health Fund (No. RGA1004) and the Israel Science Foundation (1100/11), both to I.K. and from the Ramat Hovav Council (to A.K.). We thank the reviewers for their helpful insight and comments that improved the manuscript.
Author information
Authors and Affiliations
Contributions
I.K., A.Z., E.B.D. and A.K. carried out experimental design; I.K. and H.K. carried out physical-chemical and transport measurements; E.B.D. performed DNA isolation and amplification; L.A. carried out 16S rRNA data analyses; E.B.D. and I.K. coordinated the project; all authors contributed to manuscript preparations, discussed the results and implications and commented on the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
SupplInfo
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Katra, I., Arotsker, L., Krasnov, H. et al. Richness and Diversity in Dust Stormborne Biomes at the Southeast Mediterranean. Sci Rep 4, 5265 (2014). https://doi.org/10.1038/srep05265
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep05265
This article is cited by
-
Aerial transport of bacteria by dust plumes in the Eastern Mediterranean revealed by complementary rRNA/rRNA-gene sequencing
Communications Earth & Environment (2023)
-
Bacterioplankton community assembly in migratory fish habitat: a case study of the southern East China Sea
Environmental Science and Pollution Research (2022)
-
Efficiency of bioaerosol samplers: a comparison study
Aerobiologia (2021)
-
Diversity and Sources of Airborne Eukaryotic Communities (AEC) in the Global Dust Belt over the Red Sea
Earth Systems and Environment (2021)
-
High genetic diversity and variability of microbial communities in near-surface atmosphere of Crete island, Greece
Aerobiologia (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.