Abstract
MicroRNAs (miRNAs) participate in diverse biological pathways and may act as oncogenes or tumor suppressors. Single nucleotide polymorphisms (SNPs) in miRNAs (MirSNPs) might promote carcinogenesis by affecting miRNA function and/or maturation; however, the association between MirSNPs reported and cancer risk remain inconsistent. Here, we investigated the association between nine common MirSNPs and cancer risk using data from large scale case-control studies. Eight precursor-miRNA (pre-miRNA) SNPs (rs2043556/miR-605, rs3746444/miR-499a/b, rs4919510/miR-608, rs2910164/miR-146a, rs11614913/miR-196a2, rs895819/miR-27a, rs2292832/miR-149, rs6505162/miR-423) and one primary-miRNA (pri-miRNA) SNP (rs1834306/miR-100) were analyzed in 16399 cases and 21779 controls from seven published studies in eight common cancers. With a novel statistic, Cross phenotype meta-analysis (CPMA) of the association of MirSNPs with multiple phenotypes indicated rs2910164 C (P = 1.11E-03), rs2043556 C (P = 0.0165), rs6505162 C (P = 2.05E-03) and rs895819 (P = 0.0284) were associated with a significant overall risk of cancer. In conclusion, MirSNPs might affect an individual's susceptibility to various types of cancer.
Similar content being viewed by others
Introduction
The worldwide cancer burden continues to increase; however, the precise mechanisms of carcinogenesis remain largely unknown. A number of investigators have demonstrated that genetic factors play a significant role in an individual's risk of cancer. MicroRNAs (miRNAs) are naturally occurring, small, noncoding, single-stranded RNA molecules that regulate gene expression by base pairing with the 3′ untranslated region of their target mRNAs, leading to mRNA cleavage or translational repression1. Numerous studies have demonstrated that miRNAs regulate a variety of biological processes, including cell proliferation, differentiation, apoptosis and development, thus dysregulation of these processes is closely associated with carcinogenesis2,3.
Recently, single nucleotide polymorphisms (SNPs) located in miRNAs, named as MirSNPs, have attracted increasing attention due to their possible involvement in the development of various types of cancer. Such MirSNPs may play functional roles through affecting the transcription of the primary target gene, altering pri-miRNA/pre-miRNA processing, or exerting effects on miRNA-mRNA interactions4. We performed a literature search and review of the association of common MirSNPs, including rs1834306, rs2043556, rs3746444, rs4919510, rs2910164, rs11614913, rs895819, rs2292832 and rs6505162, with the risk of cancer. However, the conclusions of the relevant studies were inconsistent, in part because of the heterogeneity of the types of cancer studied, the small sample sizes and the varied ethnicity of the patients. Therefore, there is an urgent need to further investigate the association of cancer-related MirSNPs with the risk of various types of cancer. Although the identification of cancer-related miRNAs based on gene association studies has become increasingly popular5, no study has yet investigated the association of cancer-related MirSNPs with the risk of various types of cancer based on an analysis of a large number of MirSNP association studies.
Therefore, we conducted a candidate-gene designed association study employing large numbers of cases and controls for eight kinds of cancer that commonly jeopardize human health (bladder cancer, breast cancer, esophageal squamous cell carcinoma (ESCC), gastric cancer, lung cancer, pancreatic cancer and renal cell carcinoma (RCC)) and analyzed these nine MirSNPs (either by direct genotyping or imputation) to further determine the association of these MirSNPs with the risk of developing cancer. Cross phenotype meta-analysis (CPMA) was performed to analyze the association of MirSNPs and overall cancer risk and specific cancer risk was further discussed.
Results
Patient characteristics
The risk of developing eight different types of cancer, including bladder cancer, breast cancer, lung cancer, pancreatic cancer, RCC, prostate cancer, ESCC and gastric cancer was assessed. The patients and controls in the gastric cancer and ESCC study were from Asian population, while the patients and controls in the six other cancer studies were from a Caucasian population.
Quantitative analysis
Primary analyses were conducted through unconditional logistic regression models for genotype trend effects (1 degree of freedom) and adjusted for eigenvectors, gender and cohort. The false discovery rate (FDR) method was considered to correct for multiple testing. Results revealed a significant association between rs2910164 C vs G and the risk of bladder cancer (OR = 1.12, 95% CI: 1.04–1.21, P = 2.06E-03(PFDR = 0.0297)) and gastric cancer (OR = 0.85, 95% CI: 0.77–0.93, P = 5.98E-04(PFDR = 0.0108)); rs2043556 C vs. T and the risk of bladder cancer (OR = 1.19, 95% CI: 1.10–1.28, P = 1.44E-05(PFDR = 5.18E-04)); rs6505162 C vs. A and the risk of bladder cancer (OR = 1.12, 95% CI: 1.068–1.18, P = 4.05E-05(PFDR = 9.72E-04)); rs895819 C vs. T and the risk of bladder cancer (OR = 1.19, 95% CI: 1.10–1.28, P = 6.70E-06(PFDR = 4.82E-04)). We further performed the above analyses 1000 times but randomly selected 70% cases and controls each time and results showed that the five MirSNPs mentioned above were consistently associated with specific type of cancer risk (P < 0.001).
CPMA analysis was performed to unveil the association of each MirSNP with the overall risk of cancer, which suggested that rs2910164 C (P = 1.11E-03), rs2043556 C (P = 0.0165), rs6505162 C (P = 2.05E-03) and rs895819 (P = 0.0284) involved with cancer occurrence (1Table 2). A meta-analysis using different effects model with inverse-variance weighting based on the heterogeneity existing in the results of the studies in each MirSNP was also provided (Figure 1).
Begg's test was used to investigate publication bias in the literature. The shapes of the funnel plots showed no obvious asymmetry and no statistical evidence of bias existed (Figure 2).
Begg's funnel plot with pseudo 95% confidence limits for publication bias of the MirSNPs in meta-analysis.
Begg's test offers no evidence of publication bias. (a) rs2292832 (P = 0.386); (b) rs2910164 (P = 0.536); (c) rs2043556 (P = 0.386); (d) rs4919510 (P = 1.000); (e) rs1834306 (P = 0.174); (f) rs11614913 (P = 0.536); (g) rs6505162 (P = 0.174); (h) rs895819 (P = 0.386); (i) rs3746444 (P = 1.000).
Discussion
Approximately 50% of all annotated human miRNA genes are located in fragile sites or areas of the genome that are frequently associated with cancer. SNPs, the most common type of genetic variation in the human genome, result in phenotypic differences6; such sequence variations in miRNA genes may potentially affect the processing of miRNAs, pri-miRNAs, pre-miRNAs and/or mature miRNAs, and/or target selection and may thus significantly affect an individual's risk of cancer7.
Here we evaluated the associations between nine common MirSNPs (rs1834306, rs2043556, rs3746444, rs4919510, rs2910164, rs11614913, rs895819, rs2292832 and rs6505162) and the susceptibility to cancer using data from seven published studies; each study investigated a single type of cancer, except for one study which investigated both gastric adenocarcinoma and ESCC. Therefore, this study was a large population-based and multi-cancer stratified investigation. We observed significant relations between the MirSNPs rs2910164, rs2043556, rs6505162 and the overall risk of developing cancer using FDR adjusted CPMA analysis. CPMA analysis adopts association P values and examine whether the observed P values diverge from the expected distribution of P values under the null hypothesis of no additional associations besides those already known. The CPMA analysis is especially well fitted to wide phenotypic surveys, resulting from its benefits from increased numbers of phenotypes8.
The rs2910164 G/C polymorphism of the miR-146a gene is situated in the stem structure opposite the mature miR-146 sequence and leads to a change from a G:U pair to C:U mismatch in the stem region of the miR-146a precursor. The G allele of the miR-146a precursor might influence the generation of mature miR-146a and impact on target mRNA binding9,10. Our study revealed an association between rs2910164 and the overall risk of cancer by CPMA, which is inconsistent with He et al using random effects meta-analysis11. Although random effects meta-analysis incorporates a moderate level of the effects of heterogeneity, it is not well suited for the cases in which the genetic variant produces the opposite effects on diverse phenotypes. For rs2910164, the results of the two different meta-analyses may be due to the opposite effects of the MirSNP in different types of cancer, thus the use of CPMA seems more reasonable10,11,12,13. It is of interest to learn that the amount of mature miR-146a from the C allele were 1.8-fold reduced, compared to the G allele in papillary thyroid carcinoma, while the miR-146a levels in the CC genotype were significantly increased compared with the GG genotype in gastric cancer13. The rs2910164 C allele was associated with a decreased risk of gastric cancer in the Asian population, a finding supported by Xu et al14. An increased risk of bladder cancer in the Caucasian population was observed in the rs2910164 C allele. However, a study performed by Wang et al. indicated a reduced risk of bladder cancer in the rs2910164 C allele in Asian population15. These results suggest that the rs2910164 polymorphism may have varying effects in different genetic backgrounds or patients with a different ethnicity, and/or during the pathogenesis of different types of cancer.
The rs6505162 SNP, located in the pre-miR-423, 12 base pairs 5′ of miR-423-5p offers an association with cancer development based on CPMA analysis. So far, most research on miR-423 has concentrated on expression analyses, where aberrant expression of both mature forms of the miRNA has been seen in cancer, as well as during cellular differentiation16,17,18,19. Studies have shown that pre-miRNA SNPs from miRNAs can affect the production of mature forms and the binding of nuclear factors related to miRNA processing20,21,22. We suppose that rs6505162 might affect the expression or processing of miR-423, therefore, studies evaluating the effect of this SNP in miRNA functionality are required. However, studies of the rs6505162 polymorphism on cancer risk have yielded inconsistent results23,24,25. The first of these studies was conducted in 2009 on ESCC in a population of 346 Caucasian ESCC patients and suggested the C allele of rs6505162 being significantly higher in cancer patients compared with controls23. A study performed in 2012 indicated that the C genotype of the rs6505162 SNP reduces the risk of breast cancer development, however, another study undertaken in 2009 suggested that the C genotype of rs6505162 offered an increased risk of developing both ovarian and breast cancer in Breast Cancer Associated 2 (BRCA2) mutation carriers26. Our research observed an increased risk of bladder cancer in the rs6505162 C allele using the Caucasian population, as to our knowledge, this is the first study to show a relation between this SNP and bladder cancer, thus needs further validation.
The allele C of rs2043556, located in miR-605, was marginally associated with a risk of developing cancer; this is the first study to associate this MirSNP with cancer development, which needs to be validated by more studies. Stratified analysis revealed that the miR-605 allele C was associated with an increased risk of developing bladder cancer in the Caucasian population. Recently, analysis of this SNP was conducted on gastrointestinal cancer among Asians and produced data similar to our own, with C allele being significantly lower in controls compared to cancer patients27. Researchers have found that miR-605 to be an element of the p53 network which forms a positive feedback loop in response to stress28, thus miR-605 may play a key role in carcinogenesis. It will make more sense if the association between SNP and miRNA expression have been investigated and might be an answer to the relation of SNP and cancer risk.
The allele C of the MirSNP rs895819, located in the terminal loop of the pre-miR-27a, was associated with increased risk of bladder cancer in the Caucasian population and it is the first study to address association between the MirSNP and bladder cancer. MiR-27a has been investigated in several types of cancer and comes into inconsistent results. MiR-27a functions as a tumor suppressor in ESCC and hepatocellular carcinoma, while serves as promoting factor in gastric tumorigenesis29,30. Therefore, we assume that miR-27a plays pleiotropic signaling roles in regulating tumorigenesis. The MirSNP rs895819 initially reported to relate with a reduced risk of familial breast cancer risk (P = 0.0215) in a Caucasian population31; however, no significant association of rs895819 with the risk of breast cancer was observed in Chinese population32. A previous study suggested no association between rs895819 and the risk of colorectal cancer in a Central-European Caucasian population, a population with an extremely high incidence of sporadic colorectal cancer33; this observation is supported by our results. Since the high probability of MirSNP rs895819 involved with carcinogenesis, these conflicting results may be due to the analysis of varying sample sizes and warrant further analysis of larger cohorts to clearly establish the impact of rs895819 on the risk of cancer.
The rs11614913 polymorphism of miR-196a2 has a significant impact on the expression of miR-196a2 and is associated with carcinogenesis in various types of cancer34,35. Previous, meta-analysis studies suggested a significant association between rs11614913 and the overall risk of cancer in the Asian population, which was inconsistent with our results11,36,37. Our study suggests the rs116114913 C allele might protect against lung cancer in the Caucasian population, but the significance was mitigated with P value 0.197 after FDR adjustment, while Tian et al found rs116114913 C allele associated with significantly increased risk of lung cancer in Chinese38, suggesting that the effect of the rs11614913 polymorphism may rely on the genetic background or ethnicity of the patients and/or the effects of the environment, in agreement with the reports of Chu et al.37 and Wang et al.36. The effect of rs11614913 on the risk of different types of cancer needs to be confirmed in additional studies.
No significant associations were observed for the rs1834306, rs4919510, rs2292832 and rs3746444 polymorphisms in terms of the overall risk of cancer or the risk of specific types of cancer.
Though miR-100 has been shown to suppress the expression of proteins in the insulin-like growth factor (IGF)/mammalian target of rapamycin (mTOR) signaling cascade in childhood adrenocortical tumors39 and clear cell ovarian cancer40, thus suppressing tumorigenesis, while act as a oncogene in acute myeloid leukemia41. Our results showed mir-100 polymorphism, located in the pri-miR-100 region had no relation with the risk of cancer. Rs4919510 lies within the mature miR-608 sequence and is located at the junction between the stem and canonical hairpin loop42. Rs4919510 G allele was observed to relate with increased risk in bladder cancer, gastric cancer and prostate cancer, however, P values were mitigated after FDR adjustment, which needs to be validated by further studies. Rs2292832, located in pre-miR-149, was previously reported to have no significant associations with the risk of evaluated in breast cancer43, lung cancer44 or gastrointestinal cancer45, in agreement with the results of this study. Although a sizeable number of studies have been performed to investigate the role of the miR-499 rs3746444 polymorphism in several types of cancer, including breast cancer43,46, lung cancer44, gastric cancer47 and bladder cancer48, these existing studies have yielded contradictory results. These discrepancies may be due to the study of different populations from different areas and variations in selection of the case groups; therefore, the effect of the miR-499 rs3746444 polymorphism needs to be investigated further.
One limitation of the present study that needed to be addressed is the multiple comparison problems resulting from the number of MirSNPs tested. Therefore the FDR method was used to correct for multiple testing.
Second, several MirSNPs were imputed rather than directly genotyped in this study. Although using imputed MirSNPs might lead to less accurate results, we ensured that only SNPs with high imputation confidence > 95% were included into further analysis.
Taken together, the findings of the present study have substantial scientific significance and may have implications in the clinical setting. Our results suggest that common MirSNPs may contribute to an individual's susceptibility to diverse types of cancer. Further functional characterization of MirSNPs and their influence on their target mRNAs may reveal the underlying mechanisms responsible for the associations between these polymorphisms and the etiology of cancer. Further prospective investigations of larger numbers of cases and controls are required in order to clarify the inconsistent associations between MirSNPs and the risk of cancer.
Methods
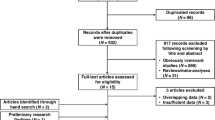
Identification of eligible studies
We evaluated the effect of nine MirSNPs on the risk of bladder cancer in 3527 cases and 5119 controls from the Maryland bladder cancer study (dbGAP number: phg0000132.v1) performed among the Caucasian population of the United States49; the risk of breast cancer in 1145 postmenopausal women of European ancestry with invasive breast cancer and 1142 controls from the Massachusetts breast cancer study (dbGAP number: phg000032.v1) performed in the United States50; the risk of lung cancer in 3782 cases and 3840 controls from the Maryland lung cancer study (dbGAP number: phg000124.v1) performed in the United States51; the risk of pancreatic cancer in 2452 affected individuals (cases) and 2461 unaffected controls from the Minnesota pancreatic cancer study (dbGAP number: phg000089.v1) performed in the United States52; the risk of prostate cancer in a nested case-control study (dbGAP number: phg000067.v1) including 659 cases and 1593 controls of European origin performed in the United States53; the risk of RCC in 1311 affected individuals and 3424 controls with a European background from the Maryland renal cell carcinoma study (dbGAP number: phg000123.v1) performed in the United States54; and the risk of gastric adenocarcinoma and ESCC in a study (dbGAP number: phg000128.v1) performed in the United States of individuals of Chinese ethnicity, including 1625 cases of gastric cancer, 1898 cases of ESCC and 2100 controls55.
This study is based on an in-silicon re-analyze of the human genotyping data downloaded form dbGAP(www.ncbi.nlm.nih.gov/gap). The data submitters have obtained the informed consent from each participant.
Selection of SNPs
We carried out a search of the PubMed and Embase databases for all relevant reports on the association of MirSNPs with the risk of cancer. The following candidate MirSNPs were selected for this study: miR-605 A/G (rs2043556), miR-499a/b A/G (rs3746444), miR-608 C/G (rs4919510), miR-146a G/C (rs2910164), miR-196a2 C/T (rs11614913), miRNA-27a T/C (rs895819), miR-149 C/T (rs2292832), miR-423 A/C (rs6505162) and miR-100 T/C (rs1834306), which are present in the pre-miRNA regions of miR-196a2, miR-146a, miR-499a/b, miR-423, miR-608, miRNA-27a, miR-149 and miR-605 and the pri-miRNA region of miR-100, respectively (Table 1).
Imputation of the MirSNPs
The SNPs not present in the original chip were imputed by the program IMPUTE2, using both HapMap (NCBI Build 36 (db126b)) CEU data and 1000 Genomes as a reference haplotype set. All SNPs showed high imputation confidence (>95%). Rs2292832, rs2043556 and rs11614913 were directly genotyped, the others were imputed.
Association testing and adjustment for covariates
All the association tests were performed by Plink v1.07 using additive logistic regression models. To account for potential population stratification or admixture in these samples, principal component analyses (PCA) was carried out using the EIGENSTRAT56. After adjustment for significant principal components (PCs) in each study based on leveling off of the PCA screen plot, there was no evidence for large scale inflation of the association test statistics by comparison of observed and expected distributions, ruling out the significance hidden population substructure. The principal component score for each individual was included as a covariate in all models along with gender and cohort in logistic regression models. Multivariate logistic regression was performed in R software package (http://www.r-project.org/). The FDR method was used to correct for multiple testing (FDR q < 0.05).
Resampling
To examine the robustness of the associations, we conduct a re-sampling analysis in accordance with Li et al.57. Using the association test mentioned above, P-values (Prandom) were obtained by performing the test 1,000 times but randomly selected 70% of population in corresponding study. Then we tested the null hypothesis, Prandom ≥ 0.05(Supplementary Table S1).
Statistical analysis
The associations of the nine MirSNPs with the risk of cancer were examined by performing meta-analysis using inverse-variance method. We examined the association of the MirSNPs with the overall risk of cancer as measured by odds ratios (ORs) and 95% confidence intervals (CIs). Moreover, stratified analyses were also performed by the type of cancer for each MirSNP. The heterogeneity of the cancer type between studies was evaluated using the Chi-square-based Q statistical test, with a heterogeneity (Ph) < 0.05 considered significant. A fixed-effect model using the Mantel–Haenszel method and a random-effects model using the DerSimonian and Laird method were used to pool the data according to the cancer types and individual MirSNPs. The random-effects model was used when heterogeneity existed in the results of the studies; otherwise the fixed-effect model was used.
Additionally, cross phenotype meta-analysis (CPMA) was performed to determine the associations of the MirSNPs with the overall cancer risk; P < 0.05 was considered significant after FDR adjustment. The CPMA statistic determines evidence for the hypothesis that single SNP has multiple phenotypic associations. The CPMA statistic is agnostic to the direction of effect in each disease. It has one degree of freedom as it measures a deviation in P value behavior instead of testing all possible combinations of diseases for association to each SNP and therefore provides high power to reject the null hypothesis58,59.
All statistical tests for the meta-analysis were performed with review manager version 5.2 (The Nordic Cochrane Centre, The Cochrane Collaboration, Copenhagen, Denmark). Begg's test was used to evaluate publication bias.
References
Lagos-Quintana, M., Rauhut, R., Lendeckel, W. & Tuschl, T. Identification of novel genes coding for small expressed RNAs. Science. 294, 853–8 (2001).
Ambros, V. MicroRNA pathways in flies and worms: growth, death, fat, stress and timing. Cell. 113, 673–6 (2003).
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature. 435, 834–8 (2005).
Ryan, B. M., Robles, A. I. & Harris, C. C. Genetic variation in microRNA networks: the implications for cancer research. Nat Rev Cancer. 10, 389–402 (2010).
Ma, Y. et al. Candidate microRNA biomarkers in human colorectalcancer: systematicreview profiling studies andexperimental validation. Int J Cancer. 130, 2077–87 (2012).
Hinds, D. A. et al. Whole-genome patterns of common DNA variation in three human populations. Science. 307, 1072–9 (2005).
Duan, R., Pak, C. & Jin, P. Single nucleotide polymorphism associated with mature miR-125a alters the processing of pri-miRNA. Hum Mol Genet. 16, 1124–31 (2007).
Solovieff, N., Cotsapas, C., Lee, P. H., Purcell, S. M. & Smoller, J. W. Pleiotropy in complex traits: challenges and strategies. Nat Rev Genet. 14, 483–95, 10.1038/nrg3461 (2013).
Xu, T. et al. A functional polymorphism in the miR-146a gene is associated with the risk for hepatocellular carcinoma. Carcinogenesis. 29, 2126–31 (2008).
Jazdzewski, K. et al. Common SNP in pre-miR-146a decreases mature miR expression and predisposes to papillary thyroid carcinoma. Proc Natl Acad Sci USA. 105, 7269–74 (2008).
He, B. et al. The association between four genetic variants in microRNAs (rs11614913, rs2910164, rs3746444, rs2292832) and cancer risk: evidence from published studies. PLoS One. 7, e49032 (2012).
Shen, J. et al. A functional polymorphism in the miR-146a gene and age of familial breast/ovarian cancer diagnosis. Carcinogenesis. 29, 1963–6 (2008).
Kogo, R., Mimori, K., Tanaka, F., Komune, S. & Mori, M. Clinical significance of miR-146a in gastric cancer cases. Clin Cancer Res. 17, 4277–84 (2011).
Xu, W. et al. Effects of common polymorphisms rs11614913 in miR-196a2 and rs2910164 in miR-146a on cancer susceptibility: a meta-analysis. PLoS One. 6, e20471 (2011).
Wang, M. et al. Genetic variants in miRNAs predict bladder cancer risk and recurrence. Cancer Res. 72, 6173–82 (2012).
Kasashima, K., Nakamura, Y. & Kozu, T. Altered expression profiles of microRNAs during TPA-induced differentiation of HL-60 cells. Biochem Biophys Res Commun. 322, 403–410 (2004).
Guled, M. et al. CDKN2A, NF2 and JUN are dysregulated among other genes by miRNAs in malignant mesothelioma – A miRNA microarray analysis. Genes Chromosomes Cancer. 48, 615–623 (2009).
Hui, A. B. et al. Comprehensive microRNA profiling for head and neck squamous cell carcinomas. Clin Cancer Res. 16, 1129–1139 (2010).
Farazi, T. A. et al. MicroRNA sequence and expression analysis in breast tumors by deep sequencing. Cancer Res. 71, 4443–4453 (2011).
Xu, B. et al. A functional polymorphism in Pre-miR-146a gene is associated with prostate cancer risk and mature miR-146a expression in vivo. Prostate. 70, 467–472 (2010).
Hu, Y. et al. Two common SNPs in pri-miR-125a alter the mature miRNA expression and associate with recurrent pregnancy loss in a Han-Chinese population. RNA Biol. 8, 861–72 (2011).
Akkız, H., Bayram, S., Bekar, A., Akgöllü, E. & Ulger, Y. A functional polymorphism in pre-microRNA-196a-2 contributes to the susceptibility of hepatocellular carcinoma in a Turkish population: a case-control study. J Viral Hepat. 18, e399–407 (2011).
Ye, Y. et al. Genetic variations in microRNA-related genes are novel susceptibility loci for esophageal cancer risk. Cancer Prev Res. 1, 460–9 (2008).
Farazi, T. A. et al. MicroRNA sequence and expression analysis in breast tumors by deep sequencing. Cancer Res. 71, 4443–53 (2011).
Smith, R. A. et al. A genetic variant located in miR-423 is associated with reduced breast cancer risk. Cancer Genomics Proteomics. 9, 115–8 (2012).
Kontorovich, T., Levy, A., Korostishevsky, M., Nir, U. & Friedman, E. Single nucleotide polymorphisms in miRNA binding sites and miRNA genes as breast/ovarian cancer risk modifiers in Jewish high-risk women. Int J Cancer. 127, 589–97 (2009).
Zhang, M. W. et al. Associations of lifestyle-related factors, hsa-miR-149 and hsa-miR-605 gene polymorphisms with gastrointestinal cancer risk. Mol Carcinog. 51, E21–31 (2012).
Xiao, J., Lin, H., Luo, X. & Wang, Z. MiR-605 joins p53 network to form a p53: miR-605:Mdm2 positive feedback loop in response to stress. EMBO J. 30, 524–32 (2011).
Zhu, L., Wang, Z., Fan, Q., Wang, R. & Sun, Y. MicroRNA-27a functions as a tumor suppressor in esophageal squamous cell carcinoma by targeting KRAS. Oncol Rep. 31, 280–6 (2013).
Chen, Z. et al. MiR-27a modulates the MDR1/P-glycoprotein expression by inhibiting FZD7/β-catenin pathway in hepatocellular carcinoma cells. Cell Signal. 25, 2693–701 (2013).
Yang, R. et al. A genetic variant inthe pre-miR-27a oncogene is associated witha reduced familial breast cancer risk. Breast Cancer Res Treat. 121, 693–702 (2010).
Zhang, M. et al. Associations of miRNA polymorphisms and female physiological characteristics with breast cancer risk in Chinese population. Eur J Cancer Care. 2, 274–80 (2012).
Hezova, R. et al. Evaluation of SNPs in miR-196-a2, miR-27a and miR-146a as risk factors of colorectal cancer. World J Gastroenterol. 18, 2827–31 (2012).
Zhan, J. F. et al. A functional variant in microRNA-196a2 is associated with susceptibility of colorectal cancer in a Chinese population. Arch Med Res. 42, 144–8 (2011).
Li, X. D., Li, Z. G., Song, X. X. & Liu, C. F. A variant in microRNA-196a2 is associated with susceptibility to hepatocellular carcinoma in Chinese patients with cirrhosis. Pathology. 42, 669–73 (2010).
Wang, F. et al. A genetic variant in microRNA-196a2 is associated with increased cancer risk: a meta-analysis. Mol Biol Rep. 39, 269–75 (2012).
Chu, H. et al. Hsa-miR-196a2 Rs11614913 polymorphism contributes to cancer susceptibility: evidence from 15 case-control studies. PLoS One. 6, e18108 (2011).
Tian, T. et al. A functional genetic variant in microRNA-196a2 is associated with increased susceptibility of lung cancer in Chinese. Cancer Epidemiol Biomarkers Prev. 18, 1183–7 (2009).
Doghman, M. et al. Regulation of insulin-like growth factor-mammalian target of rapamycin signaling by microRNA in childhood adrenocortical tumors. Cancer Res. 70, 4666–75 (2010).
Nagaraja, A. K. et al. A link between mir-100 and FRAP1/mTOR in clear cell ovarian cancer. Mol Endocrinol. 24, 447–63 (2010).
Zheng, Y. S. et al. MiR-100 regulates cell differentiation and survival by targeting RBSP3, a phosphatase-like tumor suppressor in acute myeloid leukemia. Oncogene. 31, 80–92 (2012).
Ryan, B. M. et al. Rs4919510 in hsa-mir-608 is associated with outcome but not risk of colorectal cancer. PLoS One. 7, e36306 (2012).
Hu, Z. et al. Common genetic variants in pre-microRNAs were associated with increased risk of breast cancer in Chinese women. Hum Mutat. 30, 79–84 (2009).
Vinci, S. et al. Genetic variants in miR-146a, miR-149, miR-196a2, miR-499 and their influence on relative expression in lung cancers. Clin Chem Lab Med. 49, 2073–80 (2011).
Zhang, M. W. et al. Associations of lifestyle-related factors, hsa-miR-149 and hsa-miR-605 gene polymorphisms with gastrointestinal cancer risk. Mol Carcinog. 51, E21–31 (2012).
Catucci, I. et al. Evaluation of SNPs in miR-146a, miR196a2 and miR-499 as low-penetrance alleles in German and Italian familial breast cancer cases. Hum Mutat. 31, E1052–7 (2010).
Okubo, M. et al. Association between common genetic variants in pre-microRNAs and gastric cancer risk in Japanese population. Helicobacter. 15, 524–31 (2010).
Okubo, M. et al. Investigative role of pre-microRNAs in bladder cancer patients: a case-control study in North India. DNA Cell Biol. 30, 401–6 (2011).
Rothman, N. et al. A multi-stage genome-wide association study of bladder cancer identifies multiply susceptibility loci. Nat Genet. 42, 978–84 (2010).
Hunter, D. J. et al. A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nat Genet. 39, 870–4 (2007).
Landi, M. T. et al. A genome-wide association study of lung cancer identifies a region of chromosome 5p15 associated with risk for adenocarcinoma. Am J Hum Genet. 85, 679–91 (2009).
Petersen, G. M. et al. A genome-wide association study identifies pancreatic cancer susceptibility loci on chromosomes 13q22.1, 1q32.1 and 5p15.33. Nat Genet. 42, 224–8 (2010).
Yeager, M. et al. Genome-wide association study of prostate cancer identifies a second risk locus at 8q24. Nat Genet. 39, 645–9 (2007).
Purdue, M. P. et al. Genome-wide association study of renal cell carcinoma identifies two susceptibility loci on 2p21 and 11q13.3. Nat Genet. 43, 60–5 (2011).
Abnet, C. C. et al. A shared susceptibility locus in PLCE1 at 10q23 for gastric adenocarcinoma and esophageal squamous cell carcinoma. Nat Genet. 42, 764–7 (2010).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet. 38, 904–909 (2006).
Li, J. et al. Identification of high-quality cancer prognostic markers and metastasis network modules. Nat Commun. 1, 1–8; 10.1038/ncomms1033 (2010).
Evangelou, E. & Ioannidis, J. P. Meta-analysis methods for genome-wide association studies and beyond. Nat Rev Genet. 14, 379–89 (2013).
Cotsapas, C. et al. Pervasive sharing of genetic effects in autoimmune disease. PLoS Genet. 7, e1002254 (2011).
Acknowledgements
We thank all the subjects of this study for their participation. The GWAS dataset used for the analysis described in this manuscript was obtained from the database of Genotypes and Phenotypes (dbGaP) found at http://www.ncbi.nlm.nih.gov/gap through dbGaP accession number phg0000132.v1, phg000032.v1, phg000124.v1, phg000089.v1, phg000067.v1, phg000123.v1 and phg000128.v1. This study is sponsored by Shanghai Natural Science Foundation, No. 13ZR14244000.
Author information
Authors and Affiliations
Contributions
Y.H. and C.Y.Y. contribute equally to the work. Y.H., C.Y.Y. and H.Y.C. wrote the main manuscript text, Y.H. prepared figure 1 and table 1 and C.Y.Y. prepared figure 2 and table 2. Y.H., C.Y.Y., J.L.W., J.G., H.Y.C. and J.Y.F. reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Table S1
Rights and permissions
This work is licensed under a Creative Commons Attribution 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by/3.0/
About this article
Cite this article
Hu, Y., Yu, CY., Wang, JL. et al. MicroRNA sequence polymorphisms and the risk of different types of cancer. Sci Rep 4, 3648 (2014). https://doi.org/10.1038/srep03648
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03648
This article is cited by
-
Genetic variants in miR-145 gene are associated with the risk of asthma in Taiwan
Scientific Reports (2022)
-
The miRNA 196a2 rs11614913 variant has prognostic impact on Turkish patients with multiple myeloma
BMC Research Notes (2020)
-
MicroRNAs identification and bioinformatics analysis in large yellow croaker Larimichthys crocea using an integrated comparative and ab initio approach
Journal of Oceanology and Limnology (2018)
-
SNP in microRNA sequences or binding sites of miRNAs: association with type 2 diabetes mellitus susceptibility and in silico analysis
International Journal of Diabetes in Developing Countries (2018)
-
Association of microRNA polymorphisms with the risk of head and neck squamous cell carcinoma in a Chinese population: a case–control study
Chinese Journal of Cancer (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.