Abstract
Microinjection of foreign DNA in male pronucleus by in-vitro embryo manipulation is difficult but remains the method of choice for generating transgenic animals. Other procedures, including retroviral and embryonic stem cell mediated transgenesis are equally complicated and have limitations. Although our previously reported technique of testicular transgenesis circumvented several limitations, it involved many steps, including surgery and hemicastration, which carried risk of infection and impotency. We improved this technique further, into a two step non-surgical electroporation procedure, for making transgenic mice. In this approach, transgene was delivered inside both testes by injection and modified parameters of electroporation were used for in-vivo gene integration in germ cells. Using variety of constructs, germ cell integration of the gene and its transmission in progeny was confirmed by PCR, slot blot and immunohistochemical analysis. This improved technique is efficient, requires substantially less time and can be easily adopted by various biomedical researchers.
Similar content being viewed by others
Introduction
Mouse transgenesis is an important tool for functional genetics and for creating humanized models of diseases. Transgenic animals can be produced by pronuclear DNA microinjection1,2, retroviral mediated gene transfer3, embryonic stem cell based gene transfer4 and via somatic cell nuclear transfer5,6. Despite the success of these procedures in generating genetically altered animals, these techniques are difficult to perform and hence limited to selected laboratories having facilities and skilled personnel for embryo manipulation and embryo transfer7. Intracytoplasmic sperm injection (ICSI) or male germ cell mediated techniques of gene transfer have also been reported in literature8,9,10,11. However, these techniques either require in vitro germ cell manipulation followed by surgical germ cell transplantation or ICSI followed by embryo transfer in surrogates or surgical manipulation to deliver DNA in the testis followed by electroporation of DNA in the spermatogonia. All of these methods involve surgical steps which can potentially cause infection and/or impotency. Therefore there is a need to develop relatively safer and a less time consuming procedure for transgenesis.
Testicular transgenesis has advantage over zygote or oocyte mediated transgenesis because this does not require large number of females for super ovulation and expertise for embryo manipulation. To ensure easy adaptability of our previously reported testicular transgenesis technique, we improved the technique into a two step non-surgical electroporation procedure for making transgenic mice. We standardized the technique using construct having Enhanced Green Fluorescence Protein (EGFP) as the reporter. Subsequently, we also generated transgenic lines of mice using different constructs including those for tissue specific expression and those displaying patho-physiological effect of the transgene. This is the first report of non-surgical approach for in vivo germline integration of foreign gene and efficient generation of transgenic mice without using any assisted reproductive technique (ART). This procedure can be potentially adopted for transgenesis in farm animals where surgery as well as embryo transfer are not as easy as those in mice.
Results
Establishment of the technique for non-surgical in-vivo electroporation of the testis
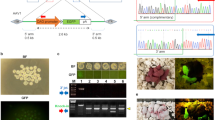
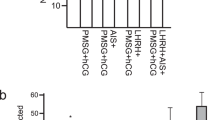
Linearised pCX-EGFP plasmid (Fig. 1a-i) was used for standardization of the technique; one testis was injected with the DNA and the contra-lateral testis was kept as control (non-injected). Various DNA injection parameters such as amount of DNA (ranging from 10 μg–30 μg per testis), volume of suspended DNA (ranging from 20–25 μl) injected per testis, electroporation parameters such as voltage (ranging from 50 V–90 V), time constant (ranging from 10 ms–60 ms) and number of pulses were varied to determine the most optimum condition for gene integration in the spermatogonial cells of testis (Table 1).
(a): Schematic diagram of the various constructs used in the study: i) pCX-EGFP plasmid. (ii) PEPCK1-IRES2-EGFP plasmid. (iii) K14-IRES2-EGFP plasmid. (iv) SMAR1-EGFP plasmid. (b): Schematic representation of the non-surgical external in-vivo testicular electroporation procedure. (i) represents the anaesthetized mice; (ii) shows the DNA injection in one of the testis from two different sites; (iii) shows the DNA injection in remaining testis; (iv) shows the electroporation in both the testes, held together, using tweezer type electrode; (v) shows the electroporation by changing the poles (red and black) of the tweezer type electrode after 4 pulses.
Desirable results were obtained in the combination of steps described in EP12 (Table 1) where efficient transgene integration and expression in the testis was achieved by injecting 20 μl of linearized DNA suspension (0.5 μg/μl) into testis of about 30 days old mice before electroporating germ cells with 4 square 60 V electric pulses (duration of 50 milli-seconds each with an inter-pulse interval of approximately one second) in one direction (forward direction) and 4 more similar pulses after changing the sides of the electrodes (reverse direction; Fig. 1b and Table 1). Electroporated testis expressed EGFP even at 80 days of age (Fig. 2a & Fig. 2b). GFP expressing germ cells were discernible in the seminiferous tubules of all 4 such pCX-EGFP electroporated mice (Fig. 2c).
EGFP expression in mouse testis, 50 days after electroporation.
(a). Image in bright field stereozoom microscope. (b). Image under FITC filter of stereozoom microscope. In the area injected with DNA, EGFP expression is marked with yellow line. (c). EGFP expression in germ cells of seminiferous tubules of electropoarated testis upon staining with anti EGFP antibody. Yellow arrow heads show staining in germ cells. Note: EP – electroporated testis; NEP – Non-electroporated testis.
Generation of transgenic mice for tissue specific expression of transgene
We also investigated whether the optimized procedure can generate transgenic progeny when tissue specific expression cassette was used. For this, we electroporated testicular germ cells with pK14-IRES2-EGFP construct in which EGFP expression was driven by promoter of skin specific protein, keratin 14 (K14) and with pPEPCK1-IRES2-EGFP construct, in which EGFP expression was driven by the promoter of liver specific phosphoenolpyruvate carboxykinase 1 (PEPCK1). In case of both constructs, transmission of transgene from electroporated mice (fore-founder) to several of their progeny was discerned by PCR analysis. PCR results (data not shown) were further confirmed by slot-blot analysis where the IRES2-EGFP region of the transgene (Fig. 1a-ii & iii) was detected with αP32 labelled probe (Fig. 3a & 3b). One of the pK14-IRES2-EGFP fore founder mice sired transgenic progeny even after 274 days of electroporation. Integration of pK14-IRES2-EGFP transgene in germ cell was also confirmed by fluorescence in situ hybridization of sperm from epididymis of electroporated mice (Fig. 3c). Expression of EGFP was detected specifically in the epidermis of the skin of pK14-IRES2-EGFP transgenic progeny (Fig. 4a). Similarly, EGFP expression was also detected in the cross section of the liver (Fig. 4b) of pPEPCK1-IRES2-EGFP transgenic progeny.
Determination of transgene integration by slot blot analysis.
(a): P4 & P6 (F1 of Ext 21); P19–P21 (F1 of Ext 22); P25–P33 (F2 of P6); P34–P49 (F2 of P4). Wt1–Wt5 gDNA from five different wild type mice. (b): K1–K8 (F1 of Ext 17); K12–21 (F1 of Ext 18); K23–K28 (F2 of K17); K33–67 (F2 of K18). Wt1–Wt5 represents gDNA from five different wild type mice. The gDNA were probed with labelled IRES2-EGFP fragment as all of the lines mentioned above contained IRES2-EGFP. The same fragment was used as positive control in both the blots. Note: K denotes transgenic animal of pK14-IRES2-EGFP line; P denotes transgenic animal of pPEPCK1-IRES2-EGFP line; WT denotes wild type mice. +ve denotes plasmid DNA fragment. Ext denotes the respective fore-founder mice. F1 and F2 denote first and second generation of progeny respectively. (c): Fluorescence in situ hybridization of the sperm from pK14-IRES2-EGFP electroporated mice. Green signal is seen in sperm nuclei due to hybridization of probe with pK14-IRES2-EGFP transgene and blue signal displays DAPI staining of nucleus. Field under observation shows two transgenic sperm and two non transgenic sperm. (i) Phase contrast image. (ii) Image under UV. Scale bar, 20 μm.
(a): EGFP expression in the skin of pK14-IRES2-EGFP transgenic mice stained with anti EGFP antibody. (i & ii) immunohistochemical images of the skin sections of the transgenic mice. (iii & iv) immunohistochemical images of the skin sections of wild type mice. Left panel shows the phase contrast image and right panel shows the image under UV with FITC filter. Scale bar: 50 μm (i–ii); 100 μm (iii–iv). (b): EGFP localization in the liver of pPEPCK1-IRES2-EGFP transgenic mice stained with anti EGFP antibody. (i & ii) immunohistochemical images of the liver sections of the transgenic mice. (iii & iv) immunohistochemical images of the liver section of wild type mice. Left panel shows the phase contrast image and right panel shows the image under UV with FITC filter. Scale bar, 50 μm (i–ii) & 20 μm (iii–iv).
Generation of transgenic mice displaying patho-physiological effect
We used pEGFP-SMAR1construct (Fig. 1a-iv) for making fore-founders which were cohabitated with wild type females. Presence of transgene in progeny was determined by PCR analysis (data not shown). Over-expression of Scaffold/Matrix attachment region binding protein1 (SMAR1) caused splenomegaly in transgenic mice (Fig. 5a & 5b). We also detected EGFP expression in the cross section of spleen (Fig. 5c) obtained from pEGFP-SMAR1transgenic mice.
(a & b): Enlarged spleen of the mice transgenic for Egfp-Smar1 (left) as compared to wild type mice (right); Tg1, Tg2 and Tg3 denote enlarged spleen of three different transgenic mice and WT1 and WT2 denote normal spleen of two different wild type mice. (c): EGFP expression in the spleen of pSMAR1-EGFP transgenic mice stained with anti EGFP antibody. (i & ii) immunohistochemical images of spleen sections from the transgenic mice. (iii & iv) immunohistochemical images of the spleen section from the wild type mice. Left panel shows the phase contrast image and right panel shows the image under UV with FITC filter. Scale bar, 20 μm (i–iv).
Discussion
Presently used techniques for generation of transgenic mice exploiting spermatogonial cells require in vitro germ cell culture and/or various surgical steps which are difficult to adopt10,12. We have substantially improved the procedure of in vivo testicular transgenesis making it an easily adaptable, user friendly technique. In the hands of a person who is less experienced in performing surgery, surgical intervention in the vicinity of the reproductive area is prone to development of impotency or post surgical infection in the animals. Unlike surgical exposure of one of the testis for DNA injection followed by removal of contra-lateral testis, as described by us before11, in the present procedure, suspension of gene was delivered in both testes of mice directly from outside followed by electroporation (Supplementary Fig. S1). Simultaneous electroporation and gene integration in both testes by holding them together at the time of electroporation precluded the need of hemicastration as required for previously reported procedure11. As compared to this new technique, the previously reported procedure where electroporation of testis followed surgical exposure had several drawbacks (Table 2). Expression of EGFP in the electroporated testis about a month and a half after electroporation with pCX-EGFP, confirmed genomic integration of the transgene. The improved procedure resulted in integration of the transgene, largely in germ cells of the seminiferous tubule. Validation of the technique was done using different mammalian gene constructs. Electroporation with this chosen set of parameters (60 v/50 ms, 4 pulses each in forward and reverse direction with inter pulse interval of 1 second) did not alter the cyto-architecture of the testis, the normalcy of which is crucial for spermatogenesis13.
To determine the suitability of this procedure for tissue specific gene expression, we designed pK14-IRES2-EGFP and pPEPCK1-IRES2-EGFP constructs for expression of EGFP in skin and liver, respectively. Fore-founder mice were generated using these constructs and cohabitated with wild type females after 35 days post electroporation. In mice, differentiation of spermatogonia into sperm occurs in about 35 days10,14,15. Generation of transgenic progeny from mating after 35 days of electroporation suggested that spermatogonial cells were successfully transfected at the time of electroporation. Detection of transgene in the epididymal sperm of electroporated male by fluorescence in situ hybridization provided substantial evidence in favour of germ line gene integration upon external electroporation of testis. F1 progeny obtained upon mating of fore-founders were checked for the presence of transgene. Several pups were found to be PCR positive for the transgene. This indicated the transmission of integrated transgene to the next generation. We confirmed these PCR results by slot blot analysis. Genomic DNA of PCR positive progeny were probed with IRES2-EGFP fragment (Fig. 1a-ii & iii) labeled with radioactive αP32. Please note that IRES2-EGFP was the part of both pK14-IRES2-EGFP and pPEPCK1-IRES2-EGFP constructs. Slot blot positive male and female mice of F1 generation were bred to generate F2 generation and so on.
Immunohistochemical analysis of skin from F1 progeny of pK14-IRES2-EGFP transgenic mice strongly displayed the specific expression of EGFP in the epidermal layer of skin. Use of K14 promoter for skin specific expression of genes is well documented and our results were similar to those reported previously for such transgenic animals bearing K14 promoter driven genes which were generated using traditionally used pronuclear DNA microinjection method16,17,18. Similarly, immunohistochemical analysis of liver sections of pPEPCK1-IRES2-EGFP transgenic mice revealed expression of transgene (EGFP) predominantly in the liver, as reported before using traditional procedure of transgenesis19. These observations suggested that this new technique is capable of generating transgenic mice with tissue specific targeted gene expression.
To explore the possibility of making animal models showing patho-physiological condition, we generated transgenic mice by this procedure using pEGFP-SMAR1 construct. SMAR1 is the MAR binding protein that functions as a candidate tumor suppressor by interacting with and activating p53, thereby regulating cell cycle20. Please note that SMAR1 is also expressed endogenously21. In this study, transgenic mice carrying pEGFP-SMAR1 had enlarged spleen which over expressed SMAR1-EGFP fusion protein; this was similar to presence of enlarged spleen in SMAR1 transgenic mice, traditionally made using male pronuclear DNA microinjection22. Observations from pEGFP-SMAR1 transgenic mice made by this new technique confirmed that transgene was stably integrated and transmitted to the next generation with visible patho-physiological effect (enlargement of spleen).
In summary, generation of transgenic mice by non-surgical, external testicular electroporation resulted in the germ line integration of the transgene. All electroporated male mice successfully produced litters among which several were transgenic. This procedure required less period of sedation and less time as compared to previously reported procedures1,8,11. The percentage of F1 progeny bearing transgene was about 57%–62%, using various types of gene constructs used in this study (Table 3). This is remarkably higher than Pronuclear DNA microinjection mediated transgenesis where 10%–20% of attempted progeny are PCR positive for the transgene7 making our technique relatively more efficient. All four animals electroporated with pCX-EGFP construct expressed EGFP in the testicular tissue. Similarly each of 4 slot blot positive animals of F1 generation showed presence of EGFP in organs for which specific promoters were used. This modified technique is simple, less time consuming and ethically superior than our previously described procedure as it uses mild anesthesia for short duration and does not require hemicastration or any other surgical interventions or sacrifice for transgenesis. Since cumbersome surgical steps or assisted reproductive techniques are not involved, several fore-founders can be generated with minimum effort using various constructs by this procedure. Right and left testes of an animal can also be independently used for transfecting two different kinds of gene constructs which can further reduce (by half) the production time for generation of two different transgenic lines. We used FVB/J strain of mice, however, minor procedural variations (for eg. different concentration of DNA or moderate variation in current used) may be required while using other strains of mice. A large variety of mice carrying different transgenes can be generated within short span of time using this 2 step short-cut method (DNA injection and external electroporation). Such a simple non-surgical technique may also potentially increase the scope of generating transgenic farm animals with desirable traits.
Methods
Animals
Mice (FVB/J), bred at the small animal facility of National Institute of Immunology, were used for the present study. All animals were kept at 24 ± 2°C under 14 h light and 10 h dark cycle and used as per the National Guidelines provided by the Committee for the Purpose of Control and Supervision of the Experiments on Animals (CPCSEA). Protocols for the experiments were approved by the Institutional Animal Ethics Committee.
Constructs
pCX-EGFP
This plasmid contains functional chicken beta actin promoter along with cytomegalovirus transcription enhancer element (CX) as a strong ubiquitous transcription regulator. A functional EGFP gene is cloned under this promoter assembly which yields a strong ubiquitous EGFP expression (Fig. 1a-i). This plasmid was linearized with Sal I restriction enzyme before used for the standardization of the technique.
pPEPCK1-IRES2-EGFP
The promoter for the cytosolic variant of phosphoenolpyruvate carboxykinase 1(PEPCK1) was PCR amplified from mice genome employing high fidelity Pfu DNA polymerase (Fermentas, Pittsburgh, PA, USA). The PCR condition included initial denaturation at 95°C for 4 min followed by 29 cycles of 95°C for 45 s, 64°C for 45 s, 72°C for 1 min with final extension of 72°C for 10 min; resulting in 1037 bp amplicon. Primers employed for amplifying the promoter were Forward: 5′-CTCCCCTCCTTTCTCCAGACAC-3′ and Reverse: 5′-GAAGGTCTGGATCCGAGATCGC-3′. The amplicon was ligated into pIRES2-EGFP vector (Clontech, Mountain View, CA, USA) to generate the plasmid pPEPCK1-IRES2-EGFP (Fig. 1a-ii). After plasmid amplification, construct was digested with EcoRI and AflII and the desired linearized fragment (∼2.6 kb), containing promoter and the gene, was purified using gel extraction kit (Qiagen, Hilden, Germany) before it was used for electroporation.
pK14-IRES2-EGFP
The promoter region for human keratinocyte specific protein (K14) was amplified using long PCR employing high fidelity Pfu DNA polymerase (Fermentas, Pittsburgh, PA, USA) from human genomic DNA isolated from blood. Primers used for amplification of 2486 bp upstream region of K14 gene were Forward: 5′-AACCCTTTAACCCCTGATGC-3′ and Reverse: 5′- GTGCAGAGGAGGGAGGTGAG-3′. PCR condition included initial denaturation of 4 min at 95°C followed by 29 cycles of 45 s at 95°C, 45 s at 63°C, 3 min at 72°C with a final extension at 72°C for 10 min. The ∼2.5 kb amplicon thus obtained was ligated into pIRES2-EGFP vector (Clontech, Mountain View, CA, USA) to generate plasmid pK14-IRES2-EGFP (Fig. 1a-iii). After plasmid amplification, this construct was digested with EcoRI and AflII and the desired linearized fragment (∼4.1 kb) was purified using gel extraction kit (Qiagen, Hilden, Germany) before it was used for electroporation.
pEGFP-SMAR1
This fusion gene construct consisted of full-length fragment of Scaffold/Matrix attachment region binding protein1 (SMAR1) as described before23 where EGFP which was cloned downstream of cytomegalovirus immediate early promoter/enhancer of pEGFP-C3 vector (Clontech, Mountain View, CA, USA). For testicular injection, the plasmid was digested with XbaI and ApaLI restriction enzyme (Fig. 1a-iv). Two fragments were generated, one of ∼4 kb and the other of ∼3.2 kb. The 3.2 kb fragment which had CMV at the 5′ end and SMAR1 at 3′ end was used for electroporation.
Non-surgical in vivo injection and electroporation of mice testis
For this purpose, about 30 days old male mice (FVB/J) were anesthetized for short duration by intraperitonial injection (100 μl) of ketamine hydrochloride (45 mg/kg) and xylazine hydrochloride (8 mg/kg). After removing hair from lower abdominal and scrotal region, area was cleaned with betadine (povidone-iodine) solution. Subsequently, sterile water was spread on scrotal area to remove excess betadine. Plasmid DNA containing 0.04% Trypan blue, which was used to monitor the accuracy of the injection, was injected slowly into the testis using the 10 μl Hamilton syringe (701N; Hamilton Bonaduz AG, Switzerland) from two different, diagonally opposite sites. DNA was delivered through one injection site at a time (Fig. 1b). Usually, needle of Hamilton syringe is long and tends to bend because of the resistance posed by the scrotal sac. Therefore, a small (0.5 inch long) 26 gauge sterile needle (usually supplied with tuberculin syringe) was slowly inserted up to middle of the testis and withdrawn to generate path for introducing the needle of a Hamilton syringe. For standardization, about 20 μl–25 μl of sterile water containing the desired plasmid DNA (concentration ranging from 0.5 μg/μl–1.2 μg/μl) was delivered unilaterally into a testis. Mild square electric pulses ranging from 50 V to 90 V were delivered to testis injected with DNA using tweezer type electrode and an electric pulse generator (Electroporator ECM2001, BTX Instrument Division, Harvard Apparatus, Inc., Holliston, Massachusetts, USA). Variation in several parameters like voltage, time constant (duration of the pulse), number of pulses and number of injection sites were done at the time of electroporation to determine the most suitable condition for achieving a successful and stable gene integration in germ cell of the testis (Table 1). Sterile normal saline was spread occasionally on scrotum to keep the skin moist for uniform electrical conductance during the procedure.
For generating transgenic lines using various constructs, DNA was injected in both testes of every animal and testes were held together between a sterile tweezer-type electrode for their simultaneous electroporation.
Analysis of electroporated testis under stereozoom microscope
pCX-EGFP electroporated male mice were sacrificed by cervical dislocation after 50 days of electroporation. Testes were observed under a stereo-zoom microscope SMZ-1500 (Nikon Corporation, Chiyoda-ku, Tokyo, Japan) fitted with epi-fluorescence attachment. Images were captured using DS-5M camera with Digital sight DS-LI software.
Propagation of the transgene & PCR based screening
The electroporated males (fore-founders) were cohabitated with wild-type females, 35 days post-electroporation and the pups generated (F1 generation) were analyzed for the presence of transgene by PCR. F2 generations of mice were generated by breeding PCR positive males and females from F1 generation.
Slot blot analysis
The results of PCR were further confirmed by Slot blot analysis24 of few PCR-positive animals from each generation. Transgene specific probe was generated with αP32dCTP using High Prime DNA labeling kit (Roche Diagnostic GmbH, Mannheim, Germany). About 1 μg of gDNA was blotted on membrane and hybridized with transgene specific probe for detection of transgene in the progeny.
Immunohistochemistry
Expression of EGFP in the cross section of the testis of pCX-EGFP transgenic mice, spleen of pEGFP-SMAR1 transgenic mice, skin of pK14-IRES2-EGFP transgenic mice and liver of pPEPCK1-IRES2-EGFP transgenic mice was detected by immuno-histochemistry. Tissue samples were fixed in Bouin's solution for 18–24 h and then embedded in paraffin. Tissue sections (4 micron) were stained with mouse anti-GFP (Clontech, Mountain View, CA, USA) antibody at 1∶250 dilution for 10 hrs at 4°C. Alexa fluor 488 conjugated goat anti mouse IgG (Molecular probes, Eugene, OR, USA) was used as secondary antibody at 1∶250 dilution for 4 hrs at room temperature for the detection of specific protein. Stained sections were analyzed under Zeiss LSM 510 META confocal laser-scanning microscope (Zeiss, Jena, Germany) and LSM 510 software was used for acquisition of images. Images were also captured using Nikon Eclipse TE2000-S inverted microscope (Nikon Corporation, Chiyoda-ku, Tokyo, Japan) attached to DS-5 M camera assisted with digital sight DS-L1 software.
Fluorescence in situ hybridization
For fluorescence in situ hybridization, sperm from pK14-IRES2-EGFP electroporated mouse were taken from cauda epididymis and washed in PBS; sperm nuclei were prepared as described previously25. pK14-IRES2-EGFP plasmid was labeled with green dUTP using the Vysis nick translation kit (Abott Molecular Inc., Des Plaines, IL, USA) and used as probe. In situ hybridization was performed as described previously25. The fluorescence was observed under Nikon Eclipse Ti inverted fluorescence microscope (Nikon Corporation, Chiyoda-ku, Tokyo, Japan). The images were captured using Nikon-digital sight DS-Ri1 camera.
References
Gordon, J. W., Scangos, G. A., Plotkin, D. J., Barbosa, J. A. & Ruddle, F. H. Genetic transformation of mouse embryos by microinjection of purified DNA. Proc. Natl. Acad. Sci. 77, 7380–7384 (1980).
Palmiter, R. D. et al. Dramatic growth of mice that develop from eggs microinjected with metallothionein-growth hormone fusion genes. Nature 300, 611–615 (1982).
Van der, P. H. et al. Efficient insertion of genes into the mouse germ line via retroviral vectors. Proc. Natl. Acad. Sci. 82, 6148–6152 (1985).
Beard, C., Hochedlinger, K., Plath, K., Wutz, A. & Jaenisch, R. Efficient method to generate single-copy transgenic mice by site-specific integration in embryonic stem cells. Genesis 44, 23–28 (2006).
Cibelli, J. B. et al. Cloned transgenic calves produced from non quiescent fetal fibroblasts. Science 280, 1256–1258 (1998).
Baguisi, A. et al. Production of goats by somatic cell nuclear transfer. Nat Biotechnol. 17, 456–461 (1999).
Wall, R. J. Pronuclear microinjection. Cloning stem cells. 3, 209–220 (2001).
Perry, A. C. et al. Mammalian transgenesis by intracytoplasmic sperm injection. Science 284, 1180–1183 (1999).
Nagano, M. et al. Transgenic mice produced by retroviral transduction of male germ-line stem cells. Proc. Natl. Acad. Sci. 98, 13090–13095 (2001).
Brinster, R. L. Germline stem cell transplantation and transgenesis. Science 296, 2174–2176 (2002).
Dhup, S. & Majumdar, S. S. Transgenesis via permanent integration of genes in repopulating spermatogonial cells in vivo. Nature Methods 5, 601–603 (2008).
Kanatsu-Shinohara, M. et al. Long-term proliferation in culture and germline transmission of mouse male germline stem cells. Biol Reprod. 69, 612–616 (2003).
Jégou, B. The Sertoli-germ cell communication network in mammals. Int. Rev. Cytol. 147, 25–96 (1993).
Russell, L. D., Ettlin, R. A., Sinha Hikim, A. P. & Clegg, E. D. [Mammalian spermatogenesis]. Histological and Histopathological Evaluation of the Testis [Russell, L. D. & Ettlin, R. A. (1st ed.)] [1–40] (Cache River Press, Clearwater, Florida, 1990).
de Rooij, D. G. & Russell, L. D. All you wanted to know about spermatogonia but were afraid to ask. J. Androl. 21, 776–798 (2000).
Wang, X., Zinkel, S., Polonsky, K. & Fuchs, E. Transgenic studies with a keratin promoter driven growth hormone transgene: prospects for gene therapy. Proc. Natl. Acad. Sci. 94, 219–226 (1997).
Fuchs, E. & Green, H. Changes in keratin gene expression during terminal differentiation of the keratinocytes. Cell 19, 1033–1042 (1980).
Stellmach, V., Leask, A. & Fuchs, E. Retinoid-mediated transcriptional regulation of keratin genes in human epidermal and squamous cell carcinoma cells. Proc. Natl. Acad. Sci. 88, 4582–4586 (1991).
McGrane, M. M. et al. Tissue specific expression and dietary regulation of a chimeric phosphoenolpyruvate carboxykinase/bovine growth hormone gene in transgenic mice. J. Biol. Chem. 263, 11443–11451 (1988).
Kaul, R. et al. Direct interaction with and activation of p53 by SMAR1 retards cell-cycle progression at G2/M phase and delays tumor growth in mice. Int. J. Cancer 103, 606–615 (2003).
Chattopadhyay, S., Kaul, R., Charest, A., Housman, D. & Chen, J. SMAR1, a novel, alternatively spliced gene product, binds the Scaffold/Matrix-associated region at the T cell receptor beta locus. Genomics 68, 93–96 (2000).
Kaul-Ghanekar, R. et al. Abnormal V(D)J recombination of T cell receptor beta locus in SMAR1 transgenic mice. J. Biol. Chem. 280, 9450–9459 (2005).
Chattopadhyay, S., Whitehurst, C. E., Schwenk, F. & Chen, J. Biochemical and functional analyses of chromatin changes at the TCR-beta gene locus during CD4-CD8- to CD4 + CD8+ thymocyte differentiation. J. Immunol. 160, 1256–1267 (1998).
Brown, T. [Dot and slot blotting of DNA]. Current Protocols in Molecular Biology. [Brown, T. ] [21: 2.9.15–2.9.20] (John Wiley and Sons, Hoboken, 2001).
Ibáñez, E., Molist, J., Vidal, F., Egozcue, J. & Santaló, J. Assessment of the proportion of transgene-bearing sperm fluorescence in situ hybridization: a novel approach for the detection of germline mosaicism in transgenic male founders. Mol Reprod Dev. 58, 166–172 (2001).
Acknowledgements
We are grateful to the Director of National Institute of Immunology for her support. We are thankful to Dr S. Chattopadhyay (National Center for Cell Science, Pune, India) for providing the SMAR1-EGFP gene construct. We are also grateful to Dr. Y. Junming (University of Tennesse, Memphis, USA) for giving us the pCX-EGFP construct. We greatly acknowledge the technical support from Birendra N. Roy, Dharamveer and Ram Singh. We acknowledge the help from staff of Small Animal Facility of National Institute of Immunology. We are grateful to Department of Biotechnology, Govt. of India, for providing the financial assistance under grants BT/PR11313/AAQ/01/376/2008 and BT/HRD/35/01/01/2010 (TATA Innovation Award).
Author information
Authors and Affiliations
Contributions
Experiments were conceived and designed by S.S.M. Experiments were performed by A.U., N.G., H.S., S.D., S.R.B., M.V., N.G., S.B. and P.N. Data of the manuscript was analyzed by A.U., N.G. and S.S.M. Manuscript was written and reviewed by A.U., N.G. and S.S.M.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Figure 1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Usmani, A., Ganguli, N., Sarkar, H. et al. A non-surgical approach for male germ cell mediated gene transmission through transgenesis. Sci Rep 3, 3430 (2013). https://doi.org/10.1038/srep03430
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03430
This article is cited by
-
Follicle-stimulating hormone-mediated decline in miR-92a-3p expression in pubertal mice Sertoli cells is crucial for germ cell differentiation and fertility
Cellular and Molecular Life Sciences (2022)
-
Intratesticular injection followed by electroporation allows gene transfer in caprine spermatogenic cells
Scientific Reports (2018)
-
Defective Wnt3 expression by testicular Sertoli cells compromise male fertility
Cell and Tissue Research (2018)
-
Robust generation of transgenic mice by simple hypotonic solution mediated delivery of transgene in testicular germ cells
Molecular Therapy - Methods & Clinical Development (2016)
-
An Efficient Method for Generation of Transgenic Rats Avoiding Embryo Manipulation
Molecular Therapy - Nucleic Acids (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.