Abstract
Early identification of causal genetic variants underlying antimalarial drug resistance could provide robust epidemiological tools for timely public health interventions. Using a novel natural genetics strategy for mapping novel candidate genes we analyzed >75,000 high quality single nucleotide polymorphisms selected from high-resolution whole-genome sequencing data in 27 isolates of Plasmodium falciparum. We identified genetic variants associated with susceptibility to dihydroartemisinin that implicate one region on chromosome 13, a candidate gene on chromosome 1 (PFA0220w, a UBP1 ortholog) and others (PFB0560w, PFB0630c, PFF0445w) with putative roles in protein homeostasis and stress response. There was a strong signal for positive selection on PFA0220w, but not the other candidate loci. Our results demonstrate the power of full-genome sequencing-based association studies for uncovering candidate genes that determine parasite sensitivity to artemisinins. Our study provides a unique reference for the interpretation of results from resistant infections.
Similar content being viewed by others
Introduction
Antimalarial drug resistance has repeatedly frustrated global efforts to limit morbidity and prevent mortality from Plasmodium falciparum malaria. Recently, landmark studies conducted in Western Cambodia in patients who had been treated with artemisinin derivatives have reported an alarming delay in parasite clearance1,2. Since then, infections with increasingly delayed clearance were also reported from Western Thailand and it was suggested that this in vivo phenotype is genetically determined3. Because artemisinin-based combination chemotherapies are the backbone of global malaria control programs, this situation constitutes a public health emergency. Historically, Southeast Asia has been the origin of global spread of drug resistance-conferring mutations. The reason for this geographical bias is only partially understood, but ecological, behavioral and biological factors that may play a role include high rates of inbreeding in the mosquito vector, which reduce competition and favor clonal expansion of emerging genetic variants under drug pressure4, indiscriminate use of poor-quality drugs5 and possibly, “hyper-mutant” parasite strains6.
High throughput whole-genome sequencing technology has revolutionized the approach for identifying genetic variants associated with phenotypes of interest in natural populations. We have harnessed this natural genetic strategy for identifying novel candidate genes that modify the susceptibility of P. falciparum to antimalarial drugs. We hypothesized that the range of phenotypic variation observed in natural populations of P. falciparum is hard-wired to naturally occurring genetic variants, termed ‘standing variation’, without necessarily reflecting “resistance” as an evolutionary adaptation to selective pressure7,8. Here we present the results of a genome-wide study in 27 isolates of Plasmodium falciparum obtained from malaria patients in Kilifi, Kenya.
Results
In vitro phenotyping of P. falciparum isolates from Kenyan malaria patients
Fully culture adapted P. falciparum isolates obtained from pediatric patients enrolled in a clinical trial in Kilifi District, Kenya9 were subjected to independently repeated growth inhibition assays to obtain reproducible drug sensitivity phenotypes (expressed as half-maximal inhibitory concentration; IC50). We focused on a panel of 8 common antimalarial drugs based on their importance as former (chloroquine, CQ; pyrimethamine, PM; and desethyl-amodiaquine, DEAQ) and current (dihydroartemisinin, DHA; lumefantrine, LM; piperaquine, PPQ; and quinine, QN) first or second line antimalarial treatments in Kenya. Mefloquine (MQ) was added because of its global importance for treating and preventing malaria. The median IC50 values (range) were 37 nM (12–310) for CQ; 22 μM (3–70) for PM; 22 nM (7–120) for DEAQ; 2 nM (0.5–4) for DHA; 17 nM (4–85) for LM; 49 nM (24–122) for PPQ; 60 nM (12–140) for QN; and 29 nM (7–99) for MQ. The intra-sample correlation between IC50 assay replicates (a measure of reproducibility) was high (median: 0.97, range: 0.90–0.99). The median correlation between drug assays was 0.18 and varied between 0.01 (lumefantrine and quinine) and 0.76 (LM and MQ) (see Figure 1 for a summary of the assays). The observed pattern of correlations (Fig. 1) was consistent with previously published data (e.g., DHA and LM, 0.46 and DHA and MQ, 0.58)10,11. We did not classify isolates into sensitive and resistant categories for several reasons. First, there is no general consensus on in vitro cutoff values and their relevance for in vivo resistance can be obscured by unrelated parameters (primarily, of pharmacokinetic and immunological nature). For the artemisinin class of drugs, only recently studies started to address the relationship between specialized novel in vitro assays (not done here) and the delayed parasite clearance phenotype observed in vivo12. Secondly, there is increased statistical power in using quantitative, as opposed to qualitative, data for association analyses.
Half-maximal inhibitory concentrations (IC50) for drug assays (phenotypes).
The diagonal is a histogram of the phenotypes, right diagonal is the Spearman’s correlation between assays; left diagonal is raw data and smoothed relationship using cubic splines; DHA dihydroartemisinin, LM lumefantrine, PPQ piperaquine, CQ chloroquine, PM pyrimethamine, MQ mefloquine, QN quinine, DEAQ desethylamodiaquine.
Whole genome sequence analysis
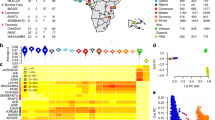
The sequencing technology yielded a median of 18.7 (range: 8.6–38.8) million 54–76 base-pair reads across the 27 samples. Mapping uniquely the reads to the reference 3D7 genome13 yielded a genome-wide average of 57.3-fold coverage and a median of 86.2% of the genome being covered, 69.6% to at least a five-fold coverage level. The average number of allelic differences to 3D7 (at an error rate of 1 per 1000) was 8899/strain and across all samples 182,357 positions were identified, leading to 75,471 high quality bi-allelic SNPs (with <10% missing alleles per position among all samples, minor allele frequency of 5%) carried forward for further analysis. The vast majority of SNPs (66,966, 88.7%) contained no heterozygous genotype calls. Overall, only 0.6% of genotype calls were heterozygous, potentially indicative that few infections/isolates were multi-clonal. Using a principal component analysis on the SNPs, there was no evidence of any samples being continental outliers or identical genetically (for instance, due to potential contamination) (Fig. S1).
Association analysis
Because of the observed deviation from a normal distribution of in vitro responses (Fig. 1) we applied a conservative non-parametric tests for the phenotype-genotype association analysis (Table 1, Table S1). We observed ten-fold differences in the range of IC50 values (the ‘effect size’) for chloroquine and DHA (Figure 1), with standard deviations of measurements of at most 30% of values14. With a sample size of 27, we would expect to have over 95% power (5% type I error) to detect a 3-fold difference using a Wilcoxon text at a minimum allele frequency of 7.4% (2/27). Similarly, we are able to detect a two-fold difference at a minimum allele frequency of 11.1% (3/27).
In a first analysis, we sought to internally validate our approach by using chloroquine resistance as reference. Indeed, we identified MAL7P1.27, which encodes the chloroquine resistance transporter (CRT), as the most significant association hit covered by four coding SNPs (P ≤ 10−4) on chromosome 7 (Table S1, Fig. S1).
Based on these reassuring results, we instituted a screen for associations with susceptibilities to 8 drugs. The following number of SNPs (genes, intergenic positions not listed) were lower than the computed significance threshold of 7 × 10−4 (Table 1): (i) 2 hits for PPQ (PF07_0019), (ii) 1 hit for LM (0), 5 hits for DHA (PFA0220w, PFB0560w, PFA0630c, PFF1445w), (iii) 21 hits for CQ 21 (MAL7P1.108 MAL7P1.27, PF11_0127, PFA0665w, PFC0690c, PFF0475w, PFI1560c, PFL2270w), (iv) 14 hits for QN (MAL13P1.333, PF13_021501 PF14_0215, PF14_0647, PF14_0726, PFD0700c, PFL0135w), (v) 1 hit for MQ (PFE1330c), (vi) 17 hits for PM (PF07_0107, PF10_0356, PF11_0271, PF11_0334, PFA0555c, PFA0650w, PFB0405w, PFC0705c, PFC0820w, PFC0970w, PFF0670w) and (vii) 19 for DEAQ (MAL8P1.157, PF07_0016, PF11_0327, PF11_0388, PF13_0104, PF13_0254, PFA0150c, PFA0510w). These hits were confirmed using the Spearman’s rank approach (Table S1). Of particular interest were two SNPs associated with DHA susceptibility on chromosome 13 at nucleotide positions 717855 and 1644675 (Wilcoxon, P = 5 × 10−5; Spearman’s rank, P = 5 × 10−5; Tables 1, S1) that represented the most significant hits across all comparisons. Equally of major interest was a coding SNP (C->G; K873R) in PFA0220w that was found to be associated with DHA sensitivity. A homolog of this gene was originally identified in P. chabaudi as determinant of parasite survival in artemisinin drug treated murine hosts (PCHAS_020720, encoding a putative deubiquitinase)15. Another SNP associated with DHA response implicated PFB0630c, a gene that has homology to stress-responsive RNA polymerase II-binding proteins16. There was some evidence for SNP associations in other candidate regions for the other tested drugs, including DHFR (PM, PFD0830w, P = 0.0173), MDR1 (MQ, PFE1150w, P = 0.0038), MRP2 (QN, PFL1410c, P = 0.0052), NHE-1 (QN, PF13_0019, P = 0.0033), but these did not exceed the stringent significance threshold (Table S2). Among the collected samples we did not find evidence of the MDR1 gene (PFE1150w) amplification, which had been found to be associated with MQ resistance17. Because it has recently been suggested that only a very limited number of genes may be involved in modifying drug susceptibility18, we studied the specificity of SNP hits for a given drug by querying the database for significant association with other drugs (Table S2). Not a single SNP hit was associated with more than one drug when using a moderate significance threshold for secondary associations (Table S2). There was a single hit for lumefantrine (MAL7P1.30) that occurred in a region highlighted by several hits for CQ on chromosome 7 (Table S2). The top 0.05% correlations (corresponding to either, rho > 0.68 or p < 0.0007) were retained from Table S1.
Signatures of recent positive selection
Drug pressure is a powerful selective force in natural Plasmodium populations19,20. It is well understood that positive selection acting on a beneficial trait gives rise to characteristic regions of low genetic diversity surrounding the causal genetic variant(s) due to the preservation of linkage disequilibrium during meiosis (recombination in regions of 17 kb is estimated to occur only in 1% of meioses during this life-cycle bottleneck in the mosquito mid-gut21). Here, we sought to identify regions of the genome under recent positive selection, as these may represent signatures of adaptation to drug pressure (Table 2Figure 3). To achieve this, we calculated the integrated haplotype score (iHS) for all 75 k SNPs across the whole genome, applying a stringent threshold (iHS > 3.6, top 0.2%). Again in an initial validation of the analytical approach using the established chloroquine resistance locus CRT as positive control, we found a large 45 kb region surrounding CRT that was characterized by lower than expected genetic diversity (PF07_0028 (2), PF07_0035 (6), PF07_0036 (1), PF07_0037 (1), MAL7P1.30 (1) and PF07_0042 (2)).
Manhattan plots of whole genome association tests.
X-axis is Chromosomes 1 to 14 in alternating colors; Y-axis is the −log10 p-value from a Wilcoxon test; points in blue indicate P-values less than 0.0007 (above horizontal dashed line); DHA dihydroartemisinin (Fig. 2A), LM lumefantrine (Fig. 2B), PPQ piperaquine (Fig. 2C), CQ chloroquine (Fig. 2D), PM pyrimethamine (Fig. 2E), MQ mefloquine (Fig. 2F), QN quinine (Fig. 2G), DEAQ desethylamodiaquine (Fig. 2H).
Evidence of recent positive selection.
We used the integrated Haplotype Score (iHS), where points above the horizontal dashed line indicated scores in excess of 3.6; x-axis is Chromosomes 1 to 14 in alternating colors; vertical lines correspond to chromosomal locations of DHFR, MDR1 and CRT, DHPS, respectively (left to right). Larger sized points indicate significant results in unique, non-telomeric and non-highly variable gene regions.
We identified the following genes located in such ‘valleys’ of low diversity (number of SNP hits) in the genome-wide scan: (i) PFA0205w (2), (ii) PFA0220w (UBP1-homologue) (1), (iii) PFC0935c (2) (coding for a putative N-acetylglucosamine-1-phosphate transferase), (iv) PFC0940c (1), (v) PFE1210c (1), (vi) PFF1350c (13) (coding for a putative member of the acetyl-CoA synthetase family22), (vii) PFF1365c (2), (viii) PFF1485w (1), (ix) PF07_0004 (3), (x) MAL7P1.207 (2), (xi) PF07_0066, (xii) a 50 kb region downstream of PfDHPS (MAL8P1.112 (1), MAL8P1.113 (5), PF08_0100 (1), (xiii) PFI0805w (1), (xiv) PF10_0015 (2), (xv) PF11_0074, (xvi) PF11_0420 (2), (xvii) PFL1525c (1), (xviii) PFL1835w (1), (xiix) PF14_0726 (3).
The |iHS| method may be insensitive to detect signatures of positive selection for polymorphisms that have reached fixation, we therefore proceeded to apply the cross-population extended haplotype score (XP-EHH) approach to compare the Kenyan to other P. falciparum populations (Burkina Faso, Cambodia, Mali, Thailand) to identify evidence for positive selection of alleles that have reached or are near fixation in individual populations23. The analysis confirms selection acting on PfCRT across all comparisons, but also at the PfDHPS locus across African populations (Fig. S2).
In our analysis of genomic regions with low diversity, we also found the current vaccine candidates MSP1 (2), AMA1 (4) (previously described by Mu et al.24) and TRAP (6). This was a surprising finding because these genes are thought to be targets of protective immunity and are known to contain extensive SNP and/or repeat polymorphisms. To obtain reassurance that our finding did not result from spurious genomic data, we implemented the Tajima’s D metric25, an approach for distinguishing between a DNA sequence evolving randomly (“neutrally”, values close to zero) and one evolving under a non-random process, including directional selection (low negative values) or balancing selection (high positive values). Indeed when calculating the Tajima’s D on a gene-by-gene basis we found 18 loci, including AMA1, MSP3, MSP3.8, MSP6 vaccine candidates (Table S3). These results could indicate the co-existence of reverse selective forces on different domains and/or upstream and downstream regulatory elements of the same gene. For instance, purifying selection could act on functional domains such as transmembrane stretches or functional motives while at the same time, diversifying selection acts on immunologically exposed extracellular loops26. Alternatively, the co-existence of hyper-variable ‘islands’ within regions of lower than expected diversity may point to a previously unrecognized feature of chromosome biology that is providing a pathway for diversification at amino acid residues or entire domains exposed to adaptive immune responses.
Association and selection by gene
Recent positive selection of survival-promoting genotypes, such as drug resistance-conferring mutations, should be detectable both by genotype-phenotype association and by ‘phenotype-free’ analysis of genomic structures (see above section on signatures of recent positive selection) as long as (i) selective pressure has had sufficient time to shape evolution or on the opposite end of the evolutionary time scale, (i) the causal genetic variant has not yet reached fixation in the population (i.e., close to 100% prevalence). We used a simple composite score (termed ‘total evidence score’, TES) calculated as the sum of the negative decadic logarithm (−log10) of the P-value for association and the iHS score for unusually large haplotypes for each of the 75,471 high-quality bi-allelic SNPs (Figure 4 and Table S4). Among the top twenty highest ranked SNPs, we found CRT (MAL7P1.27), Cg1 (immediately downstream of CRT) and UBP1. Of the 44 genes (1.7% of 2591 passing QC) identified by selection or association (Table S4), only UBP1, CRT and surrounding loci (PF07_0035) and PF14_0726 gene regions were identified by both approaches, providing stronger evidence for their role in modulating drug sensitivity. The biological relevance of a modest correlation between association P-values and selection tests (Spearman’s correlation 0.41) at a gene level in entire P. falciparum genomes is not clear and it may be an artifact stemming from limits to attain significance with low frequency variants in both tests (Fig. 4).
Scatter plot of evidence scores from genotype-phenotype association and genomic structure analyses.
Values of iHS or −log10 P-value from association testing are presented, with the gene names that exceed thresholds (blue: iHS > 3.6 or p < 0.0007; red: iHS > 3 or p < 0.001). The Spearman’s correlation is 0.417 (P < 0.00001).
The collected samples were all resistant to pyrimethamine and there is some evidence of a selective sweeps within 50 kb of the DHFR gene and in the 3’ region of DHPS (which encodes the target of sulfadoxine, involved in the combination therapy with pyrimethamine). The iHS metric is powered to detect sweeps only at intermediate frequency and prior to fixation. This could explain the failure to detect a stronger signal in our samples, all of which were resistant to pyrimethamine in vitro and carried the resistance-conferring gatekeeper mutation at codon position 108 (S108N).
Discussion
The identification of loci associated with malarial drug resistance has the potential to support disease surveillance systems and provide public health bodies with the information needed to deliver effective interventions. Here we studied associations between (i) whole-genome sequence variation at single-nucleotide resolution obtained through next-generation sequencing technology and (ii) robust drug susceptibility phenotypes obtained through repeat in vitro experiments with the aim to discover novel genes or genomic regions that modify drug susceptibility in 27 isolates of P. falciparum collected from Kenyan patients with malaria.
The power of our approach could be demonstrated in an initial proof-of-principle screen for chloroquine resistance-associated genes. The most significant association was found for the CRT gene that encodes the well-characterized chloroquine resistance transporter27,28. When extending the analysis to seven important antimalarial drugs, including dihydroartemisinin as both active metabolite and component of the current front-line artemisinin-based combination therapies, we identified several additional loci that were strongly associated with drug response phenotypes (Table 1 and Fig. 2A–H). Because of the urgency of the artemisinin resistance problem1,2,3, we focused on specific hits associated with the dihydroartemisinin response phenotype. We could confirm the previously reported association with PFA0220w, a homologue of UBP1 previously identified in a rodent malaria model and coding for a putative de-ubiquitinating protein15,29 and we identified three novel candidate genes (PFB0560w, PFB0630c, PFF0445w). Of note, our screen also identified a SNP (MAL13-1644675) located in a 35-kb segment on chromosome 13 that was recently linked to delayed in vivo clearance in P. falciparum infections from Western Thailand30. PFB0630c shares homology with the human RPAP2 protein and the yeast Rtr1 protein16 with putative regulatory roles in RNA polymerase II function. This may be of interest in the light of the reported differential expression pattern observed in isolates obtained from P. falciparum infections with delayed in vivo responses in Cambodia31. PFB0560w and PFF0445w are conserved Plasmodium protein coding genes with no assigned putative functions. PFF0445w had previously been reported to be up-regulated in response to artemisinin pressure in vitro in a comparative proteomics study32. The functional relevance of the chromosome 13 hit (MAL13-1644675; correlation rho = 0.7; P = 0.001), centered between the predicted open reading frames MAL13P1.211 (−1 kb, coding for a hypothetical protein with no predicted function) and PF13_0226 (1.7 kp, predicted to code for an inner membrane complex (IMC) protein) is not known.
We also screened for evidence of recent positive selection in the genomes of our samples. Of particular interest was a strong signal surrounding PFA0220w (UBP1-homologue) (Fig. S1). However, we did not detect a similar selection signal for MAL13-1644675 in a 35-kb segment on chromosome 13 that was recently linked to delayed in vivo clearance in P. falciparum infections from Western Thailand30. The fact that our screen identified an isolated association at this locus without a signal for recent positive selection may be explained by the evolutionary time point of sampling: artemisinin-based combination therapy was introduced as first-line treatment only 2–3 years before sampling started33. This hypothesis is supported by evidence for the chloroquine resistance gene CRT where both association and signature of selection are present in our data, most likely as a result of longstanding drug pressure19,34. The absence of significantly delayed P. falciparum infections in Kilifi after artemisinin treatment despite the moderate allele frequency of MAL13-1644675 in the local parasite population suggests that this, or yet unknown causal, genetic variants in this region on chromosome 13 are required but not sufficient for full blown in vivo artemisinin tolerance. Of note, a genome-wide analysis for associations of genotypes with the rate of parasite clearance after treatment with artemisinin-based combinations in patients who donated the P. falciparum isolates for this study (presence or absence of microscopically detectable blood stage parasites on day 29) did not reveal significant signals (Fig. S2).
In this study we used a panel of 8 commonly used antimalarial drugs to determine robust chemosensitivity phenotypes. This focus on in vitro data was motivated by a lack of correlation between the reported delayed in vivo response to artemisinins and the in vitro phenotype in most1,9, if not all2, studies. Whilst an in vivo phenotype would have been preferred, these phenotypes are difficult to measure and the outcome can be confounded by host genetic, immunity and intra-assay variation. In contrast, the IC50 values measured in 27 P. falciparum isolates obtained from pediatric patients in Kilifi District on the Kenyan Coast exhibited substantial phenotypic variation (mean >10-fold; Fig. 1) and a high degree of inter-assay reproducibility. The observed pattern of correlations between drug responses was also consistent with previously published data, reinforcing the confidence in the accuracy of the phenotypes.
In general, complex genetic traits may involve many genes, each of small effect magnitude. However, drug resistance in P. falciparum has been reported as strong single locus effects, with beneficial alleles rapidly going to fixation by selective sweeps leaving characteristic low-diversity ‘scars’ in the genomes of resistant parasites. In practice, that translates into smaller sample size requirements for detecting selection events, compared to association studies for complex traits35. To account for the potential number of false positives, we applied stringent quality control on the polymorphisms included, a conservative non-parametric testing and an adjusted statistical significance threshold. Our approach relied on natural variation in the parasite, leading to a set of strong candidates, including hitherto unexpected pathways. For instance, the associations of TRAP and of PF14_0647 (coding for a putative Rab GTPase activator) with sensitivity to quinine (a known ion channel blocker36,) (Table 1) may point to a role of membrane-associated trafficking in the mechanism of action of quinine.
Our approach is conceptually similar to a study by Mu et al.24 but with >20 times higher resolution of genetic variation and with a focus on a single local parasite population obtained from patients in a well-described cohort9,37 to reduce potential confounding by population structure. In contrast to Mu et al.24 we found one gene (PFA0655w) to be associated with chloroquine and not mefloquine or dihydroartemisinin, sensitivity and we failed to find evidence for MDR1. Another study by van Tyne et al.38 also reported on a genome-wide association study using an array-based genotyping strategy. There was partial overlap in the drugs used and specifically, we could not confirm a gene (PF14_0654) associated with artemisinin sensitivity, possibly related to a lack of power in our small sample size. A parallel study by Park et al.39 could also confirm the efficiency of massively parallel shot-gun sequencing by employing a related strategy designed to increase the resolution of an initial positive selection-based screen by using association test results39. In contrast to Park et al., however, we did not assume selection through drug pressure to be driving allele frequencies conferring tolerance to the artemisinins relatively shortly after the introduction of artemisinin-based combination chemotherapies. Consequently, we used genomic signatures of positive selection not as a primary screen but as an additional parameter for identifying genes and/or genomic regions associated with artemisinin response rates through non-parametric genotype-phenotype association tests.
In summary, our study in a limited number of P. falciparum isolates has shown that a natural genetics approach powered by whole genome sequencing using new short read technologies can identify novel chemosensitivity-determining genes, applied particularly within a robust genome-wide association and selection setting. These results show promise for geographically focused and timely sequence-based studies as a powerful and efficient tool in future disease surveillance programs. We found substantial overlap with previously reported artemisinin resistance-associated candidate genes and regions (prominently, PFA0220w, a UBP1-homologue and a 35-kb segment on chromosome 13). Because the studied isolates did not originate from artemisinin resistant infections, we hypothesize that the observed associations indicate standing variation that could serve as substrate for selection under continued drug pressure. It also provides a unique reference for the interpretation of results from resistant infections.
Methods
In vitro phenotypes
The study was approved by the National KEMRI Ethical Review Committee, Kenya; the Oxford Tropical Research Ethics Committee, UK; and the Ethics Committee, Heidelberg University School of Medicine, Germany. Parasite isolates were obtained in 2007 to 2008 from patients presenting with uncomplicated episodes of P. falciparum malaria before initiation of treatment with an artemisinin-based combination therapy (n = 13) and when patients experienced recurrence of infection during follow-up (n = 14)9. Cryo-preserved isolates were consecutively thawed and adapted to cell culture conditions. Parasites were cultured in complete medium (RPMI supplemented with L-glutamine, 2% heat-inactivated AB serum, 0.1 mM hypoxanthine, gentamicin and albumax II) in the presence of O+ or A+ blood at 5% packed cell volume and a gas mixture of 5% CO2, 5% O2 and 90% N2. Growth inhibition of parasite cultures at 0.5% packed cell volume and 0.1% parasitemia was determined on 96-well plates by exposure to serial dilutions of dihydroartemisinin (DHA, Sigma), lumefantrine (LM, Novartis), piperaquine (PPQ, SigmaTau), chloroquine (CQ, Sigma), pyrimethamine (PM, Sigma), mefloquine (MQ, Sigma), N-desethylamodiaquine (DEAQ, Sigma) and quinine (QN, Sigma). After incubation at 37°C for 72 hours, 20 nM SYBR green in lysis buffer (20 mM Tris at pH 7.5, 5 mM EDTA, 0.008% (wt/vol) saponin and 0.08% (vol/vol) Triton X-100) (doi:10.1128/AAC.01607-06) was added and fluorescence intensity measured at 20 nm (model, manufacturer). Growth inhibition experiments were repeated at least twice (mean, 3.1). Half-maximal inhibitory concentrations (IC50) were estimated by non-linear regression.
Sequencing and genetic variant analysis
All samples (n = 27) underwent whole genome sequencing, with 54 or 76-base paired end fragment sizes, using Illumina technology (see40 for a description) and processed as previously described41 to identify variation including SNPs and small insertions and deletions. In brief, we mapped all isolates to the 3D7 (version 3.0) reference genome using smalt (5) and called variants using samtools (6). Sequence polymorphisms were identified empirically using sequence coverage data as previously described (4). The internal replicability and correlation between in vitro phenotypes was assessed using Spearman’s correlation. Geographical outliers were identified using a principal component clustering approach applied to multi-continental SNP data (40, Short read archive (SRA) Study ERP000190). The integrated haplotype score (iHS, (12)) method was applied to SNPs data to identify long-range directional selection. The selection metric Tajima’s D25 was used for distinguishing between a DNA sequence evolving randomly (“neutrally”, values close to 0) and one evolving under a non-random process, including directional selection (low negative values) or balancing selection (high positive values). Because of the non-symmetry of phenotypes, the primary assessment of association between phenotypes and genetic variants (alleles) used Wilcoxon rank tests. A secondary analysis applied Spearman’s correlation. A statistical significance cut-off (P = 0.0007, −log10 P = 3.15) was inferred by simulation (phenotype-based permutation) to represent a multiple test adjustment of a nominal 5% error rate. For the final hits, we considered only genomic variants in regions that were unique (calculated by sliding 50 base pairs of contiguous sequence across the reference genome), non-subtelomeric and not in highly variable gene families (rifins, surfins, stevors and vars). Regions for follow-up were compared to publically available sequence data40,42. All raw sequencing data for this work is contained in SRA study ERP000190.
References
Dondorp, A. M. et al. Artemisinin resistance in Plasmodium falciparum malaria. N Engl J Med 361, 455–67 (2009).
Noedl, H. et al. Evidence of artemisinin-resistant malaria in western Cambodia. N Engl J Med 359, 2619–20 (2008).
Phyo, A. P. et al. Emergence of artemisinin-resistant malaria on the western border of Thailand: a longitudinal study. Lancet (2012).
Eastman, R. T. & Fidock, D. A. Artemisinin-based combination therapies: a vital tool in efforts to eliminate malaria. Nat Rev Microbiol 7, 864–74 (2009).
Nayyar, G., Breman, J., Newton, P. & Herrington, J. Poor-quality antimalarial drugs in southeast Asia and sub-Saharan Africa. Lancet Infect Dis 12, 488–496 (2012).
Rathod, P. K., McErlean, T. & Lee, P. C. Variations in frequencies of drug resistance in Plasmodium falciparum. Proc Natl Acad Sci U S A 94, 9389–93 (1997).
Bjorkman, J., Nagaev, I., Berg, O. G., Hughes, D. & Andersson, D. I. Effects of environment on compensatory mutations to ameliorate costs of antibiotic resistance. Science 287, 1479–82 (2000).
Mackinnon, M. J. & Marsh, K. The selection landscape of malaria parasites. Science 328, 866–71 (2010).
Borrmann, S. et al. Declining responsiveness of Plasmodium falciparum infections to artemisinin-based combination treatments on the Kenyan coast. PLoS ONE 6, e26005 (2011).
Sidhu, A. B. et al. Decreasing pfmdr1 copy number in plasmodium falciparum malaria heightens susceptibility to mefloquine, lumefantrine, halofantrine, quinine and artemisinin. J Infect Dis 194, 528–35 (2006).
Wongsrichanalai, C., Pickard, A. L., Wernsdorfer, W. H. & Meshnick, S. R. Epidemiology of drug-resistant malaria. Lancet Infect Dis 2, 209–18 (2002).
Witkowski, B. et al. Novel phenotypic assays for the detection of artemisinin-resistant Plasmodium falciparum malaria in Cambodia: in-vitro and ex-vivo drug-response studies. Lancet Infect Dis (2013).
Gardner, M. J. et al. Genome sequence of the human malaria parasite Plasmodium falciparum. Nature 419, 498–511 (2002).
van Schalkwyk, D. A. et al. Culture-adapted Plasmodium falciparum isolates from UK travellers: in vitro drug sensitivity, clonality and drug resistance markers. Malar J 12, 320 (2013).
Hunt, P. et al. Experimental evolution, genetic analysis and genome re-sequencing reveal the mutation conferring artemisinin resistance in an isogenic lineage of malaria parasites. BMC Genomics 11, 499 (2010).
Gibney, P. A., Fries, T., Bailer, S. M. & Morano, K. A. Rtr1 is the Saccharomyces cerevisiae homolog of a novel family of RNA polymerase II-binding proteins. Eukaryot Cell 7, 938–48 (2008).
Price, R. N. et al. Mefloquine resistance in Plasmodium falciparum and increased pfmdr1 gene copy number. Lancet 364, 438–47 (2004).
Yuan, J. et al. Chemical genomic profiling for antimalarial therapies, response signatures and molecular targets. Science 333, 724–9 (2011).
Wootton, J. C. et al. Genetic diversity and chloroquine selective sweeps in Plasmodium falciparum. Nature 418, 320–3 (2002).
Roper, C. et al. Intercontinental spread of pyrimethamine-resistant malaria. Science 305, 1124 (2004).
Su, X. et al. A genetic map and recombination parameters of the human malaria parasite Plasmodium falciparum. Science 286, 1351–3 (1999).
Bethke, L. L. et al. Duplication, gene conversion and genetic diversity in the species-specific acyl-CoA synthetase gene family of Plasmodium falciparum. Mol Biochem Parasitol 150, 10–24 (2006).
Sabeti, P. C. et al. Genome-wide detection and characterization of positive selection in human populations. Nature 449, 913–8 (2007).
Mu, J. et al. Plasmodium falciparum genome-wide scans for positive selection, recombination hot spots and resistance to antimalarial drugs. Nat Genet 42, 268–71 (2010).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–95 (1989).
Amambua-Ngwa, A. et al. SNP genotyping identifies new signatures of selection in a deep sample of West African Plasmodium falciparum malaria parasites. Mol Biol Evol 29, 3249–53 (2012).
Fidock, D. A. et al. Mutations in the P. falciparum digestive vacuole transmembrane protein PfCRT and evidence for their role in chloroquine resistance. Mol Cell 6, 861–71 (2000).
Sidhu, A. B., Verdier-Pinard, D. & Fidock, D. A. Chloroquine resistance in Plasmodium falciparum malaria parasites conferred by pfcrt mutations. Science 298, 210–3 (2002).
Hunt, P. et al. Gene encoding a deubiquitinating enzyme is mutated in artesunate- and chloroquine-resistant rodent malaria parasites. Mol Microbiol 65, 27–40 (2007).
Cheeseman, I. H. et al. A major genome region underlying artemisinin resistance in malaria. Science 336, 79–82 (2012).
Mok, S. et al. Artemisinin resistance in Plasmodium falciparum is associated with an altered temporal pattern of transcription. BMC Genomics 12, 391 (2011).
Prieto, J. H., Koncarevic, S., Park, S. K., Yates, J., 3rd & Becker, K. Large-scale differential proteome analysis in Plasmodium falciparum under drug treatment. PLoS ONE 3, e4098 (2008).
Dondorp, A. M. et al. Artemisinin resistance: current status and scenarios for containment. Nat Rev Microbiol 8, 272–80.
Mwai, L. et al. Chloroquine resistance before and after its withdrawal in Kenya. Malar J 8, 106 (2009).
Anderson, T., Nkhoma, S., Ecker, A. & Fidock, D. How can we identify parasite genes that underlie antimalarial drug resistance? Pharmacogenomics 12, 59–85 (2011).
DeCoursey, T. E., Chandy, K. G., Gupta, S. & Cahalan, M. D. Voltage-gated K+ channels in human T lymphocytes: a role in mitogenesis? Nature 307, 465–8 (1984).
Olotu, A. et al. Defining clinical malaria: the specificity and incidence of endpoints from active and passive surveillance of children in rural Kenya. PLoS ONE 5, e15569 (2010).
Van Tyne, D. et al. Identification and functional validation of the novel antimalarial resistance locus PF10_0355 in Plasmodium falciparum. PLoS Genet 7, e1001383 (2011).
Park, D. J. et al. Sequence-based association and selection scans identify drug resistance loci in the Plasmodium falciparum malaria parasite. Proc Natl Acad Sci U S A 109, 13052–7 (2012).
Manske, M. et al. Analysis of Plasmodium falciparum diversity in natural infections by deep sequencing. Nature 487, 375–9 (2012).
Robinson, T. et al. Drug-resistant genotypes and multi-clonality in Plasmodium falciparum analysed by direct genome sequencing from peripheral blood of malaria patients. PLoS ONE 6, e23204 (2011).
MalariaGen. Sequence Read Archive. (Accessed on 10 Oct 2013). http://www.malariagen.net/data/sequence-read-archive.
Acknowledgements
We thank Ines Petersen for helpful comments and Bronwyn MacInnis for assistance. This work is published with the permission of the Director of KEMRI. This work was supported by a German Research Foundation (DFG) grant to S.B. The funding source had no role in the design of the study; in the collection, analysis and interpretation of data; in the writing of the report; and in the decision to submit the paper for publication.
Author information
Authors and Affiliations
Contributions
S.B. and T.C. wrote the manuscript. S.B., A.N., D.K. and T.C. designed the study. S.B., J.S. and T.C. analysed the data. L.M., A.A., A.R., J.O., S.M., M.M.K., S.C. and S. Auburn generated data. P.S. and B.L. oversaw sample collection. S. Assefa, M.M. and G.M. provided data analysis tools and assisted data analysis. N.P. and K.M. coordinated the clinical study. All authors have reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Borrmann, S., Straimer, J., Mwai, L. et al. Genome-wide screen identifies new candidate genes associated with artemisinin susceptibility in Plasmodium falciparum in Kenya. Sci Rep 3, 3318 (2013). https://doi.org/10.1038/srep03318
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep03318
This article is cited by
-
Deaggregation of mutant Plasmodium yoelii de-ubiquitinase UBP1 alters MDR1 localization to confer multidrug resistance
Nature Communications (2024)
-
MalariaSED: a deep learning framework to decipher the regulatory contributions of noncoding variants in malaria parasites
Genome Biology (2023)
-
A systematic review on anti-malarial drug discovery and antiplasmodial potential of green synthesis mediated metal nanoparticles: overview, challenges and future perspectives
Malaria Journal (2019)
-
Are k13 and plasmepsin II genes, involved in Plasmodium falciparum resistance to artemisinin derivatives and piperaquine in Southeast Asia, reliable to monitor resistance surveillance in Africa?
Malaria Journal (2019)
-
K13-propeller gene polymorphisms of Plasmodium falciparum and the therapeutic effect of artesunate among migrant workers returning to Guangxi, China (2014–2017)
Malaria Journal (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.