Abstract
The limited locations of tRNA introns are crucial for eukaryal tRNA-splicing endonuclease recognition. However, our analysis of the nuclear genome of an early-diverged red alga, Cyanidioschyzon merolae, demonstrated the first evidence of nuclear-encoded tRNA genes that contain ectopic and/or multiple introns. Some genes exhibited both intronic and permuted structures in which the 3′-half of the tRNA coding sequence lies upstream of the 5′-half and an intron is inserted into either half. These highly disrupted tRNA genes, which account for 63% of all nuclear tRNA genes, are expressed via the orderly and sequential processing of bulge-helix-bulge (BHB) motifs at intron-exon junctions and termini of permuted tRNA precursors, probably by a C. merolae tRNA-splicing endonuclease with an unidentified subunit architecture. The results revealed a considerable diversity in eukaryal tRNA intron properties and endonuclease architectures, which will help to elucidate the acquisition mechanism of the BHB-mediated disrupted tRNA genes.
Similar content being viewed by others
Introduction
Cyanidioschyzon merolae 10D is an ultrasmall unicellular red alga that inhabits an extreme environment (pH 1–3, 40–50°C) and represents one of the most ancestral forms of eukaryotes1,2,3. A primary search of the complete 16.5 Mbp nuclear genome sequence predicted only 30 tRNA genes, which are insufficient to decode 61 codons2. This paucity of tRNA genes was partly solved by our discovery of a distinctive type of disrupted tRNA gene; we identified 11 circularly permuted tRNA genes, in which the 3′-half of the tRNA coding sequence lies upstream of the 5′-half in the genome4. Biochemical analyses defined a processing pathway in which the termini of the tRNA precursor (pre-tRNAs) are ligated to form a characteristic circular RNA intermediate, which is then processed at the acceptor stem to generate the functional termini4. Notably, the sequences adjacent to the termini in these pre-tRNAs potentially form a bulge-helix-bulge (BHB) motif, which was originally found around the intron-exon junctions of nuclear and archaeal tRNAs5. To date, permuted tRNAs have only been found in some unicellular algae and archaea6,7. These observations suggest that BHB motifs and the tRNA intron splicing system by which they are processed, must have been a prerequisite for the development of such disrupted tRNA genes in early-rooted eukaryotes and archaea.
Here, we report the identification of an alternative type of BHB-mediated disrupted tRNA gene that harbors atypical introns. Introns found in nuclear and archaeal tRNAs are cleaved by the proteinaceous tRNA-splicing endonuclease8,9,10,11. Nuclear tRNA introns are generally short, comprise a relaxed BHB motif and are located exclusively between positions 37 and 38 (37/38), which is 3′ adjacent to the anticodon (the canonical position)5,12. This limited location of the intron is considered crucial for the precise recognition of pre-tRNA by the eukaryal tRNA-splicing endonucleases8,9,10,11,13,14,15,16,17,18. However, we found that a number of C. merolae nuclear-encoded tRNA genes contain single or multiple ectopic introns, which has not been reported in eukaryotes. Some genes contained both intronic and permuted gene structures. Further analysis revealed that the BHB motifs in pre-tRNAs are processed in an order that correlates with the theoretical free energy and the position of each motif in the cloverleaf structure. It also indicated that the C. merolae tRNA-splicing endonuclease may show a non-canonical subunit composition. Our findings provide a new insight into the tRNA processing system in eukaryotes and a deeper understanding of the evolutionary aspects that governed the formation of BHB-mediated disrupted tRNA genes.
Results
Approximately 63% of tRNA genes in the C. merolae nuclear genome are disrupted by introns
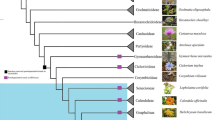
A genome-wide search for tRNA genes was performed using the SPLITS and SPLITSX programs to detect cis-spliced tRNAs (intron-containing tRNAs) and trans-spliced tRNAs (split tRNAs)19,20,21,22. A BLAST search was performed to detect conserved tRNA sequences. This identified 27 intronic tRNA genes, including nine newly identified and 12 revised genes (Fig. 1). Among them, 12 appeared to be the first examples of nuclear-encoded tRNA genes containing multiple introns: nine genes with two introns, three with three introns and 15 with a single intron. Four genes encoding tRNALeu(UAA), tRNAArg(CCU) and two copies of tRNAGly(CCC) had both intronic and permuted structures, meaning that their pre-tRNAs required removal of an intron as well as swapping of the 5′- and 3′-halves. Our search for BHB-mediated disrupted tRNA genes led to the identification of a sufficient repertoire of anticodons. C. merolae contains disrupted tRNA genes that exhibit intronic (23/43), permuted (7/43), or both (4/43) structures that account for 79.1% (34/43) of all nuclear tRNA genes. The proportion of intronic tRNA genes in C. merolae (62.8%; 27/43) is much higher than that found in other eukaryotes12, including Saccharomyces cerevisiae (21.5%, 59/275), Arabidopsis thaliana (13.2%, 83/630), Drosophila melanogaster (5.2%, 15/290) and Homo sapiens (4.3%, 28/648). However, it is comparable to the levels found in several archaea (up to 87%), which contain the highest number of introns in any organism7,22.
Anticodons in C. merolae.
Newly identified and revised tRNA genes are indicated in red and blue, respectively. All anticodons are presented in the 5′ to 3′ direction. Intron numbers are shown in parentheses. Anticodons of tRNAs encoded by a permuted gene are boxed. Two distinct genes that produce identical mature tRNA sequences encode tRNAGly(CCC) (indicated by an asterisk). iMet and eMet mean initiator and elongator tRNAMet(CAU), respectively.
The genomic intron sequences varied from 11–69 bp; tRNASer(AGA) and tRNAAsn(GTT) contained the shortest introns, 11 bp, in the anticodon-loop, while tRNAArg(TCT) contained the longest introns, 69 bp, in the D-loop (Fig. 2).
Distribution of BHB motifs in intronic and/or permuted tRNAs in C. merolae.
(A) Arrows indicate the positions of the BHB motifs. The length of the intron, number and type of BHB motifs is indicated for each tRNA. For example, “Ile(TAT) 22 3/3 hBHBh′” at the position between 53 and 54 means that the BHB is from tRNAIle(UAU), is 22 nt long and is the third of three BHB motifs. BHB motifs at the termini of the permuted pre-tRNAs are shown in double squares. The numbering of the tRNA positions follows that outlined in Reference 12. (B) The BHB motif is classified by one or two 3-nt-bulges (denoted as B = 3) separated by a central 4-bp-helix (denoted as H = 4) and flanked by two helices (denoted as h or h′), each with more than two base-pairings.
Eukaryal tRNA genes are generally transcribed by RNA polymerase III via an intragenic bipartite promoter consisting of A and B boxes, which are conserved sequences in the D-arm and TΨC-arm, respectively23. Transcription is initiated by the binding of a multi-subunit transcription factor (TFIIIC) to the promoter sequence24. However, the A and B boxes of atypical intronic tRNA genes in C. merolae are not recognized uniformly by the transcription machinery because of the interruption of either box by the intron and the variable intron sequences between the boxes, making the positional relation of the A and B boxes inadequate for binding of TFIIIC. This is also true for the permuted tRNA genes, in which the A and B boxes are not recognized by TFIIIC because they are inverted4. Additionally, no homologs of the TFIIIC components that bind to the A and B boxes have been identified in C. merolae. Instead, a TATA-like sequence and a T-stretch, which are probably the promoter and termination signal, respectively24,25, are found near most tRNA genes (see Supplementary Fig. S1 online). No promoter or termination signal is found between the exon segments of the intronic tRNA genes, suggesting that a set of putative tRNA segments is transcribed as a linear RNA. The above observations indicate that C. merolae employs the upstream TATA-like sequence and a non-canonical transcription system independent of TFIIIC, which would allow transcription of various types of disrupted (intronic, permuted and both) and non-disrupted tRNA genes.
C. merolae tRNA introns form various types of BHB motifs and are scattered throughout the cloverleaf structure
In contrast to the known nuclear tRNA genes whose introns are located exclusively at the canonical 37/38 position12, 42 introns in 27 tRNA genes of C. merolae were randomly distributed along the cloverleaf structure (Fig. 2). Although some (17 introns) were found at the 37/38 position, others were observed at novel positions for nuclear tRNA: the D-stem (five introns), the D-loop (14 introns), the variable-arm (one intron), the TΨC-stem (one intron) and the TΨC-loop (four introns). The finding that C. merolae nuclear tRNA genes contain introns at various positions is inconsistent with a requirement for introns to be located at particular positions such that they are recognized by known tRNA-splicing endonucleases in eukaryotes9,10.
The intron-exon junctions of nuclear pre-tRNAs generally form a relatively relaxed BHB motif. This is denoted as the hBH or BHL (BHB-like) motif and consists of a single 3-nt bulge and an internal loop separated by a 4-nt helix5,12. In C. merolae, the hBH motif is often located at position 37/38 (Fig. 2), as represented by tRNAAsn(GUU) (Fig. 3A). Our analysis revealed other types of BHB motifs at position 37/38, such as a strict hBHBh′ motif in tRNAIle(UAU) (Figs. 2 and 3C) and an unstructured BHB motif with no central 4-bp helix H in tRNAPhe(GAA) (Figs. 2 and 3B). Various types of BHB motifs were also identified at positions outside 37/38, including hBHBh′, BHL (hBH and HBh′) and a motif without H (no H) (Fig. 2).
RT-PCR amplification of intronic tRNAs.
Inferred pre-tRNA secondary structures for three tRNA genes are shown. (A) tRNAAsn(GUU) with a single intron at the canonical position 37/38. (B) tRNAPhe(GAA) with two introns, one in the D-stem (intron 1) and the other at 37/38 (intron 2). (C) tRNAIle(UAU) with three introns, one in the D-stem (intron 1), one at 37/38 (intron 2) and the third in the TΨC-loop (intron 3). 5′- and 3′-primers for RT-PCR are indicated as solid and broken arrows, respectively. Arrowheads indicate the positions to be processed. Intron sequences are shown in lower case. The numbering of the tRNA positions follows Reference 12. (D–F) PCR products amplified from the cDNA of processing intermediates and mature forms are indicated. Annotation of each PCR product was based on gel purification of the bands, cloning and sequencing analysis. Predicted sizes for each PCR product are described in Supplementary Table 1 online. Although I3′-primer 2 was designed to hybridize to a region within intron 3, the I5′-primer 3 and I3′-primer 2 combination produced a PCR product representing pre-tRNAIle (+intron 2) and mature tRNAIle (lane 3). This is attributable to the tendency of the I3′-primer 2 to hybridize to the TΨC-stem of tRNAIle(UAU), because of sequence similarity between the TΨC-stem and 3′ region of intron 3. The asterisk corresponds to a PCR product of pre-tRNAIle (+intron 2, 3) found in lane 3 (see details in the methods for the retardation of band mobility). Lane M: DNA molecular weight marker (ΦX174/HaeIII).
These features of tRNA genes in C. merolae – the localization of introns at positions other than 37/38 and identification of various types of BHB motifs – have never before been observed in a nuclear genome, although they do occur in some archaeal phyla, such as Crenarchaeota and Nanoarchaeota5,12,26,27,28.
Processing of the pre-tRNAs derived from disrupted genes
We performed northern blotting analysis using C. merolae total RNA to verify whether the highly disrupted tRNA genes are indeed transcribed in the cell. Northern blots using probes complementary to the 3′-halves of tRNAAsn(GUU), tRNAPhe(GAA) and tRNAIle(UAU), which harbor one, two and three introns, respectively, detected bands of the appropriate size (Supplementary Fig. S2 online), demonstrating that C. merolae disrupted tRNA genes are expressed. Their different mobilities in Fig. S2D, despite their almost identical predicted size, have been observed in PAGE and Northern blot analysis of tRNAs4,29,30,31 and it may be attributed to differences in their overall secondary structures, which may be unfolded even in the denaturing gel and to different post-transcriptional modifications. No bands for pre-tRNAs harboring unspliced introns were detected, suggesting that intron processing occurs rapidly in the cells, making the processing intermediates so scarce that they are nearly impossible to detect, even by northern blotting. In fact, it was reported that the over-expression of tRNA genes or knockdown of the processing enzymes in the cell is required to detect precursors or processing intermediates of tRNAs and non-coding RNAs by PAGE or Northern blotting32,33,34.
The introns in most C. merolae pre-tRNAs can form a BHB motif independently. These introns are not nested, as occurs in some archaea, in which the last intron can form a BHB motif only after the other introns have been processed26,35. To analyze the processing intermediates and to clarify the processing pathway in C. merolae, reverse transcription polymerase chain reaction (RT-PCR) was performed on three tRNAs and the resulting products were sequenced (Fig. 3). PCR products indicated by arrows in Fig. 3 were annotated by gel purification of the bands, cloning and sequencing analysis (Supplementary Table 1 online, Supplementary Figs. 3–5 online). While the signal representing unprocessed tRNAs was not detected in the northern blot analysis (Supplementary Fig. S2 online), RT-PCR products derived from various processing intermediates were detected with various band intensities (Fig. 3). Amplification of the RT-PCR products of tRNA processing intermediates is influenced by the amount of the target molecule and by the modification of nucleotides, which may inhibit the reverse transcription reaction36,37,38. Thus, the intensities of the PCR bands do not always reflect the amount of the target RNA molecules present in the cell. The bands for intermediates would be detectable because of significant amplification of a small amount of cDNA by PCR.
tRNAAsn(GUU) contains a single intron composed of an hBH motif at position 37/38 (Fig. 3A). RT-PCR using N5′-primer 1 and N3′-primer 1 (indicated by arrows in Fig. 3A) amplified both a pre-tRNA containing the intron and a processed form (mature form) (Fig. 3D, lane 1).
tRNAPhe(GAA) contains two introns, in the D-stem (intron 1) and at position 37/38 (intron 2), which comprise a hBHBh′ and a no H motif, respectively (Fig. 3B). RT-PCR with F5′-primer 1 and F3′-primer 1 produced PCR products representing a pre-tRNA with two introns, a processing intermediate in which intron 1 was removed but intron 2 was retained and a processed form (mature form) in which both intron 1 and 2 were removed (Fig. 3E, lane 1). No product was detected for an alternative intermediate in which intron 2 was removed but intron 1 was retained, even when using a primer set (5′-primer 2 and 3′-primer 2) specifically designed to amplify such an intermediate (Fig. 3E, lane 2). Although it remains possible that an intermediate harboring only intron 1 was produced but not detected because of technical difficulties, the above results suggest a pathway in which removal of intron 1 occurs before removal of intron 2. We then calculated the free energies of the BHB motifs to examine whether the processing pathway correlates with the stability of the BHB motifs. The free energies (ΔGs) of the BHB motifs in introns 1 and 2 from pre-tRNAPhe(GAA) were calculated as −11.1 kcal mol−1 and 1.15 kcal mol−1, respectively. This result implies that the processing of intron 1, in which the ΔG of the BHB motif is much lower, precedes that of intron 2.
Analysis of tRNAIle(UAU), which contains three introns with hBHBh′ motifs in the D-stem (intron 1), position 37/38 (intron 2) and the TΨC-loop (intron 3) (Fig. 3C), verified the sequences of a primary transcript containing all three introns (Fig. 3F, lane 2), an intermediate in which only intron 3 was removed (Fig. 3F, lane 2), an intermediate in which only intron 1 was removed (Fig. 3F, lane 3), an intermediate in which only intron 2 was retained (Fig. 3F, lane 3) and a fully processed form (mature form) in which all three introns were removed (Fig. 3F, lanes 1 and 3). No product was produced for an intermediate in which intron 2 was removed before intron 1 and 3 (Fig. 3F, lanes 1–3). These results show that the processing of introns 1 and 3 precedes that of intron 2. Processing intermediates containing introns 1 and 2 or introns 2 and 3 indicate that the removal of intron 1 or 3 can occur in either order. The ΔGs of the BHB motifs of introns 1, 2 and 3 were calculated as −11.2 kcal mol−1, −3.5 kcal mol−1 and −10.9 kcal mol−1, respectively. Thus, like tRNAPhe(GAA), the processing of pre-tRNAIle(UAU) begins with removal of introns 1 or 3, in which the ΔG of the BHB motif is lower, before intron 2 is removed. The band indicated by an asterisk (Fig. 3 F, lane 3), which was estimated to be more than 118 bp, was identified to be derived from pre-tRNAIle (+intron 2, 3) and was 116 bp in length. Such an inconsistency between the gel mobility and the actual size of PCR products was probably observed because of irregular structures of in the DNA. Annealing DNA strands with complementary repeated sequences sometimes results in DNA species such as cruciform or slipped DNA, which migrate slower in a polyacrylamide gel39,40; however, such an event in plasmid DNAs is rare41. In our analysis, PCR products consisting of tRNA sequence with stem-loop structures might have caused unknown structural abnormalities, leading to inconsistent mobility in gel electrophoresis.
The potential folding of BHB motifs is also found at the junction of the processing sites in permuted pre-tRNAs. tRNAGly(CCC), possessing both intronic and permuted structures (Fig. 4A–C, Supplementary Table 1 online, Supplementary Fig. 6 online), was analyzed to determine which BHB motif is processed first. RT-PCR amplified a product derived from a circular RNA intermediate in which the leader and trailer sequences at the RNA termini were processed, while the intervening sequence was retained (Fig. 4D). The circular intermediate lacked intron 1 in the TΨC-loop (Fig. 4E), suggesting that the BHB motif of intron 1 is processed before that of the termini in pre-tRNAGly(CCC). The ΔG of the BHB motif of intron 1 was calculated as −4.8 kcal mol−1, which is slightly lower than that of the BHB motif at the termini (−4.43 kcal mol−1). Thus, the processing of the BHB motif of permuted pre-tRNA termini may also follow the order suggested for intronic tRNAs in Fig. 3.
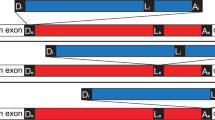
RT-PCR amplification of intronic and permuted pre-tRNAGly(CCC).
(A) Schematic representation of a permuted tRNA gene harboring an intron; tRNALeu(UAA) and tRNAArg(CCU) in the upper panel, tRNAsGly(CCC) in the lower panel. (B, C) The inferred secondary structures of pre-tRNAGly(CCC)a and pre-tRNAGly(CCC)b are shown. The sequences of the 5′-half (blue) and the 3′-half (red) of mature tRNA and the intervening sequence (green) of pre-tRNA are shown. Arrowheads indicate the processing positions. Intron sequences are shown in lower case. The numbering of the tRNA positions follows Reference 12. (D) The PCR product of cDNA generated from reverse transcription around (E), a circular intermediate of tRNAGly(CCC) without the intron, is indicated as CI-tRNAGly (Δintron 1). 5′- and 3′-primers are indicated as solid and broken arrows, respectively. Lane M: DNA molecular weight marker (ΦX174/HaeIII).
The above results implied a correlation between the processing order of introns and the theoretical free energy of BHB motifs in introns. This may explain why the removal of multiple BHB motifs from pre-tRNAs occurs in an order, even though each BHB motif can fold independently. Alternatively, it may depend on the position of the BHB motif, because the BHB motif in the anticodon arm is always the final substrate observed in tRNAPhe(GAA), tRNAIle(UAU) and tRNAGly(CCC). In the latter case, the BHB motif at position 37/38 can be recognized by the C. merolae endonuclease only after the BHB motifs at the other positions have been processed. These possibilities have to be further addressed by in vitro splicing analysis.
Unique subunit composition of C. merolae tRNA-splicing endonuclease
The tRNA-splicing endonuclease of the yeast S. cerevisiae is one of the best-studied eukaryal tRNA-processing enzymes. It possesses a heterotetrameric structure (αβδε) comprising two catalytic subunits (Sen2 and Sen34) and two accessory subunits (Sen15 and Sen54)9,10,11,16. Sen2 and Sen34 are homologs of the catalytic subunits of archaeal endonucleases. The interactions between Sen2 and Sen54 and between Sen15 and Sen34 were clarified by a yeast two-hybrid (YTH) experiment16. These four subunits function cooperatively to recognize cleavage sites via “a ruler mechanism”, in which the endonuclease measures a specified distance to where the cuts should be made in a pre-tRNA9,10,15,16,17. In addition to the localized structural fold of the BHB motif, this mechanism requires the specific recognition of the mature domain in pre-tRNAs. Thus, the yeast endonuclease would have coevolved with the introns exclusively localized at position 37/38. A search of the C. merolae genome identified three homologs of the yeast endonuclease subunits, cmSen2 (CMN231C), cmSen34 (CMH233C) and cmSen54 (CMK254C), comprising 246, 435 and 348 amino acids, respectively (see Supplementary Fig. S7 online). According to their isolated cDNA sequences, these candidates are predicted to be functionally active2. However, no apparent homolog of Sen15 was identified, which would appear to conflict with the notion that all four subunits are essential for the function of the endonuclease. In yeast, Sen15 interacts with Sen34 to aid the proper positioning of the 3′-splice site8,9,15,16,17.
To find a functional homolog of Sen15 or another potential subunit of the C. merolae endonuclease, a YTH screening was carried out. The full-length cDNA cmSen2, cmSen34, or cmSen54 was cloned into a bait protein plasmid and positive clones were screened from a C. merolae genomic cDNA library. Screening using cmSen2 (CMN231C) as a bait protein allowed the isolation of the cmSen54 (CMK254C) fragments from two independent prey clones (24 to 348 or 43 to 348 amino acids). However, no other interactions were obtained in this analysis. To confirm their specific interaction, we cloned the full-length cDNA of the three cmSen candidates and tested them using a YTH matrix experiment. The reciprocal interaction between cmSen2 and cmSen54 was clearly detected (Fig. 5). However, no pairwise interaction was detected for cmSen34, implying that the C. merolae endonuclease has a noncanonical subunit composition, which could not be clarified by YTH analysis. The C. merolae endonuclease may contain an unidentified subunit with low homology to known Sen15 proteins. It is also possible that the C. merolae endonuclease comprises a novel heterotrimeric complex. Given that Sen15 (129 amino acids) functions cooperatively with Sen34 in yeast and that cmSen34 (435 amino acids) is larger than yeast Sen34 (275 amino acids), cmSen34 may fulfill the structural and functional roles of both Sen34 and Sen15; however, there is no sequence similarity between yeast Sen15 and C. merolae Sen34. Alternatively, the C. merolae endonuclease may function as a dimer comprising catalytic subunits depending on the substrate to cleave. We are currently analyzing this process in vitro using two or three components of the recombinant endonuclease. In addition, we are planning to perform co-immunoprecipitation analysis to identify the active form of the endonuclease. Further analysis is required to understand the C. merolae tRNA-splicing endonuclease and its processing mechanism, which would be essential for the maturation of intronic and/or permuted pre-tRNAs.
Specific interactions between C. merolae Sen proteins.
A specificity test using YTH analysis was carried out on SC plates lacking Leu and Trp (-LW), Leu, Trp and adenine (-LWA), or Leu, Trp and His containing 1 mM 3AT (-LWH + 1 mM 3AT) with a 4-day incubation. Diploid strains were constructed by mating PJ69-4Aa cells harboring pGBTK or its derivatives (Baits) with PJ69-4Aα cells harboring pGAD or its derivatives (Preys); each derivative plasmid contained full-length cmSen2 (CMN231C), cmSen34 (CMH233C), or cmSen54 (CMK254C).
Discussion
In C. merolae, disrupted tRNA genes that exhibit intronic (23/43), permuted (7/43) or both (4/43) structures account for 79.1% (34/43) of all nuclear tRNA genes. Some introns of nuclear-encoded tRNAs are required for post-transcriptional modification9 or regulation of the cell cycle42, while some are dispensable43. It has also been suggested that fragmented tRNA genes may prevent the integration of mobile elements44,45. Thus, the conservation of various types of disrupted tRNA genes in the streamlined genome of C. merolae, which require more extensive processing, may suggest that the fragmentation of tRNA genes or the BHB motif itself are significant for cell function. Alternatively, disrupted tRNA genes might have developed by neutral evolution, or may be a remnant of the process for decreasing the number of tRNA genes. Even if such tRNA genes were acquired in eukaryotes, most would not have been retained because of the failure of transcription or subsequent RNA processing. However, in C. merolae, these genes could have persisted in the genome owing to the upstream promoter-dependent transcription system and, especially, the capacity to process non-canonical pre-tRNAs, including intronic and permuted pre-tRNAs, into the canonical cloverleaf structure.
BHB motifs in intronic pre-tRNAs and their corresponding splicing machinery are commonly found in eukaryotes and archaea, although their modes of splicing endonuclease recognition differ. Archaeal endonucleases exhibit symmetrical subunit architectures comprising catalytic subunits (homologs of Sen2 and Sen34) and recognize introns in pre-tRNAs in a manner that is basically dependent only on the BHB motifs11,18,27,28. Genome-wide analyses have expanded the number of archaeal tRNA genes that contain introns comprising various types of BHB motifs at various positions. Accordingly, four different types of endonuclease were identified in archaea, which indicated that the subunit compositions of archaeal endonucleases correlate with their substrate specificity toward the BHB motifs22,28,29,46,47. In eukaryotes, the processing mechanisms of tRNA-splicing endonucleases, other than those of yeast, are not fully understood. The present study showed that C. merolae contains atypical introns and an endonuclease that is substantially eukaryotic-like, but differs in subunit composition. Homology searching and YTH analysis could not identify an accessory subunit (Sen15) that is essential for functional multimerization of yeast endonuclease, indicating that the C. merolae endonuclease may contain an unidentified subunit, or is possibly comprised of a novel heterotrimeric complex. Alternatively, the C. merolae endonuclease may adopt a different type of subunit architecture. The yeast Sen54 is indicated to interact with the D-arm and the acceptor stem in the L-shaped tertiary structure of tRNA17,18; therefore, the C. merolae endonuclease containing the cmSen54 subunit is not likely to interact with pre-tRNAs harboring ectopic introns, especially in the D-arm. Thus, the C. merolae endonuclease may act on such pre-tRNAs as a dimer composed of only catalytic subunits (cmSen2 and cmSen34) to recognize atypical introns. It is tempting to speculate that the C. merolae endonuclease comprises a different subunit combination; for example, cmSen2-cmSen54-cmSen34 or cmSen2-cmSen34, depending on the positions or types of the BHB motifs in the substrates. If so, BHB motifs at non-canonical positions will be removed by cmSen2-cmSen34, making the BHB motif at canonical position (37/38) accessible to cmSen2-cmSen54-cmSen34, which interacts with the mature domain of the pre-tRNA, as it does in yeast. RT-PCR analysis (Figs. 3 and 4) showing that the BHB motif at canonical position 37/38 is always the final substrate may support this hypothesis.
An increasing number of reports on BHB-mediated disrupted tRNA genes support the hypothesis that the BHB motif and its processing system, played a central role in the development and maintenance of such tRNA genes. An evolutionary relationship has been suggested to exist between intronic tRNAs and split tRNAs in archaea7,19,26,44,45,47, while permuted tRNAs might have been generated by gene duplication, probably of an intronic tRNA through recombination to form a tandem repeat, followed by the loss of the outer segment of each copy4,6,48,49. The existence of a number of intronic tRNA genes may reflect an evolutionary background that produced a number of the permuted tRNAs in C. merolae. Even though the detailed mechanisms that govern the development of each type of disrupted tRNA gene remain unclear, the discovery of various BHB-mediated disrupted tRNA genes may imply an evolutionary relationship between them.
Some archaea, including Nanoarchaeum equitans and Caldiviga maquilingensis, harbor many split tRNAs, but few intronic tRNAs7,22. Other archaea, including Pyrobaculum and Thermophilum, harbor a number of intronic tRNAs, although Pyrobaculum have no split or permuted tRNAs22. Unicellular green algae contain some permuted tRNAs, but almost no intronic tRNAs6. Hence, C. merolae is unique in containing a number of both atypical intron-containing and permuted tRNAs. Conversely, split tRNAs have not been found in C. merolae, despite its potential ability to express them. Hitherto, there has been no report of an organism in which split tRNAs coexist with permuted tRNAs. Thus, the individual mechanisms and requisite elements necessary for the acquisition or maintenance of disrupted tRNA genes are sometimes substantially different, as previously suggested7. Our findings for C. merolae indicate that the processing system of eukaryal tRNA introns is more divergent and species-specific than previously thought; atypical introns and several types of splicing endonucleases may also be present in other eukaryotes. Previous studies may support this possibility, as evidenced by the discovery of ectopic intron-containing tRNA genes in the nucleomorph of the cryptomonad, Guillardia theta50; however, most of their introns cannot form a BHB motif. The absence of accessory subunit homologs (Sen15 and Sen54) in the A. thaliana endonuclease51 implies the possibility that plant endonucleases have evolved variations in their subunit architectures.
Considering that the origin of eukaryotes is related to that of archaea, C. merolae may retain vestigial traits derived from the archaeal lineage. However, no sequence similarity was found between the disrupted tRNA genes of C. merolae and archaea. Additionally, atypical intron-containing, permuted and split tRNAs show a discontinuous and patchy distribution among early eukaryotes and archaea6,7,22. These observations also imply a non-monophyletic origin for BHB-mediated disrupted tRNA genes and suggest the possibility that they could have arisen multiple times, independently, in each lineage. The next challenge will be to estimate the formation process of each type of disrupted tRNA gene.
The findings presented here reaffirm the unique nature of C. merolae tRNAs, which display various rearrangement patterns of RNA gene structures and tRNA processing pathways. Further comparative analysis of C. merolae tRNA genes and the splicing endonuclease with those of other organisms will help to clarify the mechanisms that govern the development of BHB-mediated disrupted tRNA genes.
Methods
Computational prediction of tRNA genes and their BHB motifs in the C. merolae nuclear genome
The 10D strain of C. merolae was used in this study1. Genome-wide prediction of tRNA genes in the C. merolae genome was conducted as described previously2,4 using tRNAscan-SE52, SPLITS20 and SPLITSX21 software with the following parameters: ‘-X 15 -I -36’, ‘-c -p 0.55 -f 3 -h 3’ and ‘-p 0.55 -f 3 -h 3’,. Initial predictions of tRNA introns and the secondary structures of splicing motifs were made by SPLITS/SPLITSX. In addition, a BLAST search was performed using BLASTN with a threshold E-value < 1. The sequences of candidate genes were manually inspected.
The free energies of the predicted introns were calculated using RNAeval, implemented within the Vienna-RNA package53 and then matched to the BHB secondary structure model. The BHB secondary structure model consisted of both intron and exon sequences and was classified into three types based on the reported archaeal BHB motifs: strict hBHBh′, relaxed HBh′ or relaxed hBH5. Some secondary structures of the predicted tRNAs harboring BHB motifs were revised by comparing them with known BHB-harboring tRNA structures.
Northern blotting analysis
Cell culture, RNA purification and northern blotting analysis followed previously described methods4. Total RNA isolated from C. merolae was separated on a 10% polyacrylamide gel containing 8 M urea and compared to RNA molecular weight markers (Dynamarker RNA Low, BioDynamics Laboratory, Tokyo, Japan). The gel was blotted onto Hybond N+ (GE Healthcare Bio-science AB, Upssala, Sweden) with TBE buffer for northern blotting analysis. The synthetic DNA oligonucleotides listed below were 5′ 32P-labeled and used as probes. Hybridization was performed at 50°C for 6 h. The membrane was then washed with 1 × SSC buffer (150 mM NaCl and 15 mM sodium citrate) at room temperature for 40 min. The predicted size of each mature molecule indicated in Fig. S2 was confirmed by sequencing analysis of the corresponding RT-PCR product (Fig. 3, Supplementary Figs. 3–5 online). The sequences of synthetic DNA oligonucleotides used as probes are listed below.
Oligo-DNA probes for northern blot analysis
N-probe 5′- cggtagtagagaggctt -3′.
F-probe 5′- tgccatcgcgcgggatc -3′.
I-probe 5′- tactcccggcgaggctc -3′.
Reverse-transcription polymerase chain reaction (RT-PCR) and DNA sequencing
Cell culture, RNA purification and RT-PCR analysis followed previously described methods4. Total RNA prepared from C. merolae cells was reverse-transcribed with ReverTraAce (TOYOBO, Osaka, Japan) and PCR-amplified with Blend Taq (TOYOBO). Total RNA was denatured for 3 min at 94°C and then incubated on ice for 5 min. Reverse transcription was performed using a reverse primer (3′-primer) at 55°C for 60 min and 5 μL aliquots of the cDNA solution were used for PCR (50 μL total reaction mixtures, using a program comprising 25 cycles) using 5′ and 3′-primers. PCR products were gel purified and cloned using a TA cloning kit (Invitrogen, Carlsbad, CA, USA), according to the manufacturer's protocol. Five clones were sequenced for each product (Supplementary Figs. 3–6 online). Not all of the PCR products could be cloned. The sequences of synthetic DNA oligonucleotides used as primers for RT-PCR shown in Fig. 3 and Fig. 4 are listed below.
Oligo-DNA primers for RT-PCR amplification of tRNAAsn(GUU)
N5′-1 5′- ggtagtatagctcagtcggttag -3′.
N3′-1 5′- cggtagtagagaggcttg -3′.
Oligo-DNA primers for RT-PCR amplification of tRNAPhe(GAA)
F5′-1 5′- gccattgtagctcagcagggag -3′.
F3′-1 5′- tgccatcgcgcgggatc -3′.
F5′-2 5′- gccattgtagctcagcagggagagga -3′.
F3′-2 5′- ccgcggacctttgcatcttca -3′.
Oligo-DNA primers for RT-PCR amplification of tRNAIle(UAU)
I5′-1 5′- actcctgtagct -3′.
I3′-1 5′- tactcccggcgaggctc -3′.
I5′-2 5′- actcctgtagctatcgg -3′.
I3′-2 5′- tactcccggcgaggctcgaacgcgca -3′.
I5′-3 5′- actcctgtagctcagctggt -3′.
Oligo-DNA primers for RT-PCR amplification of tRNAGly(CCC)a and tRNAGly(CCC)b
G5′-1 5′- tggaagattcccattcttct -3′.
G3′-1 5′- cactacaccagcagcgc -3′.
Yeast two-hybrid screening of the C. merolae tRNA-splicing endonuclease
The YTH analysis used for library screening was conducted according to a previously described method54. The full-length cDNA of cmSen2 (CMN231C), cmSen34 (CMH233C), or cmSen54 (CMK254C) was PCR-amplified and cloned into pGBTK, a GAL4 DNA-binding domain fusion vector. The C. merolae genomic library used for screening was constructed using in the pACT2 plasmid (TaKaRa). Plasmids were introduced into yeast PJ69-4Aa (for the pGBTK derivatives) and PJ69-4Aα (for the pACT2 derivatives) haploid strains, using TRP1 and LEU2, respectively, as selective markers. These yeast strains were co-incubated and became diploids. Positive protein–protein interactions between bait and prey were detected by the ability of cells to grow on SC-LWH plates containing 5 mM 3-aminotriazol and SC-LWA. The pACT2 derivatives were extracted from the diploid yeast and the insert junctions with the GAL4 activation domain were sequenced and compared with the C. merolae genome database. The synthetic DNA oligonucleotides used as primers for cloning are listed below.
Oligo-DNA primers used for cloning the C. merolae Sen2 homolog (cmSen2; CMN231C)
cmSen2Fwd 5′- ggggaattcatgagcgttcaagcgcaact -3′.
cmSen2Rev 5′- gggggatccctacttcgcggcacgctgtt -3′.
Oligo-DNA primers used for cloning the C. merolae Sen34 homolog (cmSen34; CMH233C)
cmSen34Fwd 5′- ccccccggggatgtatccaagtgtccagag -3′.
cmSen34Rev 5′- gggggggtcgactcagggtagagcctcacacc -3′.
Oligo-DNA primers used for cloning the C. merolae Sen54 homolog (cmSen54; CMK254C)
cmSen54Fwd 5′- ggggaattcatgcgaaatcagcggggtgg -3′.
cmSen54Rev 5′- gggggatcctcacagtccctgctcgtcgt -3′.
References
Kuroiwa, T. The primitive red algae Cyanidium caldarium and Cyanidioschyzon merolae as model system for investigating the dividing apparatus of mitochondria and plastids. Bioessays 20, 344–354 (1998).
Matsuzaki, M. et al. Genome sequence of the ultrasmall unicellular red alga Cyanidioschyzon merolae 10D. Nature 428, 653–657 (2004).
Nozaki, H. et al. A 100%-complete sequence reveals unusually simple genomic features in the hot-spring red alga Cyanidioschyzon merolae. BMC Biol. 5, 28 (2007).
Soma, A. et al. Permuted tRNA genes expressed via a circular RNA intermediate in Cyanidioschyzon merolae. Science 318, 450–453 (2007).
Marck, C. & Grosjean, H. Identification of BHB splicing motifs in intron-containing tRNAs from 18 archaea: evolutionary implications. RNA 9, 1516–1531 (2003).
Maruyama, S., Sugahara, J., Kanai, A. & Nozaki, H. Permuted tRNA genes in the nuclear and nucleomorph genomes of photosynthetic eukaryotes. Mol. Biol. Evol. 27, 1070–1076 (2010).
Chan, P., Cozen, A. E. & Lowe, T. M. Discovery of permuted and recently split transfer RNAs in Archaea. Genome Biol. 12, R38 (2011).
Reyes, V. M. & Abelson, J. Substrate recognition and splice site determination in yeast tRNA splicing. Cell 55, 719–730 (1988).
Shawn, K. W. & Abelson, J. in tRNA, eds Söll D., RajBhandary U. L. (ASM Press, Washington, D.C.), pp 79–92 (1995).
Abelson, J., Trotta, C. R. & Li, H. tRNA splicing. J. Biol. Chem. 273, 12685–12688 (1998).
Calvin, K. & Li, H. RNA-splicing endonuclease structure and function. Cell Mol. Life Sci. 65, 1176–1185 (2008).
Marck, C. & Grosjean, H. tRNomics: analysis of tRNA genes from 50 genomes of Eukarya, Archaea and Bacteria reveals anticodon-sparing strategies and domain-specific features. RNA 8, 1189–1232 (2002).
Baldi, M. I., Mattoccia, E., Ciafrè, S., Attardi, D. G. & Tocchini-Valentini, G. P. Binding and cleavage of pre-tRNA by the Xenopus splicing endonuclease: two separable steps of the intron excision reaction. Cell 47, 965–971 (1986).
Greer, C. L., Söll, D. & Willis, I. Substrate recognition and identification of splice sites by the tRNA-splicing endonuclease and ligase from Saccharomyces cerevisiae. Mol. Cell Biol. 7, 76–84 (1987).
Di Nicola Negri, E. et al. The eucaryal tRNA splicing endonuclease recognizes a tripartite set of RNA elements. Cell 89, 859–66 (1997).
Trotta, C. R. et al. The yeast tRNA splicing endonuclease: a tetrameric enzyme with two active site subunits homologous to the archaeal tRNA endonucleases. Cell 89, 849–858 (1997).
Fabbri, S. et al. Conservation of substrate recognition mechanisms by tRNA splicing endonucleases. Science 280, 284–286 (1998).
Xue, S., Calvin, K. & Li, H. RNA recognition and cleavage by a splicing endonuclease. Science 312, 906–910 (2006).
Randau, L., Münch, R., Hohn, M. J., Jahn, D. & Söll, D. Nanoarchaeum equitans creates functional tRNAs from separate genes for their 5′- and 3′-halves. Nature 433, 537–541 (2005).
Sugahara, J. et al. SPLITS: a new program for predicting split and intron-containing tRNA genes at the genome level. In Silico Biol. 6, 411–418 (2006).
Sugahara, J., Yachie, N., Arakawa, K. & Tomita, M. In silico screening of archaeal tRNA-encoding genes having multiple introns with bulge-helix-bulge splicing motifs. RNA 13, 671–681 (2007).
Fujishima, K. et al. A novel three-unit tRNA splicing endonuclease found in ultrasmall Archaea possesses broad substrate specificity. Nucleic Acids Res. 39, 9695–9704 (2011).
Galli, G., Hofstetter, H. & Birnstiel, M. L. Two conserved sequence blocks within eukaryotic tRNA genes are major promoter elements. Nature 294, 626–631 (1981).
Geiduschek, E. P. & Kassavetis, G. A. The RNA polymerase III transcription apparatus. J. Mol. Biol. 310, 1–26 (2001).
Hamada, M., Sakulich, A. L., Koduru, S. B. & Maraia, R. J. Transcription termination by RNA polymerase III in fission yeast. A genetic and biochemically tractable model system. J. Biol. Chem. 275, 29076–29081 (2000).
Sugahara, J. et al. Comprehensive analysis of archaeal tRNA genes reveals rapid increase of tRNA introns in the order thermoproteales. Mol. Biol. Evol. 25, 2709–2716 (2008).
Tocchini-Valentini, G. D., Fruscoloni, P. & Tocchini-Valentini, G. P. Coevolution of tRNA intron motifs and tRNA endonuclease architecture in Archaea. Proc. Natl. Acad. Sci. USA 102, 15418–15422 (2005).
Tocchini-Valentini, G. D., Fruscoloni, P. & Tocchini-Valentini, G. P. Structure, function and evolution of the tRNA endonucleases of Archaea: an example of subfunctionalization. Proc Natl Acad Sci USA 102, 8933–8938 (2005).
Komine, Y., Adachi, T., Inokuchi, H. & Ozeki, H. Genomic organization and physical mapping of the transfer RNA genes in Escherichia coli K12. J. Mol. Biol. 212, 579–598 (1990).
Miyauchi, K., Ohara, T. & Suzuki, T. Automated parallel isolation of multiple species of non-coding RNAs by the reciprocal circulating chromatography method. Nucleic Acids Res. 35, e24 (2007).
Suzuki, T. & Suzuki, T. Chaplet column chromatography: Isolation of a large set of indivisual RNAs in a single step. Methods in Enzymology 425, 25010–25014 (2007).
Li, Z. & Deutscher, M. P. The tRNA processing enzyme RNase T is essential for maturation of 5S RNA. Proc. Natl. Acad. Sci. USA 92, 6883–6886 (1995).
Li, Z. & Deutscher, M. P. Maturation pathways for E. coli tRNA precursors: a random multienzyme process in vivo. Cell 86, 503–512 (1996).
Nakanishi, K. et al. Structural basis for translational fidelity ensured by transfer RNA lysidine synthetase. Nature 461, 1144–1148 (2009).
Tocchini-Valentini, G. D., Fruscoloni, P. & Tocchini-Valentini, G. P. Processing of multiple-intron-containing pretRNA. Proc. Natl. Acad. Sci USA 106, 20246–20251 (2009).
Maden, B. E. The numerous modified nucleotides in eukaryotic ribosomal RNA. Prog. Nucleic Acid Res. Mol. Biol. 39, 241–303 (1990).
Maden, B. E., Corbett, M. E., Heeney, P. A., Pugh, K. & Ajuh, P. M. Classical and novel approaches to the detection and localization of the numerous modified nucleotides in eukaryotic ribosomal RNA. Biochimie 77, 22–29 (1995).
Kiss-László, Z., Henry, Y., Bachellerie, J. P., Caizergues-Ferrer, M. & Kiss, T. Site-specific ribose methylation of preribosomal RNA: a novel function for small nucleolar RNAs. Cell 85,1077–1088 (1996).
Pearson, C. E. & Sinden, R. R. Alternative structures in duplex DNA formed within the trinucleotide repeats of the myotonic dystrophy and fragile X loci. Biochemistry 35, 5041–5053 (1996).
Oussatcheva, E. A. et al. Structure of branched DNA molecules: Gel retardation and atomic force microscopy studies. J. Mol. Biol. 292, 75–86 (1999).
McLaren, A., Gonos, E. S., Carr, T. & Goddard, J. P. The conformation of tRNA genes. Chemical modification studies. FEBS Lett. 330, 177–180 (1993).
Ghavidel, A. et al. Impaired tRNA nuclear export links DNA damage and cell-cycle checkpoint. Cell 131, 915–926 (2007).
Mori, S., Kajita, T., Endo, T. & Yoshihisa, T. The intron of tRNA-TrpCCA is dispensable for growth and translation of Saccharomyces cerevisiae. RNA 17, 1760–1769 (2011).
Randau, L. & Söll, D. Transfer RNA genes in pieces. EMBO Rep. 9, 623–628 (2008).
Di Giulio, M. Transfer RNA genes in pieces are an ancestral character. EMBO Rep. 9, 820–821 (2008).
Randau, L. et al. The heteromeric Nanoarchaeum equitans splicing endonuclease cleaves noncanonical bulge-helix-bulge motifs of joined tRNA halves. Proc. Natl. Acad. Sci. USA 102, 17934–17939 (2005).
Fujishima, K. et al. Tri-split tRNA is a transfer RNA made from 3 transcripts that provides insight into the evolution of fragmented tRNAs in archaea. Proc. Natl. Acad. Sci. USA 106, 2683–2687 (2009).
Williams, K. P. Descent of a split RNA. Nucleic Acids Res. 30, 2025–2030 (2002).
Di Giulio, M. Permuted tRNA genes of Cyanidioschyzon merolae, the origin of the tRNA molecule and the root of the Eukarya domain. J. Theor. Biol. 253, 587–592 (2008).
Kawach, O. et al. Unique tRNA introns of an enslaved algal cell. Mol. Biol. Evol. 22, 1694–1701 (2005).
Akama, K., Junker, V. & Beier, H. Identification of two catalytic subunits of tRNA splicing endonuclease from Arabidopsis thaliana. Gene 257, 177–185 (2000).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Schuster, P., Fontana, W., Stadler, P. F. & Hofacker, I. L. From sequences to shapes and back: a case study in RNA secondary structure. Proc. Biol. Sci. 255, 279–284 (1994).
Watanabe, S. et al. Studies on the role of HtpG in the tetrapyrrole biosynthesis pathway of the cyanobacteria Synechococcus elongatus PCC 7942. Biochem. Biophys. Res. Commun. 352, 36–41 (2007).
Acknowledgements
We thank A. Muto and S. Maruyama for valuable discussions; K. Tanaka, O. Misumi, M. Matsuzaki, Y. Yoshida, N. Sato, T. Fujiwara and M. Monma for technical advice; and R. Saito, Y. Ohashi and K. Kurokawa for advice on the computational analysis. This work was supported by a grant from the Nakajima Foundation (to A.S.), a Grant-in-Aid for Young Scientists B (23770211 to A.S.), the Frontier project ‘Adaptation and Evolution of Extremophiles’ (to Y.S.) and the Strategic Research Foundation Grant-aided Project for Private Universities (S1201003 to Y.S.) from the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author information
Authors and Affiliations
Contributions
Y.S., A.K., T.K. and A.S. designed the research. A.S., J.S., A.O., S.W., N.Y. and M.O. performed the research. A.S., J.S., A.O., N.Y., H.Y. and H.K. analyzed the data. A.S., N.Y., S.W., A.K., J.S., A.O. and Y.S. wrote the main manuscript text. A.S. prepared Figures 1–4, Figures S1–7 and Table S1. J.S. and A.O. prepared Figures 2 and S1. S.W. prepared Figure 5. All authors reviewed and approved the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Info.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Soma, A., Sugahara, J., Onodera, A. et al. Identification of highly-disrupted tRNA genes in nuclear genome of the red alga, Cyanidioschyzon merolae 10D. Sci Rep 3, 2321 (2013). https://doi.org/10.1038/srep02321
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02321
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.