Abstract
How TGF-β1-mediated signaling pathways are finely tuned to orchestrate the generation of carcinoma-associated fibroblasts (CAFs) is poorly understood. Here, we demonstrate that miR-21 and the signaling of its target Smad 7 determine TGF-β1-induced CAF formation. In primary cultured fibroblasts, mature miR-21 increases after TGF-β1 treatment, whereas the Smad 7 protein level decreases. MiR-21 binds to the 3′ UTR of Smad7 mRNA and inhibits its translation, rather than causing its degradation. Most importantly, Smad 7 is bound to Smad 2 and 3, which are thought to competitively bind to TGFBR1 and prevents their activation upon TGF-β1 stimulation. The depletion of miR-21 or the overexpression of Smad 7 blocks TGF-β1-induced CAF formation, whereas the overexpression of miR-21 or the depletion of Smad 7 promotes CAF formation, even without TGF-β1 stimulation. Collectively, these findings clearly demonstrate that miR-21 and Smad7 are critical regulators of TGF-β1 signaling during the induction of CAF formation.
Similar content being viewed by others
Introduction
Carcinoma-associated fibroblasts (CAF) are activated fibroblasts and a key cellular component of tumor stroma. Through specific communication with cancer cells, CAFs not only directly promote tumor progression1,2 and metastasis3,4, but are also involved in the initiation of cancer5,6,7. It is well known that CAFs secrete growth factors and ECM-degrading proteases to promote tumor growth and invasiveness. In addition to the important role of TGF-β and VEGF in tumor progression, CAF-secreted SDF1 (stromal cell-derived factor 1) also mediates the recruitment of bone-marrow-derived endothelial cells and directly increases cancer cell proliferation2. CAF-secreted MMPs and other proteases also directly affect the motility and invasiveness of cancer cells8,9, helping cancer cells to cross tissue boundaries and escape the primary tumor site8,10,11.
Although the importance of CAFs during tumor initiation, progression and metastasis has been extensively studied in recent years, the origin of these cells is not clearly understood. Mesenchymal stem cells (MSCs) and other non-stem cells are described as the origins of CAFs. Pericytes purified from different sources of tumor samples were recently shown to have myogenic potential and to express an MSC-like phenotype in vitro12, which indicates that MSCs are one of the sources of CAFs. Although other types of cells, including smooth muscle cells, pericytes, adipocytes and inflammatory cells, have also been shown to transdifferentiate into CAFs, the recruitment and activation of peripheral resting fibroblasts to the tumor site are thought to be the main source13.
It is well known that tumor cell highly secreted transforming growth factor-β (TGF-β) is one of the main activators of resting fibroblasts, along with platelet-derived growth factor (PDGF) and basic fibroblast growth factor (bFGF)14,15. Once bound with TGF-β ligands, type II TGF-β receptors (TGFBR) form a heterodimer with type I TGFBR and activate the regulatory Smad 2/3 proteins, eventually initiating Smad 4-dependent transcription. However, the detailed mechanism by which TGF-β activates the regulatory Smad proteins during CAF formation is poorly understood.
MicroRNA-21(miR-21) is located on chromosome 17 and is highly conserved in many species, including human, rat, mouse, fish and frog16. MiR-21 is a key regulator of the oncogenic process. It is significantly elevated in the vast majority of human tumors where it has been analyzed17, such as breast, lung, colon, pancreas, prostate, liver, stomach and brain tumors and hematological cancers18,19,20,21,22,23. MiR-21 regulates cell proliferation, survival and migration in most if not all cancer cells through its downstream target proteins, such as PTEN, PDCD4, RECK and Cdc 25A24,25,26,27,28,29,30,31,32,33. In addition to its pivotal roles in cancer initiation and progression, miR-21 is also involved in other pathologic processes, for example, lung fibrosis, kidney fibrosis and cardiovascular disease34,35,36,37. However, it is not clear whether and how miR-21 and its target Smad 7 are finely tuned to orchestrate the generation of CAFs.
In this study, we demonstrate that miR-21 lowers the expression of Smad 7 through translation inhibition rather than mRNA decay and that TGF-β activates the regulatory Smad 2 and 3 proteins by reducing the association of Smad 2/3 with Smad 7. Most importantly, our in vitro and in vivo data clearly demonstrate that miR-21 and its target Smad 7 are critical regulators of TGF-β signaling during the induction of CAF formation.
Results
TGF-β1 treatment successfully transformed primary resting fibroblasts into CAFs
TGF-β1 is reported to induce the CAF formation, so TGF-β1 was used in this study to establish a CAF model to fully understand the mechanism by which TGF-β signaling is finally tuned to orchestrate the generation of CAFs. Experiments on time course and dose curve were performed to optimize the appropriate TGF-β1 concentration and treatment time for primary fibroblast activation. Primary normal fibroblasts isolated from human foreskin were placed in DMEM media containing 0.5% FBS without antibiotics at a 50% confluence. Twelve hours later, the cells were treated with TGF-β1 for 48 hours at the indicated concentration (0, 1, 2, 4 and 8 ng/ml) based on our previous study. Western blot analysis showed that the expression of FSP1 – a specific marker for CAFs – was gradually enhanced and that the TGF-β1 concentration increased from 1 to 8 ng/ml, with the highest expression at 8 ng/ml (Figure 1A). Both the quantitative PCR and immunofluorescence staining data for FSP1 were consistent with this immunoblot result (Figure 1A). Thus, 8 ng/ml was used as the optimal concentration of TGF-β1 in the remainder of the study.
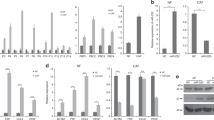
TGF-β1 treatment successfully transforms primary resting fibroblasts into CAFs.
TGF-β1 induces FSP1 expression in primary resting fibroblasts in vitro: (A). FSP1 expression is enhanced as the TGF-β1 concentration increases; (B). FSP1 expression reaches a peak at 48 hours after TGF-β1 treatment; (C). FSP1 continues to be expressed 3 weeks after TGF-β1 treatment. In parts (A), (B) and (C), the left upper panel shows the Western blot FSP1 result, the graphic shows the realtime PCR result, which is normalized to GADPH and the photos are the IF pictures (where green represents FSP1 staining). (D). TGF-β1-treated fibroblasts promote tumor growth in the xenografted mouse model. The schematic drawing shows the positions where the cells were injected; the graphic shows the tumor growth curve. The tumor size (median ± error) in the A375 plus TGF-β1-treated fibroblasts group and in the A375 group or A375 plus untreated fibroblasts at the end point is significantly different (**: P < 0.001). The picture shows the isolated tumors, where tumors in the same positions in each group are from the same mouse. The pictures in the bottom are representative fluorescence microscopy images, the greens are fibroblasts.
The duration of TGF-β1-induced fibroblast activation was then optimized. After 12 hours growth in DMEM containing 0.5% FBS, the primary fibroblasts were treated with TGF-β1 at a concentration of 8 ng/ml at different time points (0, 12, 24, 48, 72 and 96 hours). The Western blot showed that FSP1 expression increased after TGF-β1 stimulation and was highest at 48 hours after TGF-β1 treatment (Figure 1B). The quantitative PCR and immunofluorescence staining results for FSP1 confirmed this finding.
To further determine whether TGF-β1-induced primary fibroblast activation is a stable transformation, the fibroblasts were treated with two rounds of TGF-β1 at a concentration of 8 ng/ml in the first four days and then continued to grow in a regular medium without TGF-β1. The expression of FSP1 was measured 1, 2 and 3 weeks after TGF-β1 stimulation. The Western blot showed that the FSP1 continued to be expressed even at 3 weeks (Figure 1C). The data from both the quantitative PCR and immunofluorescence staining were consistent with the Western blot results (Figure 1C), suggesting that TGF-β1-induced primary fibroblast activation is a stable transformation.
To verify whether TGF-β1-activated fibroblasts are real CAFs, the tumor growth promoting effects were determined in a xenografted mouse model. Melanoma A375 cells mixed with primary fibroblasts or TGF-β1-activated fibroblasts were co-injected into the armpits of nude mice, with A375 cells alone or TGF-β1-activated fibroblasts alone taken as the controls. To avoid any individual differences among the nude mice and to make the tumor size more comparable across groups, the four groups of cells were injected into different parts of one mouse as per the schematic drawing in Figure 1D. Ten mice were used for the study. For each nude mouse, 1 × 106 of A735 cells were mixed with fibroblasts (resting or TGF-β1-activated) at a ratio of 1:3. Fifteen days after injection, the longest and shortest tumor diameters were measured every five days. The tumor volume was calculated with the formula: V = 1/2 × a × b2. As the graphic in Figure 1D shows, the tumors in the A375 mixed with TGF-β-induced CAF group grew significantly faster than those in the A375 alone or A375 mixed with primary fibroblasts (P < 0.001) groups. The tumors in the A375 mixed with primary fibroblasts group were even smaller than those in the A375 alone group. No tumors were found in the TGF-β-induced CAF group. Thirty days after injection, the tumors were isolated from the sacrificed mice after anesthesia and are presented at the third part of Figure 1D, in which the tumors in the same position in each group are from the same mouse. The representative fluorescence microscopy images are displayed in the bottom of Figure 1D. These data demonstrate that TGF-β1 successfully induces CAF formation in vitro, which promote tumor growth as reported7.
Smad 7 protects Smad 2/3 from activation through physical association
It is well known that Smad 2/3 activation is critical for Smad 4-dependent transcription regulation. Upon TGF-β1 stimulation, the Smad 7 is reported to be de-associated from the activated TGF-β receptor and then the activated TGF-β receptor activates Smad 2 and Smad 3. However, how Smad 2 and 3 are activated by TGFBR is poorly understood.
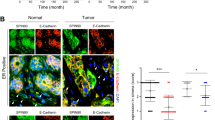
To determine whether and how Smad 7 regulates the activation of Smad 2/3 during the process of fibroblast activation, the effects of Smad 7 on Smad 2/3 activation/phosphorylation and translocation were investigated in fibroblasts with knocked down or over-expressed Smad 7. First, co-immunoprecipitation was performed using Smad 2 or Smad 3 antibodies, respectively. The Western blot in Figure 2A shows that both Smad 2 and Smad 3 bound to Smad 7, but when the cells were treated with TGF-β or overexpressed miR-21 this association ceased, which may be due to the decrease of Smad 7 (Figure S2). Furthermore, the reciprocal immunoprecipitation was performed using Flag antibody in 293 T cells overexpressing Flag-tagged Smad 7. The Western blots show Smad 7 bound to Smad 2 and Smad 3 (Figure 2A).
Smad 7 protects Smad 2/3 from activation through physical binding.
(A). Smad 7 regulates Smad 2/3 activation through physical binding. The top left panel shows Western blots of Smad 7 after Smad 2 or Smad 3 immunoprecipitation (IP), the top middle panel is the Western blots for the reciprocal IP and loading control (the full-length blots can be found in the Supplementary data Figure S1); the top right panel shows the effects of Smad 7 knock down on Smad 2/3 phosphorylation after TGF-β1 stimulation, the bottom left panel shows the phosphorylation kinetics of Smad 2 and Smad 3 after TGF-β1 stimulation and the bottom right panel shows the effects of Smad 7 overexpression on Smad 2/3 phosphorylation after TGF-β1 stimulation. (B). Immunofluorescence staining shows that Smad 2/3 nuclear translocation is affected by Smad 7. The top images are IF photos of Smad 2 (red) or Smad 3 (red); the bottom graphics show the statistical results for the nuclear density of Smad 2 and Smad 3. (C). The Western blot shows that the Smad 2/3 nuclear content is affected by Smad 7. The left part shows the data from the immunoblots; the right graphics show the quantitative analysis. (D). The effects of Smad 2, 3 and 4 on the phosphorylation of the other Smad protein. Smad 2, 3 and 4 was knocked down in fibroblasts mediated by the lentivirus. The right panel shows the knockdown efficiency of Smad 2, 3, 4 and 7.
To determine whether Smad 7 affects the activation of Smad 2 and 3, fibroblasts with knocked down or overexpressed Smad 7 were treated with TGFR1 kinase specific inhibitor (SB431542) half an hour before TGF-β stimulation. The cells were harvested half an hour after TGF-β stimulation. This time point was established based on our previous experiments (top right of Figure 2A). Our data show that TGF-β stimulation phosphorylated both Smad 2 and Smad 3 in the Smad 7 knocked down fibroblasts and control cells (bottom left of Figure 2A), but hardly at all in the Smad 7 overexpressed fibroblasts (bottom right of Figure 2A). To some extent, Smad 2/3 phosphorylation was enhanced in the Smad 7 knocked down cells. The data also reveal that TGFR kinase inhibition completely curtailed the phosphorylation induced by TGF-β (Figure 2A). Interestingly, even without TGF-β stimulation, there were some phosphorylation signals of Smad 2/3 in the Smad 7 knocked down fibroblasts, which may be due to the autocrined TGF-β. These observations suggest that Smad 7 regulates Smad 2/3 activation through physical binding, in addition to competitively binding with activated TGFBR.
To determine whether Smad 7 affects Smad 2/3 nuclear translocation, fibroblasts with over-expressed or knocked down Smad 7 were fixed and immunostained with Smad 2 or Smad 3 antibodies with or without TGF-β1 treatment. Immunofluorescence (IF) staining of the Smad 2 or Smad 3 showed that the IF density (red) of both proteins in the nucleus was significantly increased after TGF-β1 stimulation (Figure 2B). Interestingly, the IF density in the Smad 7 knocked down and overexpressed fibroblasts were lower than those in the control fibroblasts, especially in terms of the density of Smad 2. The nuclear content of Smad 2/3 after quantification and statistical analysis is shown in the graphic in Figure 2B.
To further confirm the aforementioned effects, fibroblasts overexpressing or knocking down Smad 7 were fractionated into a cytoplasmic fraction and a nuclear fraction. Smad 2 and Smad 3 in the nuclear fraction were detected by Western blot. As Figure 2C shows, both Smad 2 and Smad 3 increased in the nuclear fractions after TGF-β stimulation and the differences were more obvious in the control fibroblasts than in either the Smad 7 knocked down or Smad 7 overexpressed fibroblasts (Figure 2C), especially in terms of the difference in the Smad 2 protein levels, which is consistent with the immunofluorescence staining results. These observations suggest that the TGF-β1-induced translocation of Smad 2/3 is affected by Smad 7. However, the mechanism by which Smad 7 regulates this translocation needs to be further determined.
To determine whether Smad 2, 3 and 4 regulate one another, the phosphorylation of Smad2 and Smad 3 in Smad 2, 3, 4 or 7 knocked down cells was determined by Western blot. Figure 2D shows that Smad 2 and Smad 3 were phosphorylated upon TGF-β1 stimulation and that except for Smad 7, Smad did not stop the other Smad proteins from phosphorylating each other. The Western blots in the top panel of Figure 2D show that TGF-β1-induced Smad 2 phosphorylation was not affected by either Smad 3 or Smad 4 knockdown and that TGF-β1-induced Smad 3 phosphorylation was not affected by either Smad 2 or Smad 4 knockdown (middle panel of Figure 2D). The data also show that there were some phosphorylation signals of Smad 2 and Smad 3 in the Smad 7 knocked down cells even without TGF-β1 stimulation. These observations are consistent with the previous findings in Figure 2A. In addition, neither Smad 2 and 3 nor Smad 7 affected the Smad 4 protein level.
In brief, our observations suggest that Smad 7 regulates Smad 2 and Smad 3 activation and nuclear translocation through physical association in addition to competitively binging with TGFBR.
MiR-21 regulates the Smad 7 protein level through translation inhibition
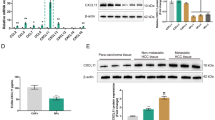
miR-21 is an upstream regulatory miRNA of Smad 7 according to web-based predictive software38 (http://starbase.sysu.edu.cn) and data from other groups35. To determine whether Smad 7 is a target of miR-21 during CAF formation, luciferase assays were performed using Smad 7 3′ UTR. As shown in Figure 3A, the miR-21 sequence partially complements the 3′ UTR sequence of Smad 7 and the miR-21 sequence at the 5′ terminal entirely complements the 3′ UTR sequence of Smad 7. The luciferase assay data show that miR-21 only reduced luciferase activity in cells containing wild type 3′ UTR, but not in cells containing mutant 3′ UTR (the mutated nucleotide is marked red; Figure 3A).
MiR-21 regulates the Smad 7 protein level by inhibiting translation.
(A). miR-21 targets 3′ UTR of Smad 7 mRNA. The luciferase assay shows that miR-21 only works on the wild type 3′ UTR of Smad 7 but not on the mutant type (the red nucleotides are mutant to cytosine). (B). The overexpression of miR-21 reduces the Smad 7 protein level, but does not affect its mRNA level; (C). miR-21 knockdown increases the Smad 7 protein level and attenuates the TGF-β-induced decrease in Smad 7. In both parts (A) and (B), the left panels show the Western blot data, the middle graphics show the realtime PCR results for Smad 7 and the right graphics show the Smad 7 knockdown or overexpression efficiency. (D). Fibroblasts with knocked down Smad 7 promote tumor growth in the xenografted mouse model; (E). Fibroblasts with overexpressed Smad 7 inhibit tumor growth in the xenografted mouse model. In both experiments, the schematic drawing shows the positions where the cells were injected and the graphic shows the tumor growth curve. The tumor size (median ± error) in the A375 plus Smad 7 knocked down or overexpressed fibroblasts group and in the A375 plus untreated fibroblasts group at the end point is significantly different (**: P < 0.01; #: P < 0.001). The picture shows the isolated tumors, where tumors in the same position in each group are from the same mouse. The pictures in the bottom are representative fluorescence microscopy images, the greens are fibroblasts.
The effects of miR-21 on Smad 7 expression were further analyzed to determine whether miR-21 regulates Smad 7 during fibroblast activation. Our data show that miR-21 over-expression significantly decreased the level of Smad 7 (Figure 3B). Interestingly, the mRNA level of Smad 7 did not change in the miR-21 over-expressed fibroblasts (Figure 3B). As expected, miR-21 knockdown minimized TGF-β1-induced Smad 7 protein reduction and there was no significant difference in the Smad 7 level without TGF-β1 treatment between the control and the miR-21 knocked down fibroblasts (Figure 3C). As in Figure 3A, there was no detectable change in the mRNA level of Smad 7 between the miR-21 knocked down fibroblasts and the control cells, either with or without TGF-1 treatment. These observations suggest that miR-21 decreases the Smad 7 level by inhibiting mRNA translation, rather than by mRNA decay.
To determine whether Smad 7 is a critical mediator of TGF-β1/miR-21 signaling in the process of fibroblast activation, Smad 7 knocked down fibroblasts were co-injected into the armpits of mice with A375 tumor cells, as per the schematic drawing in Figure 3D. Fibroblasts infected with empty viruses were used as controls. The tumor volumes were measured at the indicated time points (5, 10, 15, 20 and 25 days) as previously described. As the graphic in Figure 3C shows, the tumors in the A375 mixed with Smad 7 knocked down fibroblasts group grew faster than those in the control group, even without TGF-β1 stimulation (P < 0.05). The tumors isolated at the end point are presented in the third part of Figure 3D. The representative fluorescence microscopy images are displayed in the bottom of Figure 3D.
To determine whether Smad 7 is a critical mediator of TGF-β signaling and whether the over-expression of Smad 7 blocks TGF-β1-induced fibroblast activation, Smad 7 over-expressed fibroblasts with or without TGF-β1 stimulation were co-injected with A375 cells, as per the schematic drawing in Figure 2E, with fibroblasts infected with empty viruses used as controls. The tumor volumes were measured at the indicated time points (10, 15, 20, 25 and 30 days). As the graphic in Figure 3E shows, the tumors in the A375 mixed with Smad 7 over-expressed fibroblasts group grew significantly slower than those in the control group (3rd and 4th lines, P < 0.01) and even slower than in the control group without TGF-β1 treatment. TGF-β1 treatment only slightly promoted tumor growth in the Smad 7 overexpressed group, which suggests that Smad 7 over-expression blocks TGF-β1-induced CAF formation and that Smad 7 plays an important role in TGF-β1-induced CAF formation. The tumors isolated at the end point are presented in the third part of Figure 3E. The representative fluorescence microscopy images are displayed in the bottom of Figure 3E. These observations together demonstrate that Smad 7 is a critical mediator of TGF-β1/miR-21 signaling during fibroblast activation.
MiR-21 determines TGF-β1-induced fibroblast activation
MiR-21 was the first onco-miRNA to be identified and is over-expressed in many types of tumors. MiR-21 is also regulated by TGF-β139, but it is not clear whether miR-21 controls TGF-β1-induced CAF formation.
To determine whether miR-21 regulates TGF-β-induced fibroblast activation, mature miR-21 was detected after TGF-β1 treatment by Northern blot and quantitative PCR. The Northern blot showed that miR-21 (the signal at 22 nt) in the fibroblasts dramatically increased 48 hours after TGF-β1 treatment compared with the control, as shown in the left part of Figure 4A. The realtime PCR data were consistent with the Northern blot results. The level of pri-, pre- and mature miR-21 by realtime PCR after TGF-β1 treatment was also detected at the indicated time points (0, ¼, ½, 1, 2, 4, 8 and 24 hours) and both pre-miR-21 and mature miR-21 started to increase 15 minutes after TGF-β1 stimulation (at a concentration of 8 ng/ml) (Figure 4A, right). The pre-miR-21 level reached a peak at 30 minutes and the mature miR-21 level reached a peak in 2 hours, which is consistent with data from the Hata group39.
MiR-21 determines TGF-β1-induced fibroblast activation.
(A). TGF-β1 promotes miR-21 maturation and increases mature miR-21. The left panel and graphic show the Northern blot data and realtime PCR results for mature miR-21 48 hours after TGF treatment; the right graphic shows the realtime PCR results for pri-, pre- and mature miR-21 at different time points after TGF-β1 stimulation. (B). miR-21 over-expression triggers FSP1 expression. The left panel shows the Western blot data for FSP1 and α-SMA (another CAF marker), the middle graphic shows the realtime PCR result and the right panel shows the miR-21 overexpression efficiency. (C). miR-21 knockdown blocks TGF-β1-induced FSP1 expression. The Western blot data shows the expression of FSP1 and α-SMA, the middle graphic shows the realtime PCR result and the right panel shows the miR-21 knockdown efficiency. (D). Fibroblasts overexpressing miR-21 promote tumor growth in the xenografted mouse model. (E). Fibroblasts with knocked down miR-21 inhibit tumor growth in the xenografted mouse model. For both experiments, the schematic drawing shows the positions where the cells were injected and the graphic shows the tumor growth curve. The tumor size (median ± error) in the A375 plus miR-21 knocked down or overexpressed fibroblasts group and in the A375 plus untreated fibroblasts group at the end point is significantly different (**: P < 0.001; #: P < 0.001). The picture shows the isolated tumors, where tumors in the same position in each group are from the same mouse. The pictures in the bottom are representative fluorescence microscopy images, the greens are fibroblasts.
miR-21 was stably knocked down or over-expressed in fibroblasts. The specific marker for CAF FSP1 was detected by Western blot and realtime PCR. Both analyses showed that miR-21 over-expression induced FSP1 expression in the fibroblasts without TGF-β1 stimulation (Figure 4B), which suggests that miR-21 over-expression activates fibroblast just as TGF-β1 does. Conversely, both the Western blot and realtime PCR showed that miR-21 knockdown suppressed TGF-β1-induced FSP1 expression (Figure 4C). The Western blot showed that the TGF-β1-induced increase in FSP1 expression was reduced (left panel in Figure 4C), which suggests that miR-21 determines (or is at least a critical mediator of) TGF-β1 signaling in CAF formation.
To further confirm whether miR-21 is a critical mediator of TGF-β1 signaling in CAF formation, animal experiments were performed to determine the function of fibroblasts over-expressing miR-21 or knocking down miR-21. The miR-21 expressed fibroblasts were co-injected into the armpits of mice with A375 tumor cells, as per the schematic drawing in Figure 4D, with fibroblasts infected with empty viruses as controls. The tumor volumes were measured at the indicated time points (10, 15 and 20 days) as described previously. As the graphic in Figure 4D shows, the tumors in the A375 mixed with miR-21 over-expressed fibroblasts group grew significantly faster than those in the control group, suggesting that miR-21 activates fibroblasts just as TGF-β1 does. The tumors isolated at the end point are presented in the third of Figure 4D. The representative fluorescence microscopy images are displayed in the bottom of Figure 4D.
To determine whether miR-21 knockdown suppresses TGF-β1-induced fibroblast activation, miR-21-knocked down fibroblasts with and without TGF-β1 stimulation were co-injected with A375 cells, as per the schematic drawing in Figure 4E. The tumor volumes were measured at the indicated time points (10, 15 and 20 days). As the graphic in Figure 4E shows, the tumors in the A375 mixed with miR-21 knocked down fibroblasts group grew significantly slower than those in the control group, even when treated with TGF-β1 (middle line, P < 0.01), which suggests that miR-21 knockdown blocks TGF-β1-induced CAF formation. The tumors isolated at the end point are presented in the third part of Figure 4E. The representative fluorescence microscopy images are displayed in the bottom of Figure 4E. These observations together demonstrate that miR-21 is a critical mediator of TGF-β1 signaling in the generation of fibroblast activation.
Canonical Smad signaling mediates miR-21/Smad 7 signaling during TGF-β1-induced CAF formation
To confirm whether Smad 2/3 signaling (the canonical Smad pathway) is a critical pathway that mediates miR-21/Smad 7 signaling during TGF-β1-induced CAF formation, Smad 2, Smad 3 and Smad 4 were separately knocked down in fibroblasts and the FSP1 of the cells was detected after TGF-β1 stimulation by Western blot and quantitative PCR, with empty virus-infected fibroblasts as controls. Figure 5A shows that the expression of FSP1 increased 48 hours after TGF-β1 treatment in the control cells and the Smad 7 knocked down cells, but not in the Smad 2, 3, or 4 knocked down cells. Moreover, Smad 7 knockdown increased the basal expression of FSP1 even without TGF-β1 stimulation, which is consistent with the data shown in Figure 3B. As expected, no FSP1 was expressed after TGF-β1 stimulation in the Smad 2, Smad 3, or Smad 4 knocked down cells. The realtime PCR data were consistent with these results (Figure 5B).
Canonical Smad signaling mediates miR-21/Smad 7 signaling during TGF-β1-induced CAF formation.
(A). The Western blot shows that Smad 2, 3 and 4 knockdown curtails TGF-β-induced FSP1 expression. (B). The realtime PCR results show that Smad 2, 3 and 4 knockdown attenuates TGF-β-induced FSP1 mRNA transcription. (C). Fibroblasts with knocked down Smad 2 or Smad 3 decrease tumor growth in the xenografted mouse model. The schematic drawing shows the positions where the cells were injected and the graphic shows the tumor growth curve. The tumor size (median ± error) in the A375 plus Smad 2 or Smad 3 knocked down fibroblasts group and in the A375 plus control fibroblasts at the end point is significantly different (**: P < 0.01; #: P < 0.01). All of the fibroblasts were treated with TGF-β1 before co-injection with the A375 cells. The picture shows the isolated tumors, where tumors in the same position in each group are from the same mouse. The pictures in the bottom are representative fluorescence microscopy images, the greens are fibroblasts.
To further confirm that canonical Smad signaling is a critical pathway for TGF-β1-induced CAF formation, A375 cells were co-injected into the armpits of nude mice with fibroblasts with knocked down Smad 2, 3, or 7 as described in the schematic drawing in Figure 5C. Empty virus-infected fibroblasts treated with TGF-β1 were used as positive controls. As the graphic in Figure 5C shows, the TGF-β1-stimulated control fibroblasts significantly promoted tumor A375 cells growth (top line) and the Smad 7 knocked down fibroblasts also showed similar tumor-growth promoting effects. However, the tumors mixed with fibroblasts with knocked down Smad 2 or Smad 3 grew significantly slower compared with the TGF-β-stimulated control fibroblasts (Figure 5C). The tumors isolated at the end point are presented in the third part of Figure 5C, where tumors in the same positions in each group are from the same mouse. The representative fluorescence microscopy images are displayed in the bottom of Figure 5C. It is clear that the tumors in the control group and the Smad 7 knocked down group were much larger than those in the Smad 2 or 3 knocked down group. These observations demonstrate that Smad 2 and Smad 3 (the canonical Smad pathway) are critical for TGF-β1-induced CAF formation.
Discussion
It is well known that tumor-paracrined growth factors such as TGF-β play important roles in the induction of CAF formation and that the regulatory Smad 2 and 3 proteins mediate this signaling. However, how TGF-β signaling is finely tuned to regulate Smad 2/3 activation is poorly understood. Here, we demonstrate that miR-21 and its target Smad 7 mediates TGF-β signaling during the induction of CAF formation and that Smad 7 protects Smad 2/3 from being activated through physical association.
MiR-21 is well known as the first onco-microRNA to be identified. We show that TGF-β1 promotes miR-21 maturation in fibroblasts, which is consistent with the finding of the Hata group39 that TGF-β1 regulates miR-21 maturation in smooth muscle cells. However, the role of miR-21 in CAF formation is not known. Our in vitro and in vivo data indicate that the over-expression of miR-21 triggers fibroblast activation and promotes tumor growth, whereas miR-21 knockdown blocks TGF-β1-induced CAF formation and reduces or eliminates the tumor-growth promoting effect of fibroblasts. These observations demonstrate that miR-21 is a critical mediator of TGF-β1-induced CAF formation.
Although the Abraham group35 showed that the over-expression of miR-21 results in a decrease in Smad 7 during lung fibrosis, it is not clear how miR-21 regulates Smad 7 protein expression and whether miR-21 regulates Smad 2/3 activation through Smad 7 during TGF-β1-induced CAF formation. We show that the overexpression of miR-21 and TGF-β1 stimulation both reduce Smad 7 expression and that miR-21 knockdown reverses this TGF-β1-induced Smad 7 reduction. Moreover, in a xenografted mouse model, Smad 7 knocked down fibroblasts promote tumor growth, even without TGF-β stimulation, whereas Smad 7 overexpressed fibroblasts suppress tumor growth, even with TGF-β treatment, with the tumor size in the fibroblasts overexpressing Smad 7 group being significantly smaller than that in the control group without TGF-β treatment. These observations suggest that Smad 7 mediates miR-21 signaling during TGF-β-induced CAF formation. Both the cellular and animal data demonstrate that TGF-β1 regulates Smad 7 expression through miR-21 and that Smad 7 is a critical regulator of TGF-β during the induction of CAF formation.
It is believed that miRNA forms a RNA-induced silencing complex (RISC) with its target mRNA and decreases the expression of the target gene through mRNA degradation or the suppression of protein translation. In this study, miR-21 over-expression affected the Smad 7 protein level but not the mRNA level, which suggests that miR-21 regulates Smad 7 expression by inhibiting translation rather than by mRNA decay.
It is believed that Smad 7 and Smad 2/3 competitively bind to activated TGFBR1, which is an upstream kinase for Smad 2 and Smad 340,41. However, our data show that Smad 7 is physically bound to Smad 2 and Smad 3 and that the overexpression of Smad 7 reduces TGF-β-induced phosphorylation and the nuclear translocation of Smad 2 and Smad 3. These findings suggest that this binding may protect Smad 2 and Smad 3 from being activated by TGFBR1 (Figure 6) in addition to their competitive binding with TGFBR1. We also found that Smad 7 knockdown activated Smad 2/3, even without TGF-β1 stimulation and that both types of phosphorylation were curtailed by the TGFBR1 specific inhibitor, indicating that Smad 7 knockdown may result in autocrined TGF-β, which in turn activates Smad 2/3.
Upon activation, phosphorylated Smad 2 and Smad 3 are translocated into nuclear. Our data of fluorescence staining and fractionation Western blots show the nuclear Smad 2 and Smad 3 both increased after TGF-β stimulation in the fibroblasts with Smad 7 knocked down and with Smad 7 overexpression. However, these increases in both cells were less than in the control cells (Figure 2A and 2B). The Smad 7 overexpression-curtailed nuclear translocation of Smad 2/3 is caused by the activation inhibition, whereas the Smad 7 knockdown-minimized nuclear Smad 2/3 most likely result from the durative activation of Smad 2/3.
In brief, our data clearly demonstrate that the association with Smad 7 protects Smad 2 and 3 from activation and that miR-21 and Smad7 are critical regulators of TGF-β1 signaling during the induction of CAF formation (see the working model in Figure 6).
Methods
Reagents and antibodies
Human recombinant TGF-β1 was purchased from Biolegend (San Diego, CA). Fast start universal SYBR Green master (Rox) was purchased from Roche (Fishers, IN). The Taqman microRNA reverse transcription kit, TaqMan microRNA assay and TaqMan universal master mix II (2×) were obtained from Applied Biosystems (Foster City, CA). The BrightStar psoralen-biotin kit and the BrightStar biodect kit were purchased from Ambion (Austin, TX). The Dual-luciferase Reporter Assay System was purchased from Promega (Madison, WI). The following primary antibodies were used: mouse monoclonal to S100A4/FSP1 (Abnova, Taiwan, China); rabbit polyclonal to α-SMA (Bethyl, Montgomery, TX); mouse monoclonal to Ku86 and beta-actin (Santa Cruz Biotechnology, Santa Cruz, CA); rabbit monoclonal to Smad7 (Epitomics, Burlingame, CA); and rabbit monoclonal to Smad2, Smad3, phospho-Smad2 (Ser465/467), phospho-Smad3 (Ser423/425), rabbit polyclonal to Smad2/3 and Smad4 from Cell Signaling (Beverly, MA). The secondary antibodies used were Donkey anti-mouse IR Dye 680, Goat anti-mouse IR Dye 800CW (LI-COR Biosciences, Lincoln, NE) and Alexa flour 488 goat anti-mouse IgG1 (Invitrogen, Carlsbad, CA).
Lentivirus mediated shRNA knockdown or overexpression
Specific shRNA sequences targeting Smad 2, 3, 4, or 742,43,44 or sequences of pre-miR-21 as shown in Table 1 (supplemental data) were inserted into human GIPZ lentiviral shRNAmir individual clones (Open Biosystems, Lafayette, CO). The vectors containing shRNA were transfected together into 293 T cells with psPAX2 and PMD2G to generate the respective lentiviruses. Viral stocks were collected from the transduced 293 T cells and were used to infect the human fibroblasts. Lentivirus mediated miR-21 knockdown was generated with a pPACKH1 lentivector packaging kit (System Biosciences, Mountain View, CA) according to the manual.
Northern blot
A total of 30 μg of total mRNA per sample extracted by Trizol was mixed with an equal volume of 2 × RNA loading dye. The RNA samples were denatured by heat at 70°C for 10 min and placed on ice for at least 2 minutes before electrophoresis. After denaturing, the RNA was resolved by 15% denaturing polyacrylamide gel (with 8 M urea in Tris-borate buffer containing 90 mM of Tris, 90 mM of Boric acid and 2 mM of Na2EDTA with pH8.0 at 25°C). The gel was then transferred to a nylon H-bond membrane with a submerged apparatus at 200 mA for 1 hour. The membrane was cross-linked by Ultraviolet Crosslinkers (UVP) with 200 × 100 μJ/cm2 and then subjected to a pre-hybridization solution (7% SDS and 200 mM Na2HPO4 with pH7.2). After pre-hybridization, the membrane was hybridized with a biotin-labeled antisense miRNA probe at 42°C for 16 hours. After hybridization, the membranes were washed twice for 30 min in 2 × SSC, 0.5% SDS at 42°C. The chemiluminescent signal was detected using a BrightStar biodect kit following the manufacturer's instructions. The oligonucleotide probe used for hybridization was: 5-TCAACATCAGTCTGATAAGCTATCAACATCAGTCTGATAAGCTATCAACATCAGTCTGATAAGCTA-3.
Real-time PCR
cDNA was synthesized from total RNA using M-MuLV reverse transcriptase. Quantitative real-time PCR was performed using 2 × SYBR Green master Mix. The following primers were used: FSP1 sense: 5-GTCAGAACTAAAGGAGCTGC-3, antisense: 5-TGTTGCTGTCCAAGTTGCTC-3; Smad7 sense: 5-CGATGGATTTTCTCAAACCAA-3, antisense 5-ATTCGTTCCCCCTGTTTCA-3; pri-miR21 sense 5-TTTTGTTTTGCTTGGGAGGA-3, antisense 5-AGCAGACAGTCAGGCAGGAT-3; pre-miR21 sense 5-TGTCGGGTAGCTTATCAGAC-3, antisense 5-TGTCAGACAGCCCATCGACT-3; GAPDH sense 5-ACCCAGAAGACTGTGGATGG-3, antisense 5-CAGTGAGCTTCCCGTTCAG-3.The mRNA expression values were normalized to GAPDH. For the miRNA measurements, three independent cDNAs were synthesized from 10 ng of total RNA using the TaqMan MicroRNA Assay protocol and the Q-PCR levels were measured using the TaqMans Universal PCR Master Mix. All of the miRNA expression values were normalized to an endogenous control (RNU6B).
Luciferase assay
The full-length 3′ UTR of the human Smad7 gene was amplified by PCR using human genomic DNA as a template. The primers were sense 5-TTTTTCTAGACCGCGTGCGGAGGGGACAGA-3 and antisense 5- TTTTTCTAGAGGAGTCCTTTCTCTCTCAAAGC-3. The PCR fragment was cloned into the XbaI site downstream of the luciferase gene in the pGL3 control vector. Site-directed mutagenesis was performed to generate Smad7 3′ UTR mutants containing mutations in the conserved miR-21 binding site. In the 3′ UTR mutants, the nucleotide sequence complementary to nt 2–5 of miR-21 was mutated to the same sequence as that in miR-21 (from ATAAGCTA to TTATGCAA). 293 T cells were seeded at a density of 2 × 105 cells in 24-well plates. The cells were co-transfected with 50 ng of pGL3-SMAD7 (wild-type or mutant), 750 ng of miR-21 over-expression plasmid and Renilla plasmid using Lipofectamine 2000 Transfection Reagent, with empty vectors as controls. The cells were harvested 48 hours later and assayed using the Dual-Luciferase Reporter Assay System and a MicroLumatPlus luminometer. The firefly luciferase values were normalized to Renilla luciferase values expressed from the same pGL3 control vector.
Xenografted mouse model
Four-week-old BALB/c nude mice were obtained from a Shanghai animal laboratory (Shanghai, China). The animals were handled according to the protocol approved by the Institutional Animal Care and Use Committee of the Shanghai Jiao Tong University School of Medicine. Xenografted mice were developed by subcutaneously co-injecting human fibroblasts with A375 cells (the ratio of fibroblasts to A375 cells was 3:1 and the total cell number in each injection was 4 × 106) to the armpits bilaterally. The tumor volume was determined weekly using digital caliper measurements and the following formula: tumor volume (mm3) = ½ × longest diameter2 × shortest diameter. After 5 weeks, the mice were killed and the tumors excised.
Human normal primary fibroblast preparation
The tissue collection was approved by the Medical Ethical Committee of the Shanghai Jiao-Tong University School of Medicine. Foreskin tissue was collected after informed consent from children aged 7 to 12 years at the affiliated Xinhua Children's Hospital of the Shanghai Jiao Tong University School of Medicine. Human normal primary fibroblasts were isolated from the foreskin tissue. After posthectomy, the foreskins were transported to the laboratory as quickly as possible in sterile PBS buffer containing 1% penicillin/streptomycin on ice. After washing twice, the foreskins were minced using sterile scalpels and scissors, then digested with 0.1% type I collagenase and trypsin in a shaking water bath at 37°C for 30 minutes. After digestion, the tissue was sieved through 400-mesh filtration and centrifuged at 1000 × g for 10 min. The obtained pellets were cultured with DMEM containing 10% FBS for 2 hours. The attached cells were the fibroblasts, as verified by F-actin staining (Figure 1). After three passes, the cells were frozen in liquid nitrogen for further experiments.
Immunoprecipitation
Exponentially growing fibroblasts were harvested and lysed in 1 mL of lysis buffer (0.5% (v/v) NP-40, 5 mM of EDTA, 2 mM of EGTA, 20 mM of MOPS, 1 mM of PMSF, 20 mM of sodium pyrophosphate, 30 mM of sodium fluoride, 40 mM of β-glycerophosphatase, 1 mM of Na3VO3 and protease inhibitors). Aliquots of 1 mg total protein were mixed with 4 μg of monoclonal anti-Smad 7 antibody at 4°C overnight. The bound proteins were pulled down by binding to 25 μL of protein-A agarose (Sigma). The samples were separated by 7.5% SDS-PAGE gel electrophoresis and were subjected to mass-spec analysis or transferred to nitrocellulose membranes. The proteins of interest were detected using specific antibodies. The blots were scanned using an Odyssey infrared imaging system (LI-COR) and the proteins were quantitatively analyzed using the Odyssey software.
References
Dimanche-Boitrel, M. T. et al. In vivo and in vitro invasiveness of a rat colon-cancer cell line maintaining E-cadherin expression: an enhancing role of tumor-associated myofibroblasts. Int J Cancer 56, 512–521 (1994).
Orimo, A. et al. Stromal fibroblasts present in invasive human breast carcinomas promote tumor growth and angiogenesis through elevated SDF-1/CXCL12 secretion. Cell 121, 335–348 (2005).
Olaso, E. et al. Tumor-dependent activation of rodent hepatic stellate cells during experimental melanoma metastasis. Hepatology 26, 634–642 (1997).
Grum-Schwensen, B. et al. Suppression of tumor development and metastasis formation in mice lacking the S100A4(mts1) gene. Cancer Res 65, 3772–3780 (2005).
Kuperwasser, C. et al. Reconstruction of functionally normal and malignant human breast tissues in mice. Proc Natl Acad Sci U S A 101, 4966–4971 (2004).
Bhowmick, N. A. et al. TGF-beta signaling in fibroblasts modulates the oncogenic potential of adjacent epithelia. Science 303, 848–851 (2004).
Olumi, A. F. et al. Carcinoma-associated fibroblasts direct tumor progression of initiated human prostatic epithelium. Cancer Res 59, 5002–5011 (1999).
Boire, A. et al. PAR1 is a matrix metalloprotease-1 receptor that promotes invasion and tumorigenesis of breast cancer cells. Cell 120, 303–313 (2005).
Lochter, A. et al. Matrix metalloproteinase stromelysin-1 triggers a cascade of molecular alterations that leads to stable epithelial-to-mesenchymal conversion and a premalignant phenotype in mammary epithelial cells. J Cell Biol 139, 1861–1872 (1997).
Stetler-Stevenson, W. G., Aznavoorian, S. & Liotta, L. A. Tumor cell interactions with the extracellular matrix during invasion and metastasis. Annu Rev Cell Biol 9, 541–573 (1993).
Sternlicht, M. D. et al. The stromal proteinase MMP3/stromelysin-1 promotes mammary carcinogenesis. Cell 98, 137–146 (1999).
Crisan, M. et al. A perivascular origin for mesenchymal stem cells in multiple human organs. Cell Stem Cell 3, 301–313 (2008).
Ross, R. Aging of the intercellular matrix in relation to the fibroblast connective tissue forming cells-a retrospective view. Scand J Clin Lab Invest Suppl 65, 33–44 (1974).
Ronnov-Jessen, L. & Petersen, O. W. Induction of alpha-smooth muscle actin by transforming growth factor-beta 1 in quiescent human breast gland fibroblasts. Implications for myofibroblast generation in breast neoplasia. Lab Invest 68, 696–707 (1993).
Denk, P. O., Hoppe, J., Hoppe, V. & Knorr, M. Effect of growth factors on the activation of human Tenon's capsule fibroblasts. Curr Eye Res 27, 35–44 (2003).
Lagos-Quintana, M., Rauhut, R., Lendeckel, W. & Tuschl, T. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001).
Krichevsky, A. M. & Gabriely, G. miR-21: a small multi-faceted RNA. J Cell Mol Med 13, 39–53 (2009).
Chan, J. A., Krichevsky, A. M. & Kosik, K. S. MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res 65, 6029–6033 (2005).
Iorio, M. V. et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res 65, 7065–7070 (2005).
Volinia, S. et al. A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci U S A 103, 2257–2261 (2006).
Fulci, V. et al. Quantitative technologies establish a novel microRNA profile of chronic lymphocytic leukemia. Blood 109, 4944–4951 (2007).
Lawrie, C. H. et al. MicroRNA expression distinguishes between germinal center B cell-like and activated B cell-like subtypes of diffuse large B cell lymphoma. Int J Cancer 121, 1156–1161 (2007).
Navarro, A. et al. MicroRNA expression profiling in classic Hodgkin lymphoma. Blood 111, 2825–2832 (2008).
Pezzolesi, M. G., Platzer, P., Waite, K. A. & Eng, C. Differential expression of PTEN-targeting microRNAs miR-19a and miR-21 in Cowden syndrome. Am J Hum Genet 82, 1141–1149 (2008).
Meng, F. et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 133, 647–658 (2007).
Frankel, L. B. et al. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J Biol Chem 283, 1026–1033 (2008).
Gabriely, G. et al. MicroRNA 21 promotes glioma invasion by targeting matrix metalloproteinase regulators. Mol Cell Biol 28, 5369–5380 (2008).
Zhang, Z. et al. miR-21 plays a pivotal role in gastric cancer pathogenesis and progression. Lab Invest 88, 1358–1366 (2008).
Lu, Z. et al. MicroRNA-21 promotes cell transformation by targeting the programmed cell death 4 gene. Oncogene 27, 4373–4379 (2008).
Wang, P. et al. microRNA-21 negatively regulates Cdc25A and cell cycle progression in colon cancer cells. Cancer Res 69, 8157–8165 (2009).
de Oliveira, P. E., Zhang, L., Wang, Z. & Lazo, J. S. Hypoxia-mediated regulation of Cdc25A phosphatase by p21 and miR-21. Cell Cycle 8, 3157–3164 (2009).
Huang, T. H. et al. Up-regulation of miR-21 by HER2/neu signaling promotes cell invasion. J Biol Chem 284, 18515–18524 (2009).
Talotta, F. et al. An autoregulatory loop mediated by miR-21 and PDCD4 controls the AP-1 activity in RAS transformation. Oncogene 28, 73–84 (2009).
Thum, T. et al. Comparison of different miR-21 inhibitor chemistries in a cardiac disease model. J Clin Invest 121, 461–462; author reply 462–463 (2011).
Liu, G. et al. miR-21 mediates fibrogenic activation of pulmonary fibroblasts and lung fibrosis. J Exp Med 207, 1589–1597 (2010).
Thum, T. et al. MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature 456, 980–984 (2008).
Chau, B. N. et al. MicroRNA-21 promotes fibrosis of the kidney by silencing metabolic pathways. Sci Transl Med 4, 121ra118 (2012).
Yang, J. H. et al. starBase: a database for exploring microRNA-mRNA interaction maps from Argonaute CLIP-Seq and Degradome-Seq data. Nucleic Acids Res 39, D202–209 (2011).
Davis, B. N., Hilyard, A. C., Lagna, G. & Hata, A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature 454, 56–61 (2008).
Ito, Y. et al. Antagonistic effects of Smad2 versus Smad7 are sensitive to their expression level during tooth development. J Biol Chem 276, 44163–44172 (2001).
Zhao, J. et al. Smad7 is a TGF-beta-inducible attenuator of Smad2/3-mediated inhibition of embryonic lung morphogenesis. Mech Dev 93, 71–81 (2000).
Fuchshofer, R., Stephan, D. A., Russell, P. & Tamm, E. R. Gene expression profiling of TGFbeta2- and/or BMP7-treated trabecular meshwork cells: Identification of Smad7 as a critical inhibitor of TGF-beta2 signaling. Exp Eye Res 88, 1020–1032 (2009).
Jazag, A. et al. Smad4 silencing in pancreatic cancer cell lines using stable RNA interference and gene expression profiles induced by transforming growth factor-beta. Oncogene 24, 662–671 (2005).
Jazag, A. et al. Single small-interfering RNA expression vector for silencing multiple transforming growth factor-beta pathway components. Nucleic Acids Res 33, e131 (2005).
Acknowledgements
We are grateful to Dr. Junjie Chen (MD Anderson Cancer Center, US) and Dr. Zhaoyuan Hou (Shanghai Jiao Tong University School of Medicine) for valuable discussions and to Dr. Yeguang Chen for the reagents (Tshua University, China). Grant support was provided by the National Program on Key Basic Research Project (973 Program) (2012CB967000) (2012CB910102), Shanghai Committee of Science and Technology (11DZ2260200) and the National Science Foundation of China (81072076) to Dr. Mi.
Author information
Authors and Affiliations
Contributions
Q.L., D.Z. and Y.W. performed most of the experiments; X.H. and P.S. performed some of the experiments; W.X. provided the clinical samples; J.L. provided some of the reagents and advice; J.M. designed the project and wrote the main manuscript text; all authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Li, Q., Zhang, D., Wang, Y. et al. MiR-21/Smad 7 signaling determines TGF-β1-induced CAF formation. Sci Rep 3, 2038 (2013). https://doi.org/10.1038/srep02038
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep02038
This article is cited by
-
Cancer-associated fibroblast-derived exosome microRNA-21 promotes angiogenesis in multiple myeloma
Scientific Reports (2023)
-
Loss of microRNA-21 protects against acetaminophen-induced hepatotoxicity in mice
Archives of Toxicology (2023)
-
Hypoxic lung cancer cell-derived exosomal miR-21 mediates macrophage M2 polarization and promotes cancer cell proliferation through targeting IRF1
World Journal of Surgical Oncology (2022)
-
Signaling pathways in cancer-associated fibroblasts and targeted therapy for cancer
Signal Transduction and Targeted Therapy (2021)
-
The promising role of noncoding RNAs in cancer-associated fibroblasts: an overview of current status and future perspectives
Journal of Hematology & Oncology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.