Abstract
Recent studies show that formaldehyde participates in DNA demethylation/methylation cycle. Emerging evidence identifies that neuronal activity induces global DNA demethylation and re-methylation; and DNA methylation is a critical step for memory formation. These data suggest that endogenous formaldehyde may intrinsically link learning-responsive DNA methylation status and memory formation. Here, we report that during spatial memory formation process, spatial training induces an initial global DNA demethylation and subsequent re-methylation associated with hippocampal formaldehyde elevation then decline to baseline level in Sprague Dawley rats. Scavenging this elevated formaldehyde by formaldehyde-degrading enzyme (FDH), or enhancing DNA demethylation by a DNA demethylating agent, both led to spatial memory deficits by blocking DNA re-methylation in rats. Furthermore, we found that the normal adult rats intrahippocampally injected with excess formaldehyde can imitate the aged-related spatial memory deficits and global DNA methylation decline. These findings indicate that aging-associated excess formaldheyde contributes to cognitive decline during aging.
Similar content being viewed by others
Introduction
Formaldehyde (FA) is present in all invertebrate and vertebrate cells1,2. Using fluorescence-HPLC, we have previously shown that levels of normal physiological brain formaldehyde are about 0.2 ~ 0.4 mM, similar to levels determined by other groups using gas chromatography/mass spectrometry3,4. Enzymatic degradation of formaldehyde is mainly regulated by alcohol dehydrogenase 3 (ADH3) and aldehyde dehydrogenase 2 (ALDH2)5. ADH3 (also known as formaldehyde dehydrogenase, FDH6), is distributed in the hippocampus and cortex and helps to defend the brain against neurodegenerative processes6. ADH3-knockout mice are sensitive to formaldehyde toxicity7 and only less than 20% of these mice can survive after postnatal day 68. Moreover, knockout of ALDH2 also induces age-dependent memory loss in mice9. In our previous study, inhibition of ADH3 or ALDH2 activity results in hippocampal formaldehyde accumulation and cognitive impairments in rats10. However, over-expression of S-nitrosoglutathione reductase (GSNOR, a mammalian homolog of ADH3 that accelerates the degradation of formaldehyde11) in Drosophila markedly induces visual memory loss12. These data indicate that either an abnormal excess or a deficiency of endogenous formaldehyde leads to memory impairments.
However, under physiological condition, endogenous formaldehyde participates in DNA demethylation and methylation cycle. Recent studies have shown that DNA demethylation leads to the release of formaldehyde (as a methyl moiety) or methanol13 (methanol is reversibly transformed into formaldehyde by ADH15). In addition, S-adenosylmethionine-dependent DNMTs catalyze reversible covalent addition of exogenous aliphatic aldehydes (for instance formaldehyde) to their target residues in DNA14, because DNA-5-methyl-cytosine is the product between formaldehyde and DNA-cytosine (methyl group acceptor)2. Emerging evidence has shown that external stimuli (electroconvulsive stimulation), not only elicit initial transient DNA demethylation then re-methylation of memory-related genes15, but also induce initial global DNA demethylation and re-methylation in the hippocampus16. That is, DNA methylation (regulated by DNA methyltransferases, DNMTs17), is a critical step in neuro-epigenetic memory formation and remote memory maintenance18,19. These diverse lines of evidence strongly suggest a possible link between endogenous formaldehyde and memory-related DNA methylation status.
Accumulating evidence has shown that environmental pollutions, diet, aging and genetic factors can induce endogenous formaldehyde accumulation10. Both exogenous and endogenous excess formaldehyde can induce animal and human cognitive impairments20,21,22,23, especially in Alzheimer's disease (AD) transgenic mice and AD patients10. Recent studies have identified that DNA methylation plays a critical role in memory formation18,19. However, DNMT activity and global DNA methylation are obviously decreased in brains of rats during aging24. Moreover, global DNA methylation decline has been observed in autopsy brain samples from AD patients25. Although we have demonstrated that chronic excess formaldehyde induces cognitive deficits in rats10, the relationship between the excess formaldehyde-memory deficits and global DNA methylation decline during aging has not been clarified.
Results
Changes in global DNA methylation status and formaldehyde levels during LTP formation
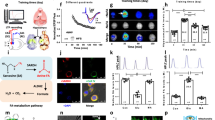
First, we investigated how external stimuli induces the dynamic changes in global DNA status and formaldehyde concentration. To explore the relationship between the elevated hippocampal formaldehyde levels and the intensity of external stimulation, an increase in the number of electrical stimuli trains or duration of spatial training were applied in this study. In anesthetized normal adult male SD rats, three trains of HFS (Three-HFS) treatments induced higher amplitudes of fEPSP than a single HFS treatment ( Fig. 1A, 1D ) and were associated with a higher elevation in formaldehyde level (~0.08 mM) than that of a single HFS treatment (~0.06 mM) after electrical stimuli 5 minutes ( Fig. 1B, 1E ). Markedly, three-HFS treatments induced a higher degree of global DNA demethylation (~3%, similar to previous report16) and formaldheyde generation (~0.08 mM) than a single HFS treatment after electrical stimuli 5 minutes ( Fig. 1C, 1F ). This result indicates that electrical stimuli induce initial global DNA demethylation and subsequent re-methylation associated with formaldheyde elevation then decline to baseline level.
Electrical stimuli induce the changes in global DNA methylation status and formaldehyde level in hippocampus of rats during LTP formation.
(A) High frequency stimulation (HFS) induces long-term potentiation (LTP) formation in vivo. (B) Changes in hippocampal FA levels after HFS treatment. (C) HFS treatment induces global DNA demethylation then methylation. (D) Three rounds of HFS treatment (Three-HFS) induce late long-term potentiation (L-LTP) formation in vivo. (E) Changes in hippocampal FA levels after Three-HFS treatment. (F) Three-HFS treatment induces global DNA demethylation then methylation. **, P < 0.01.
Changes in global DNA methylation status and formaldehyde levels during spatial memory formation
By varying the duration of training in the Morris water maze, we observed whether spatial memory was formatted in rats. We did not observe that spatial training for one day induces the changes in global DNA methylation and hippocampal formaldehyde ( Fig. 2A–2C ). However, spatial training for 3 days obviously elicited global DNA demethylation associated with an elevation in hippocampal formaldehyde concentration (~0.03 mM) on day 3 and a validate spatial memory formation on day 4 ( Fig. 2A, 2D–2E ). Furthermore, spatial training for 6 days resulted in an obvious global DNA demethylation associated with an elevation in hippocampal formaldehyde concentration (~0.08 mM) on day 6 and a stable spatial memory formation associated with global DNA re-methylation on day 7 ( Fig. 2A, 2G–2H ). These results demonstrate that learning activity also induces a dynamic change in global DNA methylatin status and formaldheyde level as well as LTP formation .
Spatial training induces the changes in global DNA methylation status and formaldehyde level in hippocampus of rats during spatial memory formation.
(A) Escape latency of the rats with training for 1, 3 and 6 days. (B) Time in target quadrant after probe trial on day 2, 4 and 7, respectively. (C) Changes in hippocampal formaldehyde in rats with training for one day and memory retrieval on day 2. (D) Changes in global DNA methylation on training for one day and memory retrieval on day 2. (E) Changes in hippocampal formaldehyde in rats with training for 3 days and memory retrieval on day 4. (F) Changes in global DNA methylation on training for 3 days and memory retrieval on day 4. (G) Changes in hippocampal formaldehyde in rats with training for 6 days and memory retrieval on day 7. (H) Changes in global DNA methylation on training for 6 days and memory retrieval on day 7. *, P < 0.05; **, P < 0.01. Sham: home caged rats for spatial training group or memory retrieval group.
Scavenging the elevated formaldehyde impairs LTP and spatial memory in rats
To explore whether this elevated formaldehyde (~0.08 mM) is indispensable for memory formation, a formaldehyde scavenger-formaldehyde dehydrogenase (FDH, a specific formaldehyde-degrading enzyme26) was used in this study. After high frequency stimulation (HFS, 100 Hz), we observed an elevation in hippocampal formaldehyde level ( : 0.06 mM), however, this elevation was suppressed in rats injected with FDH (i.c.v.). Injection with FDH also markedly impaired hippocampal LTP formation (
Fig. 3A, 3B
). Notably, FDH injection did not inhibit initial global DNA demethylation, but abolished subsequent global DNA re-methylation after HFS treatment 60 min (
Fig. 3C
).
: 0.06 mM), however, this elevation was suppressed in rats injected with FDH (i.c.v.). Injection with FDH also markedly impaired hippocampal LTP formation (
Fig. 3A, 3B
). Notably, FDH injection did not inhibit initial global DNA demethylation, but abolished subsequent global DNA re-methylation after HFS treatment 60 min (
Fig. 3C
).
Scavenging of hippocampal formaldehyde (FA) impairs LTP and spatial memory in SD rats.
(A) Intracerebroventricular injection with the FA scavenger-FDH (120 μM, 5 μL, i.c.v.) impairs hippocampal LTP in vivo. (B) Injection with FDH alone obviously reduces hippocampal FA levels detected at 65th and 125th min (##, P < 0.01). HFS (saline, 5 μL, i.c.v.) treatment induces an elevation in FA levels at 65th min (**, P < 0.01). HFS with FDH injection decreases FA levels detected at 65th min ($$, P < 0.01). (C) Injection with FDH alone blocks global DNA methylation detected at 125th min (##, P < 0.01). HFS treatment induces global DNA demethylation and re-methylation (**, P < 0.01). HFS with FDH injection blocks global DNA re-methylation detected at 125th min ($$, P < 0.01). (D, E) Intrahippocampal injection with only saline or FDH (120 μM, 1 μL, over a 5 min period) affects the ability of spatial learning and memory retrieval in rats. (F) Injection with FDH blocks the global DNA re-methylation on day 7. **, P < 0.01; $$, P < 0.01; ##, P < 0.01. Sham: home caged rats.
Furthermore, intrahippocampal injection with FDH markedly not only impaired the ability of spatial learning on day 3, 4, 5 and 6, but also damaged the ability of memory retrieval on day 7 ( Fig. 3D, 3E ). Similarly, FDH injection did not inhibit the initial global DNA demethylation on day 6, but abolished global DNA re-methylation on day 7 ( Fig. 3F ). This suggests that the elevated formaldehyde may contribute to subsequent DNA re-methylation and the deficiency of DNA re-methylation impairs spatial memory.
Blocking DNA re-methylation impairs LTP and spatial memory in rats
To further confirm whether inhibition of DNA re-methylation leads to spatial memory deficits, 5-aza-2-deoxycytidine (5-AzA, a DNA demethylating agent by inhibiting DNMT activity27,28), was used in this study. We found that injection with 5-AzA (i.c.v.) markedly suppressed hippocampal LTP formation ( Fig. 4A ). Notably, 5-AzA injection enhanced transient global DNA demethylation and formaldehyde accumulation after HFS treatment 5 min ( Fig. 4B ) and reduced global DNA re-methylation after HFS treatment 60 min ( Fig. 4C ). Furthermore, intrahippocampal injection with 5-AzA not only impaired the ability of spatial learning on day 3, 4, 5 and 6, but also damaged the ability of memory retrieval on day 7 ( Fig. 4D, 4E ). Similarly, 5-AzA enhanced global DNA demethylation on day 6 and obviously reduced global DNA re-methylation on day 7 ( Fig. 4F ). This result also indicates that blocking DNA re-methylation leads to memory deficits.
Blocking DNA methylation impairs LTP and spatial memory in SD rats.
(A) Injection with only vehicle (0.001% CH3COOH) or 5-AzA (60 μM, 5 μL, i.c.v.) affects hippocampal LTP in vivo. (B) Injection with 5-AzA alone increases hippocampal FA levels detected at 65th and 125th min (##, P < 0.01). HFS treatment induces FA levels elevation then decline (**, P < 0.01). HFS with 5-AzA injection increases FA levels detected at 65th and 125th min ($$, P < 0.01). (C) Injection with 5-AzA alone reduces global DNA methylation detected at 65th and 125th min (##, P < 0.01). HFS treatment induces global DNA demethylation and re-methylation (**, P < 0.01). HFS with 5-AzA injection reduced global DNA re-methylation detected at 65th and 125th min ($$, P < 0.01). (D, E) Intrahippocampal injection with only solvent or 5-AzA (60 μM, 1 μL, over a 5 min period) affects the ability of spatial learning and memory retrieval in rats. (F) 5-AzA injection markedly reduces global DNA re-methylation on day 7. **, P < 0.01; $$, P < 0.01; ##, P < 0.01. Sham: home caged rats.
DNA demethylation elicits formaldehyde generation in the dividing cells and post-mitotic neurons
1) External stimulation induces DNA demethylation in vitro and in vivo
Previous studies have shown that neuronal activity induced by electroconvulsive stimulation and NMDA receptor-mediated synaptic activity, can trigger DNA demethylation15,16,29. Furthermore, treatment with NMDA-receptor antagonist APV (DL-2-amino-5-phosphonovaleric acid) or Ca2+ chelator BAPTA (1,2-bis- (o-aminophenoxy)-ethane-N,N,N′,N′-tetraacetic acid) is known to inhibit DNA demethylation29,30. These data imply that activation of the NMDA-receptor-Ca2+-pathway is necessary for external stimulation-induced DNA demethylation ( Fig. 5A ). To verify this hypothesis, we used NMDA (an excitatory neurotransmitter) to treat the cultured hippocampal neurons for 30, 60 and 180 minutes, respectively. There was an obvious time-dependently elevation in formaldehyde concentration in media of cultures and BAPTA markedly reduced formaldehyde accumulation in vitro ( Fig. 5B ). After HFS (200 Hz) stimulation, there was a significant decrease (about 2.5%) in global DNA methylation (DNA demethylaiton); BAPTA treatment obviously blocked HFS-induced global DNA demethylation in hippocampus of the normal SD rats ( Fig. 5C ). In vivo study, global DNA demethylation induced by external stimuli (WFS, HFS, Three-HFS) or spatial training was positively correlated (R = 0.930) with formaldehyde generation in hippocampus of the SD rats ( Fig. 5D ).
Excess formaldehyde (FA) is accumulated in hippocampus of the SD rats during aging.
(A) The proposed model of NMDA-induced Ca2+ influx and DNA demethylation associated FA generation. (B) NMDA induces a time-dependent FA generation, BAPTA (Ca2+ chelator) inhibited NMDA-induced the production of FA. (C) BAPTA (Ca2+ chelator) blocks HFS-induced global DNA demethylaiton. (D) The formaldehyde generation was positively correlated with global DNA demethylation after electrical stimulation or spatial training in vivo. (E) The proposed model of DNA demethylation induced by 5-AzA (an inhibitor of DNMTs). (F) 5-AzA induces FA generation in the cultured medium of hippocampal neurons within 30, 60 minutes. (G, H) Intrahippocampal injection with 5-AzA obviously induces the changes including: global DNA methylation decline, FA generation and DNMT activity decline in hippocampus of rats. (I, J, K) 5-AzA treatment induces FA generation, DNMT activity decline and global DNA methylation decline in human neuroblastoma SY5Y cell lines. (L) The formaldehyde generation was negatively correlated with the decrease in global DNA methylation levels in vitro. **, P < 0.01.
2) DNA demethylating agent induces DNA demethylation in human SY5Y cell lines
As mentioned above, that 5-AzA induced global DNA demethylation associated with hippocampal formaldehyde accumulation in vivo has been observed ( Fig. 4B ). To provide the direct evidence that DNA demethylation elicits formaldehyde generation, the authorized passive demethylating agent ( Fig. 5E ), 5-AzA, was applied in post-mitotic neurons (hippocampal neurons) and the dividing cells (Human SY5Y cell lines) in vitro. We found that 5-AzA treatment induced a time-dependent elevation in formaldehyde concentrations in the cultured medium of hippocampal neurons ( Fig. 5F ) and a dose-dependent elevation in formaldehyde levels in hippocampus by inhibiting DNMT activity in vivo ( Fig. 5G, 5H ). Similarly, 5-AzA treatment for several hours also resulted in a time-dependent elevation in formaldehyde concentrations and a decrease in global DNA methylation by inhibiting DNMT activity in the cultured SY5Y cell lines ( Fig. 5I–5K ). Global DNA methylation levels affected by 5-AzA were negatively correlated (R = 0.978) with formaldehyde generation in hippocampus. The data provide evidence that DNA demethylation induces formaldehyde generation13,31.
Aging-associated with excess formaldehyde induces spatial memory decline
1) Aging induces excess formaldehyde accumulation in hippocampus
Age-related excess hippocampal formaldehyde accumulation has been observed in our previous study10. Then, to explore the relationship between global DNA methylation and formaldehyde accumulation during aging, the normal SD rats with 2, 16 and 32-months old were employed. We found that global DNA methylation levels were gradually decreased in the SD rats during aging ( Fig. 6A ). Hippocampal formaldehyde concentrations were significantly elevated to the pathological level (~0.5 mM) in the aged rats ( Fig. 6B ). This suggests that excess formaldehyde is associated with global DNA methylation decline (DNA demethylation) in hippocampus during aging.
Excess formaldehyde (FA) impairs both hippocampal LTP and spatial memory in normal adult SD rats.
(A) Global DNA methylation is decline in hippocampus of rats during aging. (B) FA is accumulated in hippocampus of rats during aging. (C, D) The effects of different concentrations of FA on DNMT activity in vitro and in vivo. (E, F) Excess formaldehyde (0.05 mM, i.c.v.) suppresses hippocampal long-term potentiation (LTP) in vivo. (G) Injection with only saline or excess FA alone affects hippocampal FA levels detected at 65th and 125th min (##, P < 0.01). HFS treatment induces an elevation in FA levels at 65th min (**, P < 0.01). HFS with FA injection increases FA levels detected at 65th and 125th min ($$, P < 0.01). (H) Injection with FA alone reduces global DNA methylation detected at 125th min (##, P < 0.01). HFS treatment induces global DNA demethylation and re-methylation (**, P < 0.01). HFS with FA injection blocks global DNA re-methylation detected at 125th min ($$, P < 0.01). (I) Intrahippocampal injection with only saline or excess FA or resveratrol (Res) (0.5 mM, 1 μL, over a 5 min period) affects the ability of spatial learning in the adult or aged rats. (J) Intrahippocampal injection with resveratrol (Res) rescues the impairments of the ability of spatial memory retrieval on day 7 in the aged rats. (K) Intrahippocampal injection with resveratrol (Res) reduces excess hippocampal FA in the aged rats. (L) Intrahippocampal injection with resveratrol rescues global DNA methylation decline in hippocampus of aged rats. T-adult: the adult rats with spatial training; T-adult-FA: the adult rats with spatial training and excess FA injection; T-aged: the aged rats with spatial training; Sham: home caged rats. T-aged-Res: the aged rats with spatial training and resveratrol injection. **, P < 0.01; $$, P < 0.01; ##, P < 0.01.
2) Excess formaldehyde blocks DNA methylation by inhibiting DNMT activity
Then, we investigated how excess formaldehyde induces global DNA methylation decline during aging. Since inhibition of DNMT activity by 5-AzA induced formaldehyde generation, this suggests that the decrease in DNMT activity may be the cause of global DNA methylation decline during aging. Therefore, we examined whether formaldehyde regulates DNMT activity in vitro and in vivo. Administration of 0.5 mM formaldehyde (i.c.v.) markedly inhibited DNMT activity in the hippocampus in vivo. Excess formaldehyde (0.5 mM) treatment obviously inhibited DNMT activity in cultured hippocampal neurons ( Fig. 6C, 6D ). Approximately, 0.08 mM formaldehyde markedly increased the DNMT activity of cultured hippocampal neurons and hippocampus in vivo ( Fig. 6C, 6D ). These results indicate that ~0.08 mM formaldehyde enhances DNMT activity; but, age-related excess formaldehyde inhibits DNMT activity.
3) Intrahippocampal injection with excess formaldheyde induces memory deficits in adult rats
To examine whether excess formaldehyde induces age-related cognitive decline, the normal adult SD rats were intrahippocampally injected with excess formaldehyde (0.5 mM, the average concentration was detected in the aged rats), then spatial memory behaviors were observed. Excess formaldehyde injection (0.5 mM, i.c.v.) markedly suppressed hippocampal LTP formation ( Fig. 6E, 6F ). Excess formaldehyde injection induced hippocampal formaldehyde accumulation associated with the initial decline in global DNA methylation after HFS treatment 5 min and abolished subsequent global DNA re-methylation after HFS treatment 60 min ( Fig. 6G, 6H ).
Furthermore, intrahippocampal injection with excess formaldehyde (0.5 mM) obviously not only impaired the ability of spatial learning in the adult rats, but also damaged the ability of memory retrieval associated with a decrease in global DNA methylation on day 7. More importantly, the changes in formaldehyde and global DNA methylation in the adult rats injected with excess formaldheyde were similar to the result of the aged rats ( Fig. 6I–6L ). These data suggest that aging-associated excess formaldehyde impairs spatial memory by blocking DNA methylation. Finally, resveratrol32, a natural formaldehyde scavenger, was applied into hippocampus of the aged rats, we found that resveratrol obviously improved the behavious performance of spatial memory in the aged rats associated with an increase in global DNA methylation ( Fig. 6I–6L ).
Discussion
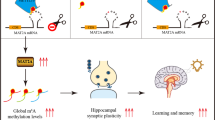
In the present study, we found that spatial training induced an initial global DNA demethylation and subsequent re-methylation and blocking DNA re-methylation led to spatial memory deficits. More importantly, aging-associated excess formaldehyde impairs spatial memory by blocking DNA methylation ( Fig. S1 ). Our results support that neuronal activity induces DNA demethylation and re-methylation during memory formation.
Emerging evidence has identified that both electroconvulsive stimulation and acute exercise induce a transient global DNA demethylation33. Electrical stimulation can induce transient DNA to RNA transcription (such as c-fos) in several seconds and DNA demethylation is required for gene activation34. This suggests that external stimuli can elicit a transient DNA demethylation. In this study, we also observed that external stimuli (WFS, HFS, Three-HFS and spatial training) elicit initial transient global DNA demethylation and formaldehyde generation in hippocampus ( Fig. 1 , 2 ). Recent studies suggest that neuronal activity induced by external stimulation can trigger DNA demethylation via NMDA-receptor pathway15,16,29. Calcium chelator BAPTA can inhibit DNA demethylation29,30. Similarly, we also found that there was a significant decrease in formaldehyde generation and a marked inhibition of global DNA demethylation after treatment with BAPTA ( Fig. 5B, 5C ). Previous studies have shown that inhibition of DNMT activity elicits DNA demethylation in the dividing cancer cells27,28 and DNA demethylation leads to formaldehyde generation13,31 ( Fig. S2A ). We also found that 5-AzA obviously induces DNA demethylation and formaldehyde accumulation in human SY5Y cell lines in several hours.
Whether DNA demethylation occurs in post-mitotic neurons is rarely reported. Recently, however, Levenson and colleagues have found that 5-AzA (a DNA demethylation reagent) obviously induces DNA demethylation of reelin gene (increasing unmethylated DNA and decreasing methylated DNA) in hippocampal slice after adminitration 40 minutes and this gene is expressed in hippocampal neurons35. In the present study, 5-AzA treatment elicited global DNA demethylation in hippocampal neurons in vitro and hippocampus of rats in vivo ( Fig. 5E–5K ). These data indicate that external stimuli can induce DNA demethylation and formaldehyde generation in post-mitotic neurons. There are multiple pathways of DNA demethylation, such as: direct active process; demethylation; indirect active process: mismatch/excision repair; and passive loss of methyl group13. However, the exact mechanism that neuroal activity induces DNA demethylation is currently unclear36.
Recent studies have shown that formaldehyde participates in DNA methylation, for DNA-5-methyl-cytosine is a product between formaldehyde and DNA-cytosine (methyl group acceptor)2. S-Adenosylmethionine-dependent DNMTs catalyze reversible covalent addition of exogenous formaldehyde (as a methyl moiety) to their target residues in DNA, yielding corresponding 5-hydroxymethylcytosine (5 hmC) and 5-methylcytosine (5 mC)14 ( Fig. S2B ). DNA methylation is required for memory formation19. In the present study, we found that external stimuli elevate hippocampal formaldehyde concentration (~0.08 mM) ( Fig. 1 ). Formaldehyde at 0.08 mM signifcantly enhanced DNMT activity both in vitro and in vivo ( Fig. 6C, 6D ). Conversely, scavenging the elevated hippocampal formaldehyde by FDH treatment did not affect DNA demethylation, but obviously blocked global DNA re-methylation and impaired spatial memory in rats ( Fig. 3 ). Over-expression of FDH (ADH3) in Drosophila can induce visual memory loss12. Especially, knockout of DNMT1a is known to impair LTP and memory17. Our data along with these reports suggest that ~0.08 mM formaldehyde derived from the initial transient DNA demethyltion induced by external stimuli, may contribute to enhance subsequent DNA re-methylation at least in part via DNMTs.
Endogenous formaldehyde is kept a relative stable level in body, but there is a chronic accumulation of formaldehyde during aging. For example, a formaldehyde generating enzyme, semicarbazide-sensitive amine oxidase (SSAO), SSAO activity is increased in human plasma during aging37 and associated with formaldehyde accumulation38. A specific formaldehyde degrading enzyme39, ADH3 (FDH), its activity is decreased in human during aging40. Recent studies show that FDH can scavenge formaldehyde by transferring S-hydroxymenthyl which is a derivative between formaldehyde with GSH5,41,42. In the present study, we also found that extracellular FDH reduced hippocampal formaldehyde ( Fig. 3 ). An unspecific formaldehyde degrading enzyme, ALDH243, its activity is also attenuated in human and diabetic rats during aging44. We also have found that inhibiting ADH3 or ALDH2 leads to an abnormal high concentration of formaldehyde accumulated in hippocampus in rats10. Especially, over-expression of ADH3 (a specific formaldehyde degrading enzyme) leads to visual memory decline in Drosophila12. Deficiency of ALDH2 (a unespecific formaldehyde degrading enzyme) induces memory loss in mice9. Age-related oxidative stress can increase in the content of reactive aldehydes45, including formaldehyde46. In this study, we found that age-related formaldehyde was over 0.5 mM in the aged rats ( Fig. 6B ). All these data support that endogenous formaldehyde is accumulated during aging.
In the present study, we found that endogenous formaldehyde participates in spatial memory formation, but during aging process, aging-associated excess formaldehyde blocks global DNA methylation and leads to memory decline. Other reports also show that DNMT activity and global DNA methylation are decreased in brains during aging24. We found that formaldehyde was over 0.5 mM in the aged rats and formaldehyde at 0.5 mM (the pathological level) obviously inhibited the DNMTs activity in vitro and in vivo, but formaldehyde at 0.3 mM (the physiological level for endogenous formaldehyde) did not affect DNMT activity ( Fig. 6C, 6D ). More importantly, the normal adult rats intrahippocampally injected with this excess formaldehyde (0.5 mM) can imitate the aged-related spatial memory deficits associated with global DNA methylation decline and formaldehyde accumulation ( Fg. 6 ). These data suggest that age-related accumulated formaldehyde (over 0.5 mM) erasues global DNA methylation by inhibiting DNMT activity and restults in spatial memory deficits. Notably, a natural formaldehyde scavenger-resveratrol32, can rescue the behavior performance of spatial memory in the aged rats by decreasing hippocampal formaldehyde concentration ( Fig. 6I–6L ). Resveratrol has been considered as a therapeutic agent for neurodegenerative diseases47. Dietary supplementation with resveratrol can reduce plaque pathology in a transgenic model of Alzheimer's disease48. Our results along with these reports suggest that aging-associated excess formaldehyde is one of the critical pathological factors for cognitive decline in neurodegenerative disease.
Methods
Reagents
All reagents were from Sigma unless otherwise indicated.
Biological samples
All animal experiments were conducted in accordance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals and were approved by the Biological Research Ethics Committee of the Institute of Biophysics, Chinese Academy of Sciences (SYXK2010-127).
Intracerebroventricular formaldehyde injection
Sprague Dawley (SD) rats were intracerebroventricularly (i.c.v.) injected with formaldehyde (dissolved in normal saline, 0.01 and 0.08 mM, 5 μl, i.c.v. over a period of 5 min), or resveratrol (Res, 0.5 mM, 5 μl). Cannulae placements in brains: 1.4 mm anterior/posterior to bregma; 1.8 mm medial/lateral to the midsagittal suture; 3.0 mm dorsal/ventral from the brain surface.
Intrahippocampal formaldehyde injection
Sprague Dawley (SD) rats were intrahippocampally injected with formaldehyde (0.08 mM, 1 μl, over a period of 5 min), or resveratrol (Res, 0.5 mM, 1 μl) 30 min before the daily behavioral experiments. Cannulae placements in the dorsal hippocampus: 3.6 mm anterior/posterior to bregma; 2.5 mm medial/lateral to the midsagittal suture; and 2.5 mm dorsal/ventral from the brain surface.
Rat hippocampal neuron and human SY5Y cell lines cultures
We prepared primary hippocampal neuron cultures from 18-day old embryos and cultured human SY5Y cell lines as described previously49. Mature hippocampal neurons or cultrued media after drugs treatment were collected in order to detect formaldehyde and the activity of DNMTs in vitro.
Detection of formaldehyde by Fluo-HPLC
Drug-treated medium from cultured hippocampal neurons was collected and immediately placed on ice before storing at −70°C. After centrifugation (3000 rpm, 4°C, 10 min), supernatant fractions of medium samples and brain homogenates (weight of brain tissue: ultrapure water = 1:4) were analyzed by high-performance liquid chromatography with fluorescence detection (Fluo-HPLC) as described10.
Electrophysiological studies in vivo
Electrophysiological recordings were performed according to our previous reports10. WFS-LTP in vivo LTP was induced using a weak frequency stimulation (WFS) protocol consisting of 20 pulses at 50 Hz.
HFS-LTP in vivo
LTP was induced using a high frequency stimulation (HFS) protocol consisting of 20 pulses at 100 Hz or 200 Hz.
L-LTP in vivo
Three pulses of HFS at 5 min intervals were applied to produce a robust late long-term potentiation (L-LTP).
Morris water maze behavioral test
Spatial memory was assessed by the Morris water maze test, as described previously10. Sprague Dawley (SD) rats were intrahippocampally injected with FDH (120 μM, 1 μl), or 5-aza-2-deoxycytidine (5-AzA, 60 μM, 1 μl, vehicle (0.001% CH3COOH)), or excess formaldehyde (0.5 mM, 1 μl), or resveratrol (Res, 0.5 mM, 1 μl), 30 min before the daily behavioral experiment.
Quantification of global DNA methylation
A MethylampTM Global DNA Methylation Quantification Ultra Kit (Epigentek, USA) was used according to the manufacturer's instructions to quantify global DNA methylation of genomic DNA in the hippocampus of SD rats.
DNA methyltransferase (DNMT) assay
Nuclei were extracted from rat hippocampi using an EpiQuik™ nuclear extraction kit (Epigentek, USA). DNA methyltransferase activity was assayed using a DNMT assay kit (Epigentek, USA) according to the manufacturer's instructions.
Statistical analysis
We determined the statistical significance of in vivo data by analysis of variance (one-way ANOVA) followed by unpaired Student's t-tests; and a repeated measures ANOVA followed by post-hoc analysis was used to determine statistical significance for the Morris water maze experiments. The statistical significance level was set at P < 0.05. Data are reported as means ± standard errors.
References
Kalapos, M. P. A possible evolutionary role of formaldehyde. Exp Mol Med 31, 1–4 (1999).
Kalasz, H. Biological role of formaldehyde and cycles related to methylation, demethylation and formaldehyde production. Mini Rev Med Chem 3, 175–192 (2003).
Heck, H. D., White, E. L. & Casanova-Schmitz, M. Determination of formaldehyde in biological tissues by gas chromatography/mass spectrometry. Biomed Mass Spectrom 9, 347–353 (1982).
Tong, Z. Q. et al. Urine formaldehyde level is inversely correlated to mini mental state examination scores in senile dementia. Neurobiol Aging 32, 31–42 (2011).
Teng, S. et al. The formaldehyde metabolic detoxification enzyme systems and molecular cytotoxic mechanism in isolated rat hepatocytes. Chem Biol Interact 130–132, 285–296 (2001).
Mori, O. et al. Histological distribution of class III alcohol dehydrogenase in human brain. Brain Res 852, 186–190 (2000).
Deltour, L., Foglio, M. H. & Duester, G. Metabolic deficiencies in alcohol dehydrogenase Adh1, Adh3 and Adh4 null mutant mice. Overlapping roles of Adh1 and Adh4 in ethanol clearance and metabolism of retinol to retinoic acid. J Biol Chem 274, 16796–16801 (1999).
Molotkov, A. et al. Stimulation of retinoic acid production and growth by ubiquitously expressed alcohol dehydrogenase Adh3. Proc Natl Acad Sci U S A 99, 5337–5342 (2002).
Ohsawa, I. et al. Age-dependent neurodegeneration accompanying memory loss in transgenic mice defective in mitochondrial aldehyde dehydrogenase 2 activity. J Neurosci 28, 6239–6249 (2008).
Tong, Z. Q. et al. Accumulated hippocampal formaldehyde induces age-dependent memory decline. AGE [Epub ahead of print] (2012).
Greenwald, R. et al. Breath formate is a marker of airway S-nitrosothiol depletion in severe asthma. PLoS One 5, e11919 (2010).
Hou, Q. et al. Nitric oxide metabolism controlled by formaldehyde dehydrogenase (fdh, homolog of mammalian GSNOR) plays a crucial role in visual pattern memory in Drosophila. Nitric Oxide 24, 17–24 (2011).
Patra, S. K. et al. Demethylation of (Cytosine-5-C-methyl) DNA and regulation of transcription in the epigenetic pathways of cancer development. Cancer Metastasis Rev 27, 315–334 (2008).
Liutkeviciute, Z., Lukinavicius, G., Masevicius, V., Daujotyte, D. & Klimasauskas, S. Cytosine-5-methyltransferases add aldehydes to DNA. Nat Chem Biol 5, 400–402 (2009).
Ma, D. K. et al. Neuronal activity-induced Gadd45b promotes epigenetic DNA demethylation and adult neurogenesis. Science 323, 1074–1077 (2009).
Guo, J. U. et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat Neurosci 14, 1345–1351 (2011).
Feng, J. et al. Dnmt1 and Dnmt3a maintain DNA methylation and regulate synaptic function in adult forebrain neurons. Nat Neurosci 13, 423–430 (2010).
Miller, C. A., Campbell, S. L. & Sweatt, J. D. DNA methylation and histone acetylation work in concert to regulate memory formation and synaptic plasticity. Neurobiol Learn Mem 89, 599–603 (2008).
Miller, C. A. et al. Cortical DNA methylation maintains remote memory. Nat Neurosci 13, 664–666 (2010).
Lu, Z. S. et al. Effect of inhaled formaldehyde on learning and memory of mice. Indoor Air 18, 77–83 (2008).
Malek, F. A., Möritz, K. U. & Fanghänel, J. A study on the effect of inhalative formaldehyde exposure on water labyrinth test performance in rats. Ann Anat 185, 277–285 (2003).
Kilburn, K. H. Neurobehavioral impairment and seizures from formaldehyde. Arch Environ Health 49, 37–44 (1994).
Perna, R. B., Bordini, E. J. & Deinzer-Lifrak, M. A case of claimed persistent neuropsychological sequelae of chronic formaldehyde exposure: clinical, psychometric and functional findings. Arch Clin Neuropsychol 16, 33–44 (2001).
Liu, L. et al. Insufficient DNA methylation affects healthy aging and promotes age-related health problems. Clin Epigenetics 2, 349–360 (2011).
Mastroeni, D. et al. Epigenetic changes in Alzheimer's disease: decrements in DNA methylation. Neurobiol Aging 31, 2025–2037 (2010).
Thompson, C. M., Ceder, R. & Grafstrom, R. C. Formaldehyde dehydrogenase: beyond phase I metabolism. Toxicol Lett 193, 1–3 (2010).
Yamagata, Y. et al. Rapid turnover of DNA methylation in human cells. Epigenetics 7, 141–145 (2012).
Hamm, C. A. et al. Global demethylation of rat chondrosarcoma cells after treatment with 5-aza-2′-deoxycytidine results in increased tumorigenicity. PLoS One 4, e8340 (2009).
Nelson, E. D., Kavalali, E. T. & Monteggia, L. M. Activity-dependent suppression of miniature neurotransmission through the regulation of DNA methylation. J Neurosci 28, 395–406 (2008).
Collas, P. Modulation of plasmid DNA methylation and expression in zebrafish embryos. Nucleic Acids Res 26, 4454–4461 (1998).
Gerken, T. et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318, 1469–1472(2007).
Tyihak, E. et al. Formaldehyde cycle and the natural formaldehyde generators and capturers. Acta Biol Hung 49, 225–238 (1998).
Barres, R. et al. Acute exercise remodels promoter methylation in human skeletal muscle. Cell Metab 15, 405–411 (2012).
Ikeda, Y. & Kinoshita, T. DNA demethylation: a lesson from the garden. Chromosoma 118, 37–41 (2009).
Levenson, J. M. et al. Evidence that DNA (cytosine-5) methyltransferase regulates synaptic plasticity in the hippocampus. J Biol Chem 281, 15763–15773 (2006).
Wu, S. C. & Zhang, Y. Active DNA demethylation: many roads lead to Rome. Nat Rev Mol Cell Biol 11, 607–620 (2010).
del Mar Hernandez, M., Esteban, M., Szabo, P., Boada, M. & Unzeta, M. Human plasma semicarbazide sensitive amine oxidase (SSAO), beta-amyloid protein and aging. Neurosci Lett 384, 183–187 (2005).
Conklin, D. J. et al. Vasoactive effects of methylamine in isolated human blood vessels: role of semicarbazide-sensitive amine oxidase, formaldehyde and hydrogen peroxide. Am J Physiol Heart Circ Physiol 286, H667–676 (2004).
Moreno, A., Pares, A., Ortiz, J., Enriquez, J. & Pares, X. Alcohol dehydrogenase from human stomach: variability in normal mucosa and effect of age, gender, ADH3 phenotype and gastric region. Alcohol Alcohol 29, 663–671 (1994).
Hines, L. M. et al. A prospective study of the effect of alcohol consumption and ADH3 genotype on plasma steroid hormone levels and breast cancer risk. Cancer Epidemiol Biomarkers Prev 9, 1099–1105 (2000).
Tulpule, K. et al. Formaldehyde induces rapid glutathione export from viable oligodendroglial OLN-93 cells. Neurochem Int 61, 1302–1313 (2012).
Tsuboi, S. et al. Purification and characterization of formaldehyde dehydrogenase from rat liver cytosol. J Biochem 111, 465–471 (1992).
Luo, H. R., Israel, Y., Tu, G. C., Eriksson, C. J. & Zhang, Y. P. Genetic polymorphism of aldehyde dehydrogenase 2 (ALDH2) in a Chinese population: gender, age, culture and genotypes of ALDH2. Biochem Genet 43, 223–227 (2005).
Wang, J. et al. Inhibition of aldehyde dehydrogenase 2 by oxidative stress is associated with cardiac dysfunction in diabetic rats. Mol Med 17, 172–179 (2011).
Maboudou, P., Mathieu, D., Bachelet, H., Wiart, J. F. & Lhermitte, M. Detection of oxidative stress. Interest of GC-MS for malondialdehyde and formaldehyde monitoring. Biomed Chromatogr 16, 199–202 (2002).
Yu, P. H., Lai, C. T. & Zuo, D. M. Formation of formaldehyde from adrenaline in vivo; a potential risk factor for stress-related angiopathy. Neurochem Res 22, 615–620 (1997).
Sun, A. Y., Wang, Q., Simonyi, A. & Sun, G. Y. Resveratrol as a therapeutic agent for neurodegenerative diseases. Mol Neurobiol 41, 375–383 (2010).
Karuppagounder, S. S. et al. Dietary supplementation with resveratrol reduces plaque pathology in a transgenic model of Alzheimer's disease. Neurochem Int 54, 111–118 (2009).
Nie, C. L., Wang, X. S., Liu, Y., Perrett, S. & He, R. Q. Amyloid-like aggregates of neuronal tau induced by formaldehyde promote apoptosis of neuronal cells. BMC Neurosci 8, 9 (2007).
Acknowledgements
This research was supported by grants from the 973-Project (2012CB911000; 2010CB912303, 2013CB531000) and QCAS Biotechnology Fund No. GJHZ1131, the Chinese Postdoctoral Fund 20090460047 and 2012T50144, the Natural Scientific Foundation of China NSFC 31171080, 30970695.
Author information
Authors and Affiliations
Contributions
T.Z. conceived and designed the experiments, analyzed the data and prepared the manuscript; T.Z., H.C., L.W., L.H., L.J., Q.M., S.T., W.B., Y.X., L.Y. and W.Y. performed the experiments. C.D. revised the manuscript. H.R. supervised the project. All authors reviewed the manuscript. T.Z., H.C. and L.W. contributed equally to this work.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Tong, Z., Han, C., Luo, W. et al. Aging-associated excess formaldehyde leads to spatial memory deficits. Sci Rep 3, 1807 (2013). https://doi.org/10.1038/srep01807
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01807
This article is cited by
-
Gut microbiome, cognitive function and brain structure: a multi-omics integration analysis
Translational Neurodegeneration (2022)
-
Amino acid dependent formaldehyde metabolism in mammals
Communications Chemistry (2020)
-
Endogenous formaldehyde is a memory-related molecule in mice and humans
Communications Biology (2019)
-
Icariin protects SH-SY5Y cells from formaldehyde-induced injury through suppression of Tau phosphorylation
Chinese Journal of Integrative Medicine (2016)
-
Effect of Support on the Activity of Ag-based Catalysts for Formaldehyde Oxidation
Scientific Reports (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.