Abstract
One of the most disputed issues in primate evolution and thus of our own primate roots, is the phylogenetic position of the Southeast Asian tarsier. While much molecular data indicate a basal place in the primate tree shared with strepsirrhines (prosimian monophyly hypothesis), data also exist supporting either an earlier divergence in primates (tarsier-first hypothesis) or a close relationship with anthropoid primates (Haplorrhini hypothesis). The use of retroposon insertions embedded in the Tarsius genome afforded us the unique opportunity to directly test all three hypotheses via three pairwise genome alignments. From millions of retroposons, we found 104 perfect orthologous insertions in both tarsiers and anthropoids to the exclusion of strepsirrhines, providing conflict-free evidence for the Haplorrhini hypothesis and none supporting either of the other two positions. Thus, tarsiers are clearly the sister group to anthropoids in the clade Haplorrhini.
Similar content being viewed by others
Introduction
Some mammalian species resist our attempts to reconstruct their true ancestry. Conflicting or weak phylogenetic signals may suggest more than one possible solution depending on marker systems, partitions of data, method and model of reconstruction, or the selected set of compared species. The reasons for controversies can be variable and often are due to polymorphisms of characters in populations before their speciation or to hybridization, but more often to lineage-specific accumulation of noise (random mutations) at the organismal and molecular levels. Lineages formally rich in both individuals and species and then reduced to a few remaining representatives are especially exposed to bottleneck effects, apparently diverging them from the general characteristics of related species.
The Southeast Asian tarsier (Tarsiiformes: Primates) is one example of such a phylogenetic history. The once species-rich Eocene tarsiers were broadly distributed over the Holarctic region, but today are restricted, with just one surviving genus and 11 species, to a few refuges in Southeast Asia. During the last decades of phylogenetic reconstructions, tarsiers have “jumped” from one phylogenetic branch to another (Figure 1): once to split off from primates before the Anthropoidea-Strepsirrhini split (tarsier-first hypothesis1,2), then close to strepsirrhines in the Prosimii clade (prosimian monophyly hypothesis3,4), then in the immediate vicinity of monkeys and apes (haplorrhine hypothesis5,6,7), or to land in an unresolvable trichotomy of strepsirrhines, tarsiers and anthropoids8. These different positions on the evolutionary tree span a phylogenetic era of some 20 million years. With more recent large-scale sequencing efforts, it seemed to be over with such “great leaps”. Compelling support fixed tarsiers on a common branch with strepsirrhines in a joined group of prosimians9. While this study, based on more than 10 kb of sequence information, provided one of the most important and influential mammalian phylogenies, subsequent data would show that it failed significantly in this particular part of the primate tree. The reason was rather simple and was later revealed after sequencing of the mitochondrial genome of Tarsius bancanus10 (a species recently renamed Cephalopachus bancanus4); the nucleotide composition of mitochondrial genes changed drastically on the lineage leading to anthropoids from high AT to a significant accumulation of GC nucleotides. Consequently, most mitochondrial phylogenies of primates artificially formed the clade Prosimii, based on their shared high mitochondrial AT nucleotide content, apart from the GC-rich genes of higher primates. The mixed nuclear/mitochondrial sequence data used in the Murphy et al.9 study, yielded the same problematic results because the major nuclear dataset contributed only weak phylogenetic signals to counter the strong mitochondrial noise. At that time most molecular evolutionists confirmed the significant mitochondrial grouping of strepsirrhines and tarsiers (i.e., the prosimian monophyly hypothesis). Molecular geneticists attempted to improve the sequence-based phylogenetic reconstruction methods to compensate for artificial sequence effects. As reviewed in Goodman et al.11, sequence-based evidence for the position of tarsiers was still ambiguous and mitochondrial supermatrix approaches continued to group tarsiers with strepsirrhines3. However, the hitherto largest comparative genomic analysis of primate species, involving 186 representatives and about 8 MB of sequence, found some support for the Haplorrhini clade (haplorrhine hypothesis12). Furthermore, a recent analysis of genomic data provided statistically significant data to reject a prosimian monophyly13. The currently popular coalescence-based genome analysis method also supports the Haplorrhini clade14,15. Zietkiewicz et al.16 compared Alu element sequences derived from strepsirrhines, tarsiers and human genomes and found that tarsier elements cluster with human Alu subfamilies and show RNA secondary structure elements absent in strepsirrhines; however, this sequence similarity analysis of Alu elements is not quite as reliable as the highly informative content of retroposon presence/absence analyses.
Three competing hypotheses of tarsier phylogeny.
Hypothesis #1, tarsiers as the first divergence of the primate tree, with Strepsirrhini and Anthropoidea on the same branch, hypothesis #2, tarsiers close to strepsirrhines, together forming the Prosimii clade and hypothesis #3, tarsiers closely related to Anthropoidea in the Haplorhini clade. The images are provided by Jón Baldur Hlíðberg.
Screening for phylogenetic diagnostic insertions of shared orthologous retroposons enabled an escape from this conflicting phylogenetic trap. Four Alu short interspersed elements (SINEs) supported a close relationship between tarsiers and anthropoids (haplorrhine hypothesis7,17). Independent of nucleotide composition effects, evolutionary rate variation, saturation of substitutions, etc., the comparison of orthologous insertions of retroposed elements provides a noise-free evaluation of phylogenetic affiliations. An exactly shared retroposon insertion is characterized by specific target site duplications (TSD) of 5–30 nucleotides flanking the insertion site produced during the insertion process18. Similar shared TSDs together with an identical retroposon with the identical length and identical orientation in two species strongly indicates an insertion event in their common ancestor and thus their close relatedness; conversely, an empty insertion site in one or the other indicates more distantly related species.
The quantitative genome-wide distribution of certain primate-specific Alu SINE elements also supported the haplorrhini hypothesis. Churakov et al.19 found hundred thousand AluJb elements well distributed in the genomes of tarsiers and anthropoids but absent in strepsirrhines.
However, our initial search for diagnostic retroposon insertions7 was biased by screening only from the human genome, the only genomic source available at the time, thus making it impossible to test all three hypotheses equally. Therefore, combined with the inherent problems associated with even the most modern sequence-based reconstructions, the placement of tarsiers was still uncertain.
Alu SINEs, long interspersed elements (LINEs) and long terminal repeats (LTRs) are exceptionally suitable as phylogenetic markers in primates because of the high number of insertions (e.g., 1.5 million Alu insertions in human20) and their broad activity covering the full diversity of primates. This makes them unrivaled as phylogenetic markers compared to other rare genomic changes in primates.
The present genome-wide approach aimed to screen for diagnostic retroposons starting from different sources of genome information, including genomes of mouse lemurs, bushbabies, tarsiers and human, to present the first genome-wide, multi-genome screening to find perfect phylogenetically informative Alu SINEs, LINEs and LTRs among all these lineages and thereby definitively test all three possible hypotheses of tarsier origin using a virtually homoplasy-free approach.
Results
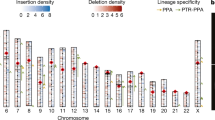
In an initial run of our PERL pipeline for the three search strategies described above, we identified genome-wide a total of 260 preliminary informative Alu and Alu-like loci, 200 preliminary LINE loci and 120 preliminary LTR loci. All of these loci were then subjected to careful manual inspections of the insertion sites, looking for only those with exact orthologous insertions according to the criteria explained below. Only 19 Alu-like, 22 LINE and 63 LTR loci with retroposed elements present in both tarsier + human fulfilled these criteria (Supplementary Information; Figure 2); none were found with shared elements in either strepsirrhines + human or tarsier + strepsirrhines. The diagnostic elements active at the time of the common ancestor of tarsiers and anthropoids were fossil Alu monomers (2 FAMs), free left Alu monomers (12 FLAMs), Alu dimers (2 AluJb, 3 AluSx), LINEs (22 L1PB4) and LTRs (62 MSTB; 1 MSTC) (Supplementary Information). According to the Waddell statistics21, with 104 conflict-free phylogenetic informative markers for haplorrhine the probability of supporting the wrong tree topology is less than 7.2 × 10−50. Thus, tarsiers are clearly the closest relatives of higher primates.
When using retroposons as phylogenetic markers, it is important to note the absolute necessity of applying a strict definition of orthology, including a careful inspection of the lengths and locations of the TSDs and a reliable definition of the type of insertion. This requires improved manual alignments and knowledge of the genomic changes introduced during the retroposon insertion process. Interestingly, none of the previously identified retroposons supporting the haplorrhine hypothesis7,17 were among the 104 new elements, as the present genome-wide search was focused on perfect insertions only. Diagnostic Alu, LINE and LTR elements were selected only if flanked by nearly perfect TSDs (maximum 30% divergence for Alu and LINE, 40% for LTR, from the solitary target site of reference species with absent state) and clear absence in both strepsirrhines and outgroup species. Furthermore, only orthologous elements with more than 70% sequence similarity were considered. Two of the previously identified haplorrhine markers were located in sequence regions not represented in the current low-coverage 2X-based assemblies of human/tarsier and human/strepsirrhines. The third marker was located in a region subsequently identified to be a misaligned part of the genome alignment and one marker exhibited more than 30% divergence between human and tarsier.
Discussion
In our current dataset, we did not find any conflicting retroposon presence/absence patterns, indicating that at the time of the divergence of haplorrhines most or all retroposons were fixed and are thus well suited for phylogenetic reconstructions and clearly do not support a trifurcation as some previous data might have suggested8. Such a clear and conflict-free pattern is not always evident from retroposon data. Ancestral polymorphisms of retroposons and subsequent speciation before fixation may lead to conflicting signals of relationships, as was previously demonstrated for the three main lineages of placental mammals (Afrotheria, Xenarthra, Boreotheria)22,23. Although such examples of conflicting retroposon patterns are rare, they are invaluable for diagnosing zones of incomplete lineage sorting or ancestral hybridization effects that influence the evolutionary patterns of species.
In contrast to the highly significant support afforded by the 104 new genome-wide-extracted, conflict-free haplorrhine retroposon markers, the ~50 million years of nuclear sequence evolution and possibly the extremely reduced number of tarsier individuals led to the fixation of much random noise, causing the previous problems to determine the correct tree based on these data.
Our analysis now provides overwhelming, conflict-free support at the genome-wide level for the haplorrhine hypothesis and a clear rejection of both the prosimian monophyly and tarsier-first hypotheses, as suggested by previous data7,14,15,16,17,18,19. The previous retroposon insertion data supporting the Haplorrhini hypothesis7 was likely biased by screening only human genome data and a clear rejection of the prosimian monophyly and tarsiers-first hypotheses were lacking; both of these limitations have now been overcome. Many molecular sequence studies demonstrated support for the prosimian monophyly based on data that appeared to be significant3,9,24,25 but were likely artificial signals connected to the dominance of mitochondrial data10. The strong but artificial mitochondrial noise and missing nuclear signals challenged the use of sequence data in certain mammalian branches. A similar example of an extreme misinterpretation based on mitochondrial DNA was the artificial placement of the order Dermoptera amidst the primate phylogenetic tree9,24,26, suggesting that primates were paraphyletic.
Ten years later with the use of genome-scale data, special reconstruction methods involving coalescence models have also correctly placed tarsiers at their natural phylogenetic branch close to Anthropoidea14,15. Also, increasingly dense fossil records bolster our current picture of the Paleogene fauna27 and will be strengthened by the highly significant evidence in favor of the haplorrhine hypothesis. This will, in turn, provide a strong base for addressing the anthropoid origin, with tarsiers as the “link” between basal primates and humans.
Methods
To test all three possible hypotheses, we used tarsier (Tarsius syrichta recently renamed Carlito syrichta4), two representatives of the Strepsirrhini lineage, mouse lemur (Microcebus murinus) and bushbaby (Otolemur garnetti) and human as representative of anthropoids.
To perform a multi-genome screening, we retrieved the following genomes, all covered by 2× genome sequences:
Tarsius syrichta (tarSyr1)28;
Microcebus murinus (micMur1)29;
Otolemur garnetti (otoGar3)30.
To find diagnostic insertions we used a combination of pairwise (2-way) alignments from the UCSC Genome Bioinformatics center in Santa Cruz:
Human/tarsier (tarSyr1)31;
Human/mouse lemur (micMur1)32;
Human/bushbaby (otoGar1)33.
Data mining
Our search strategies were designed to accomplish two concrete objectives: 1) a genome-wide screening and detection of perfect, phylogenetically informative retroposons for the phylogenetic position of tarsiers and 2) multi-genome screenings that consider all three possible phylogenetic tree topologies equally: (1) tarsiers as the first split of primates (strepsirrhines + human), (2) tarsiers on a shared branch with strepsirrhines (tarsiers + strepsirrhines), (3) tarsiers together with anthropoids (tarsiers + human).
Screening for potential markers: tarsier + human, strepsirrhines + human
To test the haplorrhine hypothesis, we developed bioinformatics tools (a) to screen available pairwise alignments of human/mouse lemur and human/bushbaby for Alu SINE elements, LINEs (L1Ps) and LTRs (MSTs), present in human but absent in both strepsirrhine species (Figure 3a). Such pairwise or 2-way alignments were organized in ‘axt alignment format’ (compiling alignment blocks with genomic coordinates for both the primary organism (first sequence) and aligning organism (second sequence))34. Alignment gaps (e.g., caused by an insertion of an Alu SINE, LINE, or LTR element in the genome of one species) were recognized by interrupted coordinates (Figure 3a; e.g., human coordinates chromosome 2_227644827-227645114 (287 nt insert) vs. bushbaby coordinates of GL873532 (reverse complement) 6388296, 6388295 (no insert). (b) We compared the coordinates of the extracted locus with the coordinates of the previously RepeatMasked human genome to detect loci with specific retroposon insertions absent in strepsirrhines (Figure 3b). (c) We inspected the pairwise alignment of human/tarsier at the selected human coordinates (Figure 3c; e.g., human coordinates on chromosome 2_227644728-227645219). (d) We used the human coordinates to extract orthologous MULTIZ sequences from the UCSC Genome server35 (Figure 3d). We extracted the resulting concatenated sequence blocks in FASTA format using a new java application (Figure 3e; UCSC2FASTA; available upon request). We manually realigned and analyzed the sequences, especially the insertion site of the diagnostic retroposed elements that were shared between human and tarsier but absent in both strepsirrhines.
Screening strategy to find genome-wide, phylogenetically informative, Alu SINE insertion markers using genome sequences and pairwise genome alignments.
(A) Pairwise genome alignment of human and bushbaby. Locus with a gap of 287 nt (insertion in human). Coordinates are abbreviated: human chromosome 2, 4728 = 227644728, 4827 = 227644827, 5114 = 227645114, 5219 = 227645219; bushbaby scaffold identifier GL873532 in reverse complement (rc) orientation, 8385 = 6388385, 8296 = 6388296, 8295 = 6388295, 8220 = 6388220. (B) The insertion is represented by the integration of an Alu SINE element. (C) Corresponding region of the human alignment with tarsier. The Alu element is present in human and tarsier. (D) MULTIZ alignment from the UCSC server. The Alu element is present in human and in tarsiers but not in strepsirrhines. (e) The MULTIZ alignment is transferred to a Fasta alignment using the UCSC2FASTA java application. The Fasta file is the basis for manual crosscheck. In analyzing gap positions, a variation of +/−30 nts is allowed. We used the UCSC axt alignment format. TSD = target site duplications. The retroposon sequence is only partially shown interrupted by a double-slash.
To test the hypothesis that tarsiers diverged early from primates (tarsier-first hypothesis), we used a similar approach to search for potential markers merging human and strepsirrhines starting from a 2-way alignment of human/tarsier and searching for retroposons specific for human. Such loci were projected/compared to a second and third 2-way alignment of human/bushbaby and human/mouse lemur. Loci with elements present in human, present in strepsirrhine, but absent in tarsier were extracted for closer inspection. Again, the complete loci were extracted via the UCSC MULTIZ alignment by querying the human coordinates and settings.
Screening for potential markers: tarsier + strepsirrhines
To test the third potential hypothesis (prosimian monophyly hypothesis), with tarsiers and strepsirrhines on one branch, we applied a slightly different strategy because tarsier/strepsirrhine pairwise alignments are not yet available. For this search we (a) started with a 2-way alignment of human/bushbaby and human/mouse lemur and derived all loci with a gap of 100-400 nt in human and an orthologous Alu SINE, LINE, or LTR insertion in strepsirrhines. (b) The pairwise alignment of human/tarsier was correspondingly screened for Alu SINE, LINE and LTR insertions in tarsier. (c) We analyzed the two sets of selected alignments for overlapping coordinates in human. (d) We extracted and merged all corresponding sequences. (e) Loci with elements present in both tarsier and strepsirrhines but absent in human and an insertion sequence similarity of more than 70% were manually inspected. With this screening we examined the potential shared evolution of tarsiers and strepsirrhines.
Derived computer pipelines for automated screening
To automatize the search for phylogenetically informative markers, we developed a PERL pipeline connected to a MySQL database (available upon request). The search criteria were (1) expected minimum size of element = 50 nt, (2) expected maximum size of element = 6,000 nt, (3) selected ‘search for retroposons', (4) pairwise distance cut off = 30% (more than 70% similarity) (5) gap expansion +/- 30 nt, (6) extracted length of flanks = 100 nt. (7) Only loci in which sequences of both mouse lemur and bushbaby were available, well alignable, free from large deletions around the insertion site and both with the same presence/absence state of the inserted element.
Manual alignment editing
We selected only cases with nearly perfect TSDs for Alu-like elements and LINEs (maximal 30% divergence from the solitary target site of reference species with absent state, considering the classical TSD length of 8–30 nt) and if available, identical truncation points in the shared orthologous markers. Because LTR elements have only very short TSDs of ~5 nt, we selected cases with maximally 40% divergence between TSDs and the solitary target sites. All Alignments are available as Supplementary Information.
Change history
28 July 2015
A correction has been published and is appended to both the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Andrews, T. D., Jermiin, L. S. & Easteal, S. Accelerated evolution of cytochrome b in simian primates: adaptive evolution in concert with other mitochondrial proteins? J Mol Evol 47, 249–257 (1998).
Gingerich, P. D. In: Recent Advances in Primatology Vol. 3 (eds D. J. Chivers & K. A. Joysey) 249–255 (Academic Press, 1978).
Chatterjee, H. J., Ho, S. Y., Barnes, I. & Groves, C. Estimating the phylogeny and divergence times of primates using a supermatrix approach. BMC Evol Biol 9, 259 (2009).
Groves, C. & Shekelle, M. The genera and species of Tarsiidae. Int J Prim 31, 1071–1082 (2010).
Goodman, M. et al. Toward a phylogenetic classification of Primates based on DNA evidence complemented by fossil evidence. Mol Phylogenet Evol 9, 585–598 (1998).
Rosenberger, A. L. & Szalay, F. S. In: Evolutionary biology of the New World monkeys and continental drift (eds R. L. Ciochon & A. B. Chiarelli) 139–157 (Plenum Press, 1980).
Schmitz, J., Ohme, M. & Zischler, H. SINE insertions in cladistic analyses and the phylogenetic affiliations of Tarsius bancanus to other primates. Genetics 157, 777–784 (2001).
Arnason, U. et al. Mitogenomic relationships of placental mammals and molecular estimates of their divergences. Gene 421, 37–51 (2008).
Murphy, W. J. et al. Molecular phylogenetics and the origins of placental mammals. Nature 409, 614–618 (2001).
Schmitz, J., Ohme, M. & Zischler, H. The complete mitochondrial sequence of Tarsius bancanus: evidence for an extensive nucleotide compositional plasticity of primate mitochondrial DNA. Mol Biol Evol 19, 544–553 (2002).
Goodman, M., Grossman, L. I. & Wildman, D. E. Moving primate genomics beyond the chimpanzee genome. Trends Genet 21, 511–517 (2005).
Perelman, P. et al. A molecular phylogeny of living primates. PLoS Genet 7, e1001342 (2011).
Jameson, N. M. et al. Genomic data reject the hypothesis of a prosimian primate clade. J Hum Evol 61, 295–305 (2011).
McCormack, J. E. et al. Ultraconserved elements are novel phylogenomic markers that resolve placental mammal phylogeny when combined with species-tree analysis. Genome Res 22, 746–754 (2012).
Song, S., Liu, L., Edwards, S. V. & Wu, S. Resolving conflict in eutherian mammal phylogeny using phylogenomics and the multispecies coalescent model. Proc Natl Acad Sci USA 109, 14942–14947 (2012).
Zietkiewicz, E., Richer, C. & Labuda, D. Phylogenetic affinities of tarsier in the context of primate Alu repeats. Mol Phylogenet Evol 11, 77–83 (1999).
Schmitz, J., Roos, C. & Zischler, H. Primate phylogeny: molecular evidence from retroposons. Cytogen Gen Res 108, 26–37 (2005).
Kapitonov, V. V., Pavlicek, A. & Jurka, J. In: Encyclopedia of Molecular Cell Biology and Molecular Medicine (ed R. A. Meyers) 251–305 (Wiley-VCH, 2006).
Churakov, G. et al. A novel web-based TinT application and the chronology of the Primate Alu retroposon activity. BMC Evol Biol 10, 376 (2010).
Mandal, P. K. & Kazazian, H. H., Jr SnapShot: Vertebrate transposons. Cell 135, 192–192 (2008).
Waddell, P. J., Kishino, H. & Ota, R. A phylogenetic foundation for comparative mammalian genomics. Genome Inform 12, 141–154 (2001).
Churakov, G. et al. Mosaic retroposon insertion patterns in placental mammals. Genome Res 19, 868–875 (2009).
Nishihara, H., Maruyama, S. & Okada, N. Retroposon analysis and recent geological data suggest near-simultaneous divergence of the three superorders of mammals. Proc Natl Acad Sci U S A 106, 5235–5240 (2009).
Arnason, U. et al. Mammalian mitogenomic relationships and the root of the eutherian tree. Proc Natl Acad Sci USA 99, 8151–8156 (2002).
Md-Zain, B. M., Lee, S. J., Lakim, M., Ampeng, A. & Mahani, M. C. Phylogenetic position of Tarsius bancanus based on partial cytochrome b DNA sequences. J Biol Sci 10, 348–354 (2010).
Schmitz, J., Ohme, M., Suryobroto, B. & Zischler, H. The colugo (Cynocephalus variegatus, Dermoptera): the primates' gliding sister? Mol Biol Evol 19, 2308–2312 (2002).
Williams, B. A., Kay, R. F. & Kirk, E. C. New perspectives on anthropoid origins. Proc Natl Acad Sci USA 107, 4797–4804 (2010).
UCSC Genome Bioinformatics. Available at: ftp://hgdownloadcse.ucsc.edu/gbdb/tarSyr1/. Accessed on February 2013.
UCSC Genome Bioinformatics. Available at: ftp://hgdownload.cse.ucsc.edu/gbdb/micMur1/. Accessed on February 2013.
UCSC Genome Bioinformatics. Available at: http://hgdownload.cse.ucsc.edu/goldenPath/otoGar1/bigZips/. Accessed on February 2013.
UCSC Genome Bioinformatics. Available at: http://hgdownload.cse.ucsc.edu/goldenPath/hg19/vsTarSyr1/. Accessed on February 2013.
UCSC Genome Bioinformatics. Available at: http://hgdownload.cse.ucsc.edu/goldenPath/hg19/vsMicMur1/. Accessed on February 2013.
UCSC Genome Bioinformatics. Available at: http://hgdownload.cse.ucsc.edu/goldenPath/hg19/vsOtoGar1/. Accessed on February 2013.
UCSC Genome Bioinformatics. Available at: http://genome.ucsc.edu/goldenPath/help/axt.html/. Accessed on February 2013.
UCSC Genome Bioinformatics. Available at: http://genome.ucsc.edu/cgi-bin/hgGateway/. Accessed on February 2013.
Acknowledgements
We thank Norbert Grundman for assistance in programming, Marsha Bundman for editing the manuscript and Jón Baldur Hlíðberg for providing the paintings. We very much thank Joachim Kurtz and the Münster Graduate School of Evolution initiative (MGSEi) for providing initial financial support for the PhD position of GH. We thank Christian Roos and Mike Montague for valuable comments on an early version of the manuscript. This paper is dedicated to the memory of Johanna Solbach, who co-organized the MGSE Symposium 2012, where these results were first presented. This work was supported by the Deutsche Forschungsgemeinschaft (SCHM1469/3-2; SCHM1469/4-1).
Author information
Authors and Affiliations
Contributions
J.S., G.C. and G.H. designed the study, developed the screening strategy and analyzed the data. G.H. performed the bioinformatic screenings of Alus in WMs group, G.C. screened for informative LINEs and LTRs. W.W. contributed special access to genome data. W.M., J.B. and W.W. discussed the results. J.S. wrote the manuscript and all authors contributed their comments.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Table 1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Hartig, G., Churakov, G., Warren, W. et al. Retrophylogenomics Place Tarsiers on the Evolutionary Branch of Anthropoids. Sci Rep 3, 1756 (2013). https://doi.org/10.1038/srep01756
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01756
This article is cited by
-
Analysis of lineage-specific Alu subfamilies in the genome of the olive baboon, Papio anubis
Mobile DNA (2018)
-
Recently integrated Alu insertions in the squirrel monkey (Saimiri) lineage and application for population analyses
Mobile DNA (2018)
-
The phylogenetic system of primates—character evolution in the light of a consolidated tree
Organisms Diversity & Evolution (2016)
-
Vγ9 and Vδ2 T cell antigen receptor genes and butyrophilin 3 (BTN3) emerged with placental mammals and are concomitantly preserved in selected species like alpaca (Vicugna pacos)
Immunogenetics (2014)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.