Abstract
The difficulty in annotating the vast amounts of biological information poses one of the greatest current challenges in biological research. The number of genomic, proteomic and metabolomic datasets has increased dramatically over the last two decades, far outstripping the pace of curation efforts. Here, we tackle the challenge of curating metabolic network reconstructions. We predict organismal metabolic networks using sequence homology and a global metabolic network constructed from all available organismal networks. While sequence homology has been a standard to annotate metabolic networks it has been faulted for its lack of predictive power. We show, however, that when homology is used with a global metabolic network one is able to predict organismal metabolic networks that have enhanced network connectivity. Additionally, we compare the annotation behavior of current database curation efforts with our predictions and find that curation efforts are biased towards adding (rather than removing) reactions to organismal networks.
Similar content being viewed by others
Introduction
Advances in high-throughput experimental biology have greatly furthered our knowledge and made it possible to interrogate cellular processes in a systematic manner. However, this data deluge is only as useful as our ability to interpret it1. The current uncertainty of data reliability hampers our efforts to understand which network topologies fulfill physical, chemical and biological constraints2,3,4,5,6. Understanding the possible network topologies is a critical condition in on-going attempts to understand the function and evolution of cellular networks.
Improving annotations of organismal metabolic networks has been an area of intense interest, especially owing to their usefulness in assessing organismal fitness in silico. There have been numerous methods proposed that attempt to solve the incompleteness of metabolic network reconstructions, ranging from gap-filling the organismal network based on what other known organismal networks possess7,8,9 to methods that rely on multiple sources of annotation information to provide an assessment of enzyme presence10.
However, even with these methods we are still far from achieving consensus on the correct metabolic network for a given organism, even one as well-studied as Escherichia coli (Fig. 1a). Indeed, there are dramatic differences in both the size and degree of overlap of the metabolic network for E. coli recorded in (i) different databases or (ii) the same database at different time snapshots (Fig. 1b). This problem is magnified in organisms that have their genomes sequenced for the first time (Fig. 1c). This last example is a perfect demonstration of both our lack of knowledge and the problem of developing computational analyses that perfectly recapitulate the known network at only the present instance.
Metabolic networks are still being actively revised.
We focus here on the metabolites and reactions belonging to the core metabolism and show network size and changes in metabolic networks for selected organisms. (a), Size of the E. coli metabolic network for four databases relative to KEGG 2009. Note the dramatic differences in both the size of the networks and the percentage of overlap with the KEGG 2009 network. (b), Number of reactions KEGG added or removed for selected organismal networks over the course of two years (2009 to 2011). Even though the networks are restricted to the intersection with the core metabolism there is appreciable turnover. (c), Number of reactions KEGG added or removed for selected organismal networks over the course of the same period for two recently sequenced organisms. To illustrate the effect of these revisions, if a model was to perfectly predict the 2009 organismal network for Taeniopygia guttata it would be less than 50% accurate when it was compared to the 2011 network.
Because data reliability is such a pressing problem for experimental and computational researchers alike, there has been a push in research to consider the analysis of metabolic networks from novel perspectives. A promising new framework is to consider metabolism in the context of a global network. This framework has been successfully applied in assessing the emergence of biological carbon fixation in phylometabolism11 and, more generally, to understand the regulation of metabolism12. A global network has also been recently used in conjunction with probabilistic methods to predict metabolic networks on a small scale with experimental verification13. While the motivation for the global network approach has been mostly pragmatic, it is reminiscent of the “Res Potentia” framework proposed by Whitehead14. Wherein he proposes that which does exist—termed the Res Extenta or in the case of metabolism the set of organismal metabolic network—are specific realizations of a “universal” framework—the Res Potentia or the global network in our analysis—that defines what is possible.
We contend that using a global network approach to the study of metabolism is comparable to what epidemiologists do when studying worldwide propagation of infection. In building the worldwide air transportation network15,16,17 all carrier flights are aggregated into a single network and analyzed. This is an important feature of its construction because it aids in the identification of and distinction between international and regional hubs, establishing their relative importance in the network. As an example, if we were to consider US Airways (a North American carrier) alone we would not have fully grasped the importance of London, since this carrier will have more flights to Los Angeles or even San Diego than to London. Even if we were to pick a group of carriers based on similarity (such as operating primarily in North America) and assess the ensemble of their individual networks, it would be difficult to assess the relative importance of the individual hub airports. Reframing the analysis of metabolic networks to a global network is an appropriate method to both assess our current network annotations and to gain an understanding of what evolutionary dynamics shape organismal metabolism into the structures that we currently know.
In the following we predict entire organismal metabolic networks using only sequence homology and the global network as a reference. We show that using an appropriate reference set, the global network, allows for more insight to be obtained from sequence homology. We compare our predicted networks to known metabolic database reconstructions and also evaluate the connectedness of the resultant graphs for predicted networks to assess the performance of our methodology. We also use our predictions to help understand how curation behavior in a metabolic database affects known organismal metabolic networks.
Results
Organismal network prediction
We constructed a global network by performing the graph union of all organismal networks (Fig. 2b). Our analyses focused on the giant component of the global network because it contains the most reliable data, as its metabolites are more conserved and have more pathway annotations. To predict individual organismal metabolic networks we assumed that a given reaction can be catalyzed within an organism if and only if, the organism synthesizes a protein that is sufficiently similar to the known enzymes for the reaction. We evaluated each reaction in the global network for its possibility of existence in any individual organism.
Construction of a global metabolic network from KEGG data.
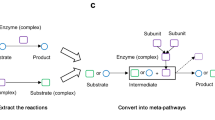
(a), Construction of the Methanococcus maripaludis metabolic network from KEGG. We show the giant component in black and the 122 small components in grey. We highlight eight metabolites (shown in green, highlighted with red circles) that will be considered in the other panel. (b), The set of all organismal networks is the extent of our current knowledge from KEGG. We construct the global metabolic network as the union of all known networks. We superimpose the eight highlighted metabolites from M. maripaludis (shown in green) on the global network making their network proximity apparent. This superimposition elucidates potential additional reactions that can bridge gaps between different components of the network. Such superimpositions can dramatically decrease the search space for new reactions that can accomplish this bridge.
We aligned the enzyme sequences associated with each reaction to each organism's protein database (Fig. 3a) and determined the expectation value (E-value) of the alignment using blastp18,19. The E-value is a measure of the number of times the match between the sequences would be expected to occur by chance; E-value = 0.0 indicates a perfect match between the queried enzyme sequence and a protein in the database, while E-value > 1.0 is interpreted as a sequence match that is not indicative of biological homology.
Organismal network prediction using the global metabolic network.
(a), We BLAST the enzyme sequences associated with Reaction 1 from the global metabolism in all organisms against the known protein sequences from open reading frames in Organism 1 in order to determine whether Organism 1 possesses an enzyme that can catalyze Reaction 1. We repeat this same procedure for Reaction 2 from the global metabolism. (b), We categorize each alignment as a hit or poor match based on the magnitude of its E-value. If an alignment has an E-value ≤ 0.01 then we classify the alignment as a hit; otherwise, if 0.01 < E-value ≤ 10, we classify it as a poor match. It is visually apparent in this example that the annotated reaction has a much larger fraction of hits. (c), The distributions of the fraction of hits for annotated and unannotated reactions show a clear difference no matter which domain of organisms is considered.
For clarity, we define several additional terms. A reaction predicted to be catalyzed in a certain organism by a certain database curation team is “annotated” in that database. Otherwise, the reaction is “unannotated” — it exists in the global network but not in the organismal one. To make our predictions of annotation status we separated the alignments associated with a reaction r into two categories, hits and poor matches, based on the magnitude of the E-values obtained. If an alignment has E-value ≤ 0.01 then we classify the alignment as a hit; otherwise, if 0.01 < E-value ≤ 10, we classify it as a poor match (Fig. 3b).
We use the fraction of alignments  that are classified as hits as a predictor of whether reaction r can be catalyzed within organism i. When the distribution of
that are classified as hits as a predictor of whether reaction r can be catalyzed within organism i. When the distribution of  is examined we find a peak at greater values for KEGG annotated reactions when all of the reactions are considered. Furthermore, we see that this behavior holds no matter which domain of organisms is considered (Fig. 3c), indicating that this is a robust behavior preserved across all organismal networks. The imperfect separation between annotated and unannotated distributions is also expected given the amount of known annotation errors in the KEGG networks (Fig. 1).
is examined we find a peak at greater values for KEGG annotated reactions when all of the reactions are considered. Furthermore, we see that this behavior holds no matter which domain of organisms is considered (Fig. 3c), indicating that this is a robust behavior preserved across all organismal networks. The imperfect separation between annotated and unannotated distributions is also expected given the amount of known annotation errors in the KEGG networks (Fig. 1).
To determine whether a given  is large enough to be considered a reaction that can be catalyzed we must set a threshold value f×; if
is large enough to be considered a reaction that can be catalyzed we must set a threshold value f×; if  , then we predict reaction r to exist in organism i. To identify an appropriate value for f×, we calculated the receiver operator characteristic (ROC) curve, accuracy and false discovery rate statistics20. The ROC curve analysis demonstrates that
, then we predict reaction r to exist in organism i. To identify an appropriate value for f×, we calculated the receiver operator characteristic (ROC) curve, accuracy and false discovery rate statistics20. The ROC curve analysis demonstrates that  can discriminate between annotated and unannotated reactions (Fig. 4a). We thus use the accuracy and false discovery rates to determine a good threshold value for
can discriminate between annotated and unannotated reactions (Fig. 4a). We thus use the accuracy and false discovery rates to determine a good threshold value for  and set f× = 0.14 (Figs. 4b). In summary, we predict organismal metabolic networks by checking whether a reaction r for organism i has a value
and set f× = 0.14 (Figs. 4b). In summary, we predict organismal metabolic networks by checking whether a reaction r for organism i has a value  (Fig. 4c). This approach allows us to predict entire organismal networks using only the global metabolic network and the associated organismal BLAST alignments.
(Fig. 4c). This approach allows us to predict entire organismal networks using only the global metabolic network and the associated organismal BLAST alignments.
Proposing a good threshold to predict reactions that will be catalyzed.
We use multiple statistics for judging the optimal threshold to differentiate between annotated and unannotated groups. The receiver operator characteristic curve demonstrates that all of the thresholds possess discriminatory power in comparison to random chance (a). To choose a precise threshold we combine this with the false discovery rate and accuracy statistics (b). Balancing the two statistics results in a threshold of 0.14 as the optimal point to separate the two groups. This value corresponds to the peak seen in the annotated distribution for all organisms (c). We use this threshold to predict whether an organism can catalyze a given reaction between two metabolites when we predict organismal metabolic networks de novo from the global metabolic network.
Comparison with consensus networks
In order to calculate the accuracy of our predictions it is necessary to compare it to a “ground truth”. However, given the significant variations in size and content that exist across different databases and in time, such a ‘true’ answer does not exist (Fig. 1). In an effort to estimate the true accuracy of our predictions we consider the metabolic network reconstructions of E. coli, a well-studied organism, from three different sources, with two of the sources having network data at two separate time points.
We constructed ten separate consensus networks as detailed in Methods leaving out three reconstructions at a time and evaluated the accuracy of the networks that were left out of the consensus, with the results shown in Table 1. While no network is 100% accurate with respect to the consensus network, we find that our predictions range in accuracy between 70 and 71% while the database networks range in accuracy between 66 and 93%.
In an effort to understand what types of reactions our method incorrectly predicts we characterize the reactions that are identified as false positives in comparison to the consensus network. First, we examine the pathway annotations associated with the metabolites in this set of false positive edges and reactions (Fig. 5a). We find that the majority of the metabolites are either unclassified or classified in pathways that are not central to metabolism (outside of the carbohydrate, amino acid, nucleotide and lipid metabolism pathways). However, even being associated with a central pathway does not mean that all of the metabolites are specifically involved in central or essential processes and these characterizations could be due only to their presence as a byproduct in a reaction.
Examination of pathway annotations and conservation for false positive reactions identified in E. coli.
We take the set of false positives from our predicted network (identified by comparison to known network reconstructions) and find that when we examine the pathway annotations the largest single group is “Unclassified” (a). Furthermore, when the reactions are binned by their conservation values we find that the bulk of the false positive assignments belong to the lowest third of conservation values (b). These two points suggest that the false positive set from our predictions is composed primarily of reactions that are relatively uncharacterized and poorly studied. This could mean that the actual amount of “true” false positives is smaller than currently suggested by comparison to other network reconstructions and has a substantial composition of present, but currently unknown reactions.
Second, we examined the conservation of the false positive reactions (Fig. 5b). Conservation is calculated as the fraction of times that the edge appears in an organismal network in comparison to the total number of organismal networks. We calculate the conservation for all edges in all organismal networks in KEGG and define three bins in the distribution (lower, middle and upper thirds of the distribution). When we bin the false positive edges into these bins we find that the overwhelming majority are in the lowest third of conservation values. If we consider the lower and middle thirds together these groups accounts for more than 90% of all edges in the false positive set.
The abundance of low conservation reactions in the “false positive” set of our method could plausibly be interpreted as suggesting that these reactions may not actually be false positives. It is likely that a majority of the edges in this set do actually exist, they just have not been incorporated into a majority of the databases due to poor characterization and understanding of the reactions themselves.
Network connectivity
Given the challenge presented by traditional validation due to the lack of a ground truth and our aim to predict an organism's true metabolic network instead of simply recapitulating the annotations in KEGG, we also use a validation scheme focused on the expected properties of metabolic networks. Specifically, we surmise that organismal metabolic networks must have a bias toward connectedness. Indeed FBA metabolic reconstructions assume that metabolic networks act as “transportation” networks that carry mass from external nutrients to biomass7,8. The possession of fewer network components implies a greater ability of the organism to exploit a broad range of incoming nutrients for disparate cellular roles and thus offers a fitness advantage over topologies where each network component must be individually fed.
We find that both our predictions and the changes made in KEGG in the period 2009-2011 close more gaps between network components than would be expected if new reactions were added at random (Fig. 6a). When we consider gaps of size one, our predictions fill almost twice as many gaps as the KEGG changes. Remarkably, we also find that our predictions introduce fewer new network components than random removals. The changes in KEGG actually cause the creation of more additional network components than would be expected if reactions were randomly removed from organisms (Fig. 6b). The fact that so many gaps are closed by both our method and by KEGG curation in the period 2009-2011 lends credence to our original hypothesis that metabolic networks should be evolutionarily biased towards minimizing the number of network components and supports the validity of our methodology.
Predicted networks are more connected than ones constructed strictly from the database.
(a), Fraction of gaps between two network components that are closed when reactions are added to a metabolic network versus gap size. Both the predicted network additions and the KEGG 2011 additions close more gaps in the KEGG 2009 network than would be expected by random chance. However, there are significantly more additions using the predicted networks when a gap size of 1 is considered. (b), Relative fraction of removed reactions that introduce an additional network component in comparison to random removals for predicted deletions and KEGG 2011 deletions. Reactions “removed” by the predictions introduce additional network components less frequently than random removals would. The opposite is true for the reactions removed in the KEGG 2011 network where the removals actually introduce additional components more frequently than random removals. The predictions significantly differ from the KEGG curation efforts in this behavior (predicted deletions zscore = −0.97, KEGG 2011 deletions zscore = 1.23). (c), When all organisms are considered there is a distinct bias for more reactions to be added to organisms in comparison to the number of reactions predicted to be added. When we consider only the set of 10 well studied organisms considered in Fig. 1 there is largely agreement between our predictions and the actual changes that occur in KEGG for both additions and removals.
It is important to note that our method takes in no information from the global network concerning reactions other than the possibility of their existence. Therefore, our method is no more biased towards closing gaps or preserving network structure than the actual changes to the database could or should be and yet it still accomplishes this goal of increased connectivity.
Biases in database curation
When we examine how our predictions compare to the corrections made to the KEGG database over time we find that there is a distinct bias towards adding instead of removing reactions to organismal networks (Fig. 6c). It would be simple to assume that our method under-predicts in comparison to the reference dataset; however, we do not observe this trend when we examine the set of well-studied organisms used in Fig. 1b and c. This suggests that the curation teams are more aggressive in adding reactions than removing them, despite the fact that both errors of omission and addition are equally detrimental. Large-scale comparison and tracking of database changes could influence curation teams' actions and help attenuate this problem.
Discussion
There are several distinct advantages to reframing the study of metabolic networks and, more broadly, metabolism to the organismal usages of the global network. As demonstrated in this study we are able to extract substantially more predictive power from sequence homology when it is used in conjunction with the global network. While most studies have moved beyond homology due to a lack of predictive power to more complicated and time consuming methods (such as Bayesian or multiple information methods), we are able to predict metabolic networks that compare favorably to the known database data and exceed them in producing connected networks. We could also easily increase the efficacy of our method by including additional network information such as whether a reaction completes a gap or not, which would be trivial to calculate and consider.
The global network also enables community detection and other graphical analyses that are unchanging in the face of organismal usage, facilitating an understanding of the true importance of a metabolite. Comparing the differences in organismal usage of metabolites and reactions can then be used to more robustly characterize the evolutionary forces that have optimized an organismal network. Specifically, when studying an organismal network we cannot fully comprehend the importance of a given metabolite because we do not have access to all the manners in which that metabolite could potentially connect to other metabolites in the network. Thus, we cannot accurately determine, for example, the centrality of the metabolite within metabolism or ascertain its true importance from an evolutionary standpoint. In contrast, the global network makes apparent these possibilities because it includes all available organismal knowledge. An increased understanding of why an organism develops certain “solutions” for its metabolic needs will aid in predicting unique features of the organism's metabolite and reaction usage that can be specifically targeted by drugs or other therapeutics and metabolic engineering.
Additionally, the global metabolism also allows us to view the metabolite and reaction usage of organisms in a general framework providing a means to identify metabolic “devices”, small groups of metabolites and reactions that have a functional purpose and other features that become apparent only when considering intermediate scales within the network21,22,23. This enables us to give greater insight into both metabolic evolution as well as ways to design synthetic metabolic “circuits” from these devices24,25.
Methods
Data acquisition
We downloaded multiple instances of the Kyoto Encyclopedia of Genes and Genomes (KEGG) LIGAND database26,27,28; the first instance on June 24, 2009 and the last on February 22, 2011. We also downloaded enzyme protein sequences from KEGG on five occasions, all between July 2010 and February 2011. All possible, unique sequences for each enzyme were used, based on the associations to reactions from KEGG. We downloaded bioreaction databases for Ma 2003 and Zeng 201129,30 from http://www.tuharburg.de and the iAF1260 Escherichia coli reconstruction31 from the BiGG database32.
We considered 998 organisms listed in the KEGG database to construct the global network. We constructed protein databases and predict metabolic networks for 874 of these 998 organisms. We did not predict the networks for 125 organisms due to a lack of sequence availability or because the time necessary to run a complete analysis for larger organisms was prohibitively long.
The bacterial domain dominates in representation due to the breakdown of organisms in KEGG itself. However, the network is not influenced by this over-representation because each reaction is only counted once in the construction of the global network. We include the domain and clade breakdown of the organisms that we tested and predicted metabolic networks for in Table 2.
Organismal and global network construction
We constructed individual metabolic networks for 998 organisms using a 2009 snapshot of the KEGG database. In these networks, each node represents a metabolite and two metabolites i and j are connected by an edge if there is a chemical reaction in which i is a substrate and j is its product, or vice versa. We established these relationships using only the main reaction pair designations on KEGG and, as in prior studies33,34, excluded transfer ions, co-factors and energy carrier molecules to maintain focus on the biomass transfer through the networks (Fig. 2a).
We constructed a global network by performing the graph union of all organismal networks (Fig. 2b). The 3,467 distinct reactions listed for the 998 organisms in the KEGG database yielded a global metabolic network comprising 6,656 metabolites and 3,328 unique edges. These metabolites are organized into a giant component comprising 2,023 metabolites and 2,729 edges and 333 smaller components typically comprising only a few metabolites each. We focused our analyses on the giant component of the global network because it contains the most reliable data, as its metabolites are more conserved and have more pathway annotations.
Metabolic networks for E. coli based on other databases were constructed in the same manner as the organismal metabolic networks constructed using the KEGG database. For the Ma 2003 and Zeng 2011 datasets the main pairs designation was included in the original dataset and it is used instead of the KEGG main pairs designation, while we used the main pairs designation from KEGG for the iAF1260 reconstruction.
Organismal network prediction
We collected 5.94 × 106 known enzyme amino acid sequences from the KEGG database that are associated with the 3,467 reactions in the global network and prepared databases of all known proteins for 874 organisms from the nr database (downloaded February 23, 2011) in accordance with the BLAST user manual35 in order to test sequence homology. We used blastp18,19, version 2.2.24, to align the known enzyme amino acid sequences to the organismal protein databases. We obtained a total of 2.6 × 1010 BLAST alignments that were subsequently used in our analysis.
Consensus network construction
We create consensus networks using a majority rule, similar to other work36. A set of the networks is selected and every edge in all of the networks is evaluated. If the edge appears in the majority of the networks in the set then it is added to the consensus network, otherwise it is not added. We then calculated the accuracy statistic for each network not used in the consensus network against the consensus metabolic network.
Network connectivity
To assess network connectivity we examine two quantities, the probability of a reaction addition closing a gap between two network components and a reaction removal creating an additional network component. We then compare the observed number of gaps versus the random chance expectation of completing a gap of a given size with the available number of additional reactions. For the random filling of gaps, we use the intersection of additional reactions between our predicted network and the KEGG 2011 network for an organism as the number of reactions that should be added. For the creation of additional network components we removed every edge individually in all organismal networks and determine if an additional network component is created. We then average the number of additional components added across all edges tested.
References
Pennisi, E. How will big pictures emerge from a sea of biological data? Science 309, 94 (2005).
Vitkup, D., Kharchenko, P. & Wagner, A. Influence of metabolic network structure and function on enzyme evolution. Genome Biology 7, R39+ (2006).
Yus, E. et al. Impact of genome reduction on bacterial metabolism and its regulation. Science 326, 1263–1268 (2009).
Sommer, M. O., Church, G. M. & Dantas, G. A functional metagenomic approach for expanding the synthetic biology toolbox for biomass conversion. Molecular systems biology 6 (2010).
Raymond, J. & Segrè, D. The effect of oxygen on biochemical networks and the evolution of complex life. Science 311, 1764–1767 (2006).
Zhao, J. et al. Modular co-evolution of metabolic networks. BMC bioinformatics 8, 311+ (2007).
Reed, J. L. et al. Systems approach to refining genome annotation. Proceedings of the National Academy of Sciences 103, 17480–17484 (2006).
Henry, C. S. et al. High-throughput generation, optimization and analysis of genome-scale metabolic models. Nat Biotech 28, 977–982 (2010).
Christian, N., May, P., Kempa, S., Handorf, T. & Ebenhoh, O. An integrative approach towards completing genome-scale metabolic networks. Mol. BioSyst. 5, 1889–1903 (2009).
Kharchenko, P., Chen, L., Freund, Y., Vitkup, D. & Church, G. M. Identifying metabolic enzymes with multiple types of association evidence. BMC bioinformatics 7, 177+ (2006).
Braakman, R. & Smith, E. The emergence and early evolution of biological Carbon-Fixation. PLoS Comput Biol 8, e1002455 (2012).
Maslov, S., Krishna, S., Pang, T. Y. & Sneppen, K. Toolbox model of evolution of prokaryotic metabolic networks and their regulation. Proceedings of the National Academy of Sciences 106, 9743–9748 (2009).
Plata, G. A., Fuhrer, T., Hsiao, T.-L., Sauer, U. & Vitkup, D. Global probabilistic annotation of metabolic networks enables enzyme discovery. Nat Chem Biol 8, 848–854 (2012).
Whitehead, A. N. Process and Reality (Gifford Lectures Delivered in the University of Edinburgh During the Session 1927-28) (Free Press, 1979), 2nd edn.
Guimerà, R., Mossa, S., Turtschi, A. & Amaral, L. A. N. The worldwide air transportation network: Anomalous centrality, community structure and cities' global roles. Proc. Natl. Acad. Sci. USA 102, 7794–7799 (2005).
Colizza, V., Barrat, A., Barthélemy, M. & Vespignani, A. The role of the airline transportation network in the prediction and predictability of global epidemics. Proc. Natl. Acad. Sci. USA 103, 2015–2020 (2006).
Colizza, V., Barrat, A., Barthelemy, M., Valleron, A.-J. & Vespignani, A. Modeling the worldwide spread of pandemic influenza: baseline case and containment interventions. PLoS Med 4, e13 (2007).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Fawcett, T. An introduction to ROC analysis. Pattern Recognition Letters 27, 861–874 (2006).
Milo, R. et al. Network motifs: Simple building blocks of complex networks. Science 298, 824–827 (2002).
Alon, U. Network motifs: theory and experimental approaches. Nature Rev. Gen. 8, 450–461 (2007).
Spirin, V., Gelfand, M. S., Mironov, A. A. & Mirny, L. A. A metabolic network in the evolutionary context: Multiscale structure and modularity. Proceedings of the National Academy of Sciences 103, 8774–8779 (2006).
Papin, J. A., Reed, J. L. & Palsson, B. Ø. Hierarchical thinking in network biology: The unbiased modularization of biochemical networks. Trends Biochem. Sci. 29, 641–647 (2004).
Lazebnik, Y. Can a biologist fix a radio?—Or, what I learned while studying apoptosis. Cancer Cell 2, 179–182 (2002).
Kanehisa, M., Goto, S., Furumichi, M., Tanabe, M. & Hirakawa, M. KEGG for representation and analysis of molecular networks involving diseases and drugs. Nucleic Acids Res. 38, D355–D360 (2010).
Kanehisa, M. et al. From genomics to chemical genomics: New developments in KEGG. Nucleic Acids Res. 34, D354–D357 (2006).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Ma, H. & Zeng, A.-P. Reconstruction of metabolic networks from genome data and analysis of their global structure for various organisms. Bioinformatics 19, 270–277 (2003).
Sun, J., Kamphans, T., Fekete, S. P. & Zeng, A.-P. An extended bioreaction database that significantly improves reconstruction and analysis of genome-scale metabolic networks. Integr. Biol. 3, 1071–1086 (2011).
Feist, A. M. et al. A genome-scale metabolic reconstruction for Escherichia coli K-12 MG1655 that accounts for 1260 ORFs and thermodynamic information. Mol. Syst. Biol. 3, 121 (2007).
Schellenberger, J., Park, J. O., Conrad, T. M. & Palsson, B. Ø. BiGG: a Biochemical Genetic and Genomic knowledgebase of large scale metabolic reconstructions. BMC Bioinformatics 11, 213 (2010).
Guimerà, R. & Amaral, L. Cartography of complex networks: modules and universal roles. J. Stat. Mech.: Theor. Exp. art. no. P02001 (2005).
Guimerà, R. & Amaral, L. Functional cartography of complex metabolic networks. Nature 433, 895–900 (2005).
Camacho, C. et al. Blast command line applications user manual. Internet (2008).
Osterman, A. Missing genes in metabolic pathways: a comparative genomics approach. Current Opinion in Chemical Biology 7, 238–251 (2003).
Acknowledgements
We thank I. Sirer, P.D. McMullen, E.N. Sawardecker, S.M.D. Seaver and P.B. Winter for comments and suggestions. ARP acknowledges support from the Northwestern Predoctoral Biotechnology Training Grant and the Chicago Biomedical Consortium with support from The Searle Funds at the Chicago Community Trust. RG acknowledges the support of the James S. McDonnell Foundation, of the Spanish Ministerio de Ciencia e Innovación grant FIS2010-18639 and of the European Union Grant PIRG-GA-2010-277166. LANA acknowledges the support of NSF award SBE 0624318 Foundation and the W.M. Keck Foundation.
Author information
Authors and Affiliations
Contributions
A.M.M. collected the initial dataset. A.R.P. and L.A.N.A. designed the analysis and A.R.P. conducted the computational analyses and collected the remaining data. A.R.P., R.G., A.M.M. and L.A.N.A. discussed the results and prepared the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Pah, A., Guimerà, R., Mustoe, A. et al. Use of a global metabolic network to curate organismal metabolic networks. Sci Rep 3, 1695 (2013). https://doi.org/10.1038/srep01695
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01695
This article is cited by
-
Robust structure measures of metabolic networks that predict prokaryotic optimal growth temperature
BMC Bioinformatics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.