Abstract
Epigenetic changes have been associated with ageing and cancer. Identifying and interpreting epigenetic changes associated with such phenotypes may benefit from integration with protein interactome models. We here develop and validate a novel integrative epigenome-interactome approach to identify differential methylation interactome hotspots associated with a phenotype of interest. We apply the algorithm to cancer and ageing, demonstrating the existence of hotspots associated with these phenotypes. Importantly, we discover tissue independent age-associated hotspots targeting stem-cell differentiation pathways, which we validate in independent DNA methylation data sets, encompassing over 1000 samples from different tissue types. We further show that these pathways would not have been discovered had we used a non-network based approach and that the use of the protein interaction network improves the overall robustness of the inference procedure. The proposed algorithm will be useful to any study seeking to identify interactome hotspots associated with common phenotypes.
Similar content being viewed by others
Introduction
Epigenetic changes, including aberrations in DNA methylation (DNAm), are a common hallmark of many complex diseases1,2. Aberrant DNA methylation has also been associated with age3,4,5,6,7,8, which is, by far, the strongest demographic risk factor for many common diseases including cancer, diabetes and Alzheimer's9,10. Thus, it is of biological and clinical interest to identify molecular pathways which may become epigenetically deregulated through age-associated DNA methylation. However, a common difficulty, shared by all genomic analyses, is the identification and interpretation of the observed molecular changes.
The most common approach to genomic analysis starts with the identification of differentially altered features (e.g. differentially expressed genes or differentially methylated regions) and subsequent biological interpretation using Gene Set Enrichment Analysis (GSEA)11. However, as shown by many studies in the gene expression field (see e.g. Ref. 12), this approach can miss important biological pathways, because the inference does not take the pathway or network structure into account and because changes affecting individual features are often of a small magnitude. Thus, a number of statistical approaches have emerged which use the pathway/network structure in the inference procedure12,13,14. These approaches directly infer network modules and altered pathways which subsequently facilitates the biological interpretation. Interestingly, while there exists a significant number of studies using these integrative network approaches in the context of gene expression12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29, there is a surprising lack of applications to the DNA methylation context30. Given that DNA methylation is implicated in the regulation of gene expression, it makes sense to also investigate the integration of a protein interactome with such epigenetic data. Indeed, the main key question we wanted to address here is whether DNA methylation changes associated with a given phenotype of interest occur randomly or not in the context of a human interactome model. In fact, we hypothesized that DNA methylation changes associated with cancer or age might cluster in the human interactome, targeting specific molecular pathways, in the same way that gene expression and copy-number changes also appear to target specific molecular pathways (see e.g. Ref. 31).
To address our hypothesis we collected DNA methylation data sets generated using the Illumina Infinium platform32, focusing our attention on gene promoter regions and on age as the phenotype of interest. Although other genomic regions may be more predictive of gene expression33, we here restrict to promoter regions since to date most of the data sets with available age information have been generated with the Infinium 27k platform, which by definition is restricted to CpGs in the promoter regions. Our focus on age is further motivated by the following. First, there is now substantial evidence that age-associated DNAm changes can be common to many different tissue types5,6,7,34. Moreover, while studies have reported individual genes and pathways that undergo age-associated changes in gene expression35,36,37,38,39,40,41, consistency of age-associated gene expression changes appears to be very weak38 in comparison to the changes observed at the DNA methylation level. Indeed, recent studies have reported tissue independent DNAm based age-predictors42,43. Second, it has already been demonstrated that age-associated DNAm changes do not happen randomly across the genome3,4,5,6,7. For instance, while most of the genome undergoes age-associated hypomethylation, promoters of high CpG density upstream of key developmental and tumour suppressor genes undergo preferential hypermethylation with age4,5,6,7. It is therefore natural to investigate age-associated DNAm changes in the context of the human interactome, since this may help identify specific molecular pathways or functional gene modules that underpin the aging process and which may indicate the risk of future disease. Thus, the second key question which we address here is whether specific molecular pathways are targeted by age-associated aberrant DNA methylation and whether this is tissue specific or not.
Results
Integration of DNA methylation with a protein interaction network
We first aimed to demonstrate that integration of DNA methylation with a protein interaction network (PIN) can make biological sense. To show this, we integrated Illumina Infinium DNA methylation beadarray data covering 27,000 CpGs32 with a human protein interaction network of over 10,000 proteins44. The main Illumina Infinium data sets used are listed in Table 1 . The integration was done by assigning to each protein in the network the methylation profile of the CpG closest to the transcription start site of the corresponding gene (Methods). The resulting networks contained on the order of 8000 genes/proteins and over 100,000 interactions. We observed that the integration led to methylation profiles of neighboring genes in the network being on average more highly correlated (as assessed over normal tissue samples) than those of non-neighbors ( SI Fig. S1 ). Interestingly, this effect was of a magnitude comparable to that seen in gene expression studies ( SI & SI Fig. S1 ). These findings indicated to us that the built integrated networks can be exploited to extract biologically meaningful information.
A spin-glass (SPG) algorithm to detected differential methylation interactome hotspots
We next wanted to assess if differential methylation associated with a given phenotype of interest targets specific regions of the protein interaction network (a property we call modularity). To address this, one must search the interactome for connected subnetworks where the association with the phenotype is stronger than in the rest of the network. These subnetworks we interchangeably call “hotspots” or “EpiMods” (Epigenetic Modules). To find hotspots associated with cancer and age, one could adapt a number of previously published algorithms including jActive Modules45, BioNet14 or DEGAS21. However, we also wanted to explore other algorithmic alternatives and so we here considered a computationally very efficient spin-glass (SPG) module detection algorithm46 (Methods, Fig. 1 ). The efficiency of the spin-glass algorithm stems from a local greedy implementation which maximises a relative weight density (called modularity) centred around specific seeds46,47 and where, in our case, the weight edges reflect the combined strength of association of each gene's DNAm profile with the phenotype of interest ( Fig. 1 , Methods). The seeding strategy used is described in Methods. The statistical significance of any inferred modules was assessed further using a permutation scheme (1000 permutations) in which the node statistics were randomly permuted over the network followed by recomputation of the modularity values (Methods). Those modules with an adjusted P-value < 0.05 were declared as hotspots/EpiMods.
The EpiMod spin-glass algorithm: algorithm for the discovery of differential methylation hotspots (“EpiMods”).
For each gene in the protein interaction network we choose the CpG closest to the transcription start site and evaluate a statistic (e.g t-statistic) of association between DNA methylation and the phenotype of interest (here we depict a binary phenotype).These statistics are then combined to assign weights to the edges of the interactome. Next, for each of the top significantly differentially methylated genes (“seeds”), a local community detection algorithm is applied to identify a module containing the seed and maximising the sum of weights. These modules are then checked to see if their modularity (i.e. total weights) are significant against distributions obtained by random permutations of the observed node statistics. Finally, the modules whose modularity is significantly larger than what would be expected by chance constitute the hotspots, called “EpiMods” and represent the candidate modules to be subsequently validated in the independent datasets.
An attractive feature of the spin-glass algorithm is its simplicity in that it is characterised by just one main free parameter, γ, which can be tuned on simulated and training data to optimize a complex trade-off between the number of inferred modules, their sizes, the average overlap between the modules and a GSEA enrichment quality metric ( SI Fig. S2 ). In fact, the sizes of the inferred modules is key since we observed that in general small modules (sizes <10) did not lead to significant modularity values, while very large modules (>200) were non-unique and highly redundant. We found that the optimal parameter choice was γ ≈ 0.5, since at this value, modules in the desired size range 10–100 genes were generated ( SI Fig. S2 ).
Benchmarking of the spin-glass algorithm
In order to further justify the use of the local greedy spin-glass (SPG) algorithm, we compared it to two other but closely related module detection methods: an agglomerative fast greedy (FG) non-local algorithm48 and a non-greedy non-local spectral decomposition (SD) algorithm49 ( SI Methods ). These other algorithms attempt to maximise a modularity score similar to that of the spin-glass algorithm46, but differ substantially in the inference procedure, allowing us to assess both the impact of “greediness” and locality. To provide an independent and objective assessment of these algorithms, we compared their ability to detect (i) biological modules on the interactome defined by gene expression modules from the Molecular Signatures Database (MSigDB)11 and (ii) modules generated by a simulation model (Methods). In both scenarios, the spin-glass algorithm at our previously determined optimal parameter (γ = 0.5) performed markedly better in finding modules that overlapped more strongly with the known biological and simulated modules ( SI Fig. S3 ).
We also decided to benchmark our algorithm on real data. We applied it to four data sets (LBC1, OVC1, CVX, ENDO, Table 1 ) in order to find cancer diagnostic (LBC1, CVX, ENDO) and prognostic (OVC1) EpiMods. Consistent with the results obtained on simulated data, we observed that the spin-glass algorithm detected, on average, smaller sized modules and a substantially higher fraction of modules with significant modularity as assessed using the random permutation scheme described earlier ( Table 2 ). In line with this, the modularities were also higher for the spin-glass algorithm ( Table 2 & SI Fig. S4 ). Thus, all these results demonstrate that the spin-glass algorithm (tuned with γ = 0.5) is more powerful than the other closely related module detection algorithms.
Existence and validation of age-EpiMods in blood tissue
Having tested and fined tuned the spin-glass algorithm, we next applied it to the problem of identifying hotspots of age-associated differential methylation (“age-EpiMods”). Given the small effect size of age on the DNA methylome we argued that a relatively large number of samples would be needed to robustly identify age-EpiMods. Thus, as a discovery set, we used a large-scale Infinium 27k study profiling DNAm levels in whole blood samples from 261 postmenopausal women, 148 from healthy women and 113 from age-matched women with ovarian cancer (UKOPS data set, Table 1 )50. Disease status was used as a covariate in the linear regression model between age and DNA methylation to ensure that results were not confounded by the presence of ovarian cancer50. We integrated the resulting regression statistics with our human interactome model resulting in a network of approximately 8000 nodes and over 100,000 interactions ( Fig. 2A ). As seeds, we chose the top ranked genes (assigned CpGs) from the linear age DNAm regression model. These seeds we also call “GAMPs” (Genes with Age-associated Methylation change in their Promoters). Importantly the previously determined choice γ = 0.5 in the spin-glass algorithm was near optimal also in this data set ( SI Fig. S5 ). Thus, the choice γ = 0.5 appears to be an optimal choice across different data sets and in relation to different phenotypes.
Discovery and validation of age-EpiMods in whole blood tissue.
(A) Integrated DNAmethylome-interactome network with illustrated differential methylation hotspots. (B) The age-modularity scores of the 12 age-EpiMods with significant modularity values, ranked according to significance. The gene names indicate the seeds/GAMPs defining the modules. (C) A zoomed-in version of one of the age-EpiMods with seed SOX8. Directionality of age-associated methylation change of nodes is as indicated, with edge colors reflecting the combined strength of age-associated methylation change of the two edge-members. (D) Validation of the age-associated modularities of 6 of the 12 age-EpiMods in the validation T1D set. y-axis labels the modularity score, with red lines indicating the observed modularities in the T1D set and the boxplots showing those obtained by the Monte Carlo sampling scheme (1000 randomisations). All 6 have modularity significance P-values < 0.05 in the test set.
Application of the EpiMod algorithm to the integrated network identified 12 fairly unique age-EpiMods (individual Monte Carlo FDR < 5%) each containing at least 10 genes ( SI Table. S1 , Fig. 2B & 2C ). Since 113 of the 261 samples were from women with active ovarian cancer, we checked that these hotspots were also retrieved in the subset of 148 samples from healthy women. Indeed, we observed that 10/12 (83%) were also retrieved in the healthy subset, indicating that the algorithm is robust to subsampling and that the majority of these EpiMods reflect changes that are observed in normal tissue. We note that fewer EpiMods (8 at a FDR < 0.05, i.e 4 fewer) would have been discovered had we applied the EpiMod algorithm to only the 148 healthy samples.
Since the spin-glass algorithm is of a greedy nature, the resulting inferred modules could represent spurious artefacts. Thus, it is key to validate the inferred age-EpiMods on independent data and thus also demonstrate that the algorithm can make robust findings. As validation, we used data from 187 independent whole blood samples from type1-diabetics (T1D data set, Table 1 ). We thus asked if the hotspot nature of the 12 age-EpiMods is also recapitulated in this independent data set. We note that this constitutes a stringent validation whereby we assess if the precise EpiMods of the discovery set are associated with age in the test set more strongly than randomly picked genes from the network. Remarkably, of the original 12 age-EpiMods, 6 (50%) validated as hotspots in the T1D data set ( Fig. 2D , Fig. 3A ). To assess the significance of this further, we randomly redefined 12 EpiMods on the network (matched for module size and local topology, see SI Methods ) and found that on average (as estimated over 100 randomisations) only 2% of the 12 random EpiMods are expected false positives. Thus, none of the 6 validated age-EpiMods are likely to be false positives. Reassuringly, these 6 age-EpiMods attained significant or marginally significant age-modularity scores in an additional whole blood data set (ALSc, Table 1 , Fig. 3B ). Importantly, we also checked that the directionality and significance of individual age-EpiMod members was validated for 6 of the 12 age-EpiMods in the independent data ( SI Fig. S6 , Fig. 3A ). We also constructed predictive age methylation scores in the test set samples, using only significant EpiMod members in the discovery set and these scores correlated with age in 10 of the 12 age-EpiMods ( SI Fig. S7 , Fig. 3A ). Using either method, 10 of the 12 age-EpiMods (83%) validated in terms of their constituent members being associated with age and with the same directionality as in the discovery set. Together, all these results indicated to us that age-EpiMods in blood tissue exist.
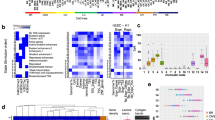
Tissue specificity and tissue-independence of the 12 age-EpiMods from the UKOPS discovery cohort in whole blood.
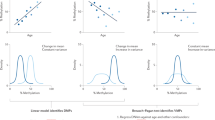
Each panel plots for each of the 12 age-EpiMods, the -log10 of the P-values (y-axis) of 3 different validation tests: “Stat” is the linear correlation test P-value testing the correlation of the age-associated t-statistics in UKOPS against the corresponding statistics in the validation set. “Score” is the linear correlation test P-value from correlating an age-predictive methylation score in the test set against age using only EpiMod gene members which were significant in the discovery UKOPS cohort. “Mod” is the P-value from the randomisation test (1000 Monte Carlo runs) testing for significance of the “hotspot” modularity values. Each panel A to H represents one different data set (see Table 1 ). (A-B) are in whole blood, (C-D) in brain tissue, (E) skin, (F) buccal, (G) CD4+ T-cells and (H) CD14+ monocytes.
Biological significance of age-EpiMods
Given that proteins involved within a given cellular process are more likely to be connected and to form communities in the network51,52,53,54, we performed a GSEA11 on each of the 6 age-EpiMods that validated in whole blood tissue to ascertain their biological significance ( SI Table S2 ). Each of these age-EpiMods was strongly enriched for specific biological functions, which were also largely unique to each module ( Table 3 , SI Table S2 ). One of the EpiMods, consisting of 19 genes, was strongly enriched for transcription factors and PolyComb Group gene Targets (PCGTs)55, which are genes that are repressed in stem cells by the PRC2 complex ( SI Table S2 , Fig. 2C ). Another two EpiMods (FZD2 & GRIA2) were highly enriched for genes in the WNT/beta-Catenin signaling and glutamate receptor pathways, respectively ( Table 3 ). Two further EpiMods (CD247 & LCK), which were significantly larger in size, were highly enriched for genes implicated in immune response pathways.
Tissue specific and tissue-independent age-EpiMods
Next, we wanted to determine the tissue-specificity of the 12 age-EpiMods identified in the original discovery set. We collected additional DNAm data sets from other normal tissue types, including skin, buccal, brain and fractionated purified blood cell types (CD4+ T-cells and CD14+ monocytes) ( Table 1 )5,34,42. Correlations of the age-EpiMods with age in these additional data sets was assessed using three different complementary evaluation frameworks, as done on the independent whole blood data sets. Two of these evaluations assess the consistency of directionality and significance of individual EpiMod members without regard to the network structure, while the third evaluation assesses specifically the network hotspot nature of the age-EpiMods. Thus, for each age-EpiMod in each data set we obtained three different P-values to assess validation at various levels. Remarkably, we observed that the three age-EpiMods with seeds SOX8, GRIA2 and FZD2 were largely tissue independent, validating across most, if not all, of the other normal tissue types ( Fig. 3C–F ). These age-EpiMods also correlated with age in purified CD4+ T-cells and more marginally so in CD14+ monocytes ( Fig. 3G–H ), further supporting the view that these associations are not driven by changes in cell-type composition. In contrast, the immune response age-EpiMods showed more variability and were not consistently validated across other normal tissue types ( Fig. 3C–F ), suggesting that these may be driven by changes in cell type composition. Supporting this view, these age-EpiMods generally did not validate in the purified blood cell types ( Fig. 3G–H ) and they exhibited strong overlaps with markers of granulocytes and lymphocytes (Binomial test P < 10−5)56. Venn diagrams summarising the degree of tissue-specificity of the age-EpiMods illustrate that most of the 12 age-EpiMods are indeed specific to blood ( Fig. 4A ), with only SOX8, GRIA2, FZD2 age-EpiMods being largely tissue independent ( Fig. 4C ). The tissue specificity of detected age-associated changes in DNAm was further confirmed at the level of individual top ranked GAMPs within each data set and tissue type ( Fig. 4B & 4D ).
Venn diagrams illustrating consistency of GAMPs and age-Epimods found within tissues and between tissues.
(A) Diagram illustrating age-EpiMods validated at the 5% level (node-weight permutation) in UKOPS, T1D and ALSc whole blood data sets. (B) Overlaps between top 100 ranked GAMPs in four separate blood datasets. (C) Overlaps in modules validated at the 5% level using Fisher combined independent probability across different tissue types. (D) Overlaps between top 100 ranked GAMPs across tissue types using UKOPS, SKIN, BUC and FCTX as representatives for whole blood, skin, buccal and brain respectively. Note the considerably lower consistency in (D) than in (B).
Although the validations demonstrate overall consistency in terms of directionality and significance of the age-associations of EpiMod members, we next wanted to determine if it is the same EpiMod members driving the significance across all data sets, or if the significance is driven by different genes in different data sets. We focused on the three age-EpiMods SOX8, GRIA2 and FZD2, since these were validated in most data sets. Heatmaps of the age-associated t-statistics revealed striking consistency ( Fig. 5 ). For instance, in the SOX8 module, UTF1, POU3F3 and FOXD3 all showed the same directional age-associated methylation patterns in the three large whole blood data sets ( Fig. 5A ). Similarly, in the FZD2 module, many genes (e.g. SFRP1, FZD2, ROR2, DKK1, FZD9, WNT3A) showed consistent associations across most data sets ( Fig. 5C ). In contrast, in the GRIA2 module, fewer genes showed consistent associations ( Fig. 5B ).
The observation that the age-EpiMods SOX8, GRIA2 and FZD2, validated across multiple tissue types including purified CD4+ T-cells which have a high-turnover rate, suggests that the observed age-associated changes must be due to underlying changes in the long-lived stem-cell populations. We first tested this in mesenchymal stem cells (MSC) from the bone marrow of 8 patients spanning a wide age-range57. As before, we tested the association with age using the 3 different aforementioned methods. The hotspot nature was recapitulated marginally for GRIA2 (P = 0.07) and FZD2 (P = 0.07), while the directionality and significance of the age-associations of individual genes was validated in the case of the SOX8 and FZD2 modules (SI Fig. S8), clearly implicating the specific genes in these modules as undergoing age-associated DNAm changes in stem cells. We also analysed 12 samples of hematopoietic progenitor (HPC) cells (7 from cord blood and 5 samples all with ages ≥ 28)58. However, in this set the hotspot nature of the three age-EpiMods was not observed, possibly due to a narrower age-range and the fact that all samples were relatively young (under the age of 41). In spite of this, the linear age correlation test P-values still showed marginal trends towards significance for the same two modules (SOX8, FZD2) that validated in mesenchymal stem cells ( SI Fig. S9 ).
EpiMod algorithm outperforms non-network based approach
Having established the existence of a number of biologically significant age-EpiMods, we next wanted to assess the importance of the EpiMod algorithm itself in making this discovery. To assess this, we compared the EpiMod method to a simple non-network based approach. First, we perfomed a simple GSEA on a number of top-ranked GAMPs from the discovery set, where the number of genes used for the GSEA was matched to the number of unique EpiMod members making up the original 12 age-EpiMods. Not unexpectedly, two of the key functional terms, namely WNT-signaling and glutamate receptors, were not picked out by the non-network based GSEA (Fisher-test P = 0.24 (WNT) and P = 0.19 (glutamate-receptors). To further demonstrate the power of the EpiMod strategy, we next selected, from the 12 age-EpiMods in the discovery cohort, those EpiMod members which were significant (P< 0.05) while also discarding the 12 GAMPs which were used to seed these EpiMods. There were a total of 75 significant EpiMod genes. We posited that these genes would constitute more robust markers of age-associated differential methylation than those we would identify using a non-network based approach. We thus compared the absolute t-statistics of these significant EpiMod genes in three independent whole blood data sets, to the corresponding statistics for a random selection of significant GAMPs from the discovery set. Across the three independent test data sets, we observed that significant EpiMod genes from the UKOPS discovery set generally attained higher absolute statistics (preserving the same directionality as in the discovery set) than the genes used for benchmarking (combined Fisher test P < 0.001, Fig. 6 ). Thus, we can conclude that the EpiMod algorithm allows improved biological interpretation of age-associated changes in DNA methylation without compromising the robustness of the identified age-associated markers and this is important in light of recent reports questioning the value of network-based methods59.
Improved robustness of the EpiMod algorithm.
Boxplots of the absolute statistics (y-axis) of the 75 significant EpiMod members from the 12 age-EpiMods discovered in the UKOPS whole blood data set, discarding the 12 GAMPs used to seed these EpiMods (“EpiMod” in green). As a benchmark, we show the corresponding absolute statistics of 75 randomly selected GAMPs (i.e. significantly associated genes). In all cases, only genes with the same directionality in discovery and test set were used. A comparison is made in the original discovery set as well as in each of three large independent whole blood test sets (T1D, ALSc, SZc). Wilcoxon rank sum test P-values are shown.
Discussion
Here we have explored the integration of DNAm profiles with a human interactome and have demonstrated that hotspots of differential methylation associated with phenotypes such as cancer and age exist. In order to identify the hotspots we adapted a powerful module detection algorithm (the spin-glass SPG algorithm). The algorithm was tested extensively, optimized and benchmarked against two closely related module detection algorithms (FG & SD), demonstrating its superiority ( SI Figs. S2–S4 ). Three key aspects of the algorithm are worth emphasizing. First, it is computationally efficient and scalable to large networks thanks to a local greedy implementation. Second, the implementation used does not seriously compromise the robustness of the inference procedure: indeed, we were able to infer hotspots of differential methylation which then validated as network hotspots in completely independent data ( Fig. 2 ), thus demonstrating that the algorithm can make robust findings. Third, the algorithm has a tunable parameter that directly controls the sizes of the inferred modules. This is important since we observed that the size of an inferred module influenced the probability of the module being a false positive ( Table 2 ).
Given the improved performance of the SPG algorithm against FG and SD, we also wondered how it would compare to some of the other module detection algorithms. Thus, we compared SPG to three popular algorithms: jActiveModules45, BioNet14 and DEGAS21 (see Methods for details of parameter choices). Like SPG, jActive Modules allows for a local search strategy centred around selected seeds, thus allowing for a direct comparison, while BioNet and DEGAS perform the module search recursively and globally, respectively. The comparison of these algorithms showed that BioNet and DEGAS only identified a relatively small number of significant modules, with high robustness observed only in the case of BioNet ( SI Fig. S10 ). jActiveModules and SPG both identified similar number of significant modules, with a similar fraction of modules identified in the training set validating in the test sets ( SI Fig. S10 ). We also point out these two algorithms inferred distinct modules. For instance, the GRIA2 module, which was validated by SPG in all data sets considered here, was not detected and not clearly validated using jActiveModules. Thus, we can see that the spin-glass algorithm used here compares favourably to some of the existing module detection algorithms.
In our main analysis, where we focused on age as the phenotype of interest and on whole blood tissue, we were able to validate age-EpiMods in several independent whole blood data sets, supporting the view that DNAm changes do not occur randomly in the context of the human interactome. Using DNAm data from other normal tissues, we showed that the majority of the hotspots/Epimods were tissue-specific ( Figs. 3 – 4 ), but also found three age-EpiMods which validated independently of tissue type ( Figs. 3 – 5 ). Two of these age-EpiMods (SOX8, FZD2) were significantly enriched for transcription factors and PCGTs, implicating specific stem cell differentiation pathways. These particular age-EpiMods were also seen to correlate with age in mesenchymal stem cells. As with the age-PCGT signature we had derived previously and which has been shown to be largely tissue-independent6 (see also Ref. 5,60), the observation made here that members of these EpiMods consistently correlate with age in a variety of different tissue and cell types is a clear indication that these age-associated epigenetic modules represent generic age-epigenetic phenomena not driven by changes in cell-type composition. The third tissue-independent age-EpiMod was highly enriched for G-protein and glutamate receptors.
It is of interest to discuss further the potential biological significance of these three age-EpiMods. One of the most consistently age-correlated genes in the SOX8 EpiMod was UTF1. A recent study61 has shown that UTF1 is a chromatin-associated protein necessary for stem cell differentiation. Hence, the observation that UTF1's promoter region becomes consistently hypermethylated with age could imply that the differentiation capacity of adult stem cells is compromised with age. A second independent observation is that UTF1 expression is a stronger marker of efficient iPSC reprogramming than other traditional markers (e.g. OCT4)62. Interestingly, it has been observed that the efficiency and safety of iPSC reprogramming decreases with the age of the adult differentiated cells used63,64. Thus, lower expression of UTF1 as a result of age-associated hypermethylation could potential contribute to this observation. To test if gene expression of UTF1 decreases with age will require access to large sample collections of CD34+ HPCs from human donors, as DNAm profiles derived from mice or other model organisms are unlikely to capture the age cumulative effects of environmental stressors which humans are exposed to naturally and which are assumed to underlie the observed age-associated changes in DNA methylation. Indeed, in a genome-wide study using MeDIP-Seq, we did not observed major changes in DNAm comparing HPCs from young and old mice (personal communication, data not shown). Our EpiMod analysis also implicated other factors which may play similar or analogous roles to UTF1, such as for instance SOX8 itself. The age-EpiMod FZD2 implicated many members of the WNT/beta-Catenin signalling pathway (e.g. SFRP1, FZD2, FZD9, WNT3A) all of which appear to become consistently hypermethylated with age. This is another striking finding, since the WNT-signalling pathway has been the subject of intensive study in ageing65,66,67. In fact, these studies indicate that WNT signalling activity is increased with age potentially increasing stem cell self-renewal function whilst blocking differentiation. However, another study reported that WNT target gene expression in osteoblast precursors decreases with age, albeit this too is an observation made in mice68. These relations are further complicated by the fact that we observe simultaneous hypermethylation in the promoters of important receptors (e.g FZD2, FZD9) and that of SFRP1, which is a key negative regulator of the WNT-pathway. Hence, although promoter hypermethylation and silencing of SFRP1 is often observed in cancer and preneoplastic lesions69,70, it is unclear what the effect of this may be in the background of promoter FZD2 & FZD9 hypermethylation. The functional significance of the FZD2 age-EpiMod is therefore unclear and analysis of large-scale gene expression data sets did not reveal a consistent pattern (data not shown). The third tissue-independent age-EpiMod involved glutamate receptors (e.g. GRIA2) and the prolactin releasing hormone receptor PRLHR. Strikingly, these two genes showed strong promoter age-associated hypermethylation across almost all of the data sets analysed. Interestingly, there are reports documenting that glutamate receptors play an important role in dietary restriction, ageing and possibly also in modulating age-associated immunosenescence71,72.
Besides the three age-EpiMods discussed above, two others (TNK1, LAMB1) ( SI Table S1 ) are of particular interest, although their validation was not consistent across the different data sets and tissues. The TNK1 age-EpiMod was a fairly large module consisting of 446 genes, hence why the age-modularity was not validated in other data sets. However, this module did validate in terms of the directional consistency and significance of individual regression statistics ( SI Fig. S6 & S7 ). Interestingly, TNK1 is a late-onset Alzheimer's disease gene as determined by a recent meta-analysis of GWAS studies73. The LAMB1 age-EpiMod was strongly enriched for laminins, including besides LAMB1, also LAMA1, LAMA4, LAMA3, LAMC3 ( SI Table S1 ), which are a family of extracellular matrix glycoproteins implicated in cell adhesion and differentiation and which we have found to be associated with early stages of cervical cancer (data not shown).
It is important to emphasize that the discovery of the three tissue-independent age-EpiMods implicating key molecular pathways was only made possible through an integrative interactome-epigenome strategy. Indeed, previous meta-analysis studies on age-associated DNAm changes (e.g. Ref. 34) have not reported the existence or importance of the SOX8, FZD2 and GRIA2 age-EpiMod members. Reinforcing this, we found that an ordinary GSEA on a matched number of top ranked GAMPs in the discovery UKOPS cohort yielded non-significant odds ratios of enrichment for the WNT-signalling and glutamate receptor pathways. We also showed that genes significantly associated with age and which were implicated by the EpiMod analysis defined more robust age-associated DNAm markers ( Fig. 6 ). These results therefore attest to the power of an integrative network strategy since it simultaneously facilitates biological interpretation while also identifying more robust markers, specially if these markers are associated with relatively small effect sizes.
In summary, we have shown that differential methylation interactome hotspots exist and specifically, have discovered tissue-independent age-epigenetically deregulated hotspots enriched for stem cell differentiation pathways. The integrative network algorithm developed here could help elucidate the biological and clinical significance of epigenetic changes associated with common diseases.
Methods
The protein interaction network
We downloaded the Human Protein Reference Database (HPRD) interaction network from Pathway Commons http://www.pathwaycommons.org44 (13th June 2012). The HPRD network used here consisted of approximately 10,000 proteins with over 200,000 documented interactions. When restricted to the genes annotated on the Infinium 27k array and with probes passing quality control criteria, the resulting networks contained typically on the order of 8000 nodes and just over 100,000 interactions (edges).
Integration and edge weighting scheme
For each gene g represented in a methylation dataset, denote the methylation state of the probe nearest to the transcription start site by βg. Given a phenotype of interest, denote by tg the statistic (usually, a t-statistic) resulting from the statistical test correlating the phenotype to βg. In the case of age, we used linear regressions t-statistics, while in normal/cancer comparisons we used a two-sided t-test. This allows us to attach a weight wgh between every pair of genes g and h in the underlying protein interaction network (PIN):

where |t|max is the maximum absolute t-statistic assigned by the test to any gene. Note that if g and h do not interact we set wgh = 0. This yields a weighting of each edge between 0 and 1, with larger weights denoting two neighbors both significantly associated with the phenotype of interest.
Identifying hotspots of alterations using a greedy spin-glass algorithm
With the choice of edge weighting regime above, the task is to search the network for subnetworks of exceptionally large average weight density (“modules” or “hotspots”) in relation to the network as a whole. The average weight density of a subnetwork we call modularity49.
In order to identify the hotspots we adopted a greedy spin-glass algorithm from46, which allows community detection in weighted networks specified by a weighted adjacancy matrix W. The justification for using this algorithm is as follows. First, we sought an algorithm that would allow flexibility in the level of mutual exclusivity of the inferred modules. Since proteins often lie in many distinct functional pathways, the algorithm must allow for some overlap between the inferred modules. On the other hand, the algorithm must also avoid inferring modules of very high overlap since this represents unwanted redundancy. One of the attractive features of the spin-glass algorithm is its simplicity in that it is characterized by only 1 main free tunable parameter, called γ with 0 ≤ γ ≤ 146, which, as we show here, directly controls the average size and hence overlap/redundancy of the inferred modules. Secondly, inference of modules in large-scale networks can be computationally demanding and unstable49. Thus, a greedy approach is attractive because it can offer the scalability and computational efficiency needed for our application. The stability of the modules inferred using a greedy approach can always be tested a posteriori using validation/test data sets, a strategy that we also adopt here. Importantly, the spin-glass algorithm allows for a greedy approach by starting the search of modules from seed nodes, defined as the nodes with the largest statistics, or alternatively as those with the largest average weights (i.e the average over nearest neighbors). Both seed choices are possible, although the former leads to less redundancy in the sense that seeds are more spread out across the network. The top number of genes ranked by largest absolute statistic were declared as seeds. In the case of the data sets and phenotypes considered here we typically focused on the top 100 seeds because these always passed an FDR correction threshold of 0.05 based on their univariate P-values. It is important to point out here that not all seeds lead to modules of higher-than-average modularity, since some seeds may represent isolated nodes of association.
In detail, the spin-glass algorithm reduces the problem of community/module detection into finding the ground state of an infinite ranged Potts spin glass46. Briefly, communities/modules are found via a Hamiltonian

where σi denotes the community that node i belongs to, Wij is the weighted adjacency matrix of the network and where pij describes the probability of an edge between nodes i and j according to some appropriate null model. In the above expression, δ (σi, σj) is the Kronecker delta and γ > 0 is the tunable parameter of the algorithm. It can be shown that this Hamiltonian rewards internal edges (i.e. those within an inferred subnetwork, or equivalently within the same spin state) as well as non-edges between inferred subnetworks, while also penalizing internal non-edges and edges between different subnetworks46. The choice of parameter γ controls the relative energy contributions of edges and non-edges occuring both internally and externally of the inferred subnetworks46. An alternative expression for the Hamiltonian is in terms of adhesion coefficients, ars,

where r and s denote the spin states (i.e communities) and where the adhesion coefficient between spin states r and s is defined by

In the above, mrs denotes the number of edges between r and s and the second term denotes the expectation of this number under the null model. Thus, minimisation of the Hamiltonian is equivalent to the minimisation of overall adhesion46. In fact, one can use a local greedy algorithm, adding a node i to the existing seed community s only if its adhesion to s, ais, is higher than the adhesion to the rest of the network (r), air, that is, if

In this work, we use as null models, random graphs with the same vertex degrees, amounting to a rewiring of the network. Under such a null model, pij = kikj/2E where E is the total number of edges.
For each seed we thus obtain a module (if present) minimising the overall adhesion using a simulated annealing procedure as implemented in the spin-glass.community function of the igraph R package. We point out again that the existence of a module associated with a given seed is not automatic since growing a module from a given seed as described above may not lead to reductions in the overall adhesion. Typically, one finds that approximately 50% of seeds are not associated with any module.
Tuning the parameter γ
The spin-glass algorithm has one main tunable parameter (γ), which determines the size of the inferred modules. We tuned γ to yield modules in the size range 10 to 100 genes. There are two justifications for focusing on this size range. First, unsupervised linear decomposition methods like PCA or ICA performed on gene expression data (see e.g74) suggest that co-regulated gene modules are typically on the order of 1% of the number of genes measured. In our network of ~8000 nodes this amounts to ~80 genes. A second more compelling justification was provided by a detailed analysis of the module size distribution, the mutual exclusivity of the resulting modules and an overall GSEA enrichment metric. This strategy allowed us to gauge the trade-off between the uniqueness of the modules (favouring smaller modules) and biological interpretation (favouring larger modules and requiring modules to be typically of size ≥10). We found that γ ≈ 0.5 was optimal in the sense that it generated a reasonable number of gene modules of size ≥10 while also exhibiting minimal overlap. In comparison, γ = 1 yielded much larger, overlapping modules, leading to significant redundancy in downstream GSEA analyses. Briefly, to estimate the GSEA quality score we first computed Q = Σ1≤i≤j max (ORi, 100) where j = 5 and ORi is the odds ratio of the i'th top ranked GSEA enriched term. The odds ratios were computed for each significantly non-overlapping module of size ≥10 and finally Q was rescaled according to the expected value for a module of the same size to get a “GSEA enrichment score” q given by q = Q/E[Q|m] with m denoting the module size.
Statistical significance and external validation of modules
It is important to establish the statistical significance of the modularity values associated with the inferred communities or modules. The modularity M of each community  is simply the total weight of all edges in
is simply the total weight of all edges in  , i.e.
, i.e.

where  has edge weights wgh and nodes
has edge weights wgh and nodes  . Permuting the edge weights over the network would yield the wrong null distribution to compare to since the edge weights themselves already contain the topological nearest neighbor information. Instead of this, we compare the distribution of modularity values with the ones expected by permuting the absolute t-statistics among nodes. By performing 1000 permutations, we can thus assess the statistical significance of the observed modularity values by counting the fraction of permutations which lead to higher modularities. We note that the modules inferred previously, i.e from the unpermuted node statistics, are kept fixed in this procedure and are not inferred de novo. In other words, only the assigned node attributes (i.e the absolute t-statistics) change between permutations, thus yielding an adjusted P-value for each separate module. We refer to the resulting modules which are significant at the 5% level, as EpiMods.
. Permuting the edge weights over the network would yield the wrong null distribution to compare to since the edge weights themselves already contain the topological nearest neighbor information. Instead of this, we compare the distribution of modularity values with the ones expected by permuting the absolute t-statistics among nodes. By performing 1000 permutations, we can thus assess the statistical significance of the observed modularity values by counting the fraction of permutations which lead to higher modularities. We note that the modules inferred previously, i.e from the unpermuted node statistics, are kept fixed in this procedure and are not inferred de novo. In other words, only the assigned node attributes (i.e the absolute t-statistics) change between permutations, thus yielding an adjusted P-value for each separate module. We refer to the resulting modules which are significant at the 5% level, as EpiMods.
To validate the EpiMods in external data we used two complementary strategies designed to test different aspects of the EpiMods. To validate the significance and directionality of the DNAm changes associated with the gene members of the EpiMods, we generated starburst scatterplots, comparing the t-statistics of differential methylation in the discovery set against the corresponding t-statistics in the validation set. To demonstrate that the EpiMods validate as hotspots of differential methylation in the network, we computed the modularities in the validation set and then tested the significance of these modularities against random permutations of the node statistics in this same validation set.
Availability
We have implemented the EpiMod algorithm as an executable R-script which is freely available on request, or from http://code.google.com/p/epimods/.
References
Jones, P. A. & Baylin, S. B. The fundamental role of epigenetic events in cancer. Nat Rev Genet 3, 415–428 (2002).
Feinberg, A. P., Ohlsson, R. & Henikoff, S. The epigenetic progenitor origin of human cancer. Nat Rev Genet 7, 21–33 (2006).
Christensen, B. C. et al. Aging and environmental exposures alter tissue-specific dna methylation dependent upon cpg island context. PLoS Genet 5, e1000602 (2009).
Maegawa, S. et al. Widespread and tissue specific age-related dna methylation changes in mice. Genome Res 20, 332–340 (2010).
Rakyan, V. K. et al. Human aging-associated dna hypermethylation occurs preferentially at bivalent chromatin domains. Genome Res 20, 434–439 (2010).
Teschendorff, A. et al. Age-dependent dna methylation of genes that are suppressed in stem cells is a hallmark of cancer. Genome Res 20, 440–446 (2010).
Heyn, H. et al. Distinct dna methylomes of newborns and centenarians. Proc Natl Acad Sci U S A 109, 10522–10527 (2012).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol Cell (2012).
Balducci, L. & Ershler, W. B. Cancer and ageing: a nexus at several levels. Nat Rev Cancer 5, 655–662 (2005).
Hoeijmakers, J. H. Dna damage, aging and cancer. N Engl J Med 361, 1475–1485 (2009).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102, 15545–15550 (2005).
Chuang, H. Y., Lee, E., Liu, Y. T., Lee, D. & Ideker, T. Network-based classification of breast cancer metastasis. Mol Syst Biol 3, 140 (2007).
Ulitsky, I. & Shamir, R. Identification of functional modules using network topology and high-throughput data. BMC Syst Biol 1, 8 (2007).
Beisser, D., Klau, G. W., Dandekar, T., Mller, T. & Dittrich, M. T. Bionet: an r-package for the functional analysis of biological networks. Bioinformatics 26, 1129–1130 (2010).
Bhardwaj, N. & Lu, H. Correlation between gene expression profiles and protein-protein interactions within and across genomes. Bioinformatics 21, 2730–2738 (2005).
Bhardwaj, N. & Lu, H. Co-expression among constituents of a motif in the protein-protein interaction network. J Bioinform Comput Biol 7, 1–17 (2009).
Pujana, M. A. et al. Network modeling links breast cancer susceptibility and centrosome dysfunction. Nat Genet 39, 1338–1349 (2007).
Dittrich, M. T., Klau, G. W., Rosenwald, A., Dandekar, T. & Mller, T. Identifying functional modules in protein-protein interaction networks: an integrated exact approach. Bioinformatics 24, i223–i231 (2008).
Taylor, I. W. et al. Dynamic modularity in protein interaction networks predicts breast cancer outcome. Nat Biotechnol 27, 199–204 (2009).
Bandyopadhyay, S. et al. Rewiring of genetic networks in response to dna damage. Science 330, 1385–1389 (2010).
Ulitsky, I., Krishnamurthy, A., Karp, R. M. & Shamir, R. Degas: de novo discovery of dys-regulated pathways in human diseases. PLoS One 5, e13367 (2010).
Schramm, G., Nandakumar, K. & Konig, R. Regulation patterns in signaling networks of cancer. BMC Syst Biol 4, 162 (2010).
Teschendorff, A. E. & Severini, S. Increased entropy of signal transduction in the cancer metastasis phenotype. BMC Syst Biol 4, 104 (2010).
Li, J. et al. Identification of high-quality cancer prognostic markers and metastasis network modules. Nat Commun 1, 34 (2010).
Califano, A. Rewiring makes the difference. Mol Syst Biol 7, 463 (2011).
Ideker, T. & Krogan, N. J. Differential network biology. Mol Syst Biol 8, 565 (2012).
Wang, J. et al. Noa: a novel network ontology analysis method. Nucleic Acids Res 39, e87 (2011).
Beisser, D. et al. Robustness and accuracy of functional modules in integrated network analysis. Bioinformatics 28, 1887–1894 (2012).
Backes, C. et al. An integer linear programming approach for finding deregulated subgraphs in regulatory networks. Nucleic Acids Res 40, e43 (2012).
Liu, H. et al. Prioritizing cancer-related genes with aberrant methylation based on a weighted protein-protein interaction network. BMC Syst Biol 5, 158 (2011).
TCGA. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Bibikova, M. et al. 2009. Epigenomics 1, 177–200 (2009).
Irizarry, R. A. et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific cpg island shores. Nat Genet 41, 178–186 (2009).
Horvath, S. et al. Aging effects on dna methylation modules in human brain and blood tissue. Genome Biol 13, R97 (2012).
Lee, C. K., Weindruch, R. & Prolla, T. A. Gene-expression profile of the ageing brain in mice. Nat Genet 25, 294–297 (2000).
Fraser, H. B., Khaitovich, P., Plotkin, J. B., Pbo, S. & Eisen, M. B. Aging and gene expression in the primate brain. PLoS Biol 3, e274 (2005).
Zahn, J. M. et al. Agemap: a gene expression database for aging in mice. PLoS Genet 3, e201 (2007).
Magalhaes, J. P., Curado, J. & Church, G. M. Meta-analysis of age-related gene expression profiles identifies common signatures of aging. Bioinformatics 25, 875–881 (2009).
Edwards, M. G. et al. Gene expression profiling of aging reveals activation of a p53-mediated transcriptional program. BMC Genomics 8, 80 (2007).
Cao, K., Chen-Plotkin, A. S., Plotkin, J. B. & Wang, L. S. Age-correlated gene expression in normal and neurodegenerative human brain tissues. PLoS One 5 (2010).
Harries, L. W. et al. Advancing age is associated with gene expression changes resembling mtor inhibition: evidence from two human populations. Mech Ageing Dev 133, 556–562 (2012).
Koch, C. M. & Wagner, W. Epigenetic-aging-signature to determine age in different tissues. Aging (Albany NY) 3, 1018–1027 (2011).
Bocklandt, S. et al. Epigenetic predictor of age. PLoS ONE 6, e14821 (2011).
Cerami, E. G. et al. Pathway commons, a web resource for biological pathway data. Nucleic Acids Res 39, D685–D690 (2011).
Ideker, T., Ozier, O., Schwikowski, B. & Siegel, A. F. Discovering regulatory and signalling circuits in molecular interaction networks. Bioinformatics 18, S233–S240 (2002).
Reichardt, J. & Bornholdt, S. Statistical mechanics of community detection. Phys. Rev. E 74, 016110 (2006).
Nepusz, T., Yu, H. & Paccanaro, A. Detecting overlapping protein complexes in protein-protein interaction networks. Nat Methods 9, 471–472 (2012).
Clauset, A., Newman, M. E. & Moore, C. Finding community structure in very large networks. Phys Rev E Stat Nonlin Soft Matter Phys 70, 066111 (2004).
Newman, M. E. Modularity and community structure in networks. Proc Natl Acad Sci U S A 103, 8577–8582 (2006).
Teschendorff, A. E. et al. An epigenetic signature in peripheral blood predicts active ovarian cancer. PLoS One 4, e8274 (2009).
Jeong, H., Mason, S. P., Barabsi, A. L. & Oltvai, Z. N. Lethality and centrality in protein networks. Nature 411, 41–42 (2001).
Jonsson, P. F. & Bates, P. A. Global topological features of cancer proteins in the human interactome. Bioinformatics 22, 2291–2297 (2006).
Palla, G., Derenyi, I., Farkas, I. & Vicsek, T. Uncovering the overlapping community structure of complex networks in nature and society. Nature 435, 814–818 (2005).
Jonsson, P., Cavanna, T., Zicha, D. & Bates, P. Cluster analysis of networks generated through homology: automatic identification of important protein communities involved in cancer metastasis. BMC Bioinformatics 7, 2 (2006).
Lee, T. I. et al. Control of developmental regulators by polycomb in human embryonic stem cells. Cell 125, 301–313 (2006).
Palmer, C., Diehn, M., Alizadeh, A. A. & Brown, P. O. Cell-type specific gene expression profiles of leukocytes in human peripheral blood. BMC Genomics 7, 115 (2006).
Bork, S. et al. DNA methylation pattern changes upon long-term culture and aging of human mesenchymal stromal cells. Aging cell 9, 54–63 (2010).
Bocker, M. T. et al. Genome-wide promoter dna methylation dynamics of human hematopoietic progenitor cells during differentiation and aging. Blood 117, e182–e189 (2011).
Staiger, C. et al. A critical evaluation of network and pathway-based classifiers for outcome prediction in breast cancer. PLoS One 7, e34796 (2012).
Winnefeld, M. & Lyko, F. The aging epigenome: Dna methylation from the cradle to the grave. Genome Biology 13, 165 (2012).
van den Boom, V. et al. UTF1 is a chromatin-associated protein involved in ES cell differentiation. J. Cell Biol. 178, 913–924 (2007).
Buganim, Y. et al. Single-cell expression analyses during cellular reprogramming reveal an early stochastic and a late hierarchic phase. Cell 150, 1209–22 (2012).
Miura, K. et al. Variation in the safety of induced pluripotent stem cell lines. Nat Biotechnol (2009).
Wang, B. et al. Reprogramming efficiency and quality of induced pluripotent stem cells (ipscs) generated from muscle-derived fibroblasts of mdx mice at different ages. PLoS Curr 3, RRN1274 (2011).
Brack, A. S. & Rando, T. A. Intrinsic changes and extrinsic influences of myogenic stem cell function during aging. Stem Cell Rev 3, 226–237 (2007).
Brack, A. S. et al. Increased wnt signaling during aging alters muscle stem cell fate and increases fibrosis. Science 317, 807–810 (2007).
Maiese, K., Li, F., Chong, Z. Z. & Shang, Y. C. The wnt signaling pathway: aging gracefully as a protectionist? Pharmacol Ther 118, 58–81 (2008).
Almeida, M., Han, L., Martin-Millan, M., O'Brien, C. A. & Manolagas, S. C. Oxidative stress antagonizes wnt signaling in osteoblast precursors by diverting beta-catenin from t cell factor-to forkhead box o-mediated transcription. J Biol Chem 282, 27298–27305 (2007).
Caldwell, G. M. et al. The wnt antagonist sfrp1 in colorectal tumorigenesis. Cancer Res 64, 883–888 (2004).
Caldwell, G. M. et al. The wnt antagonist sfrp1 is downregulated in premalignant large bowel adenomas. Br J Cancer 94, 922–927 (2006).
Chakrabarti, L., Bandyopadhyay, B. C. & Poddar, M. K. Is age-induced decline in immune response associated with hypothalamic glutamate receptor density and dietary protein? Nutr Neurosci 4, 375–387 (2001).
Magnusson, K. R. Influence of dietary restriction on ionotropic glutamate receptors during aging in c57b1 mice. Mech Ageing Dev 95, 187–202 (1997).
Belbin, O. et al. Investigation of 15 of the top candidate genes for late-onset alzheimer's disease. Hum Genet 129, 273–282 (2011).
Teschendorff, A. E., Journe, M., Absil, P. A., Sepulchre, R. & Caldas, C. Elucidating the altered transcriptional programs in breast cancer using independent component analysis. PLoS Comput Biol 3, e161 (2007).
Acknowledgements
J.W. was supported by an EPSRC/BBSRC CoMPLEX PhD studentship. A.E.T. was supported by a Heller Research Fellowship. S.B. was supported by the Wellcome Trust (WT084071) and a Royal Society Wolfson Research Merit Award (WM100023). We would like to thank Gareth Wilson of the Medical Genomics group at the UCL Cancer Institute for sharing data and Martin Widschwendter for discussions.
Author information
Authors and Affiliations
Contributions
J.W. and A.E.T. performed the statistical analyses. A.E.T. devised the study. S.B. and X.W. contributed data and/or ideas. J.W. and A.E.T. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Table S2
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
West, J., Beck, S., Wang, X. et al. An integrative network algorithm identifies age-associated differential methylation interactome hotspots targeting stem-cell differentiation pathways. Sci Rep 3, 1630 (2013). https://doi.org/10.1038/srep01630
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep01630
This article is cited by
-
A higher dysregulation burden of brain DNA methylation in female patients implicated in the sex bias of Schizophrenia
Molecular Psychiatry (2023)
-
Novel epigenetic network biomarkers for early detection of esophageal cancer
Clinical Epigenetics (2022)
-
Inference of epigenetic subnetworks by Bayesian regression with the incorporation of prior information
Scientific Reports (2022)
-
Clinical epigenetics settings for cancer and cardiovascular diseases: real-life applications of network medicine at the bedside
Clinical Epigenetics (2021)
-
Sex-differential DNA methylation and associated regulation networks in human brain implicated in the sex-biased risks of psychiatric disorders
Molecular Psychiatry (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.