Abstract
Avian influenza viruses are widespread in birds, contagious in humans and are categorized as low pathogenicity avian influenza or highly pathogenic avian influenza. Ferrets are susceptible to infection with avian and human influenza A and B viruses and have been widely used as a model to study pathogenicity and vaccine efficacy. In this report, the natural history of the H5N1 influenza virus A/Vietnam/1203/04 influenza infection in ferrets was examined to determine clinical and laboratory parameters that may indicate (1) the onset of disease and (2) survival. In all, twenty of 24 animals infected with 7 × 105 TCID50 of A/Vietnam/1203/04 succumbed. A statistical analysis identified a combination of parameters including weight loss, nasal wash TCID50, eosinophils and liver enzymes such as alanine amino transferase that might possibly serve as indicators of both disease onset and challenge survival.
Similar content being viewed by others
Introduction
Worldwide, annual influenza epidemics result in approximately 3–5 million cases of severe disease and 250,000 to 500,000 deaths1. Occasionally, type A viruses emerge and cause great mortality over several countries. Examples of pandemics include the 1957 H2N2 Asian-flu, 1968 H3N2 Hong-Kong Flu and most recently the 2009 H1N1Swine-flu. The most notable example is the virus that caused the 1918 pandemic, which affected an estimated 30% of the world's population and resulted in nearly 20 to 50 million deaths worldwide2.
Until an outbreak in Hong Kong claimed six human lives in 1997 (A/Hong Kong/156/97), H5N1 avian influenza viruses were thought to be incapable of infecting humans directly. However, the initial H5N1 outbreak and more recent H5N1, H7N7 and H9N2 epizootics have revealed that avian influenza viruses can infect humans without prior adaptation and can cause significant morbidity and mortality in infected individuals. Since 2003, there have been at least 608 confirmed human cases of H5N1 influenza virus infections with 359 deaths (approximately 59% mortality; WHO, Confirmed cases of avian influenza H5N1)3.
Ferrets (Mustela putorius furo) are susceptible to infection with human influenza viruses and their disease resembles that of human influenza. A number of studies have demonstrated the pathogenicity of H5N1 avian influenza virus strains in ferrets4,5,6,7. Additionally, the ferret is widely used as the model to evaluate influenza vaccines and therapeutics8,9,10.
Successful post-exposure prophylaxis (PEP) to viral pathogens is typically dependent upon timing of treatment, requiring either immediate or early initiation following exposure. For example, as noted in a review by the writing committee of the second world health organization consultation on clinical aspects of human infection with avian influenza A (H5N1), early treatment of human patients with Oseltamivir is recommended, with high mortality associated with late initiation of treatment12. Similarly, the Centers for Disease Control and Prevention recommends that post-exposure antiviral chemoprophylaxis with neuraminidase inhibitors be provided as soon as possible to close contacts of suspected H5N1 cases (within two days). Therefore, delineation of early parameters associated with onset of influenza disease and the association of these parameters with survival and mortality will aid in understanding the disease progression and the affects of the treatments on survival.
The objective of this natural history study was to refine the ferret model of A/Vietnam/1203/04 (A/Vn) influenza infection to determine clinical and laboratory parameters that may indicate the onset of disease and survival. These parameters may serve to better evaluate vaccines and/or therapeutics in future treatment studies. Clinical and laboratory parameters used to define the onset of disease and potential survival included virus shedding in nasal wash specimens, viremia, weight, temperature, clinical activity scores and hematology and clinical chemistry. Statistical evaluation of the data revealed differences among the parameters tested between animals that survived challenged and animals that succumbed.
Results
The effects of H5N1 challenge on ferrets
Following challenge, clinical observations and activity scores were recorded at eight-hour intervals from three different groups until 96 hours post-challenge, after which observations were recorded twice daily. Time-points within groups were 24 hours apart, but staggered such that observations from one flu-infected and one sham-infected group were available every eight hours. Common clinical observations included mucoid stools (diarrhea for 45% infected animals), inappetance, lethargy and labored breathing (Figure 1A). Observations that were suggestive of neurological involvement included ataxia, torticollis, tremors or involuntary movements/seizures and limb paralysis or paresis. Time of onset for clinical signs occurring within the first 96 hours post-challenge are presented in Figure 2B. Earliest onset for neurological symptoms included tremors, ataxia and torticollis at 8, 40 and 88 hours post-challenge, respectively. Clinical observations for sham-infected animals were limited to inappetance, mucoid stools and nasal discharge (for a single animal).
Clinical signs of disease in ferrets following infection with A/Vietnam/1203/04 virus.
Animals were monitored twice daily for clinical observations and at eight-hour intervals until 96 hours, using the following scoring system; 0 = alert and playful, 1 = alert but playful only when stimulated, 2 = alert but not playful when stimulated and 3 = neither alert nor playful when stimulated. (A) Percentage of animals exhibiting clinical observations, (B) time of onset for clinical observations presenting during the first 96 hours following infection and (C) activity score changes for influenza-infected animals.
Over the full course of the study and with increased frequency during the first 96 hours following challenge, clinical observations data included the use of a scoring system incorporating a numerical scale of 0–3 with increasing score associated with reduced activity. Sham-infected animals did not exhibit significant activity level changes at any of the time-points examined (scores remained at 0). Flu-infected animals that succumbed had significant changes in activity scores beginning 88 hours post-challenge and continuing from day 4 through day 8 (Figure 2C). In contrast, the time of onset and the degree of change for activity scores were delayed and reduced for survivors of influenza infection with activity for all surviving animals returning to normal within 14 days following infection.
Kaplan-Meier estimates were plotted to illustrate the mortality data for influenza-infected and sham-infected animals (Figure 2). Only 4 of 24 animals (16.7%) survived following intranasal challenge with approximately 7 × 105 median tissue culture infectious dose (TCID50) of A/Vn virus. All of the sham-infected animals survived. The survival proportions were significantly difference (p-value < 0.001; Fisher's exact test).
Changes in body temperatures and body weight after H5N1 challenge
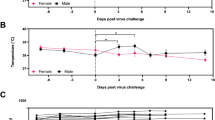
A comparison of differences among the influenza challenged and sham-infected groups in temperature changes from baseline at each study day revealed statistically significant group effects. Average temperature for the influenza-infected animals and the sham-infected animals are presented in Figure 3A. Influenza infection in all animals resulted in a disruption of the diurnal rhythm and an increase in body temperature. There were significant differences between the influenza-infected and sham-infected groups beginning 16 hours through 64 hours, 80 hours, 88 hours, study day 5 (PM), study day 6 (PM), study day 7 (PM), study day 8 (PM), study day 11 (AM), study day 13 (PM) and study day 16 (AM). There does not seem to be a correlation when comparing the amount of temperature increase, per animal, with mortality (Figure 3B). However, temperatures returned to relatively normal levels in the surviving animals nine days after challenge and a more normal diurnal temperature pattern was observed in animals that survived challenge.
Change in body temperature and body weight following infection with A/Vietnam/1203/04 virus.
Temperatures were taken twice daily prior to challenge and then at approximately eight-hour intervals from 8–96 hours by use of subcutaneous implantable temperature transponders. After 96 hours, temperatures were recorded twice daily. Body weights were taken every other day, beginning on day 0 until the end of the study (day 18) or death of the animal. Weights taken immediately prior to challenge on day 0 served as baseline. (A) Mean body temperature, (B) Mean temperature comparison between survivors and animals that succumbed to disease, (C) mean body weight, (D) mean change from baseline weight for influenza-infected animals by survival status.
Body weight data collected prior to challenge on day 0 were used as the baseline value for each animal. A/Vn-infected animals had a significant decrease in body weight when compared to baseline as early as one day post-challenge (Figure 3C). The average changes in body weight were significant on all post-challenge study days (Figure 3C). By comparison, a single significant change (decrease) in body weight for the sham-infected animals was observed on day 7, which may be due to variations within ferret weights that can occur in a relatively quick manner. It should be noted that the weights of the sham-challenged animals were back to normal by day 9. When comparing animals that survived challenge to animals that succumbed to infection (Figure 3D) animals that succumbed to infection lost more body weight in a quicker manner than animals that survived. The surviving animals lost the most average weight by day 11 post-challenge. However, these animals began to gain weight by day 13, post-challenge.
Hematology and clinical chemistry changes associated with h5n1challenge in the ferret
Blood was collected on day −22 and −7 in order to establish baseline hematology and clinical chemistry values. Following challenge, due to limitations on blood volumes and to reduce effects of repeated sedation, data collection was staggered by group pairs (Groups A+B, C+D and E+F) according to Table 1. Samples were collected at eight-hour intervals from 8 through 96 hours, after which time a final terminal blood sample was collected from moribund animals at the time of euthanasia or from survivors on day 18.
For red blood cell parameters, there were decreases from baseline in the flu-infected animals that were significantly different from those in the sham-infected animals for RDW (red cell distribution width) on 7 consecutive time points between 48–96 hours (Figure 4A). RDW is a coefficient of variation of the population of red blood cells on a volume histogram. For PLT (platelets), reductions in the flu-infected animals that were significantly different from those in the sham-infected animals were observed at 24 hours and from 40–96 hours (Figure 4B). However, platelets have a very wide range and require significant variation outside of normal range to affect bleeding. Among the white blood cell parameters from infected animals, significant reductions relative to the sham-infected animals were observed for WBC (white blood cells) from 56–96 hours, eosinophils at 8, 48, 64, 72, 88, 96 hours and on day 18, monocytes at 56, 64, 80 and 88 hours and lymphocytes from 16–96 hours (4C–F, respectively). Decreased eosinophils are clinically insignificant, while the number of WBCs in the blood may be an indicator of stress or inflammation of disease. Increases were observed for large unstained cells (LUC) at 32 hours and from 72–96 hours, neutrophils from 24–56 hours and the N/L ratio from 16–88 hours (4G–I, respectively) in infected animals that were significantly different than those in the sham-infected animals. Neutrophils are the most common cell seen in acute inflammation of disease and also as a response to stress, while LUC are either large or reactive lymphocytes, monocytes or leukemic blasts. Terminal hematology values for PLT, WBC, lymphocytes and EOS remained significantly lower and N/L ratio and LUC significantly higher than baseline pre-challenge values for infected animals that were euthanized prior to day 18. Changes in WBC, N/L and lymphocytes were less pronounced in survivors when compared to those animals that succumbed to infection.
For clinical chemistry, over the first 96 hours post-infection, noteworthy significant changes were limited to liver function parameters. These included significant increases in the flu-infected animals that were significantly different from those seen in the sham-infected animals for AST (aspartate transaminase) at 48, 64, 88 and 96 hours, ALT (alanine transaminase) at 88 and 96 hours and also for blood glucose at 32–96 hours (Figure 5A–C, respectively). Terminal clinical chemistry parameters that were significantly elevated included billirubin, AST, ALT, SDH, ALP and GGT (not shown).
Virus recovery from blood, nasal washes and post mortem animals following h5n1 challenge
Following challenge, development of viremia was monitored by qPCR. Viral shedding in nasal wash specimens and viral load in selected tissues was assessed using a combination of TCID50 analysis and quantitative polymerase chain reaction assays (qPCR).
Viremia, as measured by a positive blood qPCR result, was only observed for two animals over the first 96 hours post-challenge (one animal at 80 hours and one animal at 88 hours). The animal that was positive at 80 hours survived infection and was euthanized on day 18, at which time blood was negative for viral nucleic acid. The second animal (positive at 88 hours) succumbed to infection. For the remaining A/Vn challenged animals, blood samples were found to be positive only at the time of euthanasia/terminal blood draw (50% of the terminal blood samples from animals that were euthanized prior to the end of study were positive for virus; log mean of 3.86 gene copies/ml). All animals that survived to day 18 of the study were negative for viremia as measured by qPCR.
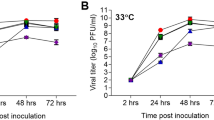
Results of TCID50 and qPCR assays on nasal wash specimens are summarized in Figure 6 A and B, respectively. For TCD50 assays, if the lower bound of the 95 percent confidence interval for the log-adjusted nasal wash was greater than 2.12 (the base-10 logarithm of the LOQ), then there was a significant increase in nasal wash TCID50for that sample. None of the animals were positive by TCID50 assay at 8 hours post-challenge. However, the percentage of animals positive for viral shedding gradually increased beginning at 16 hours (25% at 16 hours) so that by 24 hours, 6 of 8 (75%) of the samples were positive and by 48 hours all samples were positive. Animals that survived infection were all found to be positive for viral shedding by nasal wash TCID50. Significant viral nasal wash TCID50 titers were observed in the flu-infected animals at 48, 72 and 96 hours post-challenge. The qPCR assay was more sensitive in that viral nucleic acid was detected for all but a single nasal wash specimen from the influenza-infected animals at 8 hours post-challenge. All remaining nasal wash specimens collected between 16–96 hours were positive and represented significant virus recovery in the qPCR assay at all post-challenge time-points tested. As expected, none of the sham-infected animals were positive by either the TCID50 or qPCR assay.
Time-to abnormal analysis
A statistical analysis was performed in order to identify differences in time-to-abnormal between influenza infected animals that survived and those that succumbed to disease. Statistically significant differences among these groups were observed for weight (p = 0.01; t-test), nasal wash TCID50 (p = 0.03; t-test), eosinophils (p = 0.001; t-test) and ALT (p = 0.02; t-test). Animals that died had an earlier time to abnormal than those that lived (Table 2).
Discussion
Infection of ferrets with H5N1 was characterized by acute onset and rapid progression with twenty of the 24 infected animals succumbing prior to day 10 post-challenge (83.3% mortality) when animals were challenged with 7 × 105 TCID50 of A/Vietnam/1203/04. Results of complete necropsies confirmed systemic infection with extensive involvement of the lungs, liver, lymphoid tissues and brain (data not shown). Infected animals demonstrated a number of clinical signs of disease. With the exception of inappetence, infected animals that succumbed had more clinical signs, including lethargy, labored breathing, ataxia, torticollis, tremors, seizures and hind limb paralysis or paresis and demonstrated a higher percentage for those signs during the 18 day period following challenge. Mucoid stool and innappetance, while the most common observations, were present for both infected and sham-infected animals. However, it is possible that repeated administration of anesthesia provided as part of scheduled nasal wash procedures contributed to these observations. Sneezing has been reported to be a common clinical observation for animals infected with influenza virus11. However, this study found only a minority of animals demonstrating this symptom following infection with A/Vn. The discrepancy may be attributed to different influenza viruses; the current study utilized a H5N1 virus (A/Vn), while the former study used a H1N1 viral strain. While the majority of signs were observed for all infected animals, onset within the first 96 hours was more common for animals that succumbed to infection. Survivors were less likely to exhibit gastrointestinal symptoms, with vomiting and diarrhea observed exclusively for animals that died (observed at 24 hours post-challenge). Additional signs exclusively observed in animals that succumbed to disease included hind limb paralysis/paresis and involuntary movements. It should be noted that development of neurological symptoms, ataxia for example, was not predictive of eventual survival status. Pathology data supported this observation in that survivors were found to have evidence of inflammation and/or necrosis in the brain. Altogether, these data suggest that onset of disease within 96 hours after challenge may lead to mortality in a higher percentage of animals. Inclusion of a scoring system as part of the scheduled clinical observation schedule was valuable in that onset of change could be identified at 88 hours following infection. This outcome was similar for the combination of body temperature and weight change. Significant increases in temperature occurred as early as 16 hours post-infection. Weight loss, which was recorded daily, declined significantly beginning at 24 hours post-infection. Activity score and weight loss changes relative to baseline were limited exclusively to animals that succumbed to infection. Because temperature increases appears to be one of the first signs of disease following A/Vn challenge, this parameter should be considered as a “trigger to treat” in therapeutic efficacy studies and would likely guarantee product delivery within the 96 hour post-challenge window, allowing for a potential increase in survival.
Common symptoms of H5N1 infection in humans include fever, cough and sore throat, diarrhea and vomiting12. Most human deaths have been attributed to respiratory failure, but multi-organ failure with renal dysfunction, encephalitis and sepsis have been reported13. Common laboratory features of influenza A H5N1 infection in human patients include leukopenia, lymphopenia, thrombocytopenia, elevated aminotransferase levels and hyperglycemia14,15. Significant transient reductions in peripheral blood lymphocytes have also been demonstrated in H5N1 infected ferrets5. In the current study, similar to human disease, onset of changes in A/Vn-infected ferrets within the first 96 hours included significant reductions in lymphocytes, platelets, white blood cells, eosinophils, an altered N/L ratio and elevated LUC. LUC are either large lymphocytes or monocytes and in most animals, this parameter is a good indicator of either a stress response or inflammation of disease.
Lymphocyte changes are also a good indicator of the disease process, as they have been shown to transiently decrease in both ferrets and in human clinical cases. Pancytopenia is commonly observed in humans infected with H5N1, A/Vn infection in ferrets results in mixed picture with reductions in a number of cell types in the majority of animals. Animals that survived A/Vn challenge tended to have less pronounced changes for lymphocyte counts and liver function parameters over the first 96 hours post-infection.
For the clinical chemistry values, liver function parameters (ALT and glucose) correlate well with clinical pathology that demonstrated severe necrosis and inflammation of the liver. Additionally, billirubin, AST, ALT, SDH, ALP and GGT were significantly elevated in terminal samples. ALP and GGT are also indicators of liver damage and more specifically the biliary tract.
H5N1 virus shedding has been demonstrated in the nasopharynx of human patients. Between 76–100% of nose and throat swab specimens from human H5N1 infected patients were positive for viral nucleic acid by RT-PCR assays16. By comparison, ferrets infected with A/Vn had approximately 5 logs of virus in nasal wash specimens on days 3 and 5 post-infection as determined by an egg infectious dose 50 assay7. In this study, shedding of virus in the nasopharynx was assessed in nasal wash specimens by use of a combination of TCID50 and qPCR assays. The intent was to define onset of disease for each parameter in the ferret model. The eclipse period represents the time interval between infection with virus (viral penetration) and the production of infectious progeny virus. Analysis of qPCR demonstrated all, but a single sample, positive for viral nucleic acid. Thus, identifying onset of replication and production of new virus following the eclipse period using qPCR was not practical. The qPCR assay was of value however, as significant increases in viral nucleic acid were evident beginning at 16 hour post-infection. The TCID50 results, by comparison, demonstrated an eclipse period with a gradual increase in detectable virus through 48 hours. Onset of significant change occurred at 24 hours post-infection. Overall, qPCR may be advantageous for screening samples for H5N1 nucleic acid; however, the TCID50 assay should be used to detect onset of shedding of viable virus resulting from A/Vn infection.
Wang et al. found that viral RNA was frequently detected in the peripheral blood of ferrets one or two days before death17. In the current study, blood qPCR assays were positive for two samples during the first 96 hours post-infection (approximately 8% of the total number tested). The proportion increased to 50% for animals euthanized prior to day 18 of the study and was uniformly negative for animals surviving to day 18. The results are in line with clinical data in which 56% of blood samples from patients infected with A/Vn were positive for viral RNA16.
In all, the results of the current study demonstrate that the ferret is a relevant model to evaluate potential products designed to combat A/Vietnam/1203/04 exposure. The time-to-abnormal analysis showed statistically significant differences between the influenza infected animals and those that succumbed to disease for weight, nasal wash TCID50, eosinophils and ALT. Thus, these parameters may be used to predict whether animals will succumb or survive challenge. Future studies could be designed to test whether these parameters are useful as indicators of disease and disease treatment triggers when testing the efficacy of various antivirals or therapeutics.
Methods
Ethics statement
All experiments were conducted according to protocols reviewed and approved by the Institutional Animal Care and Use Committee of Battelle which adhere to the National Institutes of Health guidelines for the care and use of laboratory animals. Influenza-infected animals were housed in an ABSL-3 enhanced facility.
Animals
16-week-old male ferrets (Mustela putorius furo), purchased from Triple F Farms (Sayre, PA), were shipped to Covance (Denver, PA) for implantation of venous access ports prior to delivery to Battelle. All animals placed on study were confirmed to be seronegative by use of a hemagglutination inhibition assay for the currently circulating influenza viruses [Influenza B (Malaysia and Florida), Influenza A (Solomon Islands, Wisconsin and Brisbane-like)] and were in good health and free from malformations and signs of clinical disease at study start. Ferrets were single housed and ear tags were used to identify each animal. A computer randomization program was used to divide the animals into 3 groups of 8 ferrets and 3 groups of 3 ferrets, according to Table 1.
Intranasal challenge
Challenges occurred on day 0, after collection of weight and temperature data. Challenges were conducted by instilling approximately 600 μL of calcium and magnesium free phosphate buffered saline (CMF-PBS) containing 7 × 105 TCID50 of Influenza A/Vietnam/1203/04 (300 μL into each nostril) for each Telazol-anesthetized ferret (intramuscular at 22 mg/kg). The A/Vn virus used for challenge experiments was passed a single time in embryonated eggs prior to use. A back titer was performed via TCID50 to confirm the inoculation dose. Sham-infected groups received allantoic fluid diluted in CMF-PBS.
Body weights were taken every other day, beginning on day 0 until the end of the study (day 18) or death of the animal.
Clinical observations
Clinical observations were recorded twice daily for all animals from the time of arrival until challenge. Following challenge, observations were recorded according to the schedule defined in Table 1. Observations included the use of a scoring system based on work previously described by Reuman et al. 198918; see also Zitzow et al. 20025:
0 = alert and playful;
1 = alert but playful only when stimulated;
2 = alert but not playful when stimulated;
3 = neither alert nor playful when stimulated.
Observations were recorded twice daily for all groups after 96 hours post-challenge until the end of the study (day 18).
Temperature
Animals were quarantined for approximately 7 days, at which time they were implanted subcutaneously (under Telazol anesthesia) in the shoulder (1 chip) and rump area (1 chip) with temperature transponder chips. Twice daily temperatures (morning and evening; AM and PM, respectively) were taken from day -7 through day 0 in order to establish baseline. Following challenge, temperature was recorded according to Table 1. Temperatures were recorded twice daily for all groups after 96 hours post-challenge. Temperature readings were taken from both the chip in the rump and the chip in the shoulder. The average of these two measurements was used for determination of body temperature.
Blood collection
Bleeding was conducted by use of VAPs according to the schedule defined in Table 1. Due to limitations on blood volumes and to reduce effects of repeated sedation, data collection was staggered by group pairs (Groups A+B, C+D and E+F) so that samples were collected at approximately eight-hour intervals from 8–96 hours. Any animal found moribund underwent a terminal bleed for hematology and qPCR and for sera collection (for clinical chemistry) prior to euthanasia.
Median tissue culture infectious dose (TCID50)
A standard TCID50 assay using Madine Darby canine kidney cells (MDCK) cells was used to determine virus titer of nasal washes. Briefly, serial dilutions of samples were plated onto MDCK cell monolayers and the development of cytopathic effect (CPE) recorded. The Spearman Kärber method was used to determine the dilution at which 50% of the wells demonstrated CPE.
Quantitative polymerase chain reaction
Viral RNA was isolated from 100 µL of whole blood, nasal wash, or tissue homogenate using a NucliSens® easyMAG™ instrument (BioMérieux). A real-time one step RT-PCR assay was used for the detection of a A/Vietnam/1203/04 hemagglutinin (HA) gene using Invitrogen SuperScript III Platinum One-Step Quantitative RT-PCR kit in a 7900HT Sequence Detection System (Applied Biosystems, Foster City, CA) with forward primer 5'-GTGGCGAGCTCCCTAGCA-3', reverse primer 5'-TCTGCATTGTAACGACCCATTG-3' and probe 6'FAM-TGGCAATCATGGTAGCTGGTCTATCCTTATGG-MGBNFQ-3'. The final concentration of primers and probe were 900 nM and 250 nM, respectively. Thermal cycling parameters were as follows: 50°C for 15 minutes, 95°C for 2 minutes, followed by 40 cycles of 95°C for 15 seconds and 60°C for 1 minute. A standard curve (2 to 2 × 107 gene copies/μL) was run on each PCR plate to allow for absolute quantitation of all test samples.
Hematology and clinical chemistry
Blood was collected into EDTA tubes according to Table 1 and hematology analysis was conducted on the Advia 120 (Siemens). Blood was collected into SST tubes according to Table 1 and clinical chemistry analysis was conducted on the Advia 1200 (Siemens). The hematology parameters analyzed included: Red Blood Cell Count (RBC, 106 cells/μL), Hemoglobin (HGB, g/dL), Hematocrit (HCT, %), Mean Corpuscular Volume (MCV, fL), Mean Corpuscular Hemoglobin (MCH, pg), Mean Corpuscular Hemoglobin Concentration (MCHC, g/dL), Cell Hemoglobin Concentration Mean (CHCM, g/dL), Red Cell Distribution Width (RDW, %), Platelet Count (PLT, 103 cells/μL), Mean Platelet Volume (MPV, fL), White Blood Cell Count (WBC, 103 cells/μL), Neutrophils (103 cells/μL), Lymphocytes (103 cells/μL), Neutrophils/Lymphocytes Ratio (N/L Ratio), Monocytes (103 cells/μL), Eosinophils (103 cells/μL), Basophils (103 cells/μL) and Large Unstained Cells (LUC, 103 cells/μL). The clinical chemistry parameters analyzed included: Total Bilirubin (mg/dL), Aspartate Aminotransferase (AST, U/L), Alanine Aminotransferase (ALT, U/L), Sorbitol Dehydrogenase (SDH, U/L), Glucose (mg/dL), Alkaline Phosphatase (ALP, U/L), Gamma Glutamyl Transferase (GGT, U/L), Total Protein (g/dL), Albumin (g/dL), Globulin (g/dL), Albumin/Globulin Ratio (A/G Ratio), Blood Urea Nitrogen (BUN, mg/dL), Creatinine (mg/dL), BUN/Creatinine Ratio, Sodium (mEq/L), Potassium (mEq/L), Chloride (mEq/L), Calcium (mg/dL) and Phosphorus (mg/dL).
Nasal washes
In order to measure viral shedding, nasal washes were collected according to Table 1 by flushing the nasal cavities of ferrets with 1 mL of PBS (calcium and magnesium free; CMF) using a tuberculin syringe connected to a catheter. Nasal wash specimens from each animal were collected into individual sterile specimen cups. Specimens were transferred to cryovials by pipette and stored at ≤−70°C. Nasal wash samples were tested using a combination of real-time qPCR and TCID50 assays.
Pathology
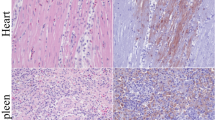
Complete gross necropsies were performed on any ferret found dead or euthanized prior to day 18 of the study. Histopathology was performed on any tissue found abnormal as well as target tissues which included but were not limited to lungs, liver, brain, spleen, mediastinal lymph nodes, nasal turbinates, heart and intestine.
Study endpoints
On day 18 surviving animals were euthanized following a terminal bleed. Blood was collected in EDTA tubes for real-time qPCR and hematology. Blood collected in SST tubes was used for clinical chemistry. Complete necropsies were performed following terminal bleeds on survivors of A/Vn challenge. Complete necropsies were also performed on a single representative (randomly selected) animal from each sham-infected control group.
Analytical and statistical plan
Survival data were analyzed using Fisher's exact test. Paired t-tests were used to test for significant changes from baseline and two-sample t-test were used to test for significant differences between flu-infected and sham-infected groups for weight, temperature, hematology and clinical chemistry parameters. The two-sample t-tests were all based on the change from baseline data (computed for each individual animal) to account for differences among the animals at baseline and a log-transformation was used if needed to make the assumption of normality more reasonable. One-sample t-tests were used to test for significant changes in nasal wash qPCR, nasal wash TCID50 and activity. For each group and time-point, qPCR was compared the assay limit of detection (LOD), TCID50 was compared to the assay limit of quantitation (LOQ) and activity was compared to zero.
The times until the first abnormal value were found for the temperature, activity, weight, nasal wash TCID50, nasal wash qPCR, clinical chemistry and hematology data. The definition of an abnormal value (significantly high or low) was different for each parameter. For nasal wash TCID50 a value greater than the LOQ was considered abnormal, for nasal wash qPCR a result greater than the LOD was considered abnormal and for activity a non-zero score was considered abnormal. For weight and temperature the baseline average and standard deviation was calculated for each individual animal. Weight was abnormal when an animal's weight fell two standard deviations below its baseline average and temperature was considered abnormal when an animal's temperature was two standard deviations above its baseline average. Analysis of variance models were fitted to each hematology and clinical chemistry parameter with effects for challenge (flu or sham) and time. A flu-infected animal was considered abnormal at a time-point if its value for a parameter was outside of the 95 percent confidence interval calculated for the sham-infected animals at the time-point.
The time from challenge until abnormal was calculated for each animal. The mean time to abnormal, 95 percent confidence intervals and the minimum and maximum values for all parameters were calculated within each challenge group and in flu-infected groups based on survival. Additionally, the proportion of animals that were abnormal was calculated for each flu-infected group and bar charts were produced. For each parameter, a two-sample t-test was used to test for significant differences between flu-infected survivors and flu-infected non-survivors.
All statistical analysis was performed using SAS® (version 9.1). Statistical comparisons are reported at the 0.05 level of significance.
References
WHO, Fact sheet on influenza. http://www.who.int/mediacentre/factsheets/fs211/en/index.html (Accessed September 11, 2012).
Taubenberger, J. K. & Morens, D. M. 1918 Influenza: the mother of all pandemics. Emerg. Infect. Dis. 12, 15–22 (2006).
WHO, Confirmed cases of avian influenza (H5N1). http://www.who.int/influenza/human_animal_interface/EN_GIP_20120412CumulativeNumberH5N1cases.pdf (Accessed September 11, 2012).
Belser, J. A., Katz, J. M. & Tumpey, T. M. The ferret as a model organism to study influenza A virus infection. Dis. Model Mech. 4, 575–579 (2011).
Zitzow, L. A. et al. Pathogenesis of avian influenza A (H5N1) viruses in ferrets. J. Virol. 76, 4420–4429 (2002).
Maines, T. R. et al. Avian influenza (H5N1) viruses isolated from humans in asia in 2004 exhibit increased virulence in mammals. J. Virol. 79, 11788–11800 (2005).
Govorkova, E. A. et al. Lethality to ferrets of H5N1 influenza viruses isolated from humans and poultry in 2004. J Virol. 79, 2191–2198 (2005).
Ellebedy, A. H. et al. Impact of prior seasonal influenza vaccination and infection on pandemic A(H1N1) influenza virus replication in ferrets. Vaccine. 29, 3335–3339 (2010).
Vela, E. M., Trodoff, K., Stark, G. V. & Bigger, J. B. Efficacy of a heterologous vaccine and adjuvant in ferrets challenged with A/Vietnam/1203/04. Influenza Other Respi. Viruses. (2011).
Boltz, D. A., Rehg, J. E., McClaren, J., Webster, R. G. & Govorkova, E. A. Oseltamivir prophylactic regeimens prevent H5N1 influenza morbidity and mortality in a ferret model. J Infect. Dis. 197, 1315–1323 (2008).
Maines, T. R. et al. Transmission and pathogenesis of swine-origin 2009 A(H1N1) influenza viruses in ferrets and mice. Science 325, 484–487 (2009).
Skeik, N. & Jabr, F. I. Influenza viruses and the evolution of avian influenza virus H5N1. International Journal of Infectious Disease 12, 233–238 (2007).
Beigel, J. H. et al. Avian influenza A (H5N1) infection in humans. New England Journal of Medicine 353, 1374–1385 (2005).
Gafar et al. Update on avian influenza A (H5N1) virus infection in humans. N. Engl. J. Med. 358, 261–273 (2008).
Tran, T. H. et al. Avian Influenza A (H5N1) in 10 patients in Vietnam. N. Engl. J. Med. 350, 1179–1188 (2004).
De Jong, M. et al. Fatal outcome of human influenza A (H5N1) is associated with high viral load and hypercytokinemia. Nat. Med. 12, 1203–1207 (2006).
Wang, X., Zhao, J., Tang, S., Ye, Z. & Hewlett, I. Viremia associated with fatal outcomes in ferrets infected with avian H5N1 influenza virus. PLoS ONE. 5, e12099 (2010).
Reuman, P. D., Keely, S. & Schiff, W. J. Assessment of signs of influenza illness in the ferret model. J. Virol. Meth 24, 27–34 (2004).
Acknowledgements
We would like to thank Michael Ethridge, Michelle Chafins, Ann Wasko, Phyllis Herr-Calomeni, Sherrie Wade, Brock Miniard and the rest of the scientific staff at the Battelle Biomedical Research Center for their time and effort with this project. This study was funded with funds from the National Institute of Allergy and Infectious Diseases (contract: HHSN266200400095I).
Author information
Authors and Affiliations
Contributions
James Long and Eric Vela wrote the majority of the manuscript. All authors reviewed the data and contributed to analysis of the data. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Long, J., Vela, E., Stark, G. et al. Early Indicators of Disease in Ferrets Infected with a High Dose of Avian Influenza H5N1. Sci Rep 2, 972 (2012). https://doi.org/10.1038/srep00972
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00972
This article is cited by
-
Clinical response to pandemic h1n1 influenza virus from a fatal and mild case in ferrets
Virology Journal (2015)
-
All Carbon-Based Photodetectors: An eminent integration of graphite quantum dots and two dimensional graphene
Scientific Reports (2013)
-
DBA/2 mouse as an animal model for anti-influenza drug efficacy evaluation
Journal of Microbiology (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.