Abstract
The cellular phenotype is described by a complex network of molecular interactions. Elucidating network properties that distinguish disease from the healthy cellular state is therefore of critical importance for gaining systems-level insights into disease mechanisms and ultimately for developing improved therapies. By integrating gene expression data with a protein interaction network we here demonstrate that cancer cells are characterised by an increase in network entropy. In addition, we formally demonstrate that gene expression differences between normal and cancer tissue are anticorrelated with local network entropy changes, thus providing a systemic link between gene expression changes at the nodes and their local correlation patterns. In particular, we find that genes which drive cell-proliferation in cancer cells and which often encode oncogenes are associated with reductions in network entropy. These findings may have potential implications for identifying novel drug targets.
Similar content being viewed by others
Introduction
Cancer cells differ from normal cells in terms of a complex landscape of genetic and epigenetic mutations (more generally aberrations), which at a systems-level cause a fundamental dynamic rewiring of the cellular interaction network, ultimately impairing normal cell physiology and allowing cells to acquire key cancer hallmarks such as uncontrolled cell-proliferation and evasion of apoptosis1. Although a number of studies have also made progress in identifying systems-level hallmarks underlying the cancer phenotype2,3, these remain largely unexplored and many more hallmarks remain to be elucidated. Elucidating these cancer-system principles represents a key challenge, not only for achieving a deeper understanding of cancer biology but also for identifying novel drug targets4. Given that cellular function is governed by a complex network of cellular interactions5, it seems natural to explore network properties which may help elucidate some of these cancer-system hallmarks.
In this work, we explore the role of network entropy in cancer, with the weights reflecting correlations in gene expression and normalized to define a random walk on a protein interaction network. While the concept of entropy has been studied in the cancer context previously6,7,8, the current study is significantly different in that here we consider a network entropy of a random walk on the graph, which was not considered in7,8. In our previous study we explored this same network entropy but only in the context of metastatic breast cancer6. A secondary motivation to focus on network entropy derives from a fluctuation theorem of dynamical systems theory9 which asserts that the macroscopic resilience of a system,  , is correlated to the level of uncertainty or entropy (disorder),
, is correlated to the level of uncertainty or entropy (disorder),  , of the underlying microsopic dynamical processes that take place within that system. More precisely, the theorem states that
, of the underlying microsopic dynamical processes that take place within that system. More precisely, the theorem states that  where
where  and
and  represent respectively the changes to the robustness and entropy of the system9,10. In11,12 this theorem was applied to protein interaction networks in yeast and C.elegans and it was demonstrated that those genes that contribute most to the network entropy are more likely to be essential genes for the organism. This important result demonstrates that network entropy can predict a gene property, i.e essentiality, which determines the system's robustness under knock-down of the respective gene.
represent respectively the changes to the robustness and entropy of the system9,10. In11,12 this theorem was applied to protein interaction networks in yeast and C.elegans and it was demonstrated that those genes that contribute most to the network entropy are more likely to be essential genes for the organism. This important result demonstrates that network entropy can predict a gene property, i.e essentiality, which determines the system's robustness under knock-down of the respective gene.
We point out that in the previous studies11,12, the stochastic matrix defining the dynamics on the network and hence the network entropy, was purely topological, i.e the stochastic matrix and entropy were completely specified by the underlying network topology. In our goal to study the role of network entropy in cancer it is key to compare to a normal reference, that is, cells of normal (healthy) physiology. Hence, in order to explore the role of network entropy in cancer, we use static gene expression data from representative samples of normal and cancer tissue to approximate a stochastic dynamics on a human protein interaction network. Thus, the dynamics we consider refers to the random walk generated by a stochastic matrix on the network and not to an underlying temporal dynamics, as time course data for individual cancer patients is not available. To clarify, the stochastic matrix on the network is specified by the gene expression data and therefore the dynamics is not entirely specified by the network topology. In fact, we assume that the network topology is unchanged between the normal and cancer phenotypes, but allow the dynamics, defining the weights in the network, to be dependent on the phenotype. Equivalently, we view the protein interaction network as providing only a backbone topological structure as to which interactions are allowed and use the phenotype-specific gene expression data (and specifically, the correlations in gene expression over the disease phenotype) to modulate and approximate the interaction probabilities. Using this perspective, cancer cells differ from normal cells due to differential weights on the same underlying network.
Therefore, our approach is based on two key concepts. First, the integration of gene expression data with protein interaction networks to yield integrated weighted networks, a methodological approach which has already proved fruitful in a variety of different applications within the cancer genomics field6,7,13,14,15,16,17,18,19,20,21,22,23,24,25. Second, we use the recent notion of “differential networks”, which attempts to better characterise disease phenotypes by studying the changes in the interaction patterns of these networks4,6,19,20,26,27, as opposed to merely analysing the changes in mean levels of some molecular quantity (e.g gene expression). As demonstrated by several studies6,19,20, differential networks can identify important gene modules implicated in cancer and also provide critical novel biological insights not obtainable using other approaches. This differential network strategy has recently received further impetus from studies of differential epistasis mapping in yeast, demonstrating that differential interactions may hold the key to understanding the systems-level responses of cells to exogenous and endogenous perturbations, including those present in cancer cells4,26.
Using network entropy defined locally for nodes in the network, we here demonstrate that cancer is characterized by an increase in network entropy. We next extend the notion of local entropy to a non-local/global one, i.e for extended subnetworks and find that non-local entropy measures are less discriminatory of the cancer phenotype. We also explore the relation between local differential entropy and differential expression and in the process, elucidate a novel cancer system hallmark. Finally, we discuss the meaning of our results in the context of the entropy-robustness theorem above and discuss the potential implications of our findings for devising novel cancer therapies with a view to future studies that will attempt to integrate drug sensitivity data with multi-dimensional (mutational, copy-number, epigenetic and transcriptomic) tumour profiles.
Results
We identified six expression data sets encompassing sufficient numbers of normal and cancer tissue samples and which passed our quality control criteria (Methods). The tissues profiled were bladder, lung, stomach, pancreas, cervix and liver. Integration of these expression data sets with our protein interaction network (PIN) (Methods) yielded sparse weighted networks of approximately 7500 nodes and 98500 edges. The average degree, median degree and diameter of these integrated networks were approximately 26, 8 and 12, respectively. An important assumption underlying any analysis on these integrated networks is that genes which are neighbors in the network are more likely to be correlated at the level of gene expression. While this has been shown for specific data sets (see e.g19), we verified that it also holds for the integrated mRNA-PIN networks considered here ( Fig. S1 ).
Increased local network entropy is a cancer system hallmark
We previously showed that primary breast cancers that metastasize exhibit an increased network entropy compared to breast cancers that do not spread6. The network entropy of a node i was defined by6

where pij defines a stochastic matrix on the graph and ki is the degree of gene i (see Methods).
Comparing distinct cancer phenotypes (e.g metastasizing cancers to non-metastasizing) to each other has the advantage that large sample collections are available, thus allowing for more reliable estimates of expression correlations. However, having identified suitable expression data sets encompassing relatively large and balanced numbers of normal and cancer samples, we here sought to determine if the network entropy also discriminates cancer from its respective normal tissue phenotype. We first compared the local entropies (equation 1) between normal and cancer, focusing on high-degree nodes (here, nodes with at least 10 neighbours) following the assumption that high degree nodes have higher relevance in cancer19. Performing this comparison across six different tissue types, using both unpaired and paired non-parametric statistics (to account for the degree and hence node dependence of differential entropy) clearly confirmed that cancer is characterised by an increased network entropy ( Fig. 1A , Table 1 ). Next, we sought to determine if this increased network entropy is also seen if all nodes are included in the analysis. The analogous analysis over all nodes of degree ≥ 2 (to define the local entropy we need a node to have at least two neighbours) confirmed that network entropy is increased in cancer ( Fig. S2 ), with the discriminatory power somewhat reduced but still highly significant ( Table 1 ).
(A) Boxplots of the local network entropies (y-axis, S) (equation 1)in cancer (C) and normal (N) tissue for all nodes with degree ≥ 10 (~ 3500 nodes) and across the six different tissue types. P-values are from a one-tailed unpaired Wilcoxon rank sum test. Network entropies have been normalised so that the maximum attainable value is 1. See Fig. S2 for the corresponding plot using all nodes with degree ≥ 2. (B) Boxplots of the z-statistics of differential entropy between cancer and normal tissue. Positive z-statistics indicate higher entropy in cancer. P-values from a one-tailed Wilcoxon rank sum test are given.
We also observed that the magnitude of differential entropy change was strongly anti-correlated to node degree ( Fig. 2A ). This dependence of network entropy and differential network entropy on the degree of the node was already explored by us previously and reflects an intrinsic bias which needs to be corrected for if meaningful rankings of genes are to be obtained6. In order to correct for this bias, we here devised a statistical framework based on the jackknife to derive z-statistics of differential entropy, which, by construction, would be degree-independent (Methods). Confirming this, we observed that absolute z-statistics did not exhibit a strong anti-correlation with degree and in fact were on the whole degree-independent ( Fig. 2B ). Supporting our previous result, we also observed that differential entropy z-statistics were significantly higher in cancer compared to normal tissue, independently of tissue type ( Fig. 1B ).
(A) Scatterplots of absolute differential network entropy changes between normal and cancer (y-axis) against log2(k) (x-axis) where k is the degree of the node, for each tissue type.(B) Scatterplots of the corresponding absolute differential entropy z-statistics (y-axis) against log2(k) (x-axis). In both cases, we provide the Spearman rank correlation coefficient (SCC).
Non-local network entropy is increased in cancer, albeit weaker than local network entropy
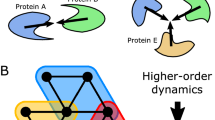
Next, we asked if the higher order network entropy, computed over paths of length larger than 1, are also discriminatory. To this end, we computed for the normal and cancer phenotypes, a higher-order network entropy

where  satisfies an approximate diffusion equation over the network allowing for paths of maximum length 2 (Methods). We point out that even when i and j are neighbors, that
satisfies an approximate diffusion equation over the network allowing for paths of maximum length 2 (Methods). We point out that even when i and j are neighbors, that  is not equal to pij, since we allow for alternative signaling paths (of maximum length 2) between genes i and j. Thus, this entropy also takes the well-known redundancy of signaling paths into account28.
is not equal to pij, since we allow for alternative signaling paths (of maximum length 2) between genes i and j. Thus, this entropy also takes the well-known redundancy of signaling paths into account28.
For S(2), we also observed a higher entropy in cancer compared to normal tissue across all tissue types, although this increase was statistically significant only for the four larger studies ( Fig. 3 ). We also computed higher order entropies up to paths of maximum length 5. However, as with S(2), higher order network entropies S(k), k ≥ 3 generally exhibited reduced discriminatory power, suggesting that the interesting changes associated with network entropy in cancer are localised to neighbors and nearest neighbors in the interaction network. This is not entirely surprising since we observed that correlations in gene expression dropped significantly beyond neighbours and second nearest neighbours ( Fig. S3 ).
z-statistics of differential non-local network entropy (x-axis) for the six different tissues (y-axis).
The network entropy considered here is the  measure (equation 2) which is defined for a stochastic diffusion matrix for maximum path lengths of order 2 (Methods). Positive z-statistics means higher entropy in cancer compared to normal. Green lines indicate the 95% confidence interval envelope and given P-values are from a normal null distribution centred at zero.
measure (equation 2) which is defined for a stochastic diffusion matrix for maximum path lengths of order 2 (Methods). Positive z-statistics means higher entropy in cancer compared to normal. Green lines indicate the 95% confidence interval envelope and given P-values are from a normal null distribution centred at zero.
Differential network entropy and differential expression are anti-correlated
We argued that if the observed changes in network entropy have a biological basis, that there should be a relationship between the changes in local entropy and gene expression. Specifically, genes which become inactivated in cancer generally exhibit lower expression and this should be reflected as an increased local entropy around these nodes. Conversely, we hypothesized that genes which become activated in cancer (i.e oncogenes) and which are thus more likely to exhibit higher expression in cancer, would be associated with a lower network entropy since the increased activity of oncogenes is normally associated with activation of specific downstream signal transduction pathways. This means that there is less uncertainty (i.e entropy) along which paths in the network the information flow proceeds. To test this hypothesis, we computed for each gene a regularized t-statistic29 that reflects the degree of differential expression between normal and cancer tissue.
Similarly, for each gene we used the previous jackknife procedure to obtain a z-statistic which is a statistical measure of the differential entropy change between the normal and cancer phenotype (Methods). Next, we selected those genes with significant changes in both differential expression and differential entropy (P < 0.05). Restricting to these genes, we first verified that differential entropy statistics were not correlated with degree or at least only marginally so ( Table 2 ). In contrast, differential entropy statistics exhibited a strong anti-correlation with differential expression independently of tissue type and these anti-correlations remained significant after adjustment for node degree ( Table 2 ). To assess the overall significance of a composite null hypothesis test of no association between differential expression and differential entropy, we used Fisher's combined probability test30 to obtain an overall P-value (P = 8e−27), which was highly significant ( Table 2 ). Confirming this analysis further, we observed that genes significantly overexpressed in cancer showed preferential reductions in network entropy compared to genes which were underexpressed and the associated odds ratios (OR) were statistically significant across all 6 tissue types ( Table 3 ). Once again, treating the 6 data sets as independent tests, Fisher's combined probability test confirmed the overall significance (P = 1e−11) of the 6 P-values in ( Table 3 ). Thus, the results in Tables 2 and 3 are consistent with each other, supporting the existence of another cancer system hallmark: that differential expression and differential network entropy are anticorrelated.
Cell-cycle/proliferation genes preferentially associate with a lower network entropy in cancer
Overexpression of cell-cycle and cell-proliferation genes is a key cancer hallmark with many of these genes representing also candidate drug targets1. Although we have seen that differential entropy changes anti-correlate with differential expression, it is important to check if (1) cell-cycle/proliferation genes are preferentially associated with a reduced network entropy and (2) whether the anti-correlation between differential entropy and differential expression is driven entirely by cell-cycle genes. To address the first point, we performed a gene set enrichment analysis (GSEA) using the Molecular Signatures Database (MSigDB31) on the top ranked genes, ranked according to the statistics of differential network entropy (separately for increased and reduced entropy). The GSEA analysis showed that genes implicated in the cell-cycle were indeed strongly enriched among genes exhibiting lower network entropy in cancer, but not so among genes exhibiting increases in network entropy ( Table 4 ).
To address the second point, we repeated the correlation analysis between differential entropy and differential expression, but removing cell-cycle genes prior to the analysis. Importantly, we still observed the anti-correlation between differential entropy and differential expression in 5 of the 6 tissue types ( Table S1 ), indicating that this anticorrelation is a general systemic feature.
It could be argued that since tumour expression profiles analyzed here are from the bulk, meaning that the measured expression profiles represent an average over epithelial tumour cells and non-epithelial stromal cells (e.g immune cells), that entropy changes are entirely confounded by changes in the tumour-stromal cell composition ratio. Therefore, it is important to point out here that the enrichment of cell-cycle/proliferation genes among those showing the largest reductions in network entropy, indicates that these differential entropy changes reflect underlying changes in the epithelial tumour cell population and not changes in the tumour-stromal cell ratio. In other words, the fact that entropy changes can retrieve known tumour cell biology (i.e increased proliferation of tumour cells) shows that interesting tumour cell biology can be extracted from the network entropy.
Discussion
In this work we have constructed a weighted network entropy and have shown that it is increased in cancer compared to normal tissue. Both local and non-local versions of the network entropy were considered, with the local entropy exhibiting the more significant increases. This partly reflects the local nature of expression correlations in the protein interaction network with correlations dropping significantly beyond neighbours and second nearest neighbours. It is of importance to discuss (i) what may cause cancer cells to exhibit the observed increase in network entropy and (ii) what it may mean for the cancer phenotype itself.
Concerning the first question, one would expect genes that become inactivated in cancer to represent foci of increased network entropy since the inactivation compromises its biological function: at the level of mRNA expression this would manifest itself as reduced expression correlations with its interacting neighbors, but more generally as an increased uncertainty as to which neighbors it may interact with. Conversely, for a gene that is overactivated in cancer its biological function is enhanced, thus confering the cell a selective advantage, which for oncogenes manifests itself as an increased flux of the associated oncogenic pathway. In terms of the local network entropy this increased flux along a particular pathway in the network corresponds to a reduced uncertainty (i.e less network entropy) along which path the information is transferred. In line with these biological expectations we did observe that genes overexpressed in cancer were significantly more likely to exhibit reductions in network entropy than underexpressed genes. Thus, the fact that cancers were characterised globally by an increased network entropy points towards a higher frequency of inactivating over activating alterations in cancer. Intuitively, this makes sense since a random mutation/alteration is more likely to inactivate than activate a gene and indeed this would be in agreement with recent reports suggesting that most genetic alterations are inactivating and affect tumour suppressors32. We should point out that to formally demonstrate that the increased network entropy is associated with an increased frequency of inactivating alterations (mutations, losses and deletions, promoter DNA methylation) in the tumours analysed here is not possible as matched mutational, copy-number and DNA methylation information is not available for these specific tumours. However, it will be interesting to explore this in the context of the matched multi-dimensional cancer genomic data from the The Cancer Genome Atlas (TCGA)33.
Concerning the second question posed above, our observation that differential network entropy and differential expression are anti-correlated is strongly suggestive of an underlying entropy robustness theorem,  . In fact, we have seen that genes driving cell-proliferation, which are known to be overexpressed in cancer34 and which often encode oncogenes1, were preferentially associated with significant reductions in network entropy. Now, it is well known that cancer cells exhibit the phenomenon of oncogene addiction, whereby they become overly dependent and reliant on activated oncogenes1. Oncogene addiction has been exploited therapeutically: indeed, in cases where the oncogene is druggable, targeting of the oncogene has proved to be an effective drug therapy strategy1. Thus, oncogenes have the paradoxical effect of making cancer cells less robust to targeted intervention. Hence, our observation that overexpressed genes and oncogenes in particular, are associated with reductions in network entropy is consistent with an entropy-robustness theorem like equation 1. Similarly, the observed increased network entropy in cancer could underpin the intrinsic robustness of cancer cells to general endogenous and exogeneous perturbations, including those caused by environmental stresses (e.g hypoxia) within the tumour microenvironment1.
. In fact, we have seen that genes driving cell-proliferation, which are known to be overexpressed in cancer34 and which often encode oncogenes1, were preferentially associated with significant reductions in network entropy. Now, it is well known that cancer cells exhibit the phenomenon of oncogene addiction, whereby they become overly dependent and reliant on activated oncogenes1. Oncogene addiction has been exploited therapeutically: indeed, in cases where the oncogene is druggable, targeting of the oncogene has proved to be an effective drug therapy strategy1. Thus, oncogenes have the paradoxical effect of making cancer cells less robust to targeted intervention. Hence, our observation that overexpressed genes and oncogenes in particular, are associated with reductions in network entropy is consistent with an entropy-robustness theorem like equation 1. Similarly, the observed increased network entropy in cancer could underpin the intrinsic robustness of cancer cells to general endogenous and exogeneous perturbations, including those caused by environmental stresses (e.g hypoxia) within the tumour microenvironment1.
It follows from the above argument that network entropy may be used to identify novel drug targets. As a specific example, we observed that the kinase AURKB exhibited the largest reductions in network entropy in bladder cancer ( Table S2 ). Importantly, AURKA, which has already a well established oncogenic role in bladder cancer (see e.g35) was also highly ranked ( Table S2 ). Thus, our analysis suggests that the closely related kinase, AURKB, which has already been implicated as an oncogene and potential drug target in other cancers36,37,38, may also play an equally important role in the pathogenesis of bladder cancer. In fact, a very recent study further showed that AURKB phosphorylates and instigates degradation of P5339, a key tumour suppressor in cancer. Given that AURKB is also druggable (by the drug rebamipide)40, this kinase therefore represents an attractive drug target for those bladder cancers that overexpress it. In cases where the oncogene is not directly druggable, we speculate that differential network entropy may be used to identify neighboring druggable targets that also exhibit significant reductions in network entropy. This novel computational strategy could therefore guide non-oncogene addiction based therapeutic strategies that aim to select drug targets within the same oncogenic pathway41,42. Moreover, it has become clear that mutational and copy-number status alone or in combination with gene expression levels are not highly predictive of drug response43,44, hence there is an urgent need for improved in-silico predictors of drug sensitivity. We leave these open and exciting questions for a future bioinformatic study that will analyze matched genomic (mutational, copy-number), epigenomic (DNA methylation), functional (e.g mRNA expression) and drug sensitivity data for large panels of drugs and cancer cell-lines43,44.
While network entropy provided a good discrimination between normal and cancer tissue, it is clear that it does not outperform raw gene expression levels, which offer significantly higher classification accuracies34. Other network measures may also provide equally good discriminators of the cancer phenotype as network entropy. Indeed, the average of the absolute correlations over neighbours of a given node provides an equally good discriminator ( Fig. S4 ), indicating that the loss of local connectivity is a key cancer characteristic. However, the loss of local connectivity (i.e reduced absolute correlations) does correspond to an increase in local network entropy. Therefore, network entropy may provide, through an entropy-robustness theorem (equation 1) a more meaningful framework in which to interpret and understand the systemic changes in gene expression between normal and cancer tissue.
In summary, in this work we have explored the notion of network entropy in cancer and have used it to elucidate two cancer system hallmarks: (1) that network entropy is increased in cancer relative to the normal phenotype and (2) that differential network entropy is anti-correlated with differential expression. Therefore, this work further supports the view that the cell's network entropy and robustness are correlated. Further investigation of the statistical mechanical principles characterising cancer gene networks is warranted as this may help rationalize the choice of drug targets.
Methods
The protein interaction network (PIN)
We downloaded the complete human protein interaction network from Pathway Commons (www.pathwaycommons.org) (Jan.2011)45, which brings together protein interactions from several distinct sources. We then built a reduced protein interaction network from integrating the following sources: the Human Protein Reference Database (HPRD)46, the National Cancer Institute Nature Pathway Interaction Database (NCI-PID) (pid.nci.nih.gov), the Interactome (Intact) http://www.ebi.ac.uk/intact/ and the Molecular Interaction Database (MINT) http://mint.bio.uniroma2.it/mint/. Protein interactions in this network include physical stable interactions such as those defining protein complexes, as well as transient interactions such as posttranslational modifications and enzymatic reactions found in signal transduction pathways, including 20 highly curated immune and cancer signaling pathways from NetPath (www.netpath.org)47. We focused on non-redundant interactions, only included nodes with an Entrez gene ID annotation and focused on the maximally conntected component, resulting in a connected network of 10,720 nodes (unique Entrez IDs) and 152,889 documented interactions. In what follows we refer to this network as the “PIN”.
Normal and cancer tissue gene expression data sets
We searched Oncomine34 for studies which (i) had profiled reasonable numbers of cancer and normal tissue samples (at least ~ 25 of each type) and (ii) which had been profiled on an Affymetrix platform. In order to reliably estimate covariance of two genes across a set of samples, at least ~ 25 samples are needed. The second criterion reflects the desire to conduct the study on a common platform and Affymetrix arrays are the most widely used. Using the same platform across studies ensured that the integrated mRNA-PIN networks were of similar size. In all cases, the intra-array normalised data was downloaded from GEO (www.ncbi.nlm.nih.gov/geo/), quantile normalized and subsequently probes mapping to the same Entrez gene ID were averaged. We then subjected each study that passed these criteria through a quality control step, which involved a Principal Component Analysis (PCA) to check that (iii) the dominant component of variation correlated with cancer/normal status. If not, this indicated to us a more pronounced source of non-biological variation, which would confound our downstream analysis. There were six studies satisfying all three criteria and the tissues profiled included bladder (48 normals and 81 cancers)48, lung (49 normals and 58 cancers)49, gastric (31 normals and 38 cancers)50, pancreas (39 normals and cancers)51, cervix (24 normals and 33 cancers52 and liver (23 normals and 35 cancers)53.
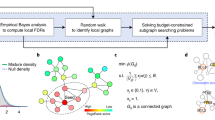
Integrated PIN-mRNA expression networks and the stochastic matrix
For a given cellular phenotype (i.e. cancer or normal), we build an integrated mRNA-PIN using the same procedure as described in6. Briefly, edge weights in the PIN are defined by a stochastic matrix pij,

with  , where
, where  denotes the neighbors of gene i in the PIN and where
denotes the neighbors of gene i in the PIN and where  denotes the transformed Pearson correlation coefficient Cij of gene expression between genes i and j across the samples belonging to the given phenotype. This definition of wij reflects our desire to treat correlations and anti-correlations differently. We also note that we enforce pij = 0 whenever (i, j) is not an edge in the PIN. Thus, the integrated mRNA-PINs with the edge weights as defined by pij, can be viewed as approximate models of signal transduction flow (as measured by positive gene-gene correlations in expression) subject to the structural constraint of the PIN. Applying this procedure to the two phenotypes yields two integrated PIN-mRNA networks, one for the cancer phenotype with stochastic matrix
denotes the transformed Pearson correlation coefficient Cij of gene expression between genes i and j across the samples belonging to the given phenotype. This definition of wij reflects our desire to treat correlations and anti-correlations differently. We also note that we enforce pij = 0 whenever (i, j) is not an edge in the PIN. Thus, the integrated mRNA-PINs with the edge weights as defined by pij, can be viewed as approximate models of signal transduction flow (as measured by positive gene-gene correlations in expression) subject to the structural constraint of the PIN. Applying this procedure to the two phenotypes yields two integrated PIN-mRNA networks, one for the cancer phenotype with stochastic matrix  and one for the normal phenotype with stochastic matrix
and one for the normal phenotype with stochastic matrix  . It is important to point out that the construction of our stochastic matrix means that the topological degrees of each node remain unchanged between the normal and cancer phenotypes: it is only the weights specifying the random walk on the network which differ between the two phenotypes.
. It is important to point out that the construction of our stochastic matrix means that the topological degrees of each node remain unchanged between the normal and cancer phenotypes: it is only the weights specifying the random walk on the network which differ between the two phenotypes.
It is important to stress that we have approximated signal transduction flux on the PIN by positive correlations in expression between interacting genes. This is obviously a crude approximation and therefore a limitation of this study, however, until other types of matched molecular data (e.g protein expression, phosphorylation and other post-translational modification states) become available on a genome-wide basis, we are restricted to the use of only gene expression data. Some further justification for the use of gene expression correlations to approximate signaling flux over the network will be provided by careful comparison of the local correlations to those which are non-local.
A heat kernel stochastic matrix
It is clear that the stochastic matrix pij above defines a (biased) random walk on the network  . One may thus compute an information (or probability) flux between any two nodes i and j in
. One may thus compute an information (or probability) flux between any two nodes i and j in  54. In fact, it is clear that the probability flux of moving from i to j over a path of length L is given by (pL)ij. It follows that the total probability flux Eij between i and j is given by
54. In fact, it is clear that the probability flux of moving from i to j over a path of length L is given by (pL)ij. It follows that the total probability flux Eij between i and j is given by

where γ is a normalisation factor and where we have introduced a set of arbitrary weights αL to allow variable contributions for paths of different lengths. One possibility is to suppress paths of longer lengths using αL = 1/L!, which also guarantees convergence of the infinite series54. Formally, defining αL = tL/L!, we obtain the stochastic matrix

where we have introduced a “temperature” parameter t55. This stochastic matrix is a modified version of the heat-kernel stochastic matrix55 and satisfies

where we have suppressed matrix indices and where I denotes the identity matrix. Since pij, Kij(t) ≤ 1 for all i, j, t, it follows that for sufficiently large temperatures (t ≥ 1), K(t) approximates a solution of the heat-diffusion equation55

Thus, the choice α = tL/L! leads to a natural interpretation in terms of a discrete approximate diffusion process on a graph56. This construction is therefore closely related to the heat kernel PageRank algorithm55,56,57.
The network entropy
Given the matrix Kij, let Q denote the number of non-zero Kij, i.e Q = Σij I(Kij > 0) where I is here the indicator function. We then define the network entropy as

where we have rescaled Kij(t) by 1/n in order to ensure that Σij Kij(t) = 1. Note that the entropy defined above can be thought of as a non-equilibrium entropy, since the stationary distribution πi of Kij, defined by πiKij = πj , was not included. Our choice to consider this non-equilibrium version is motivated by our desire to avoid biasing the entropic contribution of each node to its topological properties (e.g degree).
Suppose now that we consider diffusion/flux over paths of maximum length 1. Then, this leads to Kij = pij/n where n is the number of nodes in  (we have set t = 1 for convenience). This leads to the expression
(we have set t = 1 for convenience). This leads to the expression

In the above expression, Si is the local entropy of node i6,56,

where ki is the degree of node i and the normalisation factor ensures that the maximum attainable entropy is equal to 1, independent of the degree of the node. We note that  is in effect a network average of these local network entropies, but is distinct from the global entropy defined in equation 8.
is in effect a network average of these local network entropies, but is distinct from the global entropy defined in equation 8.
Next, we can consider flux over paths up to length two, in which case

and the corresponding entropy,

In principle, we can estimate the entropy S(h) for paths of arbitrary order h. In this case,

In this work we compute network entropies up to moments of order 5 using the R-package expm. Not going beyond h = 5 is justified for two reasons: (i) the most interesting behaviour is found for h ≤ 3, (ii) the computational cost for h = 5 is considerable, for instance, estimation of network entropy and associated sampling variance estimates for a typical data set of 30 samples and ~ 7500 nodes at h = 5 takes at least ~ 20 hours on a high-performance quad processor workstation.
Sampling variance using the jackknife
To estimate the statistical significance of observed differences in entropy between two phenotypes, we decided to use the jackknife procedure58. Briefly, the jackknife procedure removes one sample at a time from the given phenotype and recomputes the desired quantity S (here entropy). Thus, if there are n samples in the given phenotype one obtains n jackknife estimates ( ). A jackknife estimate for the mean Sµ and variance SV of S is then obtained as
). A jackknife estimate for the mean Sµ and variance SV of S is then obtained as


where  is the estimate using all n samples and
is the estimate using all n samples and  . Thus, for two phenotypes “N” and “C”, we compute the difference
. Thus, for two phenotypes “N” and “C”, we compute the difference  and obtain a z-statistic
and obtain a z-statistic

where  .
.
This jackknife procedure can be applied to the network entropy defined over the network or for each node individually. Note that in the case where we obtain z-statistics for each gene/node, the genes can then be ranked according to the significance of this z-statistic. We also note that by construction the z-statistic should be independent of the degree of the node. In fact, while both the differential entropy ΔSJ as well as the standard deviation estimate σJ will demonstrate the same degree-dependence, the ratio given by the z-statistic z = ΔSJ (k)/σJ(k) should be degree independent. We demonstrate this empirically across the six different data sets considered here.
References
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Cui, Q. et al. A map of human cancer signaling. Mol Syst Biol 3, 152 (2007).
Dutkowski, J. & Ideker, T. Protein networks as logic functions in development and cancer. PLoS Comput Biol 7, e1002180 (2011).
Califano, A. Rewiring makes the difference. Mol Syst Biol 7, 463 (2011).
Barabasi, A. L. & Oltvai, Z. N. Network biology: understanding the cell's functional organization. Nat Rev Genet 5, 101–113 (2004).
Teschendorff, A. E. & Severini, S. Increased entropy of signal transduction in the cancer metastasis phenotype. BMC Syst Biol 4, 104 (2010).
Schramm, G., Nandakumar, K. & Konig, R. Regulation patterns in signaling networks of cancer. BMC Syst Biol 4, 162 (2010).
van Wieringen, W. N. & van der Vaart, A. W. Statistical analysis of the cancer cell's molecular entropy using high-throughput data. Bioinformatics 27, 556–563 (2011).
Demetrius, L., Grundlach, V. M. & Ochs, G. Complexity and demographic stability in population models. Theo Pop Biol 65, 211–225 (2004).
Demetrius, L. & Manke, T. Robustness and network evolution-an entropic principle. Physica A 346, 682–696 (2005).
Manke, T., Demetrius, L. & Vingron, M. Lethality and entropy of protein interaction networks. Genome Inform 16, 159–163 (2005).
Manke, T., Demetrius, L. & Vingron, M. An entropic characterization of protein interaction networks and cellular robustness. J R Soc Interface 3, 843–850 (2006).
Tuck, D. P., Kluger, H. M. & Kluger, Y. Characterizing disease states from topological properties of transcriptional regulatory networks. BMC Bioinformatics 7, 236 (2006).
Pujana, M. A. et al. Network modeling links breast cancer susceptibility and centrosome dysfunction. Nat Genet 39, 1338–1349 (2007).
Platzer, A., Perco, P., Lukas, A. & Mayer, B. Characterization of protein-interaction networks in tumors. BMC Bioinformatics 8, 224 (2007).
Ulitsky, I. & Shamir, R. Identification of functional modules using network topology and high-throughput data. BMC Syst Biol 1, 8 (2007).
Chuang, H. Y., Lee, E., Liu, Y. T., Lee, D. & Ideker, T. Network-based classification of breast cancer metastasis. Mol Syst Biol 3, 140 (2007).
Milanesi, L., Romano, P., Castellani, G., Remondini, D. & Li, P. Trends in modeling biomedical complex systems. BMC Bioinformatics 10, I1 (2009).
Taylor, I. W. et al. Dynamic modularity in protein interaction networks predicts breast cancer outcome. Nat Biotechnol 27, 199–204 (2009).
Hudson, N. J., Reverter, A. & Dalrymple, B. P. A differential wiring analysis of expression data correctly identifies the gene containing the causal mutation. PLoS Comput Biol 5, e1000382 (2009).
Nibbe, R. K., Koyutrk, M. & Chance, M. R. An integrative -omics approach to identify functional sub-networks in human colorectal cancer. PLoS Comput Biol 6, e1000639 (2010).
Yao, C. et al. Multi-level reproducibility of signature hubs in human interactome for breast cancer metastasis. BMC Syst Biol 4, 151 (2010).
Komurov, K., White, M. A. & Ram, P. T. Use of data-biased random walks on graphs for the retrieval of context-specific networks from genomic data. PLoS Comput Biol 6 (2010).
Komurov, K. & Ram, P. T. Patterns of human gene expression variance show strong associations with signaling network hierarchy. BMC Syst Biol 4, 154 (2010).
Vazquez, A. Protein interaction networks. in: Alzate O, editor. Neuroproteomics, Chapter 8, CRC Press, 2010.
Bandyopadhyay, S. et al. Rewiring of genetic networks in response to dna damage. Science 330, 1385–1389 (2010).
Ideker, T. & Krogan, N. J. Differential network biology. Mol Syst Biol 8, 565 (2012).
Tieri, P. et al. Network, degeneracy and bow tie integrating paradigms and architectures to grasp the complexity of the immune system. Theor Biol Med Model 7, 32 (2010).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol 3, Article3 (2004).
Borenstein, M., Hedges, L., Higgins, J. & Rothstein, H. Introduction to Meta-Analysis (Wiley, 2009).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102, 15545–15550 (2005).
Wood, L. D. et al. The genomic landscapes of human breast and colorectal cancers. Science 318, 1108–1113 (2007).
e. t. w. o. r. k. The Cancer Genome Atlas Research Network. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Rhodes, D. R. et al. Large-scale meta-analysis of cancer microarray data identifies common transcriptional profiles of neoplastic transformation and progression. Proc Natl Acad Sci U S A 101, 9309–9314 (2004).
Park, H. S. et al. Quantitation of aurora kinase a gene copy number in urine sediments and bladder cancer detection. J Natl Cancer Inst 100, 1401–1411 (2008).
Lens, S. M., Voest, E. E. & Medema, R. H. Shared and separate functions of polo-like kinases and aurora kinases in cancer. Nat Rev Cancer 10, 825–841 (2010).
Lucena-Araujo, A. R. et al. High expression of aurka and aurkb is associated with unfavorable cytogenetic abnormalities and high white blood cell count in patients with acute myeloid leukemia. Leuk Res 35, 260–264 (2011).
Morozova, O. et al. System-level analysis of neuroblastoma tumor-initiating cells implicates aurkb as a novel drug target for neuroblastoma. Clin Cancer Res 16, 4572–4582 (2010).
Gully, C. P. et al. Aurora b kinase phosphorylates and instigates degradation of p53. Proc Natl Acad Sci U S A 109, E1513–E1522 (2012).
Ahmed, J. et al. Cancerresource: a comprehensive database of cancer-relevant proteins and compound interactions supported by experimental knowledge. Nucleic Acids Res 39, D960–D967 (2011).
Luo, J. et al. A genome-wide rnai screen identifies multiple synthetic lethal interactions with the ras oncogene. Cell 137, 835–848 (2009).
Luo, J., Solimini, N. L. & Elledge, S. J. Principles of cancer therapy: oncogene and nononcogene addiction. Cell 136, 823–837 (2009).
Garnett, M. J. et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483, 570–575 (2012).
Barretina, J. et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Cerami, E. G. et al. Pathway commons, a web resource for biological pathway data. Nucleic Acids Res 39, D685–D690 (2011).
Prasad, T. S., Kandasamy, K. & Pandey, A. Human protein reference database and human proteinpedia as discovery tools for systems biology. Methods Mol Biol 577, 67–79 (2009).
Kandasamy, K. et al. Netpath: a public resource of curated signal transduction pathways. Genome Biol 11, R3 (2010).
Sanchez-Carbayo, M., Socci, N. D., Lozano, J., Saint, F. & Cordon-Cardo, C. Defining molecular profiles of poor outcome in patients with invasive bladder cancer using oligonucleotide microarrays. J Clin Oncol 24, 778–789 (2006).
Landi, M. T. et al. Gene expression signature of cigarette smoking and its role in lung adenocarcinoma development and survival. PLoS One 3, e1651 (2008).
D'Errico, M. et al. Genome-wide expression profile of sporadic gastric cancers with microsatellite instability. Eur J Cancer 45, 461–469 (2009).
Badea, L., Herlea, V., Dima, S. O., Dumitrascu, T. & Popescu, I. Combined gene expression analysis of whole-tissue and microdissected pancreatic ductal adenocarcinoma identifies genes specifically overexpressed in tumor epithelia. Hepatogastroenterology 55, 2016–2027 (2008).
Scotto, L. et al. Identification of copy number gain and overexpressed genes on chromosome arm 20q by an integrative genomic approach in cervical cancer: potential role in progression. Genes Chromosomes Cancer 47, 755–765 (2008).
Wurmbach, E. et al. Genome-wide molecular profiles of hcv-induced dysplasia and hepatocellular carcinoma. Hepatology 45, 938–947 (2007).
Estrada, E. & Rodriguez-Velazquez, J. A. Subgraph centrality in complex networks. Phys Rev E 71 (2005).
Chung, F. The heat kernel as the pagerank of a graph. PNAS 104, 19735–19740 (2007).
Barrat, A., Barthelemy, M. & Vespignani, A. Dynamical Processes on Complex Networks (CUP, 2008).
Brin, S. & Page, L. The anatomy of a large-scale hypertextual web search engine. Comput Networks and ISDN Systems 30, 107–117 (1998).
Wu, C. F. J. Jackknife, bootstrap and other resampling methods in regression analysis. In The Annals of Statistics, vol. 14, 1261–1295 (1986).
Acknowledgements
JW is supported by a CoMPLEX PhD studentship. SS is supported by the Royal Society. AET is supported by a Heller Research Fellowship.
Author information
Authors and Affiliations
Contributions
JW and AET performed statistical analyses and devised the study. GB and SS contributed ideas. JW and AET wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information File
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
West, J., Bianconi, G., Severini, S. et al. Differential network entropy reveals cancer system hallmarks. Sci Rep 2, 802 (2012). https://doi.org/10.1038/srep00802
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00802
This article is cited by
-
Network-level analysis of ageing and its relationship with diseases and tissue regeneration in the mouse liver
Scientific Reports (2023)
-
The maximum entropy principle for compositional data
BMC Bioinformatics (2022)
-
Bakry–Émery Ricci Curvature Bounds for Doubly Warped Products of Weighted Spaces
The Journal of Geometric Analysis (2022)
-
Understanding the role of potential pathways and its components including hypoxia and immune system in case of oral cancer
Scientific Reports (2021)
-
Novel gene signatures for stage classification of the squamous cell carcinoma of the lung
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.