Abstract
Oriented immobilization of sensing molecules on solid phases is an important issue in biosensing. In case of immunosensors, it is essential to scrutinize not only the direction and shape of immunoglobulin G (IgG) in solution but also the real-time movement of IgGs, which cannot be achieved by conventional techniques. Recently, we developed bio-nanocapsules (BNCs) displaying a tandem form of the IgG Fc-binding Z domain derived from Staphylococcus aureus protein A (ZZ-BNC) to enhance the sensitivity and antigen-binding capacity of IgG via oriented-immobilization. Here, we used high-speed atomic force microscopy (HS-AFM) to reveal the fine surface structure of ZZ-BNC and observe the movement of mouse IgG3 molecules tethered onto ZZ-BNC in solution. ZZ-BNC was shown to act as a scaffold for oriented immobilization of IgG, enabling its Fv regions to undergo rotational Brownian motion. Thus, HS-AFM could decipher real-time movement of sensing molecules on biosensors at the single molecule level.
Similar content being viewed by others
Introduction
Due to the high specificity and affinity of biological molecules (e.g. immunoglobulin G (IgG), ligands, receptors, aptamers, sugar chains, lectins), biosensors are expected to be a promising technology for sensing various biological materials. The sensor surface (e.g., glass or metal) is commonly functionalized with chemical crosslinking or self-assembled monolayers (SAMs)1. Especially, long-chain (number of methylene groups n > 10) alkanethiols assemble in a crystalline-like way and can introduce uniform functional groups onto gold surface. While biological molecules were subsequently immobilized on the sensor surface, the control over the orientation of these molecules (i.e., oriented-immobilization of IgGs)2 has not so far been fully achieved. In case of immunosensors, the crosslinking molecules should neither increase steric hindrance around the antigen-recognition Fv regions nor reduce the antigen-recognition activity of IgGs and, furthermore, should align the Fv regions for efficient antigen-recognition. Specific sites within IgG molecules have been used to achieve oriented immobilization, such as the Fc region via an Fc-binding protein A or G3,4, aldehyde groups introduced into the carbohydrate moiety of the CH2 domain via an hydrazide-containing crosslinker5 and the thiol group of monovalent Fab' fragments via a thiol-containing solid phase6. However, the orientation of protein A or G itself cannot be fully controlled on the solid phase. Chemical and enzymatic treatments may affect the antigen-recognition activity of IgGs. These situations led us to develop rigid and self-assembled scaffolds for aligning IgGs at the nanoscale level, without modification.
We previously developed ZZ-BNCs of ~30 nm diameter by expressing the gene encoding hepatitis B virus (HBV) surface antigen (HBsAg) L protein with a tandem form of Fc-binding Z domain (Fig. 1a)7 in yeast8. ZZ-BNC contains about 120 molecules of ZZ-L protein (N-terminally ZZ-fused L protein) embedded in a liposome and has ability for capturing ~60 mouse IgG molecules, as well as displaying all the IgG Fv regions outwardly9. Furthermore, ZZ-BNCs can enhance the sensitivity of immunosensors9 and immunoassays10,11 not only through the oriented immobilization of antibodies but also the clustering of antibodies and labelling molecules. Thus, ZZ-BNC is a promising scaffold for a variety of conventional immunosensors and immunoassays.
HS-AFM analyses of ZZ-BNC in solution.
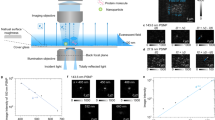
(a) Structure of ZZ-BNC. One ZZ-BNC particle consists of about 120 ZZ-L proteins embedded in a lipid bilayer. The dimeric forms of ZZ-L proteins are described in this study. Mouse IgG3 could interact with the ZZ domain through domain B between CH2 and CH3 of the Fc subunit32. (b) (upper), (c) HS-AFM images of ZZ-BNCs in 2D and 3D, respectively. Bars, 20 nm. The surface morphology of ZZ-BNC is shown in (b) (bottom). (d) Single-particle reconstructions of ZZ-BNC by EMAN software, line 1; average images, line 2; Fourier transformed images, line 3; binarized images. (e) SDS-PAGE analyses of ZZ-BNC reduced with 0–1,000 mM 2ME (0.5 μg protein/lane), followed by silver staining.
To evaluate immunosensors, it is necessary to elucidate not only the direction and shape of IgGs on the solid phase in solution but also the real-time movement of IgGs. However, conventional techniques cannot fully analyze the real-time movement of IgGs12,13,14. Particularly, it is difficult for AFM to distinguish IgGs from surrounding molecules using these static observations (snapshots). Recently, HS-AFM (high-speed atomic force microscopy) equipped with a highly sensitive, ultra-fast cantilever and efficient AFM scanning capabilities, have demonstrated biomolecule dynamics in solution15, encouraging us to analyze the behaviour of IgGs in solution for rigorous evaluation of immunosensors. Furthermore, the surface analysis using HS-AFM could contribute to the refinement not only of immunosensors but also of a variety of biosensors.
Results
Elucidation of fine surface structure of ZZ-BNC in solution
When analyzed a topological image of ZZ-BNCs on a mica surface by HS-AFM, they showed nanoparticles with rough surfaces (51.7 ± 4.8 nm in diameter; 17.6 ± 1.0 nm in height (mean ± SD, n = 19)) (Fig. 1b and c). The volume and surface area of an average ZZ-BNC were calculated to be 21,300 nm3 and 5,170 nm2, respectively, corresponding to those of a spherical ~40.6-nm particle. Because this value agreed well with the diameter obtained by dynamic light scattering (45.4 ± 2.2 nm), ZZ-BNCs were considered to adsorb onto mica surface without disrupting their capsule structure. Semi-automated, single-particle reconstructions using the EMAN software16 (10 particles from 48 frames) revealed protrusions on the surface of ZZ-BNC (Fig. 1d). The number and area of protrusions were estimated by ImageJ software to be 27 ± 3 and 28.9 ± 10.1 nm2, respectively. The diameter was 6.1 ± 3.6 nm. The distance between adjacent protrusions (centre to centre) was 8.2 ± 1.7 nm, which agrees well with the space (~6 nm) between two HBsAg proteins on the HBV virion17. One ZZ-BNC was estimated to possess 54 protrusions, while reported to contain about 120 molecules of ZZ-L protein9. Meanwhile, the HBsAg protein of the HBV virion18, as well as the HBsAg L protein of BNC8, is a transmembrane protein containing three membrane-spanning domains as dimeric19 and multimeric8 forms. ZZ-BNCs were then reduced with 0–1,000 mM 2-mercaptoethanol (2ME), separated by SDS-PAGE and stained with silver (Fig. 1e). By increasing the concentration of 2ME, two major bands of 96 and >260 kDa were gradually shifted to 96 and 48 kDa and finally converged to 48 kDa, strongly suggesting that nascent ZZ-L proteins form a dimer by intramolecular disulfide bonds which then form multimers by intermolecular disulfide bonds, as previously proposed for recombinant mammalian cell-derived HBsAg L protein20. Taken together, each protrusion on ZZ-BNCs may be a ZZ-L dimer containing six membrane-spanning domains, which is corroborated by the fact that the diameter of the protrusions (6.1 ± 3.6 nm) is consistent with that of seven membrane-spanning rhodopsin (~4.3 nm)21.
Observation of mouse IgG3 in solution
We next observed mouse IgG3 molecules in solution by HS-AFM (Fig. 2a). Conventional AFM observation of IgG molecules on a mica surface in air have shown that the hinge region of IgG is flexible, allowing two Fab subunits to rotate about their axes to present the side to the mica surface, with deposition of the Fc subunit on the opposite side in two forms (extended and folded)22. Assuming that the vertical length of Fab subunits is consistent amongst IgG forms, each IgG was identified as a Y-shaped structure including three subunits (two Fab and one Fc) in extended or folded form. By measuring the size of the extended form of IgG (n = 3), the distance between the two Fab subunits was calculated at 26.8 ± 2.4 nm and those between the Fc subunit and each Fab subunit at 32.5 ± 5.7 nm and 31.0 ± 6.3 nm, respectively. The width of the Fc subunit was 8.9 ± 1.6 nm. The greatest height of mouse IgG3 was 3.7 ± 0.5 nm and the centre of mass was estimated to be located at 19.1 ± 4.0 nm from the tip of the Fc subunit. These sizes of mouse IgG3 molecules are consistent with those of various IgG measurements obtained by conventional AFM observations23. We previously reported that one ZZ-BNC can capture a maximum of 60 IgG molecules9. When the surface of ZZ-BNC (5,170 nm2; radius, 20.3 nm) was filled with Fc subunits (base area, 32.8 ± 6.0 nm2; height, 19.1 ± 4.0 nm) in close-packed arrangement, ZZ-BNC could be surrounded by 161 ± 29 molecules of Fc subunits. On the other hand, ZZ-BNC fully covered with Fc subunits (surface area, 19,500 nm2; radius, 39.4 nm) could be surrounded by 197 ± 18 molecules of F(ab')2 subunits (base area, 99.3 ± 8.8 nm2; height, 9.6 ± 2.0 nm), suggesting that Fc subunits on ZZ-BNC contact one another intermolecularly, rather than F(ab')2 subunits. Consequently, each IgG could be tethered onto the surface of ZZ-BNC at appropriate intervals, thereby improving antigen recognition by the Fv region.
HS-AFM analyses of mouse IgG3 and ZZ-BNC in solution.
(a) HS-AFM images of mouse IgG3 molecules. Bar, 20 nm. (b) Video image of the movement of mouse IgG3 on ZZ-BNC (0.2 fps). Yellow arrow, the addition of mouse IgG3. Red arrow, first appearance of attachment. Times (s) after the start of observation are indicated in the left margin. Bar, 20 nm. (c) HS-AFM images of ZZ-BNCs on a gold surface without (10 s, left panel)/with (150 s, right panel) mouse IgG3. ZZ-BNC used for (b) is indicated by a white box. The locations where mouse IgG3 appeared are indicated by red arrowheads. Bars, 20 nm. (d) Binarized video image of the movement of mouse IgG3 on ZZ-BNC (0.2 fps). Yellow arrow, the addition of mouse IgG3. Red arrow, first appearance of mouse IgG3. Times (s) after the start of observation are indicated in the left margin. Bar, 20 nm.
Real-time movement of mouse IgG3 on ZZ-BNC in solution
For developing the next generation of immunosensors, it is a prerequisite to analyze and optimize the real-time movement of immobilized IgGs in solution at the single molecular level. ZZ-BNCs adsorbed onto gold surfaces were immersed into PBS (0 s), contacted with mouse IgG3 (55 s, yellow arrows in Figs. 2b and d) and then subjected to time-lapse imaging with HS-AFM (0.2 frames per second (fps), total 1,300 s, 260 frames (Fig. 2b and Supplementary Movie 1). After 45 s from the addition of IgG (100 s from the start), some of the ZZ-BNCs exhibited a rough surface. After 95 s from the addition of IgG (150 s from the start), the height of three ZZ-BNCs (red arrowheads in Fig. 2c) changed from 14.4, 19.6 and 12.6 nm (left panel) to 24.7, 27.7 and 18.0 nm (right panel), respectively. Next, the portion of attachments on ZZ-BNC was extracted with PaintShop Pro software by setting the height of the ZZ-BNC surface (14.4 nm) as a threshold (Fig. 2d and Supplementary Movie 2). The attachment appeared on the surface of ZZ-BNC 55 s after the addition of IgG (110 s from the start, red arrows in Figs. 2b and d), suggesting that mouse IgG3s were captured by ZZ-BNCs and retained during the observation.
Analyses of the movement of mouse IgG3 on ZZ-BNC
The positions of the centre of mass of attachments were calculated from their projected areas using ImageJ software and were found to exhibit a two-dimensional Gaussian distribution (major axis, 25.6 nm; minor axis, 20.4 nm; Fig. 3a). Because the length between the centre of mass of mouse IgG3 and the tip of the Fc subunit was estimated to be 19.1 ± 4.0 nm (see above), the deflection angle of mouse IgG3 on ZZ-BNC was estimated to be 88.1 ± 20.5 degrees. These results demonstrate that mouse IgG3s were tightly bound onto ZZ-BNC through the Fc subunit with Fab subunits swinging outwardly (Fig. 3b). The average speed of movement of mouse IgG3 was estimated as 4.6 ± 3.0 nm (13.8 ± 9.0 degrees) per 5 s (Fig. 3c). The speed fluctuated from 0.24 to 15.6 nm (0.74 ± 0.14 to 49.7 ± 10.1 degrees) per 5 s (Fig. 3d). It has been reported that the diffusion coefficient of monomeric human IgG is ~4 × 10−7 cm2 s−1, implying that one IgG molecule can move spontaneously at 20 nm per 5 second24. These results indicate that IgG molecules on ZZ-BNC undergo rotational Brownian motion. ZZ-BNC acts as a scaffold for oriented-immobilization of IgGs, being expected to enhance the sensitivity and specificity of a wide range of immunosensors and immunoassays.
Analyses of the movement of mouse IgG3 on ZZ-BNC.
(a) Positions of the centre of mass of mouse IgG3 on ZZ-BNC are shown with blue dots (n = 210). The 99% confidence level is indicated by the red circle. (b) Postulated structure of mouse IgG3 on ZZ-BNC. Mouse IgG3 was tightly bound onto ZZ-BNC through the Fc subunit with Fab subunits swinging outwardly. Centres of mass of IgGs are indicated with black asterisks. (c) Relationship between speed (nm per 5 s) and time (s) for the movement of mouse IgG3 (centre of mass). (d) Relationship between occurrence and speed (nm per 5 s) of the movement of mouse IgG3 (centre of mass). Cumulative relative occurrence (%) of each speed is indicated with a blue line.
Discussion
The surface structure of BNC (recombinant yeast-derived HBsAg L protein particle) has been analyzed by conventional transmission electron microscopy (TEM) and AFM in air8,9,25. While rugged surface was suggested by TEM observation, no protrusion has so far been identified. Recently, the surface structure of native HBV virion has been analyzed by electron cryomicroscopy17. The envelope of HBV virion was decorated with 160–200 surface protrusions that are spaced ~6 nm apart. In this study, HS-AFM observation revealed for the first time that ZZ-BNC (a derivative of BNC) in solution displays 54 surface protrusions that are spaced 8.2 ± 1.7 nm apart. The small differences in space may be attributed to either the conditions for observation (air versus solution) or the mass of displayed molecules (~14-kDa pre-S region for HBV virion versus ~20-kDa ZZ domains for ZZ-BNC). Furthermore, during the biosynthesis of HBsAg particle in recombinant mammalian cells, nascent HBsAgs were shown to translocate across endoplasmic reticulum (ER) membrane along with the formation of intramolecular disulfide bonds, dimerize by forming intermolecular disulfide bonds and then spontaneously assemble into particle structure without forming multimers19. But, while recombinant yeast-derived HBsAg L proteins (including HBsAg S and M proteins) form multimers by intermolecular disulfide bonds8, it has not been determined if they form dimers or not prior to the formation of BNC. In this study, nascent ZZ-L proteins (a derivative of HBsAg L protein) translocated across yeast ER membrane were strongly suggested to dimerize firstly, assemble to particle structures and then form multimers by intermolecular disulfide bonds. Therefore, it was considered that more oxidative conditions of protein biosynthesis in yeast cells26 rather than mammalian cells27 might contribute to the formation of multimeric HBsAg proteins in yeast cells8.
In the development of biosensors, it is indispensable to decipher the direction, shape and movement of sensing molecules on solid phase for improving sensitivity, specificity and analyte-binding capacity. Conventionally, these molecules have been analyzed by electron microscopy (EM) and AFM12,13,14. While both methods allowed us to observe IgGs at single molecule level, the former could be operable only in air. However, they could neither achieve real-time observation of IgGs in solution nor distinguish Fc from Fab clearly. Exceptionally, Garcia et al. have succeeded in identifying the position of the Fc and Fab fragments in air by tapping-mode AFM28,29. The technical defects of these methods have forced us to evaluate the sensing molecules on solid phase by indirect methods (i.e., functional analyses). For example, in case of immunosensors, the degree of oriented immobilization of IgGs has been evaluated by the reactivity to antigens30. These indirect methods are indeed suitable for the optimization of biosensors, but it is equivocal to judge if sensing molecules exert their ability as much as possible. In this study, we could directly observe real-time movement of mouse IgG3s tethered onto ZZ-BNCs in solution by HS-AFM. It was therefore confirmed that ZZ-BNCs achieved oriented immobilization of IgGs, which has been hypothesized by functional analyses9,10,11, by tethering Fc subunits in nearly close-packed arrangement and minimizing the steric hindrance of Fv regions for efficient antigen bindings. Since ZZ-BNCs are stable to heat, chemical and mechanical stresses9, ZZ-BNCs would be an ideal and practical scaffold for oriented immobilization of IgGs.
When observing the surface of biosensors by EM and AFM, it is essentially difficult to identify what molecules are adsorbed onto solid phase. For the molecular identification in EM, samples are usually pre-treated with labelling reagents (e.g., gold particle-labelled antibodies). Worse yet, since conventional AFM can detect the ruggedness of sample surface only, it is nearly impossible to identify the molecules on the surface of mixed samples. On the contrary, HS-AFM can identify the molecules on the surface of solid phase by their movements without any modification. Taken together, HS-AFM might be worthwhile for the refinement of other biosensing molecules (e.g. ligands, receptors, aptamers, sugar chains, lectins) in various biosensors at the single molecule level.
Methods
ZZ-BNCs
ZZ-BNCs were overexpressed in Saccharomyces cerevisiae AH22R− cells carrying the ZZ-BNC-expression plasmid pGLD-ZZ507,8. Following a previously described purification protocol for BNC31, ZZ-BNCs were extracted from yeast cells by disruption with glass beads and purified using an AKTA™ liquid chromatography system (GE Healthcare, Amersham, UK) by affinity chromatography on porcine IgG, followed by gel filtration. The ZZ-BNCs were analyzed by 12.5% SDS-PAGE, followed by silver staining (Wako Chemicals, Osaka, Japan).
High-speed AFM
Topology images of ZZ-BNCs and mouse IgG3s (Sigma-Aldrich, St Louis, MO, USA) were obtained on a high-speed AFM system (Nano Live Vision, RIBM, Tokyo, Japan) using a silicon nitride cantilever (BL-AC10EGS, Olympus, Tokyo, Japan). To obtain static images, either ZZ-BNCs (50 μg ml−1, 2 μl) or mouse IgG3s (1 μg ml−1, 2 μl) were adsorbed on a mica surface (Ted Pella Inc., Redding, CA, USA), incubated for 10 min, washed with phosphate-buffered saline (PBS) three times and then subjected to time-lapse imaging. To observe the real-time movement of IgGs on ZZ-BNC in solution, ZZ-BNCs (50 μg ml−1, 2 μl) were adsorbed onto an atomically flat gold surface (Auro sheet (111) HS; Tanaka Kikinzoku Kogyo K.K., Tokyo, Japan), incubated for 10 min, washed with PBS three times, subjected to time-lapse imaging (0 s) and then mixed with mouse IgG3 (50 μg ml−1, 4 μl) (55 s). Images (192 × 144 pixels) were obtained at a scan rate of 0.2 frames per second (fps) for 1,300 s. Images were analyzed by ImageJ 1.44o software (http://rsbweb.nih.gov/ij/) and PaintShop Pro software (COREL Corporation, Ontario, Canada). Semi-automated, single-particle reconstructions were performed using the EMAN software package, v1.916. Ten particles were selected from 48 frames of the HS-AFM movies (total 761 frames) using the boxer program in the EMAN package.
References
Wink, T., van Zuilen, S. J., Bult, A. & van Bennkom, W. P. Self-assembled monolayers for biosensors. Analyst 122, 43R–50R (1997).
Lu, B., Smyth, M. R. & O'Kennedy, R. Oriented immobilization of antibodies and its applications in immunoassays and immunosensors. Analyst 121, 29R–32R (1996).
Gersten, D. M. & Marchalonis, J. J. A rapid, novel method for the solid-phase derivatization of IgG antibodies for immune-affinity chromatography. J. Immunol. Methods. 24, 305–309 (1978).
Björck, L. & Kronvall, G. Purification and some properties of streptococcal protein G, a novel IgG-binding reagent. J. Immunol. 133, 969–974 (1984).
O'Shannessy, D. J. & Quarles, R. H. Labeling of the oligosaccharide moieties of immunoglobulins. J. Immunol. Methods 99, 153–161 (1987).
Prisyazhnoy, V. S., Fusek, M. & Alakhov, Y. Synthesis of high-capacity immunoaffinity sorbents with oriented immobilized immunoglobulins or their Fab' fragments for isolation of proteins. J. Chromatogr. 424, 243–253 (1988).
Kurata, N. et al. Specific protein delivery to target cells by antibody-displaying bionanocapsules. J. Biochem. 144, 701–707 (2008).
Kuroda, S., Otaka, S., Miyazaki, T., Nakao, M. & Fujisawa, Y. Hepatitis B virus envelope L protein particles. Synthesis and assembly in Saccharomyces cerevisiae, purification and characterization. J. Biol. Chem. 267, 1953–1961 (1992).
Iijima, M. et al. Nanocapsules incorporating IgG Fc-binding domain derived from Staphylococcus aureus protein A for displaying IgGs on immunosensor chips. Biomaterials 32, 1455–1464 (2011).
Iijima, M. et al. Bionanocapsule-based enzyme-antibody conjugates for enzyme-linked immunosorbent assay. Anal. Biochem. 396, 257–261 (2010).
Iijima, M. et al. Fluorophore-labeled nanocapsules displaying IgG Fc-binding domains for the simultaneous detection of multiple antigens. Biomaterials 32, 9011–9020 (2011).
Dutta, P., Sawoo, S., Ray, N., Bouloussa, O. & Sarkar, A. Engineering bioactive surfaces with Fischer carbene complex: protein A on self-assembled monolayer for antibody sensing. Bioconjug. Chem. 22, 1202–1209 (2011).
Bendikov, T. A., Rabinkov, A., Karakouz, T., Vaskevich, A. & Rubinstein, I. Biological sensing and interface design in gold island film based localized plasmon transducers. Anal Chem. 80, 7487–7498 (2008).
Lv, Z., Wang, J., Chen, G. & Deng, L. Imaging recognition events between human IgG and rat anti-human IgG by atomic force microscopy. Int. J. Biol. Macromol. 47, 661–667 (2010).
Kodera, N., Yamamoto, D. & Ando, T. Video imaging of walking myosin V by high-speed atomic force microscopy. Nature 468, 72–76 (2010).
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Dryden, K. A. et al. Native hepatitis B virions and capsids visualized by electron cryomicroscopy. Mol. Cell 22, 843–850 (2006).
Eble, B. E., Lingappa, V. R. & Ganem, D. Hepatitis B surface antigen: an unusual secreted protein initially synthesized as a transmembrane polypeptide. Mol. Cell. Biol. 6, 1454–1463 (1986).
Whunderlich, G. & Bruss, V. Characterization of early hepatitis B virus surface protein oligomers. Arch. Virol. 141, 1191–1205 (1996).
Nagaoka, T. et al. Characterization of bio-nanocapsules as a transfer vector targeting human hepatocyte carcinoma by disulfide linkage modification. J. Control. Release. 118, 348–356 (2007).
Palczewski, K. et al. Crystal structure of rhodopsin: a G protein-coupled receptor. Science 289, 739–745 (2000).
Thomson, N. H. The substructure of immunoglobulin G resolved to 25 kDa using amplitude modulation AFM in air. Ultramicroscopy 105, 103–110 (2005).
Thomson, N. H. Imaging the substructure of antibodies with tapping-mode AFM in air: the importance of a water layer on mica. J. Microsc. 217, 193–199 (2005).
Jøssang, T., Feder, J. & Rosenqvist, E. Photon correlation spectroscopy of human IgG. J. Protein Chemistry 7, 165–171 (1988).
kanno, T. et al. Size distribution measurement of vesicles by atomic force microscopy. Anal. Biochem. 309, 196–199 (2002).
Frand, A. R. & Kaiser, C. A. Ero1p oxidized protein disulfide isomerise in a pathway for disulfide bond formation in the endoplasmic reticulum. Mol. Cell 4, 469–477 (1999).
Jessop, C. E. & Bulleid, N. J. Glutathione directly reduces an oxidoreductase in the endoplasmic reticulum of mammalian cells. J. Biol. Chem. 279, 55341–55347 (2004).
San Paulo, A. & García, R. High-resolution imaging of antibodies by tapping-mode atomic force microscopy: attractive and repulsive tip-sample interaction regimes. Biophys. J. 78, 1599–1605 (2000).
Patil, S., Martinez, N. F., Lozano, J. R. & García, R. Force microscopy imaging of individual protein molecules with sub-pico Newton force sensitivity. J. Mol. Recognit. 20, 516–523 (2007).
Song, H. Y., Zhou, X., Hobley, J. & Su, X. Comparative study of random and oriented antibody immobilization as measured by dual polarization interferometry and surface plasmon resonance spectroscopy. Langmuir 28, 997–1004 (2012).
Jung, J. et al. Efficient and rapid purification of drug- and gene-carrying bio-nanocapsules, hepatitis B virus surface antigen L particles, from Saccharomyces cerevisiae. Protein Expr. Purif. 78, 149–155 (2011).
Nilsson, B. et al. A synthetic IgG-binding domain based on staphylococcal protein A. Protein Eng. 1, 107–113 (1987).
Acknowledgements
We thank Profs. K. Namba and T. Kato (Osaka University) for helpful advice and the Research Institute of Biomolecule Metrology Inc. for technical support. This work was supported in part by The Naito Foundation (to SK), the Canon Foundation (K09-00051, to SK), KAKENHI (Grant-in-Aid for Scientific Research (A) (21240052, to SK); Grant-in-Aid for Young Scientists (B) (23710143, to MI)), the Program for Promotion of Basic and Applied Researchers for Innovations in Bio-Oriented Industry (BRAIN) (to SK) and the Health Labour Sciences Research Grant from the Ministry of Health Labour and Welfare (to SK).
Author information
Authors and Affiliations
Contributions
M.I. designed and performed the experiments, analyzed the data and wrote the manuscript. M.S., N.Y. and T.N. gave helpful comments. S.K. designed and supervised the experiments, analyzed the data and wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Video 1
Supplementary Information
Supplementary Video 2
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Iijima, M., Somiya, M., Yoshimoto, N. et al. Nano-visualization of oriented-immobilized IgGs on immunosensors by high-speed atomic force microscopy. Sci Rep 2, 790 (2012). https://doi.org/10.1038/srep00790
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00790
This article is cited by
-
Capturing transient antibody conformations with DNA origami epitopes
Nature Communications (2020)
-
Nanopore extended field-effect transistor for selective single-molecule biosensing
Nature Communications (2017)
-
High efficiency penetration of antibody-immobilized nanoneedle thorough plasma membrane for in situ detection of cytoskeletal proteins in living cells
Journal of Nanobiotechnology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.