Abstract
In contrast to previous common wisdom that epidemic activity in heterogeneous networks is dominated by the hubs with the largest number of connections, recent research has pointed out the role that the innermost, dense core of the network plays in sustaining epidemic processes. Here we show that the mechanism responsible of spreading depends on the nature of the process. Epidemics with a transient state are boosted by the innermost core. Contrarily, epidemics allowing a steady state present a dual scenario, where either the hub independently sustains activity and propagates it to the rest of the system, or, alternatively, the innermost network core collectively turns into the active state, maintaining it globally. In uncorrelated networks the former mechanism dominates if the degree distribution decays with an exponent larger than 5/2 and the latter otherwise. Topological correlations, rife in real networks, may perturb this picture, mixing the role of both mechanisms.
Similar content being viewed by others
Introduction
The discernment of the mechanisms that contrive to activate spreading processes on heterogeneous substrates is a pivotal issue, with practical applications ranging from the containment of epidemic outbreaks1 to the viral spreading of rumors and beliefs2,3. The interest on the effects of heterogeneity has been brought about by the observation that social contact networks (the natural substrate for most human epidemic processes) are generally strongly heterogeneous4,5,6, observation that has led to the introduction of complex network theory in the quantitative analysis of epidemic spreading7. In this context, the nature of the activation mechanisms translates on simple epidemic models8 in setting the epidemic threshold λc for some rate of infection λ (the spreading rate), separating a phase in which the spreading affects a finite fraction of the population from a state in which only a vanishingly small fraction is hit. The research effort is thus focused on a twofold objective: The identification of the activation mechanisms as a function of the network topology and the determination of the functional form of the epidemic threshold.
For the sake of concreteness, we focus our discussion on the simplest models of disease spreading, namely the susceptible-infected-susceptible (SIS) and the susceptible-infected-recovered (SIR) models, leading, respectively, to a steady endemic state or to transient outbreaks affecting a given fraction of the population8 (see Methods). On a network substrate—statistically described, at the simplest level, by its degree distribution P(q), defined as the probability that a randomly chosen individual (vertex) is connected to q other individuals4—the application of a heterogeneous mean-field (HMF) approach9 assuming no topological correlations10 and neglecting dynamical correlations, yields epidemic thresholds inversely proportional to the second moment of the degree distribution, 〈q2〉11,12,13. Since most natural networks have a degree distribution scaling as4 P(q) ∼q−γ, 〈q2〉 takes the form, in the continuous degree approximation,  , where qmax is the maximum degree in the network5. The second moment therefore diverges in the infinite network size limit (i.e. when qmax → ∞) for γ ≤ 3, leading to a vanishing epidemic threshold, i.e. any disease can invade the system, whatever its infection rate14,15. For γ > 3, on the other hand, HMF predicts a finite threshold. This result has usually been interpreted in terms of the leading role of the hubs (the vertices with largest degree in the network) as the elements sustaining the epidemic activity in the network, whenever they have a sufficiently large degree to make the second moment 〈q2〉 diverge (i.e. when γ ≤ 3)9.
, where qmax is the maximum degree in the network5. The second moment therefore diverges in the infinite network size limit (i.e. when qmax → ∞) for γ ≤ 3, leading to a vanishing epidemic threshold, i.e. any disease can invade the system, whatever its infection rate14,15. For γ > 3, on the other hand, HMF predicts a finite threshold. This result has usually been interpreted in terms of the leading role of the hubs (the vertices with largest degree in the network) as the elements sustaining the epidemic activity in the network, whenever they have a sufficiently large degree to make the second moment 〈q2〉 diverge (i.e. when γ ≤ 3)9.
More refined approaches than HMF, incorporating the effects of the quenched topological structure of the network, but still neglecting dynamical correlations16,17,18, predict that the epidemic threshold for SIS is in general set by the largest eigenvalue ΛN of the adjacency matrix,  . This finding, combined with the scaling of ΛN (computed by Chung et al. for a class of finite graphs with degrees distributed according to a power-law19),
. This finding, combined with the scaling of ΛN (computed by Chung et al. for a class of finite graphs with degrees distributed according to a power-law19),  , leads to a threshold20
, leads to a threshold20

Equation (1) implies that the epidemic threshold vanishes in the thermodynamic limit in power-law distributed networks for any value of γ, even larger than 3, as long as qmax is a growing function of the network size N, in agreement with previous results for SIS21,22,23. In this perspective, it would be the hub, or most connected vertex, the main responsible of maintaining the epidemic activity and correspondingly setting the threshold20.
The relevance of hubs has been however recently called into question by Kitsak et al.24, who pointed out that in some real networks, the most efficient spreaders are located at the innermost, dense core of the network, as identified by means of a k-core decomposition25 (see Methods and Figure 1). In this alternative view, it is thus the nucleus of high k-core index which sustains epidemic activity, independently of the degree of the vertices it is composed of.
Visual representation of the k-core decomposition of a small network of size N = 30 and maximum degree qmax = 10.
Blue vertices belong to the k = 1 shell and green vertices to the k = 2 shell. The maximum k-core, with kS = 3, is composed by the red vertices. The hub (vertex with largest degree) is represented as a square.
Inspired by these results, here we analyze in detail the role of the hub and of the core of the network for the onset of epidemic spreading on complex topologies. By means of theoretical arguments and extensive numerical simulations, we are able to show that the leading mechanism governing the dynamics depends on the network features, in particular on the strength of the degree heterogeneity, as measured by the degree exponent γ. The analysis of real networks allows to determine additionally the critical role of degree correlations in suppressing or enhancing the relevant mechanism. The findings presented in this work represent an advancement in the understanding of the underlying mechanisms that control the behavior of epidemic processes on complex heterogeneous networks. By identifying with precision the set of vertices ultimately responsible for the epidemic activation, our results open the path for the formulation of immunization strategies1,26 specifically tailored for each particular network configuration considered. Moreover, our results can find application in other, more general spreading processes, such as rumor, behavior or information spreading in networks27,28, as well as other dynamical processes ruled by the largest eigenvalue of the adjacency matrix, such as synchronization phenomena29.
Results
Activation mechanisms for SIS in uncorrelated networks
In the case of the SIS model the expression of the threshold λc for γ > 5/2 can be understood by considering the largest hub and its neighbors as a star network of size qmax + 1. Such a system has, in isolation, a threshold  and is thus capable, all by itself and independently of the degree of the rest of the vertices, to propagate the infection to a finite fraction of the network, leading to a stable endemic state whenever
and is thus capable, all by itself and independently of the degree of the rest of the vertices, to propagate the infection to a finite fraction of the network, leading to a stable endemic state whenever  20. It is therefore the most connected vertex which singlehandedly can keep the epidemic activity alive, setting in this way the global threshold for activity in the system. The change for γ < 5/2 in equation (1) is however surprising and hints towards the possibility of different activation mechanisms for different γ values, thus challenging the belief in the preponderant role of hubs, which has become common wisdom in network science9. The results of Kitsak et al. would fit in place, pointing towards a preponderant role for the innermost core of the network. However, while the picture presented by Kitsak et al.24 is compelling for the SIR model, the case of the SIS deserves a closer look.
20. It is therefore the most connected vertex which singlehandedly can keep the epidemic activity alive, setting in this way the global threshold for activity in the system. The change for γ < 5/2 in equation (1) is however surprising and hints towards the possibility of different activation mechanisms for different γ values, thus challenging the belief in the preponderant role of hubs, which has become common wisdom in network science9. The results of Kitsak et al. would fit in place, pointing towards a preponderant role for the innermost core of the network. However, while the picture presented by Kitsak et al.24 is compelling for the SIR model, the case of the SIS deserves a closer look.
In order to shed light on this issue, we have performed extensive numerical simulations of the SIS process on synthetic uncorrelated scale-free networks with degree distribution P(q) ∼q−γ, generated via the uncorrelated configuration model (UCM)30 (see Supplementary Information online for more details). We have computed the density of infected vertices in the whole network and the same density when the dynamics takes place (in isolation) on the k-core of highest index (maximum k-core) and on the star-graph centered around the hub of the network, with degree qmax.
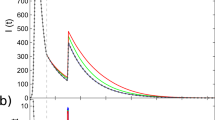
In Figure 2 we show the evolution of the recorded densities as a function of time for different values of the spreading rate λ, in networks with large and small degree exponents, namely γ = 2.75 and γ = 2.1. Our results show a remarkable dependence on the degree exponent: For large γ, the onset of a global stationary state takes place for the same values of λ for which the star-graph centered around the hub starts to be active, while the maximum k-core remains subcritical, with exponentially decaying activity. This behavior is a proof of the leading role of the hub as the main activation mechanism for large γ. For small values of γ, instead, the picture is opposite: For values of λ corresponding to a globally active network, the maximum k-core is in an active state, while the star-graph centered around the hub is inactive, indicating that now the maximum k-core is the trigger activating the whole system. Two observations are in order: The maximum k-core is the heart of the nucleus of most densely connected vertices in the network but it does not sharply coincide with it. Other nodes, belonging to k-cores of index slightly smaller, are also densely connected. The transition in the whole network is influenced also by these other nodes and therefore only approximately coincides with the transition of the maximum k-core. This explains why in Fig. 2 the whole network is fully active for λ = 0.01, while the maximum k-core is still around the transition. The second observation is that in uncorrelated networks the hub usually belongs to the maximum k-core. Yet the two activation mechanisms for epidemics are clearly distinct. In one case (hub triggered activation) the hub alone is able to sustain activity in the set of its neighbors and then propagate it to the rest of the system. In the other (maximum k-core triggered activation) the hub alone is not able to sustain activity: Only the presence of all densely connected vertices in the k-core allows them to collectively turn into the active state and propagate to the rest of the system.
Average density of infected vertices as a function of time, ρ(t), in the SIS model on uncorrelated scale-free networks generated by means of the UCM algorithm.
We consider networks with widely separated degree exponent and different size, namely γ = 2.75, N = 3×107 and γ = 2.1, N = 106. The different columns correspond to the average density computed when the dynamics runs over the whole network (left), only over the maximum k-core of the network (center) and only over the largest hub (right), considered as an isolated star network. The different colors correspond to different values of the spreading rate λ. For small γ = 2.1 (bottom row), the onset of the global steady state is correlated with the active state of the epidemics on the maximum k-core, while it corresponds to a subcritical state for the hub. This observation indicates that in this case the maximum k-core is responsible for the overall activity in the network. For large γ = 2.75, on the other hand, the global active state is linked to an active hub and a subcritical maximum k-core, signaling that it is the former mechanism the one keeping activity on a global scale.
This change of behavior with the degree exponent, which we have confirmed for different values of γ above and below 5/2 (see Supplementary Information Figure S1 online), can be made more physically transparent by linking it analytically with the different thresholds in equation (1). To do so, we estimate the threshold associated to the active maximum k-core, whose index is denoted as kS. The maximum k-core has a degree distribution which is bounded and narrow, with minimum (kS), average and maximum degree scaling with size in the same way (see Supplementary Information Figure S2 online). Hence its epidemic threshold is well approximated by  . On the other hand, in Ref.31 the maximum k-core index kS was determined as a function of the network topology, yielding for scale-free networks with 2 < γ < 3
. On the other hand, in Ref.31 the maximum k-core index kS was determined as a function of the network topology, yielding for scale-free networks with 2 < γ < 3

where qmin is the minimum degree. Introducing this result into the formula for  we obtain
we obtain  . It is most noteworthy that the scaling behavior of the maximum k-core threshold
. It is most noteworthy that the scaling behavior of the maximum k-core threshold  takes the exact same form as the eigenvalue threshold for γ < 5/2 in equation (1). This observation provides a physical interpretation of the different activation mechanisms and associated thresholds in uncorrelated scale-free networks: When γ < 5/2, the epidemic transition is collectively triggered by the vertices in the innermost core and the threshold is correspondingly given by
takes the exact same form as the eigenvalue threshold for γ < 5/2 in equation (1). This observation provides a physical interpretation of the different activation mechanisms and associated thresholds in uncorrelated scale-free networks: When γ < 5/2, the epidemic transition is collectively triggered by the vertices in the innermost core and the threshold is correspondingly given by  , as in HMF theory. On the other hand, for γ > 5/2, the hub triggers the global activity and the threshold is given by
, as in HMF theory. On the other hand, for γ > 5/2, the hub triggers the global activity and the threshold is given by  . An additional inspection of the numerical values of the different thresholds (see Supplementary Information Table S1 online) shows that the thresholds computed from the numerical estimation of the largest eigenvalue of the adjacency matrix are in very good agreement with the predictions of Eq. (1), perfectly accounting for the results observed in Figure 2.
. An additional inspection of the numerical values of the different thresholds (see Supplementary Information Table S1 online) shows that the thresholds computed from the numerical estimation of the largest eigenvalue of the adjacency matrix are in very good agreement with the predictions of Eq. (1), perfectly accounting for the results observed in Figure 2.
The SIR model
We turn now our attention to the SIR model, which, contrarily to the steady-state dynamics of the SIS model, exhibits transient outbreaks characterized by the number of infected individuals, totaling a finite fraction of the system only above the epidemic threshold. Evidence that the hub plays no special role in SIR dynamics comes from considering this process on a star network of size qmax + 1. For an epidemics starting from a randomly chosen vertex, the average final density of infected nodes takes the form (see Methods)

Hence the threshold in a star network, defined by the value of λ above which R takes a given fixed finite value, is a constant independent of qmax, in the limit of large qmax. The hub cannot therefore be the ultimate trigger of global outbreaks in the SIR model and this role must instead be played by the maximum k-core for any value of γ, in accordance with Ref.24. Additional support for this view comes from extending to the SIR case the maximum k-core threshold argument presented for the SIS model. Approximating again the maximum k-core as a narrowly distributed graph of average degree 〈q〉 ∼ kS, a threshold is obtained from MF theory of the form  . Given the form of kS in equation (2), this threshold scales in the large network limit in exactly the same form as the HMF prediction, namely
. Given the form of kS in equation (2), this threshold scales in the large network limit in exactly the same form as the HMF prediction, namely  32,33. The conclusion is that in the SIR model it is always the maximum k-core which controls epidemic spreading and sets the threshold to the HMF value. This picture is substantiated in Figure 3, where we consider the SIR model on UCM networks with different values of γ, keeping track, as a function of λ, of the density of infected individuals in the global network, in the maximum k-core and in the star-graph centered around the hub. As we can see, the position of the transition to a finite fraction of infected vertices is closely correlated in the whole network and the maximum k-core, while the behavior of the hub conforms to the prediction of equation (3).
32,33. The conclusion is that in the SIR model it is always the maximum k-core which controls epidemic spreading and sets the threshold to the HMF value. This picture is substantiated in Figure 3, where we consider the SIR model on UCM networks with different values of γ, keeping track, as a function of λ, of the density of infected individuals in the global network, in the maximum k-core and in the star-graph centered around the hub. As we can see, the position of the transition to a finite fraction of infected vertices is closely correlated in the whole network and the maximum k-core, while the behavior of the hub conforms to the prediction of equation (3).
Total density R of infected vertices, as a function of the spreading rate λ, computed after an epidemic outbreak in the SIR model on uncorrelated scale-free networks generated by means of the UCM algorithm.
The degree exponents considered are γ = 2.25 (left) and γ = 2.75 (right), with network sizes N = 106. The line colors correspond to the SIR dynamics restricted to the maximum k-core (black), to the largest hub (green) and on the whole network (red). The value of λ after which a macroscopic fraction of the network becomes infected is correlated in the whole network and in the maximum k-core, while the infection pattern on the hub conforms with the theoretical expression in equation (3). These results indicate the crucial role of the maximum k-core in keeping the SIR activity on a large scale in networks, independently of the behavior of the hub.
Effects of correlations
The scenario discussed so far applies to the case of uncorrelated networks, where the probability that a random edge is connected to a vertex of degree q is proportional to qP(q)5. Real networks, however, present in most cases some level of degree correlations10, as measured by the Pearson coefficient34 or by the average degree of the nearest neighbors (ANN) of the vertices of given degree,  35. In order to ascertain their effect on the relevant epidemic mechanisms, we have considered the SIS process on several instances of correlated real networks (see Methods): An Internet map at the autonomous system (AS) level, the social network of pretty-good-privacy (PGP) and the network of actors co-starring in Hollywood movies (Movies). All these networks have a degree distribution compatible with a power law, with an exponent close to 2 (see Supplementary Information Figure S3 online) and a range of degree correlations (see Supplementary Information Figure S4 online). In this case, according to our arguments above and neglecting correlations, one would expect the transitions to be ruled by the corresponding maximum k-cores. This fact is confirmed for the Movies and PGP networks, by the SIS simulations presented in Figure 4, showing that the transition occurs simultaneously for the maximum k-core and the whole system, while the star-graph centered around the hub remains inactive. In the case of the AS network, instead, the picture is surprisingly the opposite and it is apparently the hub the responsible of the epidemic transition, see Figure 4. The situation is still more complex when one considers other, larger AS maps (see Supplementary Information). In fact, as it turns out from the analysis of numerical simulations (see Supplementary Information Figure S5 online), the general situation in AS maps is that we can reach in the network an active, infected state for values of λ for which both the hub and the maximum k-core are apparently subcritical. This observation hints towards a mixing of activation mechanisms for the particular case of AS networks. This discrepancy between the AS and the other networks, confirmed by the inspection of the values of the different thresholds (see Supplementary Information Table S2 online) can be attributed to the presence of strong degree correlations. Measuring them by means of the auxiliary ANN function
35. In order to ascertain their effect on the relevant epidemic mechanisms, we have considered the SIS process on several instances of correlated real networks (see Methods): An Internet map at the autonomous system (AS) level, the social network of pretty-good-privacy (PGP) and the network of actors co-starring in Hollywood movies (Movies). All these networks have a degree distribution compatible with a power law, with an exponent close to 2 (see Supplementary Information Figure S3 online) and a range of degree correlations (see Supplementary Information Figure S4 online). In this case, according to our arguments above and neglecting correlations, one would expect the transitions to be ruled by the corresponding maximum k-cores. This fact is confirmed for the Movies and PGP networks, by the SIS simulations presented in Figure 4, showing that the transition occurs simultaneously for the maximum k-core and the whole system, while the star-graph centered around the hub remains inactive. In the case of the AS network, instead, the picture is surprisingly the opposite and it is apparently the hub the responsible of the epidemic transition, see Figure 4. The situation is still more complex when one considers other, larger AS maps (see Supplementary Information). In fact, as it turns out from the analysis of numerical simulations (see Supplementary Information Figure S5 online), the general situation in AS maps is that we can reach in the network an active, infected state for values of λ for which both the hub and the maximum k-core are apparently subcritical. This observation hints towards a mixing of activation mechanisms for the particular case of AS networks. This discrepancy between the AS and the other networks, confirmed by the inspection of the values of the different thresholds (see Supplementary Information Table S2 online) can be attributed to the presence of strong degree correlations. Measuring them by means of the auxiliary ANN function  , we can observe that the AS network is strongly correlated, with
, we can observe that the AS network is strongly correlated, with  decaying as q−0.5. Moreover, these correlations are so strong that they do not wash away even after randomizing the network, as they do in the PGP and Movies networks (see Supplementary Information Figure S4 online). Strong disassortative correlations reduce the interconnections of vertices of high-degree, suppressing in this way the index kS of the maximum k-core and reducing the number of vertices that compose it. This situation, i.e. a very large hub coupled with a relatively small maximum k-core, leads to a mixing of both mechanisms that does not allow to make explicit prediction about the most relevant one. Similar or opposite effects (i.e. strong assortativity enhancing the role of the maximum k-core) can be found in networks generated by means of the Weber-Porto algorithm36, a modification of the configuration model that generates graphs with prescribed degree distribution and correlations of tunable strength (see Supplementary Information Figure S6 and Table S2 online).
decaying as q−0.5. Moreover, these correlations are so strong that they do not wash away even after randomizing the network, as they do in the PGP and Movies networks (see Supplementary Information Figure S4 online). Strong disassortative correlations reduce the interconnections of vertices of high-degree, suppressing in this way the index kS of the maximum k-core and reducing the number of vertices that compose it. This situation, i.e. a very large hub coupled with a relatively small maximum k-core, leads to a mixing of both mechanisms that does not allow to make explicit prediction about the most relevant one. Similar or opposite effects (i.e. strong assortativity enhancing the role of the maximum k-core) can be found in networks generated by means of the Weber-Porto algorithm36, a modification of the configuration model that generates graphs with prescribed degree distribution and correlations of tunable strength (see Supplementary Information Figure S6 and Table S2 online).
Average density of infected vertices as a function of time, ρ(t), for the SIS model on three instances of real correlated networks: The network of actors co-starring in Hollywood movies (Movies), the social network of pretty-good-privacy (PGP) and an Internet map at the autonomous system level (AS); see Methods for further details of the networks.
As in Fig. 2, columns refer to the activity on the whole network (left) and restricted to the maximum k-core (center) or the largest hub (right). Line colors indicate different values of the spreading rate considered in the networks. In the Movies and PGP networks, the maximum k-core dominates the transition, as expected due to the small degree exponent of all three networks (γ ∼ 2). Surprisingly, in the AS network it is the activation of the hub the dominant mechanism setting in the steady state. This different behavior must be attributed to the very strong correlations present in the AS map (see main text).
Discussion
The rationalization of the different mechanisms that keep an epidemic process alive in a heterogeneous substrate turns out to be a more complex issue than previously believed. In fact, two different subsets of vertices (either the hub or the innermost core of the network) can take the role of “super-spreaders” of the infection, depending on the nature of the epidemic process and on the topological features of the underlying network. In processes with no steady state, such as the SIR model, the innermost core is the main trigger activating infection and setting the value of the epidemic threshold. On the contrary, in processes allowing an endemic steady-state, the actual activation mechanism depends essentially on the degree of heterogeneity of the network. This simple picture, valid for uncorrelated networks, can be however modified by the presence of strong degree correlations, which can shift the weight towards one or the other mechanism. These observations call for further theoretical research on epidemic processes on strongly correlated networks. On the other hand, our results might find practical applications in the implementation of optimized immunization strategies, which can be designed to target the actual “super-spreaders” of a given contact network. Moreover, our work could turn out to be relevant to other types of spreading processes, such as information, behavior or rumor spreading and more in general, to dynamical processes whose behavior and possible transitions are ruled by the value of the largest eigenvalue of the adjacency matrix, such as, for example, synchronization phenomena. Also, for these latter dynamics, our results call for additional investigations on the existence and interplay of the relevant activation mechanisms.
Methods
The SIS and SIR epidemic models
In the SIS model, individuals can be in one of two states, either susceptible or infected. Susceptibles become infected by contact with infected individuals, with a rate equal to the number of infected contacts times a given spreading rate λ. Infected individuals on the other hand become healthy again with a rate µ that can be taken arbitrarily equal to unity, thus setting the characteristic time scale. This model allows thus individuals to contract the infection time and again, making possible, in the infinite population limit, a sustained infected steady state (endemic state). This occurs for values of λ larger than the epidemic threshold λc, while for λ < λc the epidemics lasts only for a finite time and asymptotically all individuals are healthy.
In the SIR model, on the other hand, individuals can be in one of three different states: susceptible, infected and recovered (or removed). The dynamical rule for susceptible individuals is the same as for SIS. With a rate µ (again set to unity) infected individuals change their state and recover. Recovered individuals are completely inert and cannot become infected again. With this dynamics the system always reaches asymptotically an absorbing state with only susceptible or removed individuals and no infected ones. A threshold λc separates a regime where outbreaks reach a finite fraction of the individuals (i.e. the final density of removed individuals is finite) from a regime λ < λc where only an infinitesimal fraction of individuals is hit.
The k-core decomposition
The k-core decomposition is an iterative procedure to classify vertices of a network in layers of increasing density of connections. Starting with the full graph one removes the vertices with only one connection (degree q = 1). This procedure is then repeated until only nodes with degree q ≥ 2 are left. The removed nodes constitute the k = 1-shell and those remaining are the k = 2-core. At the next step all vertices with degree q = 2 removed, thus leaving the k = 3-core. The procedure is repeated iteratively. The maximum k-core (of index kS) is the set of vertices such that one more iteration of the procedure removes all of them. Notice that all vertices of the k-core of index k have degree larger than or equal to k. Figure 1 shows an example of the k-core decomposition performed on a small network of size N = 30 and largest degree qmax = 10.
SIR model on a star-graph
Let us consider the SIR process on a star network of size qmax + 1. The process starts from a randomly infected vertex and proceeds till all infected vertices become eventually removed. We want to compute the average final density of removed vertices at the end of an outbreak, which is given by R = r/(qmax + 1), where r is the total number of removed vertices. Let us define r = 1 + r*, where r* is the number of removed vertices from secondary infections. The outcome of the process will depend on whether the initial infected site is the hub or a leaf (a vertex of degree 1). If the infection starts in the hub, which will happen with probability ph = 1/(qmax + 1), the average number of secondary infected vertices will be 〈r*〉h = λqmax. On the contrary, the infection can start in a leaf with probability pl = qmax/(qmax + 1). In this case, the hub can become infected with probability λ and from there, spread the infection to the remaining qmax − 1 susceptible leaves. Therefore, in this case the average number of secondary infections is 〈r*〉l = λ[1 + λ(qmax − 1)]. The average total number of removed sites at the end of the spreading process will therefore be

where the last expression is valid in the limit of large qmax. From here it follows the expression for the average final density of infected vertices, equation (3), valid also in the limit of large qmax.
Quantitative features of the real networks considered
We consider in our analysis the following three real networks datasets:
Internet map at the Autonomous System level (AS). Map of the Internet collected at the Oregon route server. Vertices represent autonomous systems (aggregations of Internet routers under the same administrative policy), while edges represent the existence of border gateway protocol (BGP) peer connections between the corresponding autonomous systems37.
Pretty-good-privacy network (PGP). Social network defined by the users of the pretty-good-privacy (PGP) encryption algorithm for secure information exchange. Vertices represent users of the PGP algorithm. An edge between two vertices indicates that each user has signed the encryption key of the other38.
Actor collaboration network (Movies). Network of movie actor collaboration obtained from the Internet Movie Database (IMDB). Each vertex represents a movie actor. Two actors are joined by an edge if they have co-starred at least one movie39.
The relevant topological features of the different maps are summarized in Table 1.
References
Cohen, R., Havlin, S. & ben Avraham, D. Efficient immunization strategies for computer networks and populations. Phys. Rev. Lett. 91, 247901 (2003).
Leskovec, J., Adamic, L. A. & Huberman, B. A. The dynamics of viral marketing. ACM Trans. Web 1 May (2007).
Daley, D. J. & Kendall, D. G. Epidemics and rumours. Nature 204, 1118 (1964).
Albert, R. & Barabási, A.-L. Statistical mechanics of complex networks. Rev. Mod. Phys. 74, 47–97 (2002).
Dorogovtsev, S. N. & Mendes, J. F. F. Evolution of networks. Advances in Physics 51, 1079–1187 (2002).
Newman, M. The Structure and Function of Complex Networks. SIAM Review 45, 167 (2003).
Keeling, M. J. & Eames, K. T. D. Networks and epidemic models. J. R. Soc. Interface 2, 295–307 (2005).
Anderson, R. M. & May, R. M. Infectious diseases in humans (Oxford University Press, Oxford, 1992).
Barrat, A., Barthélemy, M. & Vespignani, A. Dynamical Processes on Complex Networks (Cambridge University Press, Cambridge, 2008).
Serrano, M. A., Boguñá, M., Pastor-Satorras, R. & Vespignani, A. Correlations in complex networks. In Large scale structure and dynamics of complex networks: From information technology to finance and natural sciences, Caldarelli, G. & Vespignani, A., editors, 35–66 (World Scientific, Singapore, 2007).
Boguñá, M., Pastor-Satorras, R. & Vespignani, A. Epidemic spreading in complex networks with degree correlations. In Statistical Mechanics of Complex Networks, Pastor-Satorras, R., Rubí, J. M. & Díaz-Guilera, A., editors, volume 625 of Lecture Notes in Physics. Springer Verlag, Berlin (2003).
Lloyd, A. L. & May, R. M. How viruses spread among computers and people. Science 292, 1316–1317 (2001).
Newman, M. E. J. Spread of epidemic disease on networks. Phys. Rev. E 66, 016128 (2002).
Pastor-Satorras, R. & Vespignani, A. Epidemic spreading in scale-free networks. Phys. Rev. Lett. 86, 3200–3203 (2001).
Boguñá, M., Pastor-Satorras, R. & Vespignani, A. Absence of epidemic threshold in scale-free networks with degree correlations. Phys. Rev. Lett. 90, 028701 (2003).
Wang, Y., Chakrabarti, D., Wang, C. & Faloutsos, C. Epidemic spreading in real networks: An eigenvalue viewpoint. In 22nd International Symposium on Reliable Distributed Systems (SRDS'03), 25–34. IEEE Computer Society, Los Alamitos, CA, USA (2003).
Gómez, S., Arenas, A., Borge-Holthoefer, J., Meloni, S. & Moreno, Y. Discrete-time markov chain approach to contact-based disease spreading in complex networks. Europhys. Lett. 89(3), 38009 (2010).
Van Mieghem, P. The N-intertwined SIS epidemic network model. Computing 93, 147–169 (2011).
Chung, F., Lu, L. & Vu, V. Spectra of random graphs with given expected degrees. Proc. Natl. Acad. Sci. USA 100, 6313–6318 (2003).
Castellano, C. & Pastor-Satorras, R. Thresholds for epidemic spreading in networks. Phys. Rev. Lett. 105, 218701 (2010).
Ganesh, A., Massoulié, L. & Towsley, D. The effect of network topology on the spread of epidemics. In IEEE INFOCOM, 1455–1466 (2005).
Chatterjee, S. & Durrett, R. Contact processes on random graphs with power law degree distributions have critical value 0. Annals of Probability 37, 2332–2356 (2009).
Durrett, R. Some features of the spread of epidemics and information on a random graph. Proc. Natl. Acad. Sci. USA 107, 4491–4498 (2010).
Kitsak, M. et al. Identification of influential spreaders in complex networks. Nat. Phys. 6, 888–893 (2010).
Seidman, S. B. Network structure and minimum degree. Social Networks 5, 269–287 (1983).
Pastor-Satorras, R. & Vespignani, A. Immunization of complex networks. Phys. Rev. E 65(3), 036104 Feb (2002).
Centola, D. The spread of behavior in an online social network experiment. Science 329(5996), 1194–1197 09 (2010).
Lü, L., Chen, D.-B. & Zhou, T. The small world yields the most effective information spreading. New Journal of Physics 13(12), 123005 (2011).
Arenas, A., Díaz-Guilera, A., Kurths, J., Moreno, Y. & Zhou, C. Synchronization in complex networks. Physics Reports 469(3), 93–153 (2008).
Catanzaro, M., Boguñá, M. & Pastor-Satorras, R. Generation of uncorrelated random scale-free networks. Phys. Rev. E 71, 027103 (2005).
Dorogovtsev, S. N., Goltsev, A. V. & Mendes, J. F. F. k-core organization of complex networks. Phys. Rev. Lett. 96, 040601 (2006).
Cohen, R., Erez, K., ben Avraham, D. & Havlin, S. Resilience of the internet to random breakdowns. Phys. Rev. Lett. 85(21), 4626–4628 Nov (2000).
Newman, M. E. J. The spread of epidemic disease on networks. Phys. Rev. E 66, 016128 (2002).
Newman, M. E. J. Assortative mixing in networks. Phys. Rev. Lett. 89, 208701 (2002).
Pastor-Satorras, R., Vázquez, A. & Vespignani, A. Dynamical and correlation properties of the Internet. Phys. Rev. Lett. 87, 258701 (2001).
Weber, S. & Porto, M. Generation of arbitrarily two-point-correlated random networks. Phys. Rev. E 76, 046111 (2007).
Pastor-Satorras, R. & Vespignani, A. Evolution and structure of the Internet: A statistical physics approach (Cambridge University Press, Cambridge, 2004).
Boguñá, M., Pastor-Satorras, R., Díaz-Guilera, A. & Arenas, A. Models of social networks based on social distance attachment. Phys. Rev. E 70, 056122 (2004).
Barabási, A.-L. & Albert, R. Emergence of scaling in random networks. Science 286, 509–512 (1999).
Acknowledgements
RP-S acknowledges financial support from the Spanish MICINN, under project No. FIS2010-21781-C02-01 and the Junta de Andalucía, under project No. P09-FQM4682, as well as additional support through ICREA Academia, funded by the Generalitat de Catalunya.
Author information
Authors and Affiliations
Contributions
Both authors designed the study, analyzed the data and contributed to writing the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Castellano, C., Pastor-Satorras, R. Competing activation mechanisms in epidemics on networks. Sci Rep 2, 371 (2012). https://doi.org/10.1038/srep00371
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00371
This article is cited by
-
Hyper-cores promote localization and efficient seeding in higher-order processes
Nature Communications (2023)
-
Spread of variants of epidemic disease based on the microscopic numerical simulations on networks
Scientific Reports (2022)
-
Influence control method on directed weighted signed graphs with deterministic causality
Annals of Operations Research (2022)
-
Patterns and risk of HIV-1 transmission network among men who have sex with men in Guangxi, China
Scientific Reports (2021)
-
The relative importance of structure and dynamics on node influence in reversible spreading processes
Frontiers of Physics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.