Abstract
Exposure of human populations to chronically elevated levels of ambient particulate matter air pollution < 2.5 μm in diameter (PM2.5) has been associated with an increase in lung cancer incidence. Over 70% of lung cancer cell lines exhibit promoter methylation of the tumor suppressor p16, an epigenetic modification that reduces its expression. We exposed mice to concentrated ambient PM2.5 via inhalation, 8 hours daily for 3 weeks and exposed primary murine alveolar epithelial cells to daily doses of fine urban PM (5 µg/cm2). In both mice and alveolar epithelial cells, PM exposure increased ROS production, expression of the DNA methyltransferase 1 (DNMT1) and methylation of the p16 promoter. In alveolar epithelial cells, increased transcription of DNMT1 and methylation of the p16 promoter were inhibited by a mitochondrially targeted antioxidant and a JNK inhibitor. These findings provide a potential mechanism by which PM exposure increases the risk of lung cancer.
Similar content being viewed by others
Introduction
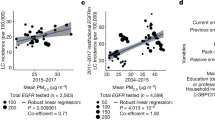
Each year, more than 160,000 people in the US and 1.4 million people worldwide die from lung cancer, which is the leading cause of cancer related death1. Exposure to cigarette smoke remains the most important cause of lung cancer, however, approximately 15% of lung cancers occur in never-smokers and lung cancer in non-smokers as a separate entity remains a leading cause of cancer mortality2,3. Epidemiologists studying the link between exposure to particulate matter air pollution (PM) and lung cancer have consistently observed a positive association3,4,5. In one study, Pope et al. reported that for every 10 µg/cm3 elevation in PM2.5 concentration there was an approximately 8% increased risk of lung cancer related mortality5.
Lung cancer is associated with several characteristic epigenetic changes; one of the most common is the methylation of the promoter for the tumor suppressor p16, which has been reported in >70% cell lines derived human non-small cell lung cancers6,7. Methylation of the p16 promoter is thought to play a critical role in lung cancer development by allowing the uncontrolled clonal expansion of premalignant lesions to cancer8,9. In sputum or cellular samples from smokers without lung cancer, smokers without malignancy, never smokers and lung cancer survivors, Belinsky and colleagues have identified hypermethylation of CpG islands in the promoter of p16 as an early event in the development of lung cancer, particularly in patients with a history of exposure to cigarette smoke8. Methylation of the p16 promoter is frequently associated with widespread changes in the methylation of other genes suggesting that promoter methylation is regulated by a common upstream pathway10.
DNA methylation in mammalian cells is catalyzed by members of the (cytosine-5)-DNA methyltransferase (DNMT) family. DNMT1 is thought to play a major role in the changes in DNA methylation observed in human cancer cells11 and an increase in DNMT1 abundance has been linked to cigarette smoke exposure induced lung carcinogenesis in mice and humans12. The c-jun-n-terminal protein kinase (JNK), a member of the mitogen activated protein kinase family is induced by oncogenes frequently observed in human lung cancers and upregulates the transcription of DNMT13,14,15. As we have previously found that exposure to PM induces apoptosis in alveolar epithelial cells through the mitochondrial oxidant-dependent activation of JNK16,17, we hypothesized that the PM induced activation of JNK might enhance DNMT1 transcription and p16 promoter methylation via a similar pathway.
RESULTS
Exposure to concentrated ambient PM2.5 results in methylation of the p16 promoter in the lungs of mice
We exposed mice to concentrated ambient PM2.5 or filtered air 8 hours daily, 5 days per week 3, 6 or 9 weeks (Supplementary Figure S1) after which we harvested the lungs for isolation of whole lung genomic DNA and measured methylation of the promoter for p16. Mean particle concentrations in the PM2.5 and filtered air chambers (measured daily at the beginning of the exposure) were 5.5×105 and 6.47×102 particles/cm3 respectively (Figure 1A). During the exposure, daily PM2.5 concentrations reported from a nearby Environmental Protection Agency Monitor averaged 11.55 µg/m3. We observed a similar increase in methylation of the p16 promoter in mice exposed to concentrated ambient PM2.5 at all three time points (Figure 1B, combined data in Figure 1C). We observed a similar increase in methylation of the promoter for the matrix metalloproteinase-2 (MMP2) gene (Figure 1C). Promoter methylation of the p16 and MMP promoters, along with those of 4 other genes in the sputum of a high risk smoking cohort was shown to increase the risk for developing lung cancer18.
Inhalation of concentrated ambient PM 2.5 results in hypermethylation of the p16 promoter in the lungs of mice.
A VACES system was used to generate concentrated ambient PM2.5, which was delivered to two identical murine exposure chambers, one of which was equipped with a teflon filter at the chamber inlet. (A) Mean particle concentrations over the duration of the exposure in the ambient air (inlet to the VACES), the inlet to the concentrated ambient PM2.5 chamber and the inlet to the filtered air chamber distal to the filter. (B) Mice were placed in either the concentrated ambient PM2.5 or filtered air chamber 8 hours daily, five days per week. At the indicated times the ratio of metyhlated/unmethylated p16 was measured in DNA isolated from whole methylated lung homogenates using methylation-specific PCR. Each bar represents 3–4 animals exposed to concentrated ambient PM2.5 or filtered air. (C) Fold change ratios in the metyhlated/unmethylated p16 and MMP2 genes of the 10 animals exposed to concentrated ambient PM2.5 or filtered air. P values (unpaired two tailed t-tests) are indicated in italics above the bars.
Exposure to PM causes a dose-dependent increase in mitochondrial Reactive Oxygen Species generation and cell death in primary alveolar epithelial cells from mice
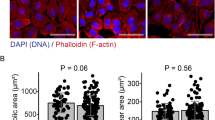
We and others have reported an increase on cellular ROS production after PM exposure in alveolar epithelial cells including primary human alveolar type II cells16,17. We isolated primary alveolar type II cells from mice and cultured them at air-liquid interfaces. Because these cells differentiate in culture, we refer to them hereafter as alveolar epithelial cells. After 48 hours in culture, we infected them with an adenovirus encoding a mitochondrially localized oxidant sensitive probe (mito-Ro-GFP). Twenty-four hours after infection, we pretreated the cells with a mitochondrially targeted antioxidant (Mito-CP, 50 µM) 30 minutes prior to treatment with increasing doses of PM and measured oxidation of the probe 4 hours later using flow cytometry (Figure 2A). To confirm our previous finding that mitochondrially generated ROS are required for PM induced cell death in alveolar epithelial cells16,17, we treated uninfected cells with increasing doses of PM in the presence or absence of Mito-CP (50 µM) and measured cell death 24 hours later. Cell death in response to high dose PM (50 µg/cm2) was prevented by treatment with Mito-CP, however, at a dose of 5 µg/cm2, PM did not cause significant cell death even in untreated cells (Figure 2B). To determine the lowest dose of Mito-CP that would scavenge mitochondrial ROS generated in response to this nonlethal dose of PM, we measured mito-Ro-GFP oxidation in cells treated with PM (5 µg/cm2) in the presence of increasing doses of Mito-CP. Mito-CP at a dose of 5 µM prevented oxidation of the mitochondrially localized probe, while the control cation used to deliver Mito-CP to the mitochondria, TPP, had no effect even at a dose of 50 µM (Figure 2C).
PM exposure induces mitochondrial ROS production and apoptosis in cultured mouse lung epithelial cells.
(A) Primary murine alveolar epithelial cells were infected with an adenovirus (5 pfu/cell) encoding a mitochondrially-localized-oxidant-sensitive GFP probe (mito-Ro-GFP) and 48 hours later were pretreated Mito-CP (50 µM) or vehicle (DMSO) followed 30 minutes later by PBS or PM (5 or 50 µg/cm2) and oxidation of the probe was measured using flow cytometry 4 hours later. (B) Primary alveolar epithelial cells were pretreated with Mito-CP (5 µM) or the control cation 30 minutes before treatment with PM (5 or 50 µg/cm2) and cell death was measured 24 hours later using an ELISA that detects DNA fragmentation. (C) Cells were infected with the Mito-Ro-GFP probe as in (A) and 48 hours later were pretreated with increasing doses of Mito-CP or the control cation TPP 30 minutes before exposure to PM (5 µg/cm2) and oxidation of the probe was measured 4 hours later. N ≥ 3, P values (unpaired two tailed t-tests) are indicated in italics above the bars.
Long-term exposure to low dose PM induces methylation of the p16 promoter in primary murine alveolar epithelial cells
We treated primary murine alveolar epithelial cells daily with Mito-CP (5 µM) followed 30 minutes later by PM (5 µg/cm2). After 10 days of treatment, we isolated the DNA from the cells for measurement p16 promoter methylation. Long-term treatment with PM was associated with methylation of the p16 promoter, which was prevented by daily treatment with Mito-CP but not the control cation TPP (Figure 3). PM-induced methylation of the p16 promoter was also prevented by daily treatment with the DNA-methyltransferase inhibitor 5-azacytidine (5-aza) (1µM) 30 minutes before PM administration (Figure 3).
Exposure of primary murine alveolar epithelial cells to repeated doses of PM results in mitochondrial-oxidant-dependent methylation of the p16 promoter.
Primary mouse alveolar epithelial cells were cultured at an air-liquid interface. After 2 days in culture, the cells were treated daily with Mito-CP (5 µM), 5-azacytidine (5-aza) (1 µM) or the cation control TPP (5 µM) followed 30 minutes later by PM (5 μg/cm2/day). After 10 days of treatment, DNA was isolated from the cells and methylation of the p16 promoter was measured using methylation specific PCR. N = 4 for all measures, P values (ANOVA with Bonferroni correction for multiple comparisons) are indicated in italics above the bars.
Exposure to PM is associated with mitochondrial-oxidant and JNK-dependent transcription and expression of DNMT1
Promoter methylation in mice and humans is catalyzed by three DNA methyltransferases, DNMT1, 3a and 3b. We measured the levels of mRNA encoding these three proteins in alveolar epithelial cells treated daily for 10 days with PM (5 μg/cm2). Treatment with PM increased the level of mRNA encoding DNMT1 but had no effect on the levels of DNMT3a and DNMT3b (Figure 4A). Treatment with PM also increased the levels of DNMT1 protein in whole cell lysates (Figure 4B). The daily administration of Mito-CP (5 µM) 30 minutes before PM treatment prevented the PM-induced increase in DNMT1 transcription and abundance (Figure 4A,B, TPP (5 µM) as control).
Exposure of primary murine alveolar epithelial cells to repeated doses of PM increases DNMT1 mRNA and protein.
Primary mouse alveolar epithelial cells were cultured at an air-liquid interface. After 2 days in culture, the cells were treated daily with PM (5 μg/cm2/day). After 10 days of treatment the cells were lysed and (A) mRNA encoding DNMT1, DNMT3a and DNMT3b (RT-qPCR) and (B) DNMT1 protein (immunoblot) were measured. N≥4 for all measures, P values (ANOVA with Dunnett correction for multiple comparisons) are indicated in italics above the bars.
Exposure to PM is associated with mitochondrial oxidant-mediated activation of JNK, which is required for methylation of the p16 promoter
We exposed primary murine alveolar epithelial cells to PM and measured JNK activation 30 minutes after exposure using an antibody that recognizes phosphorylated and total JNK. Treatment with PM increased the phosphorylation and kinase activity of JNK (Figure 5A,B). Pretreatment of the cells with the JNK inhibitor SP600125 (20 µM) or Mito-CP (5 µM) prevented the PM-induced activation of JNK (Figure 5A,B). Treatment with the control cation (TPP) had no effect on JNK activation. We then measured DNMT1 mRNA expression and p16 methylation in primary murine alveolar epithelial cells treated daily with SP600125 (20 μM) followed 30 minutes later by PM. Inhibition of JNK with SP600125 prevented the PM-induced increase in expression of DNMT1 protein (figure 5C) and p16 methylation in these cells (Figure 5D). Consistent with our findings in vivo, inhibition of either mitochondrial ROS with mito-CP or JNK with SP600125 also prevented the PM-induced methylation of the MMP2 promoter (Figure 5E, TPP (5 µM) as control for Mito-CP). Treatment with an inhibitor of the extracellular-signal related kinase pathway U0126 did not affect the increase in DNMT1 protein abundance or p16 promoter methylation (Supplementary Figure S2).
JNK is required for the PM-induced hypermethylation of the p16 promoter in primary murine alveolar epithelial cells.
(A) Primary mouse alveolar epithelial cells were cultured for 2 days at an air-liquid interface and then treated with Mito-CP (5 µM) or vehicle followed 30 minutes later by PM (5 μg/cm2) and 30 minutes later the levels of phosphorylated and total JNK were evaluated by immunoblot (densitometry below). (B) Primary mouse alveolar epithelial cells were cultured for 2 days at an air-liquid interface and then treated with Mito-CP (5 µM), SP600125 (20 µM) or the appropriate vehicle controls followed 30 minutes later by PM (5 μg/cm2) and 2 hours later JNK kinase activity in cell lysates was measured, (C,D) Cells were treated daily for 10 days with SP600125 (20 µM) or vehicle followed 30 minutes later by PM (5 μg/cm2/day). After 10 days, the levels of DNMT1 were measured in cell lysates by immunoblotting and methylation of the p16 and MMP2 promoters was measured in cellular DNA using methylation-specific PCR. N≥4 for all measures, P values (ANOVA with Dunnett correction for multiple comparisons) are indicated in italics above the bars.
Exposure of mice to concentrated ambient PM2.5 increases lung oxidant stress and the levels of DNMT1
We exposed mice to concentrated ambient PM2.5 or filtered air 8 hours daily for 3 days and looked for evidence of lung oxidant stress by immunostaining lung sections for nuclear 7,8-Dihydro-8-Oxo-2′-deoxyguanosine (8-oxo-DG). Exposure to concentrated ambient PM2.5 was associated with an increase in the number of nuclei staining positively for 8-oxo-DG (Figure 6A). In identically treated mice, we measured mRNA encoding DNMT1, DNMT3a and DNMT3b (Figure 6B). Similar to our observations in alveolar epithelial cells, we found an increase in DNMT1 mRNA and protein (Figure 6B and 6C) but no change in DNMT3a or DNMT3b in the lungs of mice exposed to CAPs.
Exposure to Concentrated Ambient PM 2.5 (CAPS) induces oxidant stress and increases DNMT1 mRNA and protein in the lungs of mice.
Mice were exposed to concentrated ambient PM2.5 (PM2.5) or to filtered air for 3 days and (A) lung sections were stained for the presence of 8-oxodG (representative positive nuclei indicated by arrows, left panel), the bar graph represents the number of positive cells in 10 random fields from each section (200X) (B) In lung homogenates from mice exposed to concentrated ambient PM2.5 (PM2.5) or to filtered air for 9 weeks, mRNA levels of DNMT1,DNMT3a or DNMT3b (RT-qPCR) and (C) DNMT1 protein (immunoblot) were measured. N = 4 for all measures, P values (two tailed t-test) are indicated in italics above the bars.
Discussion
We found that exposure of mice to concentrated ambient PM2.5 resulted in lung oxidant stress, increased transcription of DNMT1 and caused hypermethylation of the p16 promoter in the lungs of mice. In primary murine alveolar epithelial cells, we found that enhanced transcription of DNMT1 and methylation of the p16 promoter in response to PM was inhibited by a mitochondrially targeted antioxidant and an inhibitor of the JNK pathway. Our data suggest that PM-induced mitochondrial oxidant stress activates JNK to transcriptionally upregulate the expression of DNMT1. These findings provide a potential mechanism explaining the observed link between exposure to PM and the development of lung cancer.
Lung cancer is associated with characteristic epigenetic changes, which include global hypomethylation accompanied by selective hypermethylation of CpG islands in tumor suppressor genes19. Methylation of the p16 promoter can be detected in >70% cell lines derived from human NSCLCs and in the sputum of smokers before the development of lung cancer. In the cell, p16 sequesters the cyclin D-dependent kinases CDK4 and CDK6 causing a G1-phase cell-cycle arrest and senescence6,7,9. Therefore, methylation-induced suppression of p16 allows for the clonal expansion of malignant cells9. In mice, genetic loss of p16Ink4a results in spontaneous carcinogenesis and an increased incidence of carcinogen induced cancers21. However, the partial loss of function induced by promoter methylation we observed in mice exposed to concentrated ambient PM2.5 is likely insufficient to induce lung cancer in the absence of an additional genetic or environmental stimulus9. In support of this hypothesis, Belinsky and colleagues have identified the detection of hypermethylation of CpG islands in the promoter of p16 in the sputum as a risk factor for the development of lung cancer, particularly in patients with a history of cigarette smoke exposure8,10,18,20.
Mammalian DNA is modified by methylation of 60–80% of the cytosine residing in the dinucleotide sequence CpG through a reaction catalyzed by the DNMTs11,22. Mammalian cells express three DNMTs, DNMT1, DNMT3a and DNMT3b11. DNMT1 is widely expressed in human and murine cells and is required for the maintenance of methylation patterns during cellular replication23,24. In contrast, DNMT3a and DNMT3b participate in de novo methylation during early development22,25,26. Studies in transgenic and knockout mice suggest that DNMT1 plays an essential role in tumorigenesis11,27. Blunted expression of DNMT1 using hemizygous knockout mice is associated with delayed and less severe tumor formation in murine models of cancer driven by the loss of tumor suppressor genes27,28. DNMT1 expression has been reported to be increased in patients with lung cancer, with particularly high levels observed in patients who were smoking at the time of their resection12. We found a relative increase in DNMT1 expression in mice exposed to concentrated ambient PM2.5 and in primary alveolar epithelial cells from mice following prolonged exposure to PM. If PM exposure increases p16 promoter methylation by inducing the expression of DNMT1, then methylation of CpG islands should occur synchronously and early in the development of lung cancer. In support of this hypothesis, Belinsky and colleagues discovered that CpG island hypermethylation of three genes: p16, MMP2 and Basic Helix-loop-helix Family Member e23 (BHLHB4) in lung cancer cell lines was associated with a pattern of widespread CpG island hypermethylation10. We found evidence for promoter methylation in the MMP2 gene in mice and alveolar epithelial cells exposed to PM.
The c-Jun NH(2)-terminal kinase (JNK) is a member of an evolutionarily conserved subfamily of mitogen-activated protein (MAP) kinases29,30. In lung epithelial cells, we reported that exposure to PM2.5 resulted in the generation of ROS from site III in the mitochondrial electron transport chain16,17. These ROS activated apoptosis signaling kinase 1 (ASK1), a MAP kinase kinase kinase (MAPKKK) that is activated when freed from its normal binding partner thioredoxin upon its oxidation to activate the JNK and p38 MAPKs31. The ASK1-mediated activation of JNK, resulted in the phosphorylation and activation of p53, which increased the transcription of the proapoptotic protein NOXA to activate BAX/BAK and induce cell death16,17,32. Here we found that exposure to a lower dose of PM acted via a similar pathway to induce the transcription of DNMT1. These observations are consistent with the known regulation of the DNMT1 gene, which contains several conserved Activator Protein 1 (AP1) binding sites in its promoter14,33,34.
We and others have reported that mitochondrially-derived ROS play diverse and critical roles in the metabolic adaptation of cells to environmental stress. Sometimes these signaling events are maladaptive. For example, we recently reported that oncogene induced tumors require mitochondrial ROS to drive biosynthetic pathways involved in cellular proliferation35 and in this report we show that the inappropriate generation of mitochondrial ROS in response to particulate matter air pollution activates stress kinase pathways to suppress the transcription of tumor suppressors. In contrast, we have also found that mitochondrial ROS initiate adaptive signaling pathways that may prevent cancer, for example the terminal differentiation of human mesenchymal stem cells36. Collectively, these data suggest that while mitochondrially targeted antioxidants might be useful as therapeutics for cancer they may not be effective for cancer prevention.
A major strength of our study is our observation that mice exposed to concentrated ambient PM2.5 show increased p16 promoter methylation in the lung and increased transcription of DNMT1. The concentrations of PM in the concentrated ambient PM2.5 exposure chamber are about 10 fold higher than ambient levels outside our laboratories which are located in an urban area near several major roadways. Based on data reported from EPA monitors near our laboratories during the period of exposure, we estimate chamber PM concentrations between 100–120 µg/m3. Even these elevated levels are substantially less than those that would be encountered in many world cities1. Furthermore, the mice in this study were exposed to PM for 8 hours daily, 5 days per week, a schedule that would mimic a typical exposure for an outdoor worker.
There are several important limitations of our study, which we hope will prompt further investigation. Firstly, while we were able to validate some of the key mechanistic findings of our cell culture model in the lungs of mice, loss of function studies in components of the mito-ROS-JNK-DNMT1 pathway will be required to confirm the importance of this pathway in vivo. Secondly, we are unable to definitively localize the DNMT1 and p16 promoter methylation signal to the alveolar epithelium in the intact lung. Thirdly, we observed that exposure to PM activated the JNK MAPK pathway within minutes of exposure and that increased levels of DNMT1 were present in cells within a day of exposure. However, we were only able to detect methylation of the p16 promoter after 10 consecutive days of exposure to PM. We speculate that additional inhibitory mechanisms prevent promoter methylation, which are only overcome after repeated and prolonged exposure. Finally, individuals in the population vary widely in terms of their genetic risk for cancer and previous exposure to environmental carcinogens. In our murine model, we examined a relatively small number of animals in a single inbred strain of mice and we limited our analysis to male mice as female sex has been shown to be protective in murine models of gastrointestinal and lung cancer where inflammation plays an important role37.
We conclude that exposure to PM results in a mitochondrial-oxidant and JNK-mediated increase in the transcription and abundance of DNMT1 and increased methylation of the p16 promoter in the lung epithelium. Aberrant upregulation of DNMT1 could result in early and coordinated hypermethylation of key tumor suppressor genes in PM exposed patients, increasing the risk for the development of lung cancer. As these epigenetic changes are reversible and precede the development of lung cancer, they offer a novel target to prevent its development in high risk individuals.
Methods
Animals and alveolar type-2 cell isolation and cell culture
The protocol for the use of mice was approved by the Animal Care and Use Committee at Northwestern University. Six- to 8-wk-old (weighing 20–25 g)-male-C57BL/6 mice (wild-type mice) were purchased from Charles River Labs (Roanoke, Illinois). Type II alveolar epithelial cells (AECs) were isolated from pathogen-free male mice as previously described38. Briefly, the lungs were perfused via the pulmonary artery, lavaged and digested with elastase (3 U ml−1; Worthington Biochemical, Freehold, NJ). Alveolar type II cells were purified by differential adherence to immunoglobulin G and cell viability was assessed by trypan-blue exclusion (>95%). Cells were resuspended in Dulbecco's modified Eagle's medium (Sigma-Aldrich Inc., St. Louis, MO) containing 10% fetal bovine serum (Hyclone, Logan, UT) with 2 mM glutamine, 100 U ml−1 penicillin, 0.25 μg ml−1 amphotericin B and 100 μg ml−1 streptomycin. Cells were seeded in permeable transwell supports and maintained in air-liquid interface from the second day of culture until the end of the experiment while incubated in a humidified atmosphere of 5% CO2 and 95% air at 37°C. The purity of the AECs was determined to be 90±5% by immunostaining for surfactant protein C.
Reagents
Chemicals: The mitochondrially targeted anti-oxidant Mito-carboxy proxyl (Mito-CP) is a synthetic superoxide dismutase mimetic that accumulates in the mitochondria and has been previously described35,39. SP600125 was purchased from Calbiochem (Darmstadt, Germany). 5-azacytidine (5-aza) was purchased from Sigma-Aldrich (St Louis. MO). Antibodies: Anti-phosphorylated and total JNK antibodies were purchased from Cell Signaling (Boston, MA, catalogue numbers 4668 and 9252, respectively) Anti-c-Jun was purchased from Sigma-Aldrich (catalogue number J4399). Anti 8-OxodG was purchased from Trevigen (Gaithersburg, MD, catalogue number 4354-MC-50). Anti-DNMT1 was purchased from ABCAM (Cambridge, MA. catalogue number AB11891).
Particulate Matter (PM)
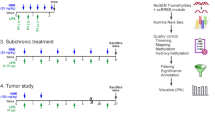
The PM for in vitro experiments was obtained from the National Institutes of Standards and Technology (Gaithersburg, MD, USA; NIST SRM 1649: Filter-Based Fine Particulate Material). The characteristics of the PM have been previously described40. For in vivo experiments, concentrated ambient PM2.5 (CAPs) was obtained using a Versatile Aerosol Concentration and Enrichment System (VACES) as previously described41,42,43. The VACES concentrates PM2.5 collected from the ambient air outside our laboratory, which is located in an urban environment with high levels of traffic. Wild-type mice were placed in the exposure chamber for 8 hours daily, five days per week for up to 9 weeks in the autumn of 2010. Control mice were identically treated except that a teflon filter was placed in the chamber inlet. The mean particle concentration in the chamber was monitored in real-time using a TSI 3775 particle counter.
Mitochondrial ROS production
We employed a mitochondrial matrix localized oxidant-sensitive ratiometric probe (mito-Ro-GFP) as previously described17. Primary murine alveolar epithelial cells were infected with 5 PFU/cell of an adenovirus containing the probe 48 h before exposure to PM. Oxidation of the mito-Ro-GFP probe was assessed by removing the cells from the plate using trypsin and transferring equal aliquots of the resulting suspension to tubes containing medium alone or medium containing 1 mM dithiothreitol (DTT) or 1 mM hydrogen peroxide (H2O2). The ratio of fluorescence (emission of 535 nm) at excitations of 405 and 488 nm was measured by flow cytometry in 5,000 cells/condition using a DakoCytomation CyAn high speed multilaser droplet cell sorter. The oxidation state was calculated as the completely reduced ratio (dithiothreitol) less the untreated value divided by the difference in the ratio observed with 1 mM dithiothreitol and 1 mM H2O2.
Apoptosis
Apoptosis was measured with a commercially available double sandwich colorimetric ELISA that detects nucleosomal fragmentation (Roche Applied Science, Cat # 11774425001) according to the manufacturer's instructions.
JNK activation and JNK Kinase assay
The phosphorylation of JNK was assessed by protein immunoblot as previously described17. Kinase assays to measure the activity of c-jun were performed using a commercially available assay (Cell Signaling; catalog number 9810)44. Briefly, cell lysates were obtained and incubated with agarose beads linked to a c-jun fusion protein. The resulting c-jun/JNK complexes were precipitated by centrifugation and sequential washing of the beads. The beads were then incubated with 2 μg of myelin basic protein and 100 μM ATP for 60 min at 30°C in 35 mM Tris, pH 7.5, 10 mM MgCl2, 5 mM EGTA, 1 mM CaCl2 and 10 mM β-glycerolphosphate. The reactions were stopped and c-jun was released from the beads by adding SDS sample buffer and boiling the samples. The proteins were separated by SDS-PAGE and detected by immunoblot.
8-oxo-dG Immunostaining
After 9 weeks of exposure to concentrated ambient PM2.5, isolated mouse lungs were inflated with 4% paraformaldehyde (15 cm H2O) then fixed for 24 hours and paraffin embedded and cut in 5 µm sections. Tissues were deparaffinized and treated with Proteinase K for 20 minutes and the slides were placed in citrate buffer on a hot plate for 2–5 minutes for antigen retrieval. The slides were then left to cool at room temperature, rinsed with deionized water and blocked for 1 hr with 1% goat serum and then incubated overnight with Anti-8-oxo-dG Monoclonal Antibody (1:100). The slides were then treated with goat anti-mouse IgG conjugated to biotin at 37°C for 60 minutes and then incubated with AP-conjugated streptavidin at 37°C in a light-tight box in a humid chamber for 60 minutes. Slides were incubated with 3,3′ Diaminobenzidine (DAB) for 15 minutes, rinsed with TBS-T and mounted. The tissues were imaged with a Nikon Eclipse T2000 microscope and the number of positive nuclei per field was counted with the ImageJ software (NIH).
Nucleic Acid Isolation and Methylation-Specific PCR
Lung DNA was isolated from paraffin embedded samples by hydrating a 20 µm section with xylene followed by two passes of 100% ethanol, one of 90%, 80% and 70% ethanol followed by digestion with proteinase K in 1% SDS. AT2 cell DNA was directly extracted by affinity columns using the Nucleospin tissue kit, following the manufacturer's recommendations (Clontech, Mountain View. CA). The methylation state of the p16 gene was determined by methylation-specific PCR (MSP) as described elsewhere8,45. Briefly, genomic DNA was modified by treatment with sodium bisulfite, which converts all unmethylated cytosines to uracil, then to thymidine during the subsequent PCR step. Before PCR, 200 ng of purified DNA was treated 15 minutes with 0.3 M NaOH and treated with a solution containing 5 M sodium bisulphite, 2.5 M sodium metabisulphite and 100 mM hydroquinone for 2.5 hr at 50°C. The modified DNA was desalted, precipitated with ethanol and finally resuspended in 20 μl of Tris-EDTA buffer. Two sets of primers were used to amplify each region of interest: one pair recognizes a sequence in which CpG sites are unmethylated (bisulfite modified to UpG) and the other recognizes a sequence in which CpG sites are methylated (unmodified by bisulfate). Treated DNA (200 ng) was used as the template and DNA amplification was measured using SYBR Green qPCR by the delta-delta Ct (ΔΔCt) method using the following primer sequences45:
Unmethylated p16:
Forward: AAT TTG AGG AGA GTT ATT TG.
Reverse: AAA TCA AAA TAC AAC CAA AAA A.
p16:
Forward: AAT TCG AGG AGA GTT ATT TG.
Reverse: AAA TCG AAA TAC GAC CGA AA.
Unmethylated MMP2:
Forward: AGAGATTTTAGGGTGATATGTGG
Reverse: CACCAAAAAACTTAATAATAAACAAT
Methylated MMP2:
Forward: AGTTAGAGATTTTAGGGTGATACGC
Reverse: GCCGAAAAACTTAATAATAAACGAT
Real-time reverse transcriptase PCR measurement of RNA
Wild-type mice were exposed to concentrated PM2.5 or filtered air for 9 weeks and the lungs were harvested for isolation of total RNA using a commercially available system (TRIzol, Invitrogen, Carlsbad, CA, USA), according to the manufacturer's instructions. Primary alveolar epithelial cells were exposed for 10 days to PBS or 5 μg/cm2 of PM2.5 and The resulting mRNA was reverse transcribed using Moloney murine leukemia virus reverse transcriptase and random decamers, using a commercially available system, according to the manufacturer's instructions (Ambion, Austin TX, USA). RT-qPCR reactions were performed using IQ SYBR Green-superscript (Bio-Rad Laboratories) with the primers listed below and analyzed on an IQ5 Real-Time PCR Detection System. Cycle threshold (Ct) values were normalized to Ct values for ribosomal protein 18 s and data were analyzed using the ΔΔCt method46. The primer sequences used were previously described32 and are the following:
DNMT1:
Forward: CCA AGC TCC GGA CCC TGG ATG TGT;
Reverse: TGA CAC CAT CTG TTC TTT CAG CTT CAC.
DNMT3a:
Forward: CGA GGC CGG TAG TAG TCA CAG TAG;
Reverse: GCA CCT ATG GGC TGC TGC GAA GAC G.
DNMT3b:
Forward: CTG CCT CCA ATC ACC AGG TCG ATT G;
Reverse: CAA GGA GGG CGA CAA CCG TCC ATT.
18 s: forward:
TGG CTC ATT AAA TCA GTT ATG GT,
Reverse: GTC GGC ATG TAT TAG CTC TAG
Statistical Analysis
Data are presented as means ± SEM. Student's t tests were performed to compare experimental data with appropriate controls (as indicated in each figure legend). For comparisons involving more than two groups, we performed an analysis of variance (ANOVA). When the ANOVA indicated a significant difference, individual differences between groups were explored with Dunnett or Bonferroni corrected t-tests. Statistical significance was determined at a value of P≤0.05.
References
Murray, C. et al. The World Health Report 2002 Reducing Risks, Promoting Healthy Life. (eds. Campanini, B., & Haden, A. ) (World Health Organization, Geneva, Switzerland, 2002).
Jemal, A. et al. Annual Report to the Nation on the Status of Cancer, 1975–2005, Featuring Trends in Lung Cancer, Tobacco Use and Tobacco Control. Journal of the National Cancer Institute 100, 1672–1694 (2008).
Samet, J. M. et al. Lung Cancer in Never Smokers: Clinical Epidemiology and Environmental Risk Factors. Clinical Cancer Research 15, 5626–5645 (2009).
Nicolich, M. J. et al. Urban Air Pollution and Lung Cancer in Stockholm. Epidemiology 12, 590–592 (2001).
Pope, C. A., 3rd et al. Lung cancer, cardiopulmonary mortality and long-term exposure to fine particulate air pollution. .Jama 287, 1132–1141 (2002).
Kamb, A. et al. A cell cycle regulator potentially involved in genesis of many tumor types. Science 264, 436–440 (1994).
Merlo, A. et al. 5[prime] CpG island methylation is associated with transcriptional silencing of the tumour suppressor p16/CDKN2/MTS1 in human cancers. Nat Med 1, 686–692 (1995).
Belinsky, S. et al. Aberrant methylation of p16INK4a is an early event in lung cancer and a potential biomarker for early diagnosis. Proceedings of the National Academy of Sciences of the United States of America 95, 11891 (1998).
Gil, J. & Peters, G. Regulation of the INK4b-ARF-INK4a tumour suppressor locus: all for one or one for all. Nat Rev Mol Cell Biol 7, 667–677 (2006).
Tessema, M. et al. Concomitant promoter methylation of multiple genes in lung adenocarcinomas from current, former and never smokers. Carcinogenesis 30, 1132–1138 (2009).
Robert, M.-F. et al. DNMT1 is required to maintain CpG methylation and aberrant gene silencing in human cancer cells. Nat Genet 33, 61–65 (2003).
Lin, R.-K. et al. The tobacco-specific carcinogen NNK induces DNA methyltransferase 1 accumulation and tumor suppressor gene hypermethylation in mice and lung cancer patients. The Journal of Clinical Investigation 120, 521–532 (2010).
Takahashi, H., Ogata, H., Nishigaki, R., Broide, D. H. & Karin, M. Tobacco Smoke Promotes Lung Tumorigenesis by Triggering IKK[beta]- and JNK1-Dependent Inflammation. Cancer Cell 17, 89–97 (2010).
Bigey, P., Ramchandani, S., Theberge, J., Araujo, F. D. & Szyf, M. Transcriptional regulation of the human DNA Methyltransferase (dnmt1) gene. Gene 242, 407–418 (2000).
Cellurale, C. et al. Requirement of c-Jun NH2-Terminal Kinase for Ras-Initiated Tumor Formation. Mol. Cell. Biol. 31, 1565–1576 (2011).
Soberanes, S. et al. p53 mediates particulate matter-induced alveolar epithelial cell mitochondria-regulated apoptosis. Am J Respir Crit Care Med 174, 1229–1238 (2006).
Soberanes, S. et al. Mitochondrial complex III-generated oxidants activate ASK1 and JNK to induce alveolar epithelial cell death following exposure to particulate matter air pollution. J Biol Chem 284, 2176–2186 (2009).
Belinsky, S. A. et al. Promoter Hypermethylation of Multiple Genes in Sputum Precedes Lung Cancer Incidence in a High-Risk Cohort. Cancer Res 66, 3338–3344 (2006).
Esteller, M. Epigenetics in Cancer. New England Journal of Medicine 358, 1148–1159 (2008).
Belinsky, S. A. et al. Aberrant Promoter Methylation in Bronchial Epithelium and Sputum from Current and Former Smokers. Cancer Research 62, 2370–2377 (2002).
Serrano, M. et al. Role of the INK4a Locus in Tumor Suppression and Cell Mortality. Cell 85, 27–37 (1996).
Szyf, M. The role of DNA hypermethylation and demethylation in cancer and cancer therapy. Curr Oncol 15, 72–75 (2008).
Leonhardt, H., Page, A. W., Weier, H.-U. & Bestor, T. H. A targeting sequence directs DNA methyltransferase to sites of DNA replication in mammalian nuclei. Cell 71, 865–873 (1992).
Pradhan, S., Bacolla, A., Wells, R. D. & Roberts, R. J. Recombinant Human DNA (Cytosine-5) Methyltransferase. Journal of Biological Chemistry 274, 33002–33010 (1999).
Okano, M., Xie, S. & Li, E. Cloning and characterization of a family of novel mammalian DNA (cytosine-5) methyltransferases. Nat Genet 19, 219–220 (1998).
Aoki, A. et al. Enzymatic properties of de novo-type mouse DNA (cytosine-5) methyltransferases. Nucleic Acids Research 29, 3506–3512 (2001).
Cormier, R. T. & Dove, W. F. Dnmt1N/+ Reduces the Net Growth Rate and Multiplicity of Intestinal Adenomas in C57BL/6-Multiple Intestinal Neoplasia (Min)/+ Mice Independently of p53 but Demonstrates Strong Synergy with the Modifier of Min 1AKR Resistance Allele. Cancer Research 60, 3965–3970 (2000).
Laird, P. W. et al. Suppression of intestinal neoplasia by DNA hypomethylation. Cell 81, 197–205 (1995).
Chang, L. & Karin, M. Mammalian MAP kinase signalling cascades. Nature 410, 37–40 (2001).
Kyriakis, J. M. & Avruch, J. Mammalian mitogen-activated protein kinase signal transduction pathways activated by stress and inflammation. Physiol Rev 81, 807–869 (2001).
Ichijo, H. et al. Induction of apoptosis by ASK1, a mammalian MAPKKK that activates SAPK/JNK and p38 signaling pathways. Science 275, 90–94 (1997).
Urich, D. et al. Proapoptotic Noxa is required for particulate matter-induced cell death and lung inflammation. FASEB J 23, 2055–2064 (2009).
Rouleau, J., MacLeod, A. R. & Szyf, M. Regulation of the DNA Methyltransferase by the Ras-AP-1 Signaling Pathway. Journal of Biological Chemistry 270, 1595–1601 (1995).
Bakin, A. V. & Curran, T. Role of DNA 5-Methylcytosine Transferase in Cell Transformation by fos. Science 283, 387–390 (1999).
Weinberg, F. et al. Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proceedings of the National Academy of Sciences 107, 8788–8793 (2010).
Tormos Kathryn, V. et al. Mitochondrial Complex III ROS Regulate Adipocyte Differentiation. Cell Metabolism 14, 537–544 (2011).
Li, N., Grivennikov Li, N., Grivennikov Sergei, I. & Karin, M. The Unholy Trinity: Inflammation, Cytokines and STAT3 Shape The Cancer Microenvironment. Cancer Cell 19, 429–431 (2011).
Budinger, G. R. et al. Proapoptotic Bid is required for pulmonary fibrosis. Proc Natl Acad Sci U S A 103, 4604–4609 (2006).
Dhanasekaran, A. et al. Mitochondria superoxide dismutase mimetic inhibits peroxide-induced oxidative damage and apoptosis: Role of mitochondrial superoxide. Free Radical Biology and Medicine 39, 567–583 (2005).
Huggins, F. E., Huffman, G. P. & Robertson, J. D. Speciation of elements in NIST particulate matter SRMs 1648 and 1650. Journal of Hazardous Materials 74, 1–23 (2000).
Sun, Q. et al. Long-term air pollution exposure and acceleration of atherosclerosis and vascular inflammation in an animal model. Jama 294, 3003–3010 (2005).
Sioutas, C., Koutrakis, P. & Burton, R. M. A technique to expose animals to concentrated fine ambient aerosols. Environ Health Perspect 103, 172–177 (1995).
Maciejczyk, P. et al. Effects of subchronic exposures to concentrated ambient particles (CAPs) in mice. II. The design of a CAPs exposure system for biometric telemetry monitoring. Inhal Toxicol 17, 189–197 (2005).
Vadasz, I. et al. AMP-activated protein kinase regulates CO2-induced alveolar epithelial dysfunction in rats and human cells by promoting Na,K-ATPase endocytosis. J Clin Invest 118, 752–762 (2008).
Cui, X., Wakai, T., Shirai, Y., Hatakeyama, K. & Hirano, S. Chronic Oral Exposure to Inorganic Arsenate Interferes with Methylation Status of p16INK4a and RASSF1A and Induces Lung Cancer in A/J Mice. Toxicol. Sci. 91, 372–381 (2006).
Bustin, S. A. et al. The MIQE Guidelines: Minimum Information for Publication of Quantitative Real-Time PCR Experiments. Clin Chem 55, 611–622 (2009).
Acknowledgements
This work was supported by National Institute of Health ES015024, ES013995, HL071643, HL092963 and Training Grant T32HL076139, the Northwestern University Clinical and Translational Sciences Institute (NUCATS) Center for Translational Innovation (CTI) Pilot Award (UL1 RR025741 from the National Center for Research Resources (NCCR), a component of the National Institutes of Health (NIH) and NIH Roadmap for Medical Research), The Veterans Administration and the American Lung Association.
Author information
Authors and Affiliations
Contributions
SS, AG, DU, KAR and SEC performed the experimental work. KMR provided the primary alveolar epithelial cells and ensured their quality. JJ and BK provided key reagents for the study (Mito-CP and TPP). AOV, NSC, assisted in study design and data analysis. SS, GM, AVR and GRSB designed the study and performed the analysis. SS and GRSB prepared the manuscript. All authors reviewed and commented on the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Materials
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Soberanes, S., Gonzalez, A., Urich, D. et al. Particulate matter Air Pollution induces hypermethylation of the p16 promoter Via a mitochondrial ROS-JNK-DNMT1 pathway. Sci Rep 2, 275 (2012). https://doi.org/10.1038/srep00275
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00275
This article is cited by
-
Relationship between particulate matter exposure and female breast cancer incidence and mortality: a systematic review and meta-analysis
International Archives of Occupational and Environmental Health (2021)
-
Combustion-derived particles from biomass sources differently promote epithelial-to-mesenchymal transition on A549 cells
Archives of Toxicology (2021)
-
Air pollution-induced epigenetic changes: disease development and a possible link with hypersensitivity pneumonitis
Environmental Science and Pollution Research (2021)
-
AHRR cg05575921 methylation in relation to smoking and PM2.5 exposure among Taiwanese men and women
Clinical Epigenetics (2020)
-
In utero exposure to diesel exhaust is associated with alterations in neonatal cardiomyocyte transcription, DNA methylation and metabolic perturbation
Particle and Fibre Toxicology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.