Abstract
Here we report the generation and characterization of 84 mouse ES cell lines with doxycycline-controllable transcription factors (TFs) which, together with the previous 53 lines, cover 7–10% of all TFs encoded in the mouse genome. Global gene expression profiles of all 137 lines after the induction of TFs for 48 hrs can associate each TF with the direction of ES cell differentiation, regulatory pathways and mouse phenotypes. These cell lines and microarray data provide building blocks for a variety of future biomedical research applications as a community resource.
Similar content being viewed by others
Introduction
Mammalian genomes encode 1,500–2,000 transcription factors (TFs)1, which cross-regulate one another to form the network of TFs. The network controls the transcriptome of cells, thereby defining the identity of cells. A powerful approach to deciphering such a complex network is the systematic perturbation of individual TFs followed by global gene expression profiling2.
Results
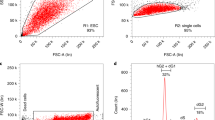
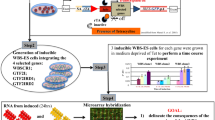
Here we report the generation of mouse embryonic stem (ES) lines, each of which has been engineered by integrating an expression cassette of a specific transcription factor (TF) into the ubiquitously expressing Rosa26 locus (Fig. 1a)2. The Rosa26 locus3 drives relatively uniform expression of the exogenous copy (transgene) of a TF, which is repressed by doxycycline (Dox) and can be induced in Dox- cell culture conditions (Fig. 1b)4. Combined with the 53 ES lines reported previously2, we present a total 137 ES cell lines. The majority of the manipulated genes were TFs, which were selected from a set of high-priority genes involved in critical functions in mouse ES cells and their differentiation5. To ensure the quality of these ES cell lines, we implemented vigorous QC steps that have been described previously in detail2. As a part of the characterization of these ES cell lines, we carried out global gene expression profiling by DNA microarrays 48 hours after TF induction (Fig. 1c; GEO accession number, GSE31381). The induction of a TF was confirmed by qRT-PCR (Fig. 1d, Supplementary Table 1 for primer pairs). The effect of TF induction on the transcriptome of mouse ES cells was highly variable (Fig. 1e; Supplementary Table 2). On a scale of the number of genes significantly changed in expression (FDR ≤ 0.05, fold change ≥1.5), the top 10% of studied TFs changed 4676 genes on average (e.g., Dmrt1), whereas the bottom 50% of TFs caused significant changes in expression in only 54.5 genes on average (e.g., Mbd3) (Fig. 1c, d).
Induction of transcription factors (TFs) in ES cells:
(a) plasmid structure that includes loxP recombination sites, puromycin resistance gene, open reading frame (ORF) of a TF with hCMV promoter followed by His6-FLAG tag; (b) schematic diagram showing the expression of transgenic TF induced in Dox- conditions; (c) examples of scatterplots of gene expression in Dox- versus Dox+ condition. Green and red dots indicate genes that are differentially expressed with statistical significance (FDR<0.05, change >1.5 fold); (d) Increase of transcription factor expression after the induction of a transgene, as measured by qPCR (Dox- vs. Dox+); results from two biological replicates (3 technical replicates each); error bars (S.E.M.; ANOVA); and dashed line = 2 fold change; (e) a list of TFs and the number of genes up- or down-regulated by the induction of the TF (FDR<0.05, change >1.5 fold) (Supplementary Table S2).
To further characterize the transcriptome alterations caused by each TF, we compared our microarray data with 3 public databases: the gene expression profiles of many mouse organs/tissues at The Genomics Institute of the Novartis Research Foundation (GNF) (ver. 2 & 3)6,7, the Genetic Association Database (GAD) on gene sets associated with mouse phenotypes8 and the MSigDB database (ver. 3) of gene sets associated with signaling pathways and cellular functions9. Because the GNF database is quantitative and the two other databases are qualitative, we used different methods to quantify association: correlation of median-subtracted log-transformed gene expression values for the GNF database and Parametric Analysis of Gene Expression (PAGE)10 for the GAD and msigdb databases (see Supplementary Methods).
A comparison of our microarray data with the GNF database showed that the induction of a TF in ES cells often initiates the differentiation of ES cells into specific cell types as soon as 48 hr later, when cells do not yet exhibit any overt phenotypes (Fig. 2 for GNF ver. 3; Supplementary Fig. 1 for GNF ver. 2). For example, the transcriptome of ES cells shifted toward a neural profile after the induction of Sox9, Foxg1, Klf3, or Pou5f1; toward endoderm after the induction of Hnf4a, Gata2, Gata3, or Esx1; and toward skeletal muscle and heart after the induction of Myod1 or Mef2c. Similarly, the transcriptome of ES cells shifted toward hematopoietic cell lineages after the induction of Sfpi1, Elf1, or Irf2; and toward T-cells and thymocytes after the induction of Elf5 or Tgif1. Interestingly, TFs associated positively with transcriptome changes toward specific lineages showed a negative association with those toward different cell lineages (Fig. 2). For example, TFs associated with transcriptome changes toward neural tissues were negatively associated with those toward hematopoietic lineages (e.g., Sox9 and Foxg1 in Fig. 2) and vice versa (e.g., Irf2, Elf1, Sfpi1 in Fig. 2). These data suggest that TF networks are organized to cross-regulate as if different tissue lineages are mutually exclusive.
Correlation of gene expression response to the induction of TFs with tissue-specific gene expression from the GNF ver. 3 database7.
A comparison of our microarray data with the GAD database identified associations of TF's with mouse phenotypes (Fig. 3). Many newly identified associations are consistent with published data. For example, Hoxa2 was associated with the pancreatic alpha and beta cells11; Foxc1, with hair follicle/shaft12,13; and Sox11 with skeletal defects14. A comparison of our microarray data with the msigdb database identified the association of each TF with specific cells and pathways (Fig. 4). For example, Smad6 was associated with keratinocytes15; Myod1, with alveolar rhabdomyosarcoma16; and Hnf4a, with lipoproteins17.
Enrichment of gene sets associated with mouse phenotypes from GAD database 8 among genes that were upregulated (positive) or downregulated (negative) after the induction of various TFs.
Enrichment of gene sets associated with various functions and signaling pathways from msigdb ver. 3 database 9 among genes that were upregulated (positive) or downregulated (negative) after the induction of various TFs.
Discussion
The collection of mouse ES cell lines reported here are freely available to the research community (http://esbank.nia.nih.gov/index.html). The analysis presented here can help researchers select ES cell lines suitable for their own research programs. For example, these TF-manipulable ES cell lines can be used to study the complex mechanisms of ES cell differentiation toward specific lineages. These ES cell lines are also adaptable to a variety of experiments and analyses, as shown in our previous report2. For example, each TF is C-terminally tagged with His6-FLAG, which simplifies studies of TF localization, protein-protein interactions and protein-DNA interactions2. Further mining of the microarray results reported here as well as additional experiments with provided ES cell lines and their derivatives will yield more insight into gene regulatory networks. Carrying out similar experiments for more regulatory proteins (ideally for all TFs and additional signaling proteins) should give increasingly complete information to comprehend gene regulation in mammalian cells and organs.
Methods
Derivation of transgenic ES cell lines
ES cell lines with inducible TF transgenes were derived from MC1 mouse ES cells (129S6/SvEvTac), passage 17. Cells were cultured in DMEM with 15% FBS and LIF on feeder cells. Cells were electroporated with a linearized pMWROSATcH vector and selected by hygromycin B. Knock-in for ROSA-TET locus was confirmed by southern blotting. For exchange vectors, PCR amplified ORFs were subcloned into pZhcSfi that was modified to express a His6-FLAG tagged protein and puromycin resistance gene. ES cells were co-transfected with a sequence verified exchange vector and pCAGGS-Cre and selected by puromycin in the presence of doxycycline (Dox). Isolated clones were tested for Venus expression, hygromycin B susceptibility, transgene RNA expression, genotyping for Cre mediated integration and mycoplasma contamination.
Gene expression analysis of cells with induced TFs
ES cells (passage 25) were cultured in the standard LIF+ medium with Dox+ on a gelatin-coated dish throughout the experiments. Cells from each cell line were split into 6 wells and the media was changed 24 hr after cell plating: 3 wells with Dox+ medium and 3 wells with Dox- medium to induce transgenic TFs. Dox was removed via washing 3 times with PBS at 3 hour intervals. Total RNA was isolated by TRIzol (Invitrogen) after 48 hr and two replications were used for real time qPCR (see primers in Supplementary Table S1) and for microarray hybridization. RNA samples were labeled with total RNA by the Low RNA Input Fluorescent Linear Amplification Kit (Agilent). For most TFs, we hybridized Cy3-CTP labeled sample from Dox- medium together with a Cy5-CTP labeled sample from Dox+ medium. But for 7 TFs we labeled samples from Dox- and Dox+ with Cy3 and hybridized them independently with a Cy5-labeled reference target, which is a mixture of Stratagene Universal Mouse Reference RNA and MC1 cells RNA (this method requires a double number of arrays). Analysis showed that both methods produce results of comparable quality. Targets were hybridized to the NIA Mouse 44K Microarray v3.0 (Agilent, design ID 015087)18. Slides were scanned with Agilent DNA Microarray Scanner. All DNA Microarray data are available in Supplementary Table S2, at GEO/NCBI19 (http://www.ncbi.nlm.nih.gov/geo; accession number GSE31381) and at NIA Array Analysis software20 (http://lgsun.grc.nia.nih.gov/ANOVA).
Normalization of microarray data and detection of outliers
Two methods of array hybridizations were used in this study: (1) RNA extracted from cells with induced transcription factors (TFs) (cultured in Dox- conditions) and from controlled cells (cultured in Dox+ conditions) were Cy3 labeled and all hybridized on separate arrays together with reference RNA labeled with Cy5; and (2) RNA extracted from cells with induced TFs (Dox-) were labeled with Cy3 and hybridized together with RNA from control cells (Dox+) which were labeled with Cy5. The second method does not use reference RNA. Data processing depended on the method of hybridization. Potential Cy3/Cy5 bias in microarrays with the hybridization of Dox- vs. Dox+ samples was removed by normalization to the median logratio of gene expression change in all TF-manipulation experiments. The details of the method are available in Supplementary Information.
Statistical analysis of microarray data
For statistical analysis we used NIA Array Analysis, which estimates the False Discovery Rate (FDR) to account for multiple hypothesis testing20. Response of genes to the knockdown of TFs was measured as a logratio (i.e., difference between means of log-transformed intensities) between manipulated (Dox-) and control (Dox+) cells. We considered gene expression change as significant if logratio was significantly different from zero (FDR < 0.05) and the change of expression was >1.5 fold.
Correlation with tissue-specific gene expression
Association of gene expression changes induced by TF manipulation with tissue-specific gene expression was evaluated based on the correlation between our microarray results with the GNF database7. Correlation was estimated between gene expression responses to TF manipulation (logratio of Dox- vs. Dox+) and median-centered log-transformed gene expression in various tissues from GNF database (ver. 2 and 3). Because the importance of genes in ES cells and adult tissues may be different and different platforms of microarrays used in these studies are not 100% compatible, we applied correlation analysis to a subset of genes that are highly expressed and dynamic in both data sets. We selected 10,000 genes in each database with the highest score equal to the product of average log-expression and standard deviation of expression (after induction of various TFs or in different tissues) and then took the intersecting portion of 5,595 genes for GNF ver. 3 (5,295 genes for ver. 2). Then, correlation values and corresponding z-values were estimated based on this subset of genes. The matrix was sorted using hierarchical clustering, TMEV, ver 3.121.
Analysis of gene set enrichment
Enrichment of target genes in subsets of genes that are upregulated or/and downregulated following the manipulation of the TF is quantified using a modified Parametric Analysis of Gene Enrichment (PAGE)10. PAGE is based on the comparison of the average expression change in a specific subset of genes, xset, with the average expression change in all genes, xall:

where nset is the size of the gene set and SDall is standard deviation of expression change among all genes. We modified this method by applying equation (1) to the subset of N top upregulated and another subset of N top downregulated genes rather than to all genes combined, which allowed us to detect the enrichment of the same gene set among both upregulated and downregulated genes. The value of N = 5000 was selected experimentally because it appeared that the enrichment of genes with TF binding sites is always limited to the top 5000 upregulated or downregulated genes. The probability distribution of expression change within subsets of N upregulated and downregulated genes is not normal; however, because we compare averages for large sets of genes (usually, nset is >50), the probability distribution of these averages is close to normal based on the central limit theorem22. Thus, it is reasonable to use equation (1) as an approximation. In the case when both up-regulated and down-regulated genes were enriched in a specific functional gene set, we subtracted the smaller z-value from both z-values. The matrix of z-values was first sorted using hierarchical clustering, TMEV, ver 3.121 and then manually converted to a semi-diagonal form.
References
Kanamori, M. et al. A genome-wide and nonredundant mouse transcription factor database. Biochem Biophys Res Commun 322, 787–793 (2004).
Nishiyama, A. et al. Uncovering early response of gene regulatory networks in ES cells by systematic induction of transcription factors. Cell Stem Cells 5, 420–433 (2009).
Soriano, P. Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat Genet 21, 70–71 (1999).
Masui, S. et al. An efficient system to establish multiple embryonic stem cell lines carrying an inducible expression unit. Nucleic Acids Res 33, e43 (2005).
Matoba, R. et al. Dissecting Oct3/4-regulated gene networks in embryonic stem cells by expression profiling. PLoS One 1, e26 (2006).
Wu, C. et al. BioGPS: an extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol 10, R130 (2009).
Su, A. I. et al. Large-scale analysis of the human and mouse transcriptomes. Proc Natl Acad Sci U S A 99, 4465–4470 (2002).
Zhang, Y. et al. Systematic analysis, comparison and integration of disease based human genetic association data and mouse genetic phenotypic information. BMC Med Genomics 3, 1 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102, 15545–15550 (2005).
Kim, S. Y. & Volsky, D. J. PAGE: parametric analysis of gene set enrichment. BMC Bioinformatics 6, 144 (2005).
Mizusawa, N. et al. Differentiation phenotypes of pancreatic islet beta- and alpha-cells are closely related with homeotic genes and a group of differentially expressed genes. Gene 331, 53–63 (2004).
Berry, F. B., Tamimi, Y., Carle, M. V., Lehmann, O. J. & Walter, M. A. The establishment of a predictive mutational model of the forkhead domain through the analyses of FOXC2 missense mutations identified in patients with hereditary lymphedema with distichiasis. Hum Mol Genet 14, 2619–2627 (2005).
Kunisada, M., Cui, C. Y., Piao, Y., Ko, M. S. & Schlessinger, D. Requirement for Shh and Fox family genes at different stages in sweat gland development. Hum Mol Genet 18, 1769–1778 (2009).
Sock, E. et al. Gene targeting reveals a widespread role for the high-mobility-group transcription factor Sox11 in tissue remodeling. Mol Cell Biol 24, 6635–6644 (2004).
Yu, H., Mrowietz, U. & Seifert, O. Downregulation of SMAD2, 4 and 6 mRNA and TGFbeta receptor I mRNA in lesional and non-lesional psoriatic skin. Acta Derm Venereol 89, 351–356 (2009).
Krskova, L. et al. Rhabdomyosarcoma: Molecular analysis of Igf2, MyoD1 and Myogenin expression. Neoplasma 58, 415–423 (2011).
Krapivner, S. et al. DGAT1 participates in the effect of HNF4A on hepatic secretion of triglyceride-rich lipoproteins. Arterioscler Thromb Vasc Biol 30, 962–967 (2010).
Carter, M. G. et al. Transcript copy number estimation using a mouse whole-genome oligonucleotide microarray. Genome Biol 6, R61 (2005).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets--10 years on. Nucleic Acids Res 39, D1005–1010 (2011).
Sharov, A. A., Dudekula, D. B. & Ko, M. S. A web-based tool for principal component and significance analysis of microarray data. Bioinformatics 21, 2548–2549 (2005).
Saeed, A. I. et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003).
Rice, J. A. Mathematical statistics and data analysis, 1 v. (Thomson/Brooks/Cole, Belmont, CA, 2007).
Acknowledgements
This research was supported entirely by the Intramural Research Program of the NIH, National Institute on Aging.
Author information
Authors and Affiliations
Contributions
LSC, YP, AN, JSC, HY, LVS, LX, HGH, MT, EM, BYB, GM and UB carried out the experiments. AAS, YQ, DD and MSHK carried out the data analysis. MSHK conceived the project. DLL, DS and MSHK supervised the project. AAS and MSHK wrote the manuscript with inputs from all authors. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Supplementary Information
Supplementary Table S2
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareALike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Correa-Cerro, L., Piao, Y., Sharov, A. et al. Generation of mouse ES cell lines engineered for the forced induction of transcription factors. Sci Rep 1, 167 (2011). https://doi.org/10.1038/srep00167
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00167
This article is cited by
-
Atlas of regulated target genes of transcription factors (ART-TF) in human ES cells
BMC Bioinformatics (2022)
-
Metabolic control of DNA methylation in naive pluripotent cells
Nature Genetics (2021)
-
Induction of human pluripotent stem cells into kidney tissues by synthetic mRNAs encoding transcription factors
Scientific Reports (2019)
-
Bioinformatics Approaches to Stem Cell Research
Current Pharmacology Reports (2018)
-
Rapid differentiation of human pluripotent stem cells into functional neurons by mRNAs encoding transcription factors
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.