Abstract
Shewanellae are microbial models for environmental stress response; however, the sequential expression of mechanisms in response to stress is poorly understood. Here we experimentally determine the response mechanisms of Shewanella amazonensis SB2B during sodium chloride stress using a novel liquid chromatography and accurate mass-time tag mass spectrometry time-course proteomics approach. The response of SB2B involves an orchestrated sequence of events comprising increased signal transduction associated with motility and restricted growth. Following a metabolic shift to branched chain amino acid degradation, motility and cellular replication proteins return to pre-perturbed levels. Although sodium chloride stress is associated with a change in the membrane fatty acid composition in other organisms, this is not the case for SB2B as fatty acid degradation pathways are not expressed and no change in the fatty acid profile is observed. These findings suggest that shifts in membrane composition may be an indirect physiological response to high NaCl stress.
Similar content being viewed by others
Introduction
Shewanellae are models for studying the genomic and phenotypic response to environmental stress because of their ability to inhabit and adapt to a wide range of environmental conditions1. Shewanellae are capable of both aerobic and anaerobic respiration utilizing diverse electron acceptors (fumarate, thiosulfate, nitrite, nitrate, iron, chromium, manganese and uranium)2,3,4,5,6. Consequently, Shewanellae have robust sensing and regulatory systems that allow rapid response and acclimation to changing environments2. Their versatility and plasticity provides an opportunity to investigate how microorganisms respond to changes in typical environmental conditions, such as temperature, dessication and osmotic stress7,8,9. Although they are typically marine-associated, Shewanella have been detected in environments ranging from fresh water6,10 to hypersaline environments11,12,13 and tolerate a wide range of salt concentrations9.
The genotypic and phenotypic response of Shewanella amazonensis SB2B (SB2B) to osmotic stress has not been described. This bacterium was isolated from shallow-water marine deposits derived largely from the Amazon River delta14. The physical mixing of these deposits by wave action combined with pore water transport exposes these deposits to periodic fluctuations in natural salinity14. Environmental fluctuations in salinity of its native habitat suggest that S. amazonensis is able to mitigate the effects of salt toxicity and hyperosmolarity. To experimentally determine the response mechanisms of S. amazonensis SB2B, we analyzed the dynamics of the proteome from initial response to acclimation (resumption of growth) during sodium chloride stress. The genome of SB2B is sequenced, providing peptide sequence information for conducting high throughput proteomics analyses using the accurate mass and time (AMT) tags approach15,16,17. We use novel approaches in time-course proteomics to gain insight to the sequence of events and coordination of mechanisms involved in salt tolerance such as initial signal transduction and regulation, turgor maintenance, motility and metabolism.
Bacterial genomic and proteomic responses to sudden changes in osmolarity play an important role in the biology and ecology of microorganisms because hyper- and hypo- osmolarity are stresses most commonly encountered in natural environments18,19. Microbes colonizing almost any environment face fluctuations in osmolarity19 and must be equipped with response mechanisms in order to survive the initial change as well as prolonged exposure to osmotic stress. The fluctuations in salinity are a key determinant in the composition of microbial communities in estuarine and brackish water ecosystems20 such as those from whence SB2B was isolated14. Bacterial response to osmotic stress is a complex, distinct sequence of cellular events21 and elucidation of dynamic cellular responses to salt stress benefit from the use of global techniques that allow high throughput analysis of proteins involved in the process. Proteomic methods are ideal for the analysis of global changes and have widely been used to enhance the knowledge of differentially expressed functional and regulatory proteins. However, until recently, proteomic technology and analytical protocols lacked the quantitative foundation to provide a platform for performing detailed time-course studies22,23. The aim of this study is to characterize the acclimation of Shewanella amazonensis str. SB2B to high sodium chloride stress using novel time-course proteomic techniques.
Results
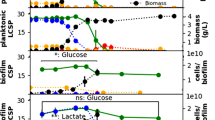
To determine SB2B’s salt tolerance, we exposed cultures to concentrations of salt ranging from 0.3M to 1.37M. The growth rate of SB2B was inhibited by approximately 50% at a salt concentration of 0.85M in growth cultures (Fig. 1a). Upon exposure of SB2B to 0.85M salt, growth is arrested between approximately 60 to 90 min following sodium chloride pulse (Fig. 1b). Accordingly, the response frame used throughout this study ranged from 0 to 90 min and was broken down to measure the initial response (up to 15 min), the intermediate response (15 to 60 min) and the sequence of events in longer-term acclimation (60 to 90 min) of SB2B to salt stress.
A total of 845 (out of 3,645 protein coding sequences; 23.2%) proteins were identified from SB2B as expressed throughout all five time points measured in this study. One hundred forty-three of the detected proteins demonstrated more than a two-fold change in expression (relative to time 0) for at least one subsequent time point. Principal component analysis of the expression patterns of all differentially expressed proteins suggest two separate responses to high salt (Fig. 2). Most of the variation within expression is described by proteins corresponding to arrested growth of SB2B and a shift to branched chain amino acid degradation.
Within the first 15 min following 0.85 M salt exposure, SB2B induces a number of genes involved in protein turnover and transcription. The first 15 min of arrested growth of SB2B correlates to a significant decrease in the abundance of DNA polymerase (Sama1310), which was also confirmed by RT-qPCR (Table 2). A significant decrease in ribosomal proteins (Sama2548, 3033 and 1999) detected, also suggests arrested cell growth and a reduction in translation, as illustrated under ‘primary repressed’ in figure 3 (although the trend is similar for other ribosomal proteins, Sama0202-0239 show no significant change in expression). Other replication (TopA; Sama3041, MutS; Sama1045 and DNA helicase; Sama3058) and cell division (FtsK; Sama1772, FtsZ; Sama0358) proteins were also lower following salt addition. TopA and FtsK levels were also confirmed using RT-qPCR (Table 2). In addition, we found significant expression of a methyltransferase (thiopurine S-methyltransferase - Sama0543). Reverse transcriptase-quantitative PCR confirms decreased expression of DNA replication and translation proteins that correspond to the diminished growth rate. One of the most highly differentially expressed proteins (8.4-fold increase) within the first 15 minutes of salt exposure is a diguanylate cyclase (Sama2418).
Cellular model of differentially expressed pathways and proteins in response to NaCl stress in SB2B.
Pathways and proteins are represented with expression values of 15 min, 30 min, 60 min and 90 min following exposure to NaCl relative to no exposure as bars next to each respective protein. Immediate induction (primary induced) proteins on the left suggest a signal cascade that influences cellular motility. Immediate repression (primary repressed) proteins in the middle suggest the arrest of DNA replication and cell division. Both the primary induced and primary repressed proteins approach pre-stressed expression levels at 90 min after NaCl pulse. Secondary induced proteins (right) involved in branched chain amino acid degradation are induced between 15 and 30 min following addition of NaCl; their expression value remains high past the 90 min time frame of this study.
In addition to a pause in cell division proteins, SB2B induces (or represses) a pulse of proteases (proteases; Sama2056 and Sama3074) and nucleases (endonucleases; Sama0514 and Sama0278, exonuclease SbcC; Sama1268 and ribonucleases; Sama1933 and Sama3067: see Supplementary Table 1). Immediately following the NaCl pulse, expression of the protease/chaperone ClpA (Sama2056) is lower and remains so until growth resumes between 60 to 90 minutes following the applied stress.
Although some fatty acid degradation proteins (Sama2071; oxidoreductase and Sama2167; fatty acid oxidation alpha subunit) were expressed, we did not observe a change in the fatty acid content of SB2B following exposure to 0.85 M NaCl (described below). While proteins involved in branched-chain amino acid degradation increased in relative abundance 15 to 30 min following NaCl exposure, proteins involved in the synthesis of branched chain amino acids did not significantly change in relative abundance. Fatty acid analysis of SB2B throughout acclimation to high salt conditions showed no significant difference in branched chain fatty acid composition (Table 2). The only significant decline for fatty acids once SB2B was exposed to high salt concentrations was 16∶0 and 16∶1.
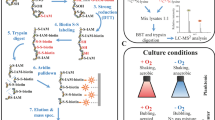
Three malonyl acyl carrier proteins were detected (Sama1978, 1982 and 2154) but none showed any difference in expression due to high salt. Sama1362 encodes an isovaleryl-CoA dehydrogenase (E.C. 1.3.99.10) specific for 3-methylbutanoyl-CoA in the complete degradation of leucine. We also observed an increased abundance of two protein homologs (Sama1377 and Sama 1870) that are butyryl-CoA dehydrogenases (E.C. 1.3.99.2) specific for isobutyryl-CoA and 2-methylbutyryl-CoA in the complete degradation of valine and isoleucine, respectively. The alternative to degradation of these amino acids at this junction in the pathway results in synthesis of branched chain fatty acids. From this point in the degradation of branched chain amino acids, each step is expressed in a similar fashion and to a similar extent (Fig. 3). Both Sama1378 and 1379 encode an enoyl-CoA hydratase (E.C. 4.2.1.17) that can be used in all three amino acid pathways. To complete the degradation of leucine to acetoacyl-CoA, we found expressed proteins for the 3-methylcrotonyl-CoA carboxylase (Sama1359; E.C. 6.4.1.4), hydroxymethylglutaryl-CoA lyase (Sama1358; E.C. 4.1.3.4) and two expressed homologs for 3-oxoacid CoA-transferase (Sama1357 and Sama1457; E.C. 2.8.3.5). For the continued degradation of valine, we found an expressed 3-hydroxyisobutyrate dehydrogenase (Sama1380; E.C. 1.1.1.31) and an NAD-dependent aldehyde dehydrogenase (Sama2648; E.C. 1.2.1.3) that can shunt to succinyl-CoA as well as a methylmalonate-semialdehyde dehydrogenase (Sama1376; E.C. 1.2.1.27) that can shunt to propanoyl-CoA. For the continued degradation of isoleucine, an acetyl-CoA acyltransferase (Sama0031; E.C. 2.3.1.16) was expressed that yields propanoyl-CoA. We found that aconitase (Sama3296; E.C. 4.2.1.3), 2-methylcitrate synthase (Sama3295; E.C. 2.3.3.5) and 2,3-dimethylmalate lyase (Sama3294; E.C. 4.1.3.30) are grouped together on the chromosome and were highly expressed between 15 and 30 minutes following salt addition. In addition, we found similar expression of an acetyl-CoA synthetase (Sama2079; E.C. 6.2.1.1).
By 90 min following salt exposure, resumption of growth in SB2B correlates with the abundance of proteins involved in DNA replication and cell division to pre-osmotic shock levels. The methyltransferase (Sama0543), diguanylate cyclase (Sama2418), endonuclease (Sama0278) and exonuclease SbcC (Sama1268) that were expressed as described earlier, decreased in abundance well below pre-perturbed levels. Conversely, DNA polymerase (Sama1310) and the DNA mismatch repair protein MutS (Sama1045) go from decreased abundance following exposure to salt (15 min) to >2-fold increase (90 min) relative to pre-salt exposure.
Physical measurements of motility confirmed that SB2B decreases motility upon exposure to high salt concentrations (Fig. 4). However, the protein expression patterns for decreased motility appear late in the acclimation to salt stress.
Discussion
Time course proteome analyses of the response of SB2B suggest an immediate, but brief motile osmotactic response to salt stress. This motility response corresponds with a pause in growth and cell division observed in growth curves (Fig. 1) as well as in the low expression of replication and division proteins. The secondary response of SB2B is a shift to branched-chain amino acid metabolism prior to resuming growth. Figure 3 illustrates the proposed model of the primary response and secondary response of SB2B to salt stress. Principal component analysis of all differentially expressed proteins shows that 90% of the variation in expression is accounted for by two components (60% for component 1, 30% for component 2; Fig. 2). Most of the proteins involved in the variation of expression fall into the two categories mentioned above (signal transduction and arrested replication/division and branched chain amino acid degradation).
Signal transduction systems play a major role in the acclimation of microorganisms to changes in osmolarity and histidine kinases have been identified as osmosensors in both prokaryotic24,25 and eukaryotic cells26. Sama2418 is a diguanylate cyclase response regulator containing a CheY-like receiver domain and a GGDEF domain that may affect the direction of rotation and of the flagellar motor to control the motile behavior of the bacterium25. In several species, diguanylate cyclases (GGDEF domain) have been implicated as signal molecules that trigger changes in the bacterial cell surface and motility21,27. Some evidence suggests that cyclases also serve as sensors for osmotic pressure in eukaryotes28. Signal transduction domains, the weak homology of Sama2418 to a flagellar hook-associated protein and the timing of expression of this regulator suggest that this protein may serve as an osmotic sensor in SB2B. In addition, protein Sama1653 is a potential methyl-accepting chemotaxis sensory transducer protein with transmembrane characteristics, a signal peptide, a kinase domain and a HAMP domain that ultimately results in the regulation of rotation of the flagellar motor29. The clustering of expression patterns of the regulator (Sama2418) and a methyltransferase (Sama0543) (Fig. 5) suggest osmotaxis may be driven by methylation of a methyl-acceptaing chemotaxis protein30.
Because cell volume and ion regulation are not instantaneous processes, osmotic stress may damage cellular macromolecules and impair cell function until compensatory acclimations counteract the stress. Damage to DNA or changes in transcription caused by negative supercoiling at high salt31,32,33 leads to impairment of cell function and to the induction of repair processes and protection systems34. In SB2B, SbcC (Sama1268), an exonuclease that removes unusual DNA structure(s), such as hairpins, that may be generated upon DNA damage35 or through supercoiling is more abundant with salt stress. To reset the proteome to function in an environment of high salt, ClpA has been shown to have similar function to the DnaK and DnaJ chaperones during replication36. However, under osmotic stress induced by exposure to salt, the binding of ClpA to helicases involves a switch from a salt-sensitive to a salt-insensitive protein complex37. Although DnaK and DnaJ are often induced immediately following osmotic stress, our observations suggest that ClpA is associated with long-term acclimation and not the immediate stress response.
As reported from a transcriptome analysis of S. oneidensis9, salt stress response and branched-chain amino acids (isoleucine, leucine and valine) metabolism seem to be linked, prompting a hypothesized decrease in branched-chain fatty acid content at high salinity38 as is found in other organisms39. We found that between 15 and 30 minutes following salt exposure, SB2B expressed proteins that degraded branched chain amino acids completely to central metabolic pathways (Fig. 3). However, we found no repression of any of the proteins involved in the synthesis of branched chain fatty acids that would be expected with a shift in the branched chain fatty acid content of the cell. Furthermore, fatty acid profiles of SB2B in response to high salt show no significant change in the concentration of branched chain fatty acids (Table 3), nor do original studies on the compositional changes of the fatty acid profile in Shewanellae during exposure to salt when grown on nutrient rich media40. Change in branched chain fatty acid profile during salt stress for S. putrefaciens only occurs with the removal of all exogenous sources of amino acids41 suggesting a shift to branched chain amino acid metabolism in the media and at the expense of fatty acid content when necessary. Expression profile similarities illustrated in (Fig. 5) suggest that Sama3141, a putative adhesion protein, may be involved in the metabolic shift to degrade branched chain amino acids for Shewanella at high salt.
Despite previous claims for Shewanella38, SB2B does not selectively degrade branched chain fatty acids as a response to high salt concentrations. The role of branched chain amino acid degradation in the adaptation of SB2B remains unclear, however the evidence presented here, in context with confirmed reports of changes in branched chain fatty acid content of Bacillus39 raises the question whether membrane fatty acid changes in other organisms are a response to high salt or the result of a metabolic shift to branched chain amino acid degradation. The advantage of the time-course approach taken in this study is that it provides expression profiles that can be compared to determine co-expression patterns using multivariate statistical algorithms. The temporal expression patterns of the SB2B proteome following osmotic shock suggest an immediate arrest of transcription and division, followed by a shift to branched chain amino acid degradation and the resumption of cell division. High expression of an adhesion protein that groups with branched chain amino acid degradation may suggest that the down-regulation of flagellar genes9 and proteins reported here may coincide with a lifestyle change in Shewanella at high salt concentrations.
Methods
Media and growth conditions
Initial experiments were carried out on S. amazonensis SB2B (ATCC: BAA-1098) to determine at what concentration of sodium chloride limits SB2B growth in LB broth. Initial cultures were inoculated to 15 mL LB broth with NaCl concentrations ranging from 1% (0.17M NaCl; normal salt concentration of LB) to 8% (w/v; 1.37M) at 30°C on an orbital shaker set at 200 rpm. Growth rates were determined spectrophotometrically (OD600 nm) in triplicate over a 24 hour period. We selected a NaCl concentration corresponding to a 50% reduction in the growth rate for further proteomic analyses.
To induce an osmotic stress response, cells were cultured in LB broth until logarithmic phase growth (OD600 nm ∼0.4) was achieved, followed by addition of NaCl to a final concentration of 0.85M salt (5%). The acclimation time was determined from the point of salt addition until resumption of growth. Growth rate dynamics were monitored by optical density and confirmed using standard plate count techniques.
Sample preparation
Based on the acclimation time of SB2B to salt, samples were harvested at five time points following the addition of salt; 0 (control immediately prior to addition of NaCl), 15, 30, 60 and 90 min. Five replicate samples were harvested by centrifugation (5,000 rpm, 4°C, 10 min) and the cells were snap frozen using liquid nitrogen and stored at −80°C.
For protein extraction, cell pellets were thawed on ice, washed and suspended in 100 mM NH4HCO3 (pH 8.4) then lysed via bead beating. From the cell lysis material, proteins were extracted and digested using trypsin into peptides according to established protocols42. Resulting peptide samples were pooled to create a composite sample for each time point to best utilize allotted instrument resources.
Total RNA was extracted from cell pellets collected from the same time points using an RNeasy mini kit (Qiagen, Valencia, CA) and the RNA was quantified spectrophotometrically. Approximately 0.1 to 1 μg of total RNA was incubated with genomic DNA elimination buffer and converted to cDNA using a QuantiTect reverse transcription kit (Qiagen, Valencia, CA).
Proteomics data generation
Proteomics data was generated using the accurate mass and time (AMT) tag proteomics approach16. In brief, this approach utilizes tandem mass spectrometry to generate a reference peptide database (accurate mass and time tag database; AMT tag database) of observed peptides and their associated theoretical masses and elution times (normalized). This database is utilized to assign peptide sequences to ion current (relative abundance) information to peptides measured using high-resolution, high mass measurement accuracy mass spectrometry (LC-MS). The AMT tag database for S. amazonensis SB2B is maintained at the Pacific Northwest National Laboratory (Richland, WA) and was constructed as part of research efforts on Shewanella by the Shewanella Federation2,42,43, For the study reported here, we utilized the normalized elution times and computed masses of peptides within the database to match sequences to peptides and their abundances measured using LC-MS from the NaCl perturbed and control culture samples.
As part of this approach, samples for LC-MS analysis were first randomized and blocked; where, a single block contained one proteomics analysis per time point for a total of five blocks. Each block of samples was analyzed using an LTQ-Orbitrap™ (ThermoFisher Scientific, San Jose, CA) mass spectrometer interfaced with a reverse phase HPLC system for peptide separation (LC-MS). Peptides were reverse-phase separated on in-house manufactured columns (60 cm × 360 µm o.d. × 75 µm i.d. fused silica capillary tubing) packed with 3 µm Jupiter C18 stationary phase (Phenomenex, Torrence, CA). The HPLC system was equilibrated with 100% mobile phase A (0.2% acetic acid and 0.05% TFA in water). Fifty min. after peptide injection, mobile phase B (0.1% TFA in 90% acetonitrile/10% water) displaced mobile phase A, generating an approximate exponential gradient. Split flow controlled the gradient speed operating under constant pressure (10 kpsi). Separated peptides were ionized (positive) using an electrospray ionization (ESI) interface (manufactured in-house) with chemically etched electrospray emitters (150 mm o.d. × 20 mm i.d).
Separated peptides were ionized (positive) using an electrospray ionization interface (manufactured in-house) that consisted of chemically etched electrospray emitters (150 mm o.d. 20 mm i.d). The LTQ-Orbitrap™ mass spectrometer was operated using a heated capillary temperature and spray voltage of 200°C and 2.2 kV, respectively. Data was acquired for 100 min, beginning 60 min after sample injection (10 min into gradient). LTQ-Orbitrap™ spectra (AGC 1×106) were collected from 400–2000 m/z at a resolution of 100k followed by data dependant ion trap generation of MS/MS spectra (AGC 3×104) of the six most abundant ions using a collision energy of 35%. A dynamic exclusion time of 60 sec was used to discriminate against previously analyzed ions. Generated MS/MS spectra were searched using the SEQUEST algorithm44 against the publicly available S. amazonensis SB2B translated genome sequence (www.jgi.doe.gov). Identified peptides of at least six amino acids in length, having 1+, 2+ and 3+ charge states and SEQUEST cross-correlation (Xcorr; an indicator of the statistical match between the observed and theoretically generated MS/MS spectra) scores of ≥ 1.9, 2.2, or 3.5, respectively, were added to the existing S. amazonensis SB2B AMT tag database.
Orbitrap™ spectra were deisotoped using the software tool Decon2LS45 and the tool VIPER44 was used to identify mass and elution time features within mass measurement accuracy and elution time accuracy cut-offs of 5 ppm and 1%, respectively. Abundances for these features were calculated by integrating the signal strength under each peak of the LC-MS spectra17. To assign peptide sequences to this quantitative data, mass and elution time features were matched to peptide sequences within the AMT tag database (identified by LC tandem mass spectrometry) for S. amazonensis SB2B. Peptide sequences were filtered according to a PeptideProphet™46 probability value of ≤ 0.90 prior to the matching of mass and elution time features. Matched features from each Orbitrap™ analysis (dataset) were then filtered on a false discovery rate (FDR) of less than or equal to 5%; the FDR associated with the AMT tag proteomics approach, calculated using STAC (Statistical Tools for AMT tag confidence). STAC is the latest statistical algorithm for assigning confidence to matched mass and elution time features and is an evolution of an earlier algorithm referred to as SMART47. A protein was considered positively identified if a minimum of 2 unique peptides from its sequence were detected by LC-MS.
Proteomics data analysis
The computational strategy set forth by Du, et al.22 for analysis of label-free temporal proteomics data was followed. In brief, this strategy encompasses peptide filtering across replicates, abundance normalization, missing-value imputation, protein abundance estimation from peptides (protein rollup) and trend analysis. The associated MatLab™ (MathWorks, Inc., Natick, MA) code developed by Du, et al. (http://omics.pnl.gov/software/) was utilized22; with the exception that central tendency normalization was performed using DANTE (http://omics.pnl.gov/software/)23. A final trend for each protein was calculated as the median of trends observed from the replicate LC-MS measurements.
Gene expression analysis
Primers for genes involved in branched chain amino acid degradation, TCA cycle, motility and DNA replication/repair were designed using the Primer3 application (http://frodo.wi.mit.edu/primer3/) with a targeted amplicon size of 80–100 bp (Table 1). One ng of cDNA was run in triplicate for 40 cycles, using a two step PCR in an Opticon 2 DNA engine (MJ Research, Waltham, MA). cDNA was labeled using a QuantiFast SYBR PCR kit (Qiagen, Valencia, CA) and 125 nM of each primer. Amplicon size and reaction specificity were confirmed by agarose gel electrophoresis and product dissociation curves. The number of target copies in each sample was interpolated from its detection threshold (CT) value using a purified PCR product standard curve. The standard curve consisted of a serial dilution of a purified 16S rRNA gene sequence from SB2B of known concentration. A representative amplicon for each gene tested was sequenced to confirm the target.
Fatty acid analysis
In order to measure the membrane acclimation of SB2B to osmotic stress, cell samples used for gene expression and proteomic analyses were analyzed for fatty acid content. Triplicate samples collected for each time point were pooled and saponified using 45 g NaOH, 150 ml methanol and 150 ml water followed by methylation using 6.0 N hydrochloric acid and methanol. Fatty acids were extracted using a 1∶1 solution of hexane and methyl tert-butyl ether and analyzed for composition as per the MIDI protocol (Microbial ID, Newark, DE).
Motility
Cell motility was qualitatively tested in triplicate using soft agar inoculations on LB plates. We prepared solid (1% agar) and soft (0.3% agar) plates both at salt concentrations of 0.17M (normal LB media salt concentration) and 0.5M for motility assessments. Each plate was inoculated with SB2B (5μl; OD600 = 0.5) and incubated at 25°C for 72 h. Colony diameter was measured for comparison.
References
Tiedje, J. M. Shewanella – the environmentally versatile genome. Nat. Biotechnol. 20, 1093–1094 (2002).
Fredrickson, J. K., Romine, M. F., Beliaev, A. S., Auchtung, J. M., Driscoll, M. E., Gardner, T. S., Nealson, K. H., Osterman, A. L., Pinchuk, G., et al. Towards environmental systems biology of Shewanella . Nat. Rev. Microbiol. 6, 592–603 (2008).
Liu, Y., Gorby, Y., Zachara, J. M., Fredrickson, J. K. & Brown, C. F. Reduction kinetics of Fe (III), Co(III), U (VI), Cr (VI) and Te (VII) in cultures of dissimilatory metal-reducing bacteria. Biotechnol. Bioeng. 80, 637–649 (2002).
Myers, C. R. & Myers, J. M. Fumarate reductase is a soluble enzyme in anaerobically grown Shewanella putrefaciens MR-1. FEMS Microbiol. Lett. 98, 13–20 (1992).
Myers, C. R. & Myers, J. M. Ferric reductase is associated with the outer membrane of anaerobically grown Shewanella putrefaciens MR-1. FEMS Microbiol. Lett. 108, 15–22 (1993).
Myers, C. R. & Nealson, K. H. Bacterial manganese reduction and growth with manganese oxide as a sole electron acceptor. Science 240, 1319–1321 (1998).
Leblanc, L., Leboeuf, C., Leroi, F., Hartke, A. & Auffray, Y. Comparison between NaCl tolerance response and acclimation to cold temperature in Shewanella putrefaciens. Curr. Microbiol. 46, 157–62 (2003).
Li, S., Xiao, X., Li, J., Luo, J. & Wang, F. Identification of genes regulated by changing salinity in the deep-sea bacterium Shewanella sp. WP3 using RNA arbitrarily primed PCR. Extremophiles 10, 97–104 (2006).
Liu, Y., Gao, W., Wang, Y., Wu, L., Liu, X., Yan, T., Alm, E., Arkin, A. P., Thompson, D. K. et al. Transcriptome analysis of Shewanella oneidensis MR-1 in response to elevated salt conditions. J. Bacteriol. 187, 2501–2507 (2004).
Pinhassi, J. & Berman, T. Different growth response of colony-forming α- and γ-proteobacteria in dilution culture and nutrient addition experiments from Lake Kinneet (Israel), the Eastern Mediterranean Sea and the Gulf of Eilat. Appl. Environ. Microbiol. 69, 199–211 (2003).
Jiang, H., Dong, H., Yu, B., Liu, X., Li, Y., Ji, S. & Zhang, C. L. Microbial response to salinity change in Lake Chaka, a hypersaline lake on Tibetan plateau. Environ. Microbiol. 9, 2603–2621 (2007).
Kulp, T. R., Hoeft, S. E., Miller, L. G., Saltikov, C., Murphy, J. N., Han, S., Lanoil, B. & Oremland, R. S. Dissimilatory arsenate and sulfate reduction in sediments of two hypersaline, arsenic –rich soda lakes: Mono and Searles Lakes, California. Appl. Environ. Microbiol. 72, 6514–6526 (2006).
Lefebvre, O., Vasudevan, N., Thanasekaran, K., Moletta, R. & Godon, J. J. Microbial diversity in hypersaline wastewater: the example of tanneries. Extremophiles 10, 505–513 (2006).
Venkateswaran, K., Dollhopf, M. E., Aller, R., Stackebrandt, E. & Nealson, K. H. Shewanella amazonensis sp. nov., a novel metal-reducing facultative anaerobe from Amazonian shelf muds. Int. J. Sys. Bacteriol. 48, 965–972 (1998).
Smith, R. D., Anderson, G. A., Lipton, M. S., Masselon, C., Pasa-Tolic, L., Shen, Y. & Udseth, H. R. (2002). The use of accurate mass tags for high-throughput microbial proteomics. OMICS 6, 61–90.
Smith, R. D., Anderson, G. A., Lipton, M. S., Pasa-Tolic, L. Shen, Y., Conrads, T. P., Veenstra, T. D. & Udseth, H. R. An accurate mass tag strategy for quantitative and high-throughput proteome measurements. Proteomics 2, 513–523 (2002).
Smith, R. D., Shen, Y. F. & Tang, K. Q. Ultrasensitive and quantitative analyses from combined separations-mass spectrometry for the characterization of proteomes. Accounts Chem. Res. 37, 269–278 (2004).
Kempf, B. & Bremer, E. Uptake and synthesis of compatible solutes as microbial stress responses to high-osmolarity environments. Arch. Microbiol. 170, 319–330 (1998).
Wood, J. M. Osmosensing by bacteria: signals and membrane-based sensors. Microbiol. Mol. Biol. Rev. 63, 230–262 (1999).
Morita, R. Y., Jones, L. P., Griffiths, R. P. & Staley, T. E. Salinity and temperature interactions and their relationship to microbiology of the estuarine environment. In Estuarine Microbial Ecology, pp 221–232. Edited by L. H. Stevenson & R. R. Colwell. University of South Carolina Press, Columbia (1973).
Ryjenkov, D. A., Tarutina, M., Moskvin, O. V. & Gomelsky, M. Cyclic diguanylate is a ubiquitous signaling molecule in bacteria: insights into biochemistry of the GGDEF protein domain. J. Bacteriol. 187, 1792–1798 (2005).
Du, X., Callister, S. J., Manes, N. P., Adkins, J. N., Alexandridis, R. A., Zeng, X., Roh, J. H., Smith, W. E., et al. A computational strategy to analyze label-free temporal bottom-up proteomics data. J. Proteome Res. 7, 2595–2604 (2008).
Polpitiya, A. D., Qian, W. J., Jaitly, N., Adkins, J. N., Camp II, D. G., Anderson, G. A. & Smith, R. D. DAnTE: a statistical tool for quantitative analysis of –omics data. Bioinformatics 24, 1556–1558 (2008).
Perraud, A-L., Weiss, V. & Gross, R. Signaling pathways in two-component phosphorelay systems. Trends Microbiol. 7, 115–120 (1999).
Vaknin, A. & Berg, H. C. Osmotic stress mechanically perturbs chemoreceptors in Escherichia coli . Proc. Natl. Acad. Sci. U.S.A. 103, 592–596 (2005).
Browse, J. & Xin, Z. Temperature sensing and cold acclimation. Curr. Opin. Plant Biol. 4, 241–246 (2001).
D’Argenio, D. A. & Miller, S. I. Cyclic di-GMP as a bacterial second messenger. Microbiology 150, 2479–2502 (2004).
van Es, S., Virdy, K. J., Pitt, G. S., Meima, M., Sands, T. W., Devreotes, P. N., Cotter, D. A. & Schaap, P. . Adenylyl cyclase G, an osmosensor controlling germination of Dictyostelium spores. J. Biol. Chem. 271, 23623–23625 (1996).
Khursigara, C. M., Wu, X., Zhanj, P., Lefman, J. & Subramaniam, S. Role of HAMP domains in chemotaxis signaling by bacterial chemoreceptors. Proc. Natl. Acad. Sci. U.S.A. 105, 16555–16560 (2008).
Wong, L. S., Johnson, M. S., Sandberg, L. B. & Taylor, B. L. Amino acid efflux in response to chemotactic and osmotic signals in Bacillus subtilis . J. Bacteriol. 177, 4342–4349 (1995).
Conter, A., Menchon, C. & Gutierrez, C. Role of DNA supercoiling and rpoS ς factor in the osmotic and growth phase-dependent induction of the gene osmE of Escherichia coli K12. J. Mol. Biol. 273, 75–83 (1997).
Graeme-Cook, K. A., May, G., Bremers, E. & Higgens, D. F. Osmotic regulation of porin expression: A role for DNA supercoiling. Mol. Microbiol. 3, 1287–1294 (1989).
Jordi, B. J. & Higgins, C. F. The downstream regulatory element of the proU operon of Salmonella typhimurium inhibits open complex formation by RNA polymerase at a distance. J. Biol. Chem. 275, 12123–12128 (2000).
Naegeli, H. Mechanisms of DNA damage recognition in mammalian cells. Georgetown: R.G. Landes Company (1997).
Darmon, E., Lopez-Vernaza, M. A., Helness, A. C., Borking, A., Wilson, E., Thacker, Z., Wardrope, L. & Leach, D. R. F. SbcCD regulation and localization in Escherichia coli . J. Bacteriol. 189, 6686–6694 (2007).
Wickner, S., Gottesman, S., Skowyra, D., Hoskins, J. & McKenney, K. A molecular chaperone, ClpA, functions like DnaK and DnaJ. Proc. Natl. Acad. Sci. U.S.A. 91, 12218–12222 (1994).
Kilstrup, M., Jacobsen, S., Hammer, K. & Vogensen, F. K. Induction of heat shock proteins DnaK, GroEL and GroES by salt stress in Lactococcus lactis . Appl. Environ. Microbiol. 63, 1826–1837 (1997).
Kazakov, A. E., Rodionov, D. A., Alm, E., Arkin, A. P., Dubchak, I. & Gelfand, M. S. . Comparative genomics of regulation of fatty acid and branched-chain amino acid utilization in proteobacteria. J. Bacteriol. 191, 52–64 (2009).
Lopez, C. S., Heras, H., Ruzal, S. M., Sanchez-Rivas, C. & Rivas, C. A. Variations of the envelope composition of Bacillus subtilis during growth in hyperosmotic medium. Curr. Microbiol. 36, 55–61 (1998).
Nichols, D. S., Olley, J., Garda, H., Brenner, R. R. & McMeekin, T. A. Effect of temperature and salinity stress on growth and lipid composition of Shewanella gelidimarina . Appl. Environ. Microbiol. 66, 2422–2429 (2000).
Nichols, D. S., McMeekin, T. A. & Nichols, P. D. Manipulation of polyunsaturated, branched-chain and trans-fatty acid production in Shewanella putrefaciens strain ACAM 342. Microbiology 140, 577–584 (1994).
Lipton, M. S., Pasa-Tolic, L., Anderson, G. A., Anderson, D. J., Auberry, D. L., Battista, J. R., Daly, M. J., Fredrickson, J., Hixson, K. K. et al. Global analysis of the Deinococcus radiodurans proteome by using accurate mass tags. Proc. Natl. Acad. Sci. U.S.A. 99, 11049–11054 (2002).
Turse, J. E., Marshall, M. J., Fredrickson, J. K., Lipton, M. S., Callister, S. J. An empirical strategy for characterizing bacterial proteomes across species in the absence of genomic sequences. PLoS ONE 12;5(11),e13968 (2010).
Eng, J. K., McCormack, A. L., Yates, J. R., III An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Jaitly, N., Mayampurath, A., Littlefield, K., Adkins, J. N., Anderson, G. A. & Smith, R. D. . Decon2LS: An open-source software package for automated processing and visualization of high resolution mass spectrometry data. BMC Bioinformatics 10, 87 (2009).
Monroe, M. E., Tolić, N., Jaitly, N., Shaw, J. L., Adkins, J. N. & Smith, R. D. VIPER: an advanced software package to support high-throughput LC-MS peptide identification. Bioinformatics 23, 2021–2023 (2007).
Jaitly, N., Adkins, J. N., Monroe, M. E., Norbeck, A. D., Mottaz, H. M., Dabney, A. R., Lipton, M. S., Anderson, G. A., Smith, R. D. A statistical approach to quantifying uncer- tainties in the AMT tag pipeline. 58th ASMS Conference on Mass Spectrometry and Allied Topics, Denver, CO (2008).
Acknowledgements
Funding for this research was provided by NSF grant DEB-021212487 to MEP, a grant from USDA CSREES 2006-34526-17001 and supported by the Utah Agricultural Experiment Station at Utah State University as journal paper number 8199. A portion of the research described in this paper was performed in the Environmental Molecular Sciences Laboratory (EMSL), a DOE/BER national scientific user facility located at Pacific Northwest National Laboratory (PNNL) in Richland, Washington. PNNL is a multi-program national laboratory operated by Battelle for the DOE under Contract DE-AC05-76RLO 1830.
Author information
Authors and Affiliations
Contributions
JP and SC wrote the paper, experiments were designed by JP, SC and MP. PCR validation was performed by AW and GR.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Table 1
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Parnell, J., Callister, S., Rompato, G. et al. Time-course analysis of the Shewanella amazonensis SB2B proteome in response to sodium chloride shock. Sci Rep 1, 25 (2011). https://doi.org/10.1038/srep00025
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep00025
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.