Abstract
Although hair cells are the sensory receptors of the auditory and vestibular systems in the ears of all vertebrates, hair cell properties are different between non-mammalian vertebrates and mammals. To understand the basic biological properties of hair cells from non-mammalian vertebrates, we examined the transcriptome of adult zebrafish auditory and vestibular hair cells. GFP-labeled hair cells were isolated from inner-ear sensory epithelia of a pou4f3 promoter-driven GAP-GFP line of transgenic zebrafish. One thousand hair cells and 1,000 non-sensory surrounding cells (nsSCs) were separately collected for each biological replicate, using the suction pipette technique. RNA sequencing of three biological replicates for the two cell types was performed and analyzed. Comparisons between hair cells and nsSCs allow identification of enriched genes in hair cells, which may underlie hair cell specialization. Our dataset provides an extensive resource for understanding the molecular mechanisms underlying morphology, function, and pathology of adult zebrafish hair cells. It also establishes a framework for future characterization of genes expressed in hair cells and the study of hair cell evolution.
Design Type(s) | parallel group design • cell type comparison design |
Measurement Type(s) | transcription profiling assay |
Technology Type(s) | RNA sequencing |
Factor Type(s) | biological replicate • cell type |
Sample Characteristic(s) | Danio rerio • otolith organ |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Hair cells are the sensory receptors of the auditory and vestibular systems in the ears of all vertebrates. Hair cells transduce mechanical stimuli, such as movement in their environment, into electrical activity1. The site of such mechanoelectrical transduction is the hair bundle, the hallmark of all hair cells. In addition to the apical stereocilia bundle, hair cells also contain structural and functional specializations in the basolateral and synaptic membranes, which are responsible for electrical activities and synaptic transmission2–4.
Although hair cells have some common properties, hair cells in non-mammalian vertebrates are substantially different from their mammalian counterpart. For example, hair cells of non-mammals are electrically tuned to specific frequencies and possess an active process in the stereocilia bundle to amplify sound signals5,6. Mammalian hair cells, in contrast, are not electrically tuned7. Mammals enhance their auditory sensitivity and frequency selectivity by outer hair cell electromotility8–10, a mammalian innovation. Hair cells from non-mammalian vertebrates such as bullfrog and chicken, are able to spontaneously repair and regenerate their damaged or lost stereocilia bundles11–13. In contrast, mammalian cochlear hair cells no longer retain that capability14. To understand the molecular mechanism underlying their differences, analysing transcriptomes of hair cells is an important first step. The transcriptome represents all the RNA transcripts in a cell and can vary depending on cell type, physiological condition, and development stage15. The adult zebrafish hair cell transcriptome was examined by the first generation of DNA microarray technique consisting of 15,000 transcript probes16. That study was completed before a complete zebrafish reference genome became available17. Transcriptomic analysis using RNA-seq allows assessment of the quantitative expression over 31,000 genes of the GRCz10 zebrafish genome, as well as possible identification of new splice variants. Several recent studies examined hair cell transcriptomes from zebrafish larval with RNA-seq18–21. Those studies have led to the identification of some hair cell-specific genes. However, since hair cells in those studies were obtained from zebrafish larvae, whose gene expression profile is known to be different from that of adult hair cells, we examined the inner ear hair cell-specific transcriptome of adult zebrafish using RNA-seq.
Our goal of sequencing the transcriptome of pristine populations of adult zebrafish hair cells was accomplished by using GFP-labeled hair cells isolated from the utricle, saccule, and lagena, the three inner-ear sensory epithelia of a pou4f3 promoter-driven GAP-GFP line of transgenic zebrafish22. In addition to GFP-labeling, hair cell identification was further aided by the visible morphological feature of all hair cells, the stereocilia bundle, to exclude any ambiguous cells from collection. A suction pipette was used to individually collect hair cells23,24. This technique has some advantages over the fluorescence-activated cell sorting (FACS) technique25. In our study, cells were individually collected based on the presence of both GFP expression and stereocilia bundles. Thus, cell identity was unambiguous and potential contamination by other cell types was mitigated. Another advantage is that the average time for collection of 300 to 350 hair cells from each zebrafish after hair cells were isolated was less than 50 min. Because cells were maintained in cold solution during collection, and individually collected cells were immediately fixed in RNALater solution, this shorter time of cell sorting allows isolation of high quality RNA and minimizes transcriptomic changes that may occur during FACS, which may take up to a few hours25.
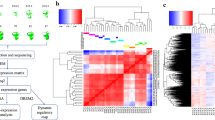
Here, we describe transcriptome-wide profiling of hair cells and non-sensory surrounding cells (nsSCs) from the adult zebrafish inner ear. Three biological replicates, each containing 1,000 individually collected hair cells, were prepared. Each of our three control samples consisted of 1,000 isolated nsSCs from the inner ear that did not exhibit fluorescence and stereocilia bundles. An overview of the study design is depicted in Fig. 1a. Careful and stringent technical design at each experimental stage has allowed generation of a high-quality RNA-seq dataset which has tremendous value for future characterization of all genes expressed in zebrafish hair cells. Our dataset is also expected to provide valuable information for the study of hair cell regeneration and evolution. Hair cell-specific transcriptomes from mouse cochleae and vestibule have been analyzed20,24,26–28. Thus, the present dataset is also useful for comparative studies of hair cells between zebrafish and mouse.
Schematic drawing of zebrafish is modified from Fig. 1 of a previous publication37 (with permission from Frontier in Cellular Neuroscience). (a) Workflow of experimental design for RNA-seq and transcriptome analysis for 1,000 individually collected hair cells and nsSCs. (b) GFP-expressing hair cells in saccule and lagena of zebrafish inner ear. (c) Suction pipette technique used to manually collect individual hair cells and nsSCs. (d) Examples of GFP-expressing hair cells. Only those cells that had both GFP expression and stereocilia bundles were selected. (e) Example of GFP-expressing cells without visible stereocilia bundles. The identify of these cells was unknown, so they were not collected. (f) An example of a nsSC with no GFP expression. An equal number of nsSCs was individually collected for comparison with hair cells. Bars: 20 μm (b), 10 μm (c), and 10 μm (d–f).
Methods
Hair cell isolation and collection
Adult female transgenic Tg(pou4f3:GAP43-GFP)s273t zebrafish22 at 11 to 13 months of age were used for the study. Animals were euthanized by submersion in ice water (0–4 °C) for ten minutes after cessation of opercular movement. The utricle, saccule, and lagena were isolated using a method described by Liang and Burgess29. Hair cells in the inner ear structures of this transgenic zebrafish line express GFP and an example of GFP-expressing hair cells in the isolated saccule and lagena is shown in Fig. 1b. The inner ear tissues then underwent an enzymatic digestion at room temperature for 20 min in the medium containing 1 ml of L-15 medium and 1 mg of Collagenase IV (Sigma-Aldrich, St. Louis, MO, USA). The inner ear tissues were transferred to Leibovitz’s L-15 medium at 300 mOsm, 7.35 pH. Hair cells and nsSCs were separated by gentle trituration. The chamber (with inlet and outlet), placed on the stage of an Olympus inverted microscopy with fluorescence capability, was perfused with fresh L-15 medium to wash out debris for 5 min. To collect solitary cells, the suction pipette technique was used19,20. Two pickup pipettes with a diameter of ~30 μm were used to separately pick up GFP-positive hair cells and GFP-negative nsSCs. The pipette was fabricated from 1.5 mm thin-wall glass tubing pulled by a two-stage electrode puller. The pickup pipettes were mounted in two separate electrode holders mounted on two Leitz 3-D micromanipulators (Leitz, Germany). By moving the pickup pipette and the stage of the microscope, cells were positioned near the tip of the pipette. The suction port of the pipette holder was connected to a micrometer-driven syringe to provide positive or negative pressure to draw in or expel the cells. An image of a GFP-positive hair cell before being drawn into a pickup pipette is shown in Fig. 1c a video showing a mouse outer hair cell being drawing into a pickup pipette is provided in Supplementary Video 1 (Data Citation 1). After ~10 to 20 cells were drawn into the pipette, the pipette was lifted out from the bath and transferred to a microcentrifuge tube containing 50 μl RNAlater (Thermo Fisher Scientific, Waltham, MA). Cells were expelled from the pipette by applying positive pressure. We repeated this step until approximately 300 to 350 hair cells and nsSCs were collected from each zebrafish. 1,000 hair cells were collected from approximately four zebrafish for each of the three samples. 15 zebrafish were used for the collection of three biological samples of hair cells and three biological samples of nsSCs. During cell collection, two pickup pipettes were used, each designated for only one cell type, to prevent contamination. To obtain highly specific hair cells, several steps were taken to avoid contamination by other cell types. First, each cell being collected was identified under direct visual observation. Since hair cells used in our study expressed GFP, presence of both GFP and the stereocilia bundle were used to identify hair cells (Fig. 1d). Any ambiguous cells that exhibited only GFP expression without the visible stereocilia bundle (Fig. 1e) were excluded. Second, only solitary cells not attached to any other cell types were collected. Finally, we were careful about the suction pressure applied to the pipette to avoid drawing unwanted cells. The suction pipette (to deposit hair cells) was withdrawn only when the pressure was balanced and no more fluid or cells were being drawn into the pipette. 1,000 nsSCs lacking GFP expression and stereocilia (Fig. 1f) from the inner ear epithelium were also collected for each of the three samples using the same procedures.
RNA isolation, amplification
Total RNA, including small RNAs (>~18 nucleotides), from approximately 1,000 each cell type separately suspended in RNALater (~100 μl in total volume), were extracted and purified using the Qiagen miRNeasy mini plus Kit (Qiagen Sciences Inc, Germantown, MD). On-column DNase digestion was performed to eliminate DNA contamination in the collected RNA. After purification, the quality and quantity of RNA was determined using an Agilent 2,100 BioAnalyzer (Agilent Technologies, Santa Clara, CA) and compared to examples of pure RNA results found in the Agilent 2,100 Bioanalyzer 2,100 Expert User’s Guide. Total RNA from each sample ranged from 5 to 10 ng/μl. These purified RNA samples were reverse transcribed into cDNA and amplified using the SMART-Seq V4 Ultra Low Input RNA kit (Clontech Laboratories, Inc., Mountain View, CA) to prepare the samples for RNA sequencing.
RNA-sequencing and bioinformatic analyses
Genome-wide transcriptome libraries were produced from triplicate biological replicates. Libraries were prepared using the SMART-Seq V4 Ultra Low Input RNA kit (Clontech) to generate cDNA in combination with the Nextera Library preparation kit (Illumina, Inc., San Diego, CA). Library size and concentration were assessed using an Agilent 2,100 Bioanalyzer and a Qubit fluorometer (Invitrogen) to ensure the inserts were the appropriate size and determine concentration prior to sequencing. Transcriptome libraries were sequenced using the HiSeq 2,500 Sequencing System (Illumina). Libraries were multiplexed and three samples per 100 bp paired-end lane were sequenced, generating approximately 100 million reads per sample. The cut-off for RNA integrity number was set at the value of 9. The files from the multiplexed RNA-seq samples were demulitplexed and fastq files representing each library and quality control data were generated.
Code availability and bioinformatics analyses
No custom code was used in these analyses. CLC Genomics Workbench software (CLC bio, Waltham, MA, USA) and UCSC genome were used to map the reads to the GRCz10 zebrafish genome and generate gene expression values in the normalized form of reads per kilobase of transcript per million mapped reads (RPKM) values. Reads were mapped to exonic, intronic, and intergenic sections of the genome. Gene expression estimates were derived from the mapped reads using HTSeq count30. Expression data was exported from CLC Genomics Workbench in the form of Microsoft Excel Spreadsheets. Clustering analyses were carried out using one of the following algorithms: CLICK, K-means and SOM, which are implemented in the EXPANDER integrative package for analysis of gene expression. The differentially expressed genes and clusters were examined for enrichment of known biological processes using DAVID31 and Ingenuity IPA program (www.ingenuity.com). Entrez Gene, HGNC, OMIM, and Ensembl database were used for verification, reference, and analyses. Additional resources such as the gEAR (http://umgear.org/index.html) and SHIELD (https://shield.hms.harvard.edu/index.html)32 websites were also used for transcriptome profile comparisons.
Quantitative reverse transcription PCR (qPCR)
Quantitative-PCR experiments were run on an Applied Biosystems 7,500 Fast Real-Time PCR system. Ten microliters of Powerup SYBR Green Master Mix (Thermo Fisher Scientific, Waltham, MA, USA) was used in each 20 microliter reaction. Primer concentrations were 450 nM. The original cDNA samples were diluted twenty-fold with two microliters for every reaction. The fast thermal cycling mode of the Applied Biosystems 7,500 instrument was used, with an initial stage of 2 min at 50 °C followed by a 2 min period at 95 °C.
The sequences of the oligonucleotide primers for reverse transcription and amplification of representative gene transcripts in real-time quantitative PCR were designed using A plasmid Editor (ApE) software (http://biologylabs.utah.edu/jorgensen/wayned/ape/), and BLAST searches (http://blast.ncbi.nlm.nih.gov/Blast.cgi) to find unique and appropriate sequences with melting temperatures above 60 °C that had predicted low rates of homodimerization. Oligonucleotide primers were acquired from Integrated DNA Technologies, Coralville, Iowa. The sequences of oligonucleotide primers are shown in Table 1.
Data Records
Raw FASTQ sequencing files comprised of the three biological replicates of hair cells and nsSCs, have been deposited to the NCBI Sequence Read Archive (Data Citation 2), and sample metadata can be found at the NCBI Gene Expression Omnibus (Data Citation 3) with series number of GSE101693. Individual accession numbers for each biological sample are also provided in Table 2. The most important file for readers not interested in re-analyzing the data is ‘GSE101693_Zebrafish_ Inner_Ear_Hair_Cell_Transcriptome_Table.xlsx’ (Data Citation 3).
The RPKM gene expression values of three biological replicates of hair cells and nsSCs are included in sheet 1 of ‘zFishhaircell.xlsx’ (Data Citation 4). RNA-seq raw data of zebrafish glia (Data Citation 5, Data Citation 6 and Data Citation 7) and liver cells (Data Citation 8), obtained from previously published studies33,34, were normalized to our datasets and reanalyzed using CLC Genomics Workbench software. The gene expression RPKM values of glia and liver cells are also included in sheet 1 of ‘zFishhaircell.xlsx’ (Data Citation 4). These data can be used for comparison with the expression pattern of hair cells and nsSCs.
Technical Validation
RNA quality control
Samples of isolated RNA were analyzed for quality of intact RNA and concentration to determine their suitability for RNA-sequencing using an Agilent 2,100 BioAnalyzer. Figure 2a,b represent two examples of quality results of isolated RNA of the hair cells and nsSCs. Samples were considered to be of high quality if the ribosomal peaks had minimal (<5% of the total peak area) shoulders. All of the samples selected for RNA for sequencing met this criterion.
Sequencing accuracy
The quality of the reads is often evaluated using the Illumina Basespace bioinformatic cloud-computing interface (https://basespace.illumina.com/ home/index). The FastQC app (version 1.0.0) on the Illumina cloud computing interface examined the quality of the reads by comparing the read signals to the probability of accurate base-reading. This number is called the Phred quality score35, with a score of 40 indicating a 99.99% accuracy of the base call, a score of 30 having an accuracy of 99.9%, and a score of 20 with an accuracy of 99%. The quality of the reads of our fastq.gz files generated from RNA-sequencing were analyzed for base-reading accuracy. A mean Phred score above 28, indicative of high accuracies of the correct base at a given nucleotide in the sequence, was used as the high-quality cutoff. Examples of these analyses are shown in Fig. 2c,d. All of the sequencing runs exceeded that mean, which suggests that the RNA-sequencing performed was of high quality and unambiguous.
Reproducibility of biological samples
We used correlation coefficient to examine reproducibility of three replicates of hair cells and nsSCs. Figure 3a show six plots of comparison between replicates from the same population of cells. As shown, the data points are all concentrated near the line (replicate 1) with small deviation. The mean correlation coefficient between hair cell replicates is 0.9984±0.0003 (mean±s.d.), while the mean correlation coefficient between nsSC replicates is 0.994±0.0045, suggesting that the results were highly reproducible.
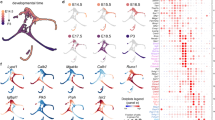
(a) Correlation coefficient of biological replicates. Replicate 1 of hair cells and replicate 1 of nsSCs were separately used as reference (red lines). (b) PCA analysis of the gene expression profiles of hair cells and nsSCs compared with microglia (MGC)33 and liver cells34, all from adult zebrafish. The top five expressed genes in PCA analysis were: s100s, mt-co2, AC024175.9, pvalb9, mt-co3 for hair cells; mt-co2, mt-co3, AC024175.9, AC024175.4, mt-cyb for nsSCs; AC024175.4, mt-co2, mt-co1, mt-co3, tmsb4x for microglial cells; and, apoa1b, apoa2, tfa, fabp10a, and apoc1l for liver cells, respectively. (c) Clustering analysis of gene expression of hair cells, nsSCs, MGC, and liver cells.
Gene clustering analysis was also used to examine reproducibility of biological replicates and hierarchy of gene expression of different cell populations. Figure 3b shows Principal Component Analysis (PCA) of the gene expression profiles of adult zebrafish hair cells, nsSCs, microglia, and liver cells. As shown, three biological replicates of hair cells are highly reproducible as the data points are clustered all together with small variability. The three replicates of nsSCs are also highly reproducible. However, PCA analysis shows that hair cells and nsSCs are separated by a large distance, suggesting that their gene expression profiles are quite different. The gene expression profiles of microglia and liver cells are even more distinct from that of hair cells, as both of them are further away from hair cells in the graph. Gene clustering analysis displayed in Fig. 3c shows the similar results as demonstrated in PCA analysis.
qPCR
We separately prepared three biological replicates of hair cells and nsSCs from 15 additional zebrafish for qPCR to validate the expression of 17 genes, with 13 being highly expressed in hair cells and 4 highly expressed in nsSCs. The expression values were all normalized to the cycle threshold (Ct) value of actb2, which was used as the reference gene for hair cells and nsSCs. The normalized Ct numbers were then inverted (1/Ct)36 and graphed in Fig. 4a. We compared the patterns of differential expression of these genes between hair cells and nsSCs using expression values from qPCR and RNA-seq. As shown, 13 genes that were differentially expressed in hair cells based on RNA-seq analysis show larger inverted Ct scores in hair cells than in nsSCs, suggesting that their expression in hair cells is greater than in nsSCs. In contrast, four genes that had higher expression in nsSCs than in hair cells in RNA-seq analysis also show larger inverted Ct scores in nsSCs than in hair cells (Fig. 4a). Figure 4b shows such a comparison after the expression values were normalized to fold changes. Although the level of expression is normally not comparable between the two techniques (due to different amplification and quantification procedures), the trend of differential expression of these genes is consistent with our RNA-seq data.
(a) Comparison of the expression of 18 genes in hair cells and nsSCs using q-PCR. The difference in the inverted Ct scores for each gene between hair cells and nsSCs was statistically significant (P≤0.01, n=3). Note that the ordinate is the inverse of the number of PCR cycles necessary to reach fluorescence threshold. So even a small difference (such as 0.01) reflects a large difference between the two cell populations36. (b) Fold differences in expression of these 17 genes between hair cells and nsSCs using q-PCR and RNA-seq. Positive values indicate higher gene expression in hair cells while negative values indicate higher expression values in nsSCs. Fold differences are all calculated in log2 base.
Validation by comparison with published studies
Several studies have examined the gene expression profiles of hair cells from inner ear and lateral line from larval or adult zebrafish16,18,20,21. Enriched genes in hair cells and nsSCs identified in these studies can be used for comparison with enriched genes identified in current study. Table 3 shows the RPKM values of enriched transcripts of hair cells and nsSCs from current study. A list of enriched transcripts in hair cells (top 21 transcripts in Table 3) and nsSCs (bottom 5 transcripts in the table) was compiled from previous studies16,18,21. The differential expression of these transcripts in hair cells or nsSCs was validated by in situ hybridization in those studies. The differential expression pattern shown in our dataset is highly consistent with the pattern demonstrated by in situ hybridization in previous studies. Comparison with published data offers another way for validation of our dataset. In Table 4, the top 30 enriched zebrafish hair cells transcripts and the expression value of their mammalian orthologs in the adult mouse inner ear are presented. The expression values of mouse inner hair cell and outer hair cell were from our previous study24. Since the expression values from zebrafish and mouse hair cells cannot be normalized for direct comparison, the abundance ranking of the mammalian orthologs detected in adult inner hair cells and outer hair cells is also presented in Table 4 for comparison. Note that except for three genes (Ocm, Atp1b2 and Hspa8), most genes or transcripts that are highly expressed in zebrafish hair cells are only expressed at modest to low levels in adult mouse hair cells. Six of the zebrafish genes have no known mouse ortholog, and two of the genes (Ocm and Gm42674) have at least two zebrafish orthologs for a single mouse gene.
Usage Notes
Transcriptome analysis is often used to identify differentially expressed genes that may underpin unique biological properties of cells. The transcriptomes of hair cells and nsSCs presented in this study can be used to identify enriched genes or transcripts of hair cells and nsSCs in adult zebrafish inner ear. Sheet 2 of ‘zFishhaircell.xlsx’ (Data Citation 4) presents the uniquely (in red fonts) and differentially (in blue fonts) expressed genes in hair cells and nsSCs. Uniquely expressed genes were those whose expression levels were below background in only one cell type, whereas differentially expressed genes were categorized as those whose RPKM values were above 0.1 and at least 2-fold different between the two cell types with statistical significance (Student’s t-test with P≤0.05). These uniquely and differentially expressed genes may provide valuable information to understand unique structures and function of hair cells and nsSCs in the adult zebrafish inner ear.
Additional information
How to cite this article: Barta, C. L. et al. RNA-seq transcriptomic analysis of adult zebrafish inner ear hair cells. Sci. Data 5:180005 doi: 10.1038/sdata.2018.5 (2018).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Fettiplace, R. & Kim, K. X. The physiology of mechanoelectrical transduction channels in hearing. Physiol. Rev. 94, 951–986 (2014).
Kros, C. J. in The Cochlea (eds Dallos P. & Fay R. R. ) 318–385 (Springer, 1996).
Sewell, W. F. in The Cochlea (eds Dallos P. & Fay R. R. ) 503–533 (Springer, 1996).
Safieddine, S., El-Amraoui, A. & Petit, C. The auditory hair cell ribbon synapse: from assembly to function. Annu. Rev. Neurosci. 35, 509–528 (2012).
Fettiplace, R. & Fuchs, P. A. Mechanisms of hair cell tuning. Annu. Rev. Physiol. 61, 809–834 (1999).
Hudspeth, A. J. Integrating the active process of hair cells with cochlear function. Nat. Rev. Neurosci. 15, 600–614 (2014).
Housley, G. D. & Ashmore, J. F. Ionic currents of outer hair cells isolated from the guinea-pig cochlea. J. Physiol. 448, 73–98 (1992).
Brownell, W. E., Bader, C. R., Bertrand, D. & de Ribaupierre, Y. Evoked mechanical responses of isolated cochlear outer hair cells. Science 227, 194–196 (1985).
Liberman, M. C., Gao, J., He, D. Z., Wu, X., Jia, S. & Zuo, J. Prestin is required for electromotility of the outer hair cell and for the cochlear amplifier. Nature 419, 300–304 (2002).
Dallos, P. et al. Prestin-based outer hair cell motility is necessary for mammalian cochlear amplification. Neuron 58, 333–339 (2008).
Baird, R. A., Steyger, P. S. & Schuff, N. R. Mitotic and nonmitotic hair cell regeneration in the bullfrog vestibular otolith organs. Ann. N. Y. Acad. Sci. 781, 59–70 (1996).
Baird, R. A., Burton, M. D., Lysakowski, A., Fashena, D. S. & Naeger, R. A. Hair cell recovery in mitotically blocked cultures of the bullfrog saccule. Proc. Natl. Acad. Sci. USA 97, 11722–11729 (2000).
Gale, J. E., Meyers, J. R., Periasamy, A. & Corwin, J. T. Survival of bundleless hair cells and subsequent bundle replacement in the bullfrog’s saccule. J. Neurobiol. 50, 81–92 (2002).
Jia, S., Yang, S., Guo, W. & He, D. Z. Fate of mammalian cochlear hair cells and stereocilia after loss of the stereocilia. J. Neurosci. 29, 15277–15285 (2009).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Mcdermott, B. M., Baucom, J. M. & Hudspeth, A. J. Analysis and functional evaluation of the hair-cell transcriptome. Proc. Natl. Acad. Sci. USA 104, 11820–11825 (2007).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Steiner, A. B., Kim, T., Cabot, V. & Hudspeth, A. J. Dynamic gene expression by putative hair-cell progenitors during regeneration in the zebrafish lateral line. Proc. Natl. Acad. Sci. USA 111, E1393–E1401 (2014).
Jiang, L., Romero-Carvajal, A., Haug, J. S., Seidel, C. W. & Piotrowski, T. Gene-expression analysis of hair cell regeneration in the zebrafish lateral line. Proc. Natl Acad. Sci 111, E1383–E1392 (2014).
Elkon, R. et al. RFX transcription factors are essential for hearing in mice. Nat. Commun. 6, 8549 (2015).
Erickson, T. & Nicolson, T. Identification of sensory hair-cell transcripts by thiouracil-tagging in zebrafish. BMC Genomics 16, 842 (2015).
Erkman, L. et al. Role of transcription factors a Brn-3.1 and Brn-3.2 in auditory and visual system development. Nature 381, 603–606 (1996).
He, D. Z., Zheng, J., Edge, R. & Dallos, P. Isolation of cochlear inner hair cells. Hear. Res. 145, 156–160 (2000).
Liu, H. et al. Characterization of transcriptomes of cochlear inner and outer hair cells. J. Neurosci. 34, 11085–11095 (2014).
Schwarz, J. M. Using fluorescence activated cell sorting to examine cell-type-specific gene expression in rat brain tissue. J. Vis. Exp. 99, e52537 (2015).
Cai, T. et al. Characterization of the transcriptome of nascent hair cells and identification of direct targets of the Atoh1 transcription factor. J. Neurosci. 35, 5870–5883 (2015).
Scheffer, D. I., Shen, J., Corey, D. P. & Chen, Z. Y. Gene expression by mouse inner ear hair cells during development. J. Neurosci. 35, 6366–6380 (2015).
Burns, J. C., Kelly, M. C., Hoa, M., Morell, R. J. & Kelley, M. W. Single-cell RNA-Seq resolves cellular complexity in sensory organs from the neonatal inner ear. Nat. Commun 6, 8557 (2015).
Liang, J. & Burgess, S. M. Gross and fine dissection of inner ear sensory epithelia in adult zebrafish (Danio rerio). J. Vis. Exp. 27, pii: 1211 (2009).
Anders, S., Theodor, Pyl, P. & Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Dennis, G. et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome Biol. 4, P3 (2003).
Shen, J., Scheffer, D. I., Kwan, K. Y. & Corey, D. P. SHIELD: an integrative gene expression database for inner ear research. Database (Oxford) bav071 (2015).
Oosterhof, N. et al. Identification of a conserved and acute neurodegeneration-specific microglia transcriptome in the zebrafish. Glia 65, 138–149 (2017).
Baumgart, M. et al. Longitudinal RNA-Seq analysis of vertebrate aging identifies mitochondrial complex I as a small-molecule-sensitive modifier of lifespan. Cell Syst 2, 122–132 (2016).
Ewing, B., Hillier, L., Wendl, M. C. & Green, P. Base-calling of automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 8, 175–185 (1998).
Li, Y. et al. Transcription factors expressed in mouse cochlear inner and outer hair cells. PLoS ONE 11, e0151291 (2016).
Monroe, J. D., Rajadinakaran, G. & Smith, M. E. Sensory hair cell death and regeneration in fishes. Front. Cell. Neurosci 9, 131 (2015).
Data Citations
Barta, C. L. Figshare http://doi.org/10.6084/m9.figshare.5625313 (2017)
NCBI Sequence Read Archive SRP113243 (2017)
NCBI Gene Expression Omnibus GSE101693 (2017)
Barta, C. L. Figshare http://doi.org/10.6084/m9.figshare.5625403 (2017)
NCBI Gene Expression Omnibus GSM2310340 (2016)
NCBI Gene Expression Omnibus GSM2310344 (2016)
NCBI Gene Expression Omnibus GSM2310348 (2016)
NCBI Gene Expression Omnibus GSM1620956-GSM1620958 (2016)
Acknowledgements
This work has been supported by the NIH grant R01 DC004696 to DH from the NIDCD and by the Béllucci Depaoli Family Foundation. Y.L. has been supported by a grant from National Natural Science Foundation of China (#81600798). Zebrafish husbandry was supported by grants from the National Center for Research Resources (5P20RR01878809) and the National Institute of General Medical Sciences (8 P20 GM10347109). We acknowledge the use of the University of Nebraska DNA Sequencing Core Facility for performing RNA-seq. The University of Nebraska DNA Sequencing Core receives partial support from the NCRR (1S10RR027754-01, 5P20RR016469, RR018788-08) and the National Institute for General Medical Science (NIGMS) (8P20GM103427, P30GM110768). This publication’s contents are the sole responsibility of the authors and do not necessarily represent the official views of the NIH or NIGMS. The study is based on experiments by Mr Cody L. Barta as part of the requirements for the Degree of Master of Science in the Department of Biomedical Sciences of Creighton University, which was awarded on February 16, 2016.
Author information
Authors and Affiliations
Contributions
D.H. designed the experiments. C.B., L.C., Y.L., K.G., K.C., and K.B. analysed the data, CB, H.L., L.C., and Y.L. performed the experiments, C.B. and D.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Barta, C., Liu, H., Chen, L. et al. RNA-seq transcriptomic analysis of adult zebrafish inner ear hair cells. Sci Data 5, 180005 (2018). https://doi.org/10.1038/sdata.2018.5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2018.5