Abstract
Under ongoing climate change and increasing anthropogenic activity, which continuously challenge ecosystem resilience, an in-depth understanding of ecological processes is urgently needed. Lakes, as providers of numerous ecosystem services, face multiple stressors that threaten their functioning. Harmful cyanobacterial blooms are a persistent problem resulting from nutrient pollution and climate-change induced stressors, like poor transparency, increased water temperature and enhanced stratification. Consistency in data collection and analysis methods is necessary to achieve fully comparable datasets and for statistical validity, avoiding issues linked to disparate data sources. The European Multi Lake Survey (EMLS) in summer 2015 was an initiative among scientists from 27 countries to collect and analyse lake physical, chemical and biological variables in a fully standardized manner. This database includes in-situ lake variables along with nutrient, pigment and cyanotoxin data of 369 lakes in Europe, which were centrally analysed in dedicated laboratories. Publishing the EMLS methods and dataset might inspire similar initiatives to study across large geographic areas that will contribute to better understanding lake responses in a changing environment.
Design Type(s) | database creation objective • data analysis objective • observation design |
Measurement Type(s) | nutrient • biological pigment • cyanotoxin • thermocline |
Technology Type(s) | data acquisition system • high performance liquid chromatography • high performance liquid chromatography-mass spectrometry |
Factor Type(s) | lake • Country |
Sample Characteristic(s) | Austria • Switzerland • Cyprus • Czech Republic • Germany • Kingdom of Denmark • Estonia • Kingdom of Spain • Finland • French Republic • Greece • Croatia • Hungary • Republic of Ireland • Italy • Lithuania • Macedonia • The Netherlands • Kingdom of Norway • Poland • Portuguese Republic • Russia • Sweden • Slovenia • Slovak Republic • Turkey • United Kingdom • lake |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Eutrophication still is the primary process threatening lakes and reservoirs and the services they provide, like good quality drinking water, irrigation, fisheries and recreational opportunities. Anthropogenic eutrophication is responsible for massive algal blooms. Cyanobacteria have diverse functional traits that allow them to proliferate under various environmental conditions1. The frequency and size of cyanobacterial blooms is increasing globally2,3. Excessive cyanobacterial biomass reduces light penetration and enhances anoxia in the hypolimnion, thus reducing species habitats and biodiversity4. Moreover, the toxins produced by bloom-forming cyanobacteria present a considerable risk to drinking water5 and pose a substantial economic cost6,7.
Climate change, through direct and indirect effects of warming, increasingly plays a role in changing physico-chemical and biological properties of aquatic ecosystems8–11, contributing to the global increase in cyanobacterial blooms. Although optimal growth temperatures vary widely between cyanobacterial strains and species, as well as for their eukaryotic competitors, their growth rate increases faster with temperature than for other phytoplankton groups9,12, Longer ice-free seasons, reduced winter overturn and enhanced water column stability in summer may all indirectly favour cyanobacterial blooms. In deep peri-alpine lakes for instance, water column stability has increased by 20% in a response to warming of the atmosphere13.
Interactions between nutrients and temperature-related changes are expected14–16. However, it is still uncertain to what extent and following which mechanisms nutrients and temperature will interact to amplify blooms. Climate forcing of blooms will differ among regions17. For example, at high latitudes and equatorial areas, intense precipitation events are expected (IPCC7) to increase nutrient enrichment of water bodies from enhanced surface runoff and groundwater discharge4, whereas at low and mid-latitude continental interiors, droughts18 may reduce river flow rates, increase lake residence times and thereby promote cyanobacteria. In hyper-eutrophic systems, high algal biomass may even increase local temperature due to enhanced light absorption19. Thus, high nutrient concentrations may promote warmer temperatures, giving a competitive advantage to buoyant cyanobacteria over non-buoyant algal species4. All of these interactions between nutrients and temperature may vary with lake depth. Shallow lakes respond more directly to nutrients and temperature than deep lakes, which, in contrast, are more typically subjected to the indirect effects of the aforementioned drivers20.
Hence, a complex interplay of regional (climate), local (nutrients, lake morphometry) and biological (species, functional groups) lake variables determines cyanobacterial bloom formation. Consequently, studies covering different regions and lake types are needed to disentangle the relative importance of the environmental predictors and their interactions. The European Multi Lake Survey (EMLS) obtained a deeper insight into cyanobacterial dynamics under different ecosystem variables across Europe. A space-for-time substitution, where contemporary spatial phenomena are studied in many lakes, instead of long-term temporal studies in a limited number of lakes21, was used. The survey took place in summer 2015, Europe’s third hottest summer on record, and comprised scientists from 27 countries that sampled 369 lakes only once. In this way, environmental gradients across wide geographic scales were covered with relatively little effort and with higher cost efficiency. We sampled in summer as cyanobacterial blooms are a distinct feature of summer phytoplankton22, during the locally warmest period, in order to test for temperature effects on cyanobacteria. In EMLS, standardized sampling procedures were strictly followed to ensure data homogeneity and eliminate site or operator related observation effects. Finally, lake samples for nutrients, algal pigments and toxins were analysed in dedicated laboratories by one person on one machine, minimizing variation in analytical errors.
Apart from providing a solid research dataset on which several analyses are being conducted, the EMLS helped to enhance the standards for limnological data collection and stimulate international collaboration. A subset of the EMLS toxin dataset has already been used in a recent publication to show how the distribution in cyanobacterial toxins and toxin quota was determined by both direct and indirect effects of temperature23. Here we additionally present the data for all lakes, including those without toxins for the full set of data. The intention is that this publication of the EMLS dataset will further demonstrate the feasibility and value of snapshot surveys, and encourage similar programs in a continuously changing environment, i.e. at times when datasets covering large geographical gradients are in great demand.
Methods
Organization
To make EMLS a robust survey we bridged two European COST Actions24, CyanoCOST25,26 (Cyanobacterial blooms and toxins in water resources: occurrence impacts and management) and NETLAKE27 (Networking Lake Observatories in Europe), planting the idea and promoting the benefits of an extended collaboration amongst researchers from all over Europe. This research was expanded to many other scientists not directly involved in these COST Actions.
The EMLS protocols required that each group of data providers collected and handled the samples following the same standards. Therefore, the steps outlined below had to be followed by all participant groups to ensure standardized sampling, sample processing and analyses, resulting in a homogenous dataset. Any deviations from these protocols were recorded in the metadata spreadsheets, and these data were handled with care after contacting the data collectors. To ensure that protocols were fit-for-purpose and understandable to everyone, we invited representatives of each country involved in the CyanoCOST and NETLAKE actions to a three-day training workshop in Evian-Les-Bains, France. During the workshop, participants discussed all aspects of EMLS and considered limitations in the financial and logistic means for given countries, without compromising research quality. They finalized the protocols and obtained hands-on experience in using them. To increase the number of studied lakes and achieve adequate spatial variation, representatives of each country acted as EMLS-ambassadors, reaching-out to further collaborators within their own countries and disseminating the decisions and protocols of the EMLS.
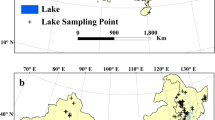
The EMLS was a collective effort, which means that each participating group used their own financial means to conduct their sampling as well as provided the personnel and facilities needed. Since the EMLS was a zero-funding effort, individual countries mainly contributed with samples from lakes that they routinely sample anyway, especially lakes with a history of eutrophication, given the implications for lake management. Although this results in a bias towards productive lakes, unfortunately it also reflects reality, with many lakes in Europe still suffering from eutrophication. A total of 369 lakes were sampled, spanning from Cyprus to Finland, and from the Asian part of Turkey to the Portuguese Azores islands (Fig. 1).
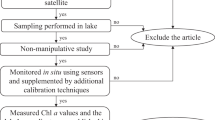
The workflow for organizing the EMLS consortium is presented as an infographic (Fig. 2). It illustrates the logistics from organizing the local surveys (1) through obtaining the samples (2), to processing and shipping them to the analytical laboratories (3-5). The data obtained from the field as well as laboratory-analyses was quality controlled and integrated into a unique dataset before making available to the EMLS network and to the rest of the scientific community. The methods of data acquisition and laboratory analyses that are described below are expanded versions of descriptions in our related work23.
CyanoCOST and NETLAKE members performed the sampling routine by: 1. Preparing the sampling material to meet the standardized procedures 2. Accessing the lake site and acquiring integrated water samples from the bottom of the thermocline up to the water surface 3. Processing and preserving the water samples based on requirements for subsequent analysis 4. Shipping samples to a central receiving laboratory 5. Analysing nutrient, pigment and toxin concentrations in dedicated laboratories. After quality control checks and data integration the dataset returns to the European network and becomes publically available for further research. Created by Sarah O’Leary and Eilish Beirne.
Data acquisition
Date and location (in situ)
The field methods for the EMLS were designed to be completed within one field day for each lake, but typically the sampling itself could be completed within two hours. Remote, poorly accessible lakes required more time to reach and field crews had to plan accordingly. To optimize time investment, lakes in close proximity were covered in one sampling trip. Each sampling group was responsible to organize and prepare the sampling material and equipment for their sampling campaign. In several cases, sampling groups of different areas or even countries collaborated and shared material such as instruments and boats.
Each EMLS data collector team had to identify the right sampling period, defined as the warmest two-week period in summer, based on long-term air temperature data in each region, covering the last 10 or more years. This predefined time-period served to minimize confounding effects of seasonality. Each lake was sampled within this time window (Date; Table 1).
The sampling location for each lake was defined as the central point of the lake. If a particular lake had been previously sampled at a specific location for long-term monitoring, the sample location from the long-term monitoring was used instead of the lake centre. For lakes with more than one relatively isolated basin, individual basins were sampled separately when possible, and indicated as such in the dataset (e.g. TR_BEY_I and TR_BEY_II). The latitude and longitude of the sampling location were recorded with a GPS device and provided in decimal degrees according to the WGS84 coordinate system.
If cyanobacterial surface blooms (defined as the presence of a visible surface scum) were present close to where the team entered the lake or in close proximity to the sampling point, a second sampling location was considered. A scoop surface sample, using a small sealable container like a Falcon tube, of the cyanobacterial scum was acquired along with the water column sample. The location of the scum sample was noted. Data collectors also provided - when available – maximum depth, mean depth and altitude (Table 1) of the lake.
Temperature and Secchi depth (in situ)
Temperature profiles were measured with available probes, such as various CTDs or a Fluoroprobe (BBE Moldaenke). If no profiling instrument was available, water samples were taken at 0.5 m intervals from the lake surface to the bottom of the thermocline (top of the hypolimnion) and water temperature was measured using hand-held thermometers directly after sampling. Here, the thermocline range was defined as the depth interval at which the rate of temperature decreased at least 1 °C per metre. The bottom of the thermocline (where temperature no longer decreased by 1 °C per meter), defined the sampling depth in the case of stratified lakes. For lakes where a thermocline was not observed (mostly shallow lakes), the temperature profile was measured until the lake bottom. In this case, the sampling depth was determined at 0.5 m above the lake bottom. From the temperature profiles, we obtained surface and epilimnetic temperature (Table 1) as the average temperature from surface until the bottom of the thermocline. The temperature profiles were also used during data analysis, to calculate the location where the thermocline lies even between two temperature measurement depths, which corresponds to where the thermocline is the most stable (point of maximum buoyancy frequency). To calculate this thermocline depth (ThermoclineDepth_m, Table 1) we used the command thermo.depth from the R package rLakeAnalyzer28. The Secchi depth (Table 1) was also recorded using a Secchi disk to the nearest 0.05 m.
Integrated water sample (in situ)
All data collectors constructed a simple device, known as the “Anaconda”, using a stoppered hose of the correct length in order to acquire the epilimnetic sample. The hosepipe was lowered with the bottom end open into the water column until the right depth (see above). When the hosepipe was vertical and the water level was visible at the surface layer of the hosepipe then the stopper was inserted to create hydrostatic pressure. The bottom end of the hosepipe was pulled-up with a rope to the surface to collect the sample in a bucket. The diameter of the hosepipe was appropriate to sample the required water volume (about 5–10 L for hypertrophic and eutrophic, 15–30 L for mesotrophic and oligotrophic lakes) for the analyses, in an acceptable number of runs. The first three sampling runs served the purpose of rinsing the hosepipe, the sampling bucket and the plastic rod. The subsequent runs were the water sample taken for analysis. The water sample in the bucket was mixed adequately before being divided into different bottles for further processing prior to analysis.
All samples were shipped frozen using dry ice in Styrofoam boxes. Shipping and storage of the EMLS samples was centralized at the University of Wageningen (The Netherlands). There, samples were sorted and sent to the dedicated laboratories for further analysis. Each of the nutrients, pigments and toxins analyses were done in one dedicated laboratory, by one operator on one machine, to minimize analytical errors and maximize integration of the datasets. Specifically, the nutrients, microcystins and nodularin analyses were done at the University of Wageningen, the pigment analysis at the University of Amsterdam and the cylindrospermopsin and anatoxin analysis at the German Environment Agency.
Total and Dissolved Nutrients (laboratory)
For analyses of total phosphorus (TP_mgL, Table 1) and nitrogen (TN_mgL, Table 1), unfiltered water subsamples of 50 mL were stored in -20 °C until shipping. For dissolved nutrients: orthophosphate (PO4_ugL, Table 1), nitrite/nitrate (NO3NO2_mgL, Table 1) and ammonium (NH4_mgL, Table 1), a volume of 250 mL was filtered through 47mm Glass fibre filters (GF/C or GF/F or similar), the filtrate was sampled in a PE bottle and stored at −20 °C. Before the collection of the nutrient samples, all polyethylene collection bottles with their screw caps were acid washed overnight in 1M HCl and rinsed with demineralized water and lake water before collection. Nutrients were measured according to Dutch NEN standards, using a Skalar SAN+ segmented flow analyser (Skalar Analytical BV, Breda, NL) with UV/persulfate digestion integrated in the system. The total phosphorus and orthophosphate were analysed conforming NEN29, the ammonium and total nitrogen according to NEN30 and the nitrite/nitrate following NEN31. The limit of detection was 0.02 mg/L for total phosphorus and ammonium, 0.2 mg/L for total nitrogen, 0.01 mg/L for nitrite/nitrate and 0.004 mg/L for orthophosphate.
Pigment analysis (laboratory)
For pigment analysis, a volume of 50–250 mL for hypertrophic and eutrophic lakes and 500–1000 mL for mesotrophic to oligotrophic lakes was filtered through 47mm glass fibre filters (GF/C or GF/F or similar) using a filtration device. Filters were stored at −20 °C in the dark until shipping. The analysis of pigments was modified from the method described by Van der Staay et al.32. All filters were freeze dried for 6 h. Filters were cut in half, placed in separate Eppendorf tubes, and kept on ice until the end of the extraction procedure. In each tube, 600 μl of 90% acetone were added with a small amount of 0.5 mm glass beads. To extract the pigments from the phytoplankton cells, filters were placed on a bead-beater for one minute. To increase the extraction yields, samples were placed in an ultrasonic bath for ten minutes. This procedure was repeated twice to ensure a complete extraction of the total pigment content of the filters. To achieve binding of the pigments during the High-Performance Liquid Chromatography (HPLC) analysis, 300 μl of a Tributyl Ammonium Acetate (1.5%) and Ammonium Acetate (7.7%) mix were added to each tube. Lastly, samples were centrifuged at 15, 000 rpm at 4 °C for ten minutes. 35 μl of the supernatant from both Eppendorf tubes of a filter were transferred into an HPLC glass vials. Pigments were separated on a Thermo Scientific ODS Hypersil column (250 mm × 3 mm, particle size 5 μm) in a Shimadzu HPLC machine and using a KONTRON SPD-M2OA diode array detector. We identified 12 different pigments (chlorophyll-a, chlorophyll-b, zeaxanthin, diadinoxanthin, fucoxanthin, diatoxanthin, alloxanthin, peridinin, chlorophyll-c2, echinenone, lutein and violaxanthin, Table 1) based on their retention time and absorption spectrum and quantified by means of pigment standards. The limit of detection (LOD) and for a 250 mL sample was: 0.094 μg/L for chlorophyll-a, 0.061 μg/L for chlorophyll-b, 0.034 μg/L for zeaxanthin, 0.053 μg/L for diadinoxanthin, 0.067 μg/L for fucoxanthin, 0.029 μg/L for diatoxanthin, 0.023 μg/L for alloxanthin, 0.085 μg/L for peridinin, 0.028 μg/L for chlorophyll-c2, 0.031 μg/L for echinenone, 0.027 μg/L for lutein and 0.075 μg/L for violaxanthin. The limit of quantification (LOQ) and for a 250 mL sample was: 0.315 μg/L for chlorophyll-a, 0.202 μg/L for chlorophyll-b, 0.115 μg/L for zeaxanthin, 0.177 μg/L for diadinoxanthin, 0.224 μg/L for fucoxanthin, 0.098 μg/L for diatoxanthin, 0.077 μg/L for alloxanthin, 0.284 μg/L for peridinin, 0.093 μg/L for chlorophyll-c2, 0.104 μg/L for echinenone, 0.089 μg/L for lutein and 0.250 μg/L for violaxanthin. In the cases where no pigment signal was detected, the respective pigment was considered absent and noted as 0 μg/L in the dataset. If the calculated pigment concentration in the dataset is above the limit of detection (qualitatively detected signal) but below the quantification limit (too small to quantify), we suggest the assignment of a very small value of half the detection limit to enable the inclusion of these samples in statistical analyses (if applicable). Alternatively, other statistical approaches that account for data censoring can be followed based on the research question and the statistical analysis followed (for suggestions see33).
Cyanotoxin analysis (laboratory)
For toxin analyses, a volume of 50–250 mL for hypertrophic and eutrophic lakes and 500-1000 mL for mesotrophic to oligotrophic lakes, was filtered through 47 mm Glass fibre filters (GF/C or GF/F or similar) using a filtration device. Filters were stored at −20 °C until shipping. In the laboratory, frozen filters were transferred to 8 mL glass tubes and freeze-dried (Alpha 1-2 LD, Martin Christ Gefriertrocknungsanlagen GmbH, Osterode am Harz, Germany). The freeze-dried filters were used for the Liquid Chromatography with tandem Mass Spectrometry detection (LC-MS/MS) analysis of microcystins, nodularin, cylindrospermopsin and anatoxin as described below. In the cases where no toxin signal was detected, the respective toxin was considered absent and noted as 0 μg/L in the dataset. A similar approach as in the section “pigment analysis” can be followed for toxin concentrations in the dataset that fall above the detection limit but below the quantification limit, as we did in23.
Microcystins and nodularin analysis (laboratory)
For the extraction of microcystins and nodularin, 2.5 mL of 75% hot methanol – 25% ultrapure water (v/v) was added to the freeze-dried filters, which were then sealed with a screw cap and placed for half an hour at 60 °C. Subsequently, the extract was transferred to a clean 8 mL glass tube. This extraction procedure was performed three times for each filter. The supernatants of the repeated extraction procedure were combined to a final volume of 7.5 mL and then dried in a Speedvac (Thermo Scientific Savant SPD121P, Asheville, NC, USA). After that, the extracts were reconstituted in 900 μL 100% MeOH. The reconstituted samples were transferred into 2 mL Eppendorf vials with a 0.22 μm cellulose-acetate filter and centrifuged for 5 min at 16, 000×g (VWR Galaxy 16DH, Boxmeer, Netherlands). Filtrates were transferred to amber glass vials for the analysis.
The LC-MS/MS analysis was performed on an Agilent 1200 LC and an Agilent 6410A QQQ (Waldbronn, Germany). The extracts were separated using a 5 μm Agilent Eclipse XDB-C18 (4.6 mm, 150 mm column, Agilent Technologies, Waldbronn, Germany) at 40 °C. The mobile phase consisted of Millipore water (v/v, eluent A) and acetonitrile (v/v, eluent B) both containing 0.1% formic acid at a flow rate of 0.5 mL/min with the following gradient program: 0–2 min 30% B, 6–12 min 90% B, with a linear increase of B between 2 and 6 min and a 5 min post run at 30% B (as described in32). The injection volume was 10 μl. Identification of the eight MC variants (MC_dmRR, MC_RR, MC_YR, MC_dmLR, MC_LR, MC_LY, MC_LW and MC_LF, Table 1) and nodularin (NOD) was performed in the positive Multiple Reaction Monitoring (MRM) with the following transitions: MC_dmRR 512.8 m/z [M + H]+ to 135.1 quantifier, MC_RR 519.8 m/z [M + H]+ to 135.1 quantifier, MC_YR 523.3 m/z [M + H]+ to 135.1 quantifier, MC_dmLR 491.3 m/z [M + H]+ to 847.6 quantifier, MC_LR 498.3 m/z [M + H]+ to 135.1 quantifier, MC_LY 868.4 m/z [M + H]+ to 163.0 quantifier, MC_LW 891.5 m/z [M + H]+ to 163.0 quantifier, MC_LF 852.5 m/z [M + H]+ to 163.0 quantifier and NOD 825.5 m/z [M + H]+ to 135.1 quantifier34. Mass spectrometric settings are given in35. Each MC variant was quantified against a calibration curve. The calibration curves were made using certified calibration standards obtained from DHI LAB Products (Hørsholm, Denmark). The limit of detection (LOD) and quantification (LOQ) for a 250 mL sample was: 0.0489 μg/L for MC_dmRR, 0.0358 μg/L for MC_RR, 0.0050 μg/L for MC_YR, 0.0054 μg/L for MC_dmLR, 0.0086 μg/L for MC_LR, 0.0817 μg/L for MC_LY, 0.0531 μg/L for MC_LW, 0.0206 μg/L for MC_LF and 0.0048 μg/L for NOD.
Cylindrospermopsin and anatoxin analysis (laboratory)
For the extraction of cylindrospermopsin (CYN, Table 1) and anatoxin-a (ATX, Table 1), 1.5 mL of 0.1% formic acid was added to the freeze-dried filters. Filters were sonicated for 10 min, shaken for 1 hour and then, centrifuged. This extraction procedure was repeated two more times and the combined supernatants were dried in a Speedvac (Eppendorf, Germany). Prior to analysis the dried extracts were re-dissolved in 1 mL 0.1% formic acid and filtered (0.2 μm, PVDF, Whatman, Maidstone, UK).
LC-MS/MS analysis was carried out on an Agilent 2900 series HPLC system (Agilent Technologies, Waldbronn, Germany) coupled to a API 5500 QTrap mass spectrometer (AB Sciex, Framingham, MA, USA) equipped with a turbo-ion spray interface. The extracts were separated using a 5 mm Atlantis C18 (2.1 mm, 150 mm column, Waters, Eschborn, Germany) at 30 °C. The mobile phase consisted of water (v/v, eluent A) and methanol (v/v, eluent A) both containing 0.1% formic acid, and was delivered as a linear gradient from 1% to 25% B within 5 min at a flow rate of 0.25 mL/min. The injection volume was 10 μL. Identification of CYN and ATX was performed in the positive MRM mode with the following transitions: CYN m/z 416.1 [M + H]+ to 194 (quantifier) and 416.1/176, and ATX m/z 166.1 [M + H]+ to 149, 166.1/131, 166.1/91, 166.1/43 (quantifier). Mass spectrometric settings are given in36. Certified reference standards were purchased from National Research Council (Ottawa, ON, Canada). The limit of detection (LOD) for both ATX and CYN was 0.0001 μg/L and the limit of quantification (LOQ) was 0.0004 μg/L for a 250 mL sample.
Code availability
Custom-made codes in R 3.3.337. were used to combine the datasets, trace missing data and inconsistencies. The codes are available in Zenodo: https://zenodo.org/record/1219878#.Wtcc4S5ubRZ
The GeoNode open source platform version 2.0 has been used for sharing the EMLS datasets among the partners. GeoNode is a web-based application that facilitates the visualization, download, sharing, and collaborative use of geospatial data through web services. GeoNode can be easily obtained at http://geonode.org/ as it is freely available under a GNU General Public License.
QGIS 2.18 Las Palmas was used for creating, managing and uploading the ESRI shapefile layers into the GeoNode platform.
Data Records
The final dataset includes all lake, environmental, nutrient, pigment and toxin data in one data table. The description of each feature in the table can be found in Table 1. The data table is made freely available as a static copy, through direct download from the online Environmental Database Initiative (EDI) and it is provided under the name “EMLSdata_10Aug_afterRev_dateformated.csv” (Data Citation 1).
The data table is also available at the GeoNode platform (http://gleon.grid.unep.ch/), where it can be downloaded through the provided web services that secure accessibility to data by using interoperable standards as provided by the Open Geospatial Consortium (OGC). The OGC- compliant web services available are 1. the Web Map Service (WMS 1.1.1) and 2. the Web Map Tile Service (WMTS 1.0.0) for accessing the maps; 3. the Web Feature Service (WFS 1.1.0) for accessing vector data and 4. Catalogue Service for the Web (CSW 2.0.2) to access the metadata. These interoperable web service endpoints enable the user to easily access and/or integrate these datasets in their desktop, web-based client, or own workflows.
The user can find and download several features of the data table as vector layers under the tab “Layers”. In the interactive tab “Maps”, the user can visualise and download datasets of combined “Layers”, or create their own maps using the available layers. The vector layers are provided in several formats such as: ESRI shapefile, Geography Markup Language (GML) and Keyhole Markup Language (KML), JPEG, pdf etc.
Both the database and the GeoNode platform can be easily updated with the expected data from DNA and flowcytometry analysis (to follow). As new surveys may be organized following the protocols of this paper, this data can also be easily included in the database and GeoNode platform.
Technical Validation
All data received from field observers (i.e. data in the tables: Lake Data & Metadata, Sampled data and depth profiles) were checked by a data curator before uploaded into the database.
Tracing missing data
Participating data collectors provided a field datasheet and a metadata sheet for each lake. All sheets for each sampling event and lake metadata which did not have matching records were double checked, and errors were corrected when found. A custom-made code generated proofing reports for each table, highlighting which lakes or basins had missing data.
Location data
All latitude and longitude records for each lake were verified by checking visually the provided locations on google maps. Lakes were marked as verified under the following conditions: (1) the location on google maps, using either satellite or map view, was found for a lake which matched the provided name, (2) the location on google maps was located close to a water body (approximately 1 km or closer) with a matching name, (3) the location was in or beside a lake which had no name or a different name, but the name provided matched with names of regions in the area (i.e. this represents cases where lakes were named according to their closest city, or region), or (4) the provided location was near an unnamed lake in the correct country where there were no other lakes nearby. All other cases were considered unverified and the data collectors were contacted to provide the right location (Latitude and longitude in decimal degrees; WGS84).
Laboratory Analyses
The nutrient, pigment and toxin concentrations were analysed centrally by certified laboratories that have optimized those specific analytical methods. Given that, these data are assumed to be correct.
Additional information
How to cite this article: Mantzouki, E. et al. A European Multi Lake Survey dataset of environmental variables, phytoplankton pigments and cyanotoxins. Sci. Data. 5:180226 doi: 10.1038/sdata.2018.226 (2018).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Carey, C. C. et al. Occurrence and toxicity of the cyanobacterium Gloeotrichia echinulata in low-nutrient lakes in the northeastern United States. Aquat. Ecol. 46, 395–409 (2012).
Huber, V., Wagner, C., Gerten, D. & Adrian, R. To bloom or not to bloom: contrasting responses of cyanobacteria to recent heat waves explained by critical thresholds of abiotic drivers. Oecologia 169, 245–256 (2012).
Sinha, R. L. et al. Increased incidence of Cylindrospermopsis raciborskii in temperate zones - is climate change responsible? Water Res 46, 1408–1409 (2012).
Schindler, D. W. & Vallentyne, J. R. In The algal bowl, overfertilization of the world's freshwaters and estuaries, (University of Alberta Press, Earthscan, 2008).
Codd, G. A., Meriluoto, J. & Metcalf, J. S. In Handbook of Cyanobacterial Monitoring and Cyanotoxin Analysis, (John Wiley & Sons, Ltd, 2017).
Dodds, W. et al. Eutrophication of U.S. freshwaters: analysis of potential economic damages. Environ. Sci. Technol. 43, 12–19 (2009).
Steffensen, D. A. In Cyanobacterial Harmful Algal Blooms: State of the Science and Research Needs (ed. Kenneth Hudnell 855–865 (Spinger Science, 2008).
Elliott, J. A. Is the future blue-green? A review of the current model predictions of how climate change could affect pelagic freshwater cyanobacteria. Water Res 46, 1364–1371 (2012).
Kosten, S. et al. Warmer climates boost cyanobacterial dominance in shallow lakes. Global Change Biol 18, 118–126 (2012).
Paerl, H. W. & Huisman, J. Blooms like it hot. Climate 320, 57–58 (2008).
Schwefel, R., Gaudard, A., Wuest, A. & Bouffard, D. Effects of climate change on deepwater oxygen and winter mixing in a deep lake (Lake Geneva): Comparing observational findings and modeling. Water Resour. Res. 52, 8811–8826 (2016).
Reynolds, C. S. In The ecology of phytoplankton. (Cambridge University Press, 2006).
Livingstone, D. M. Impact of secular climate change on the thermal structure of a large temperate central European lake. Climatic Change 57, 205–225 (2003).
Lurling, M., van Oosterhout, F. & Faassen, E. Eutrophication and warming boost cyanobacterial biomass and microcystins. Toxins 9, 64 (2017).
Moss, B. Allied attack: climate change and eutrophication. Inland Waters 1, 101–105 (2011).
Rigosi, A., Carey, C. C., Ibelings, B. W. & Brookes, J. D. The interaction between climate warming and eutrophication to promote cyanobacteria is dependent on trophic state and varies among taxa. Limnol. Oceanogr. 59, 99–114 (2014).
Salmaso, N. et al. Influence of atmospheric modes of variability on the limnological characteristics of large lakes south of the Alps: a new emerging paradigm. Hydrobiologia 731, 31–48 (2013).
Dai, A., Trenberth, K. E. & Qian, T. T. A global dataset of Palmer Drought Severity Index for 1870-2002: Relationship with soil moisture and effects of surface warming. Journal of Hydrometeorology 5, 1117–1130 (2004).
Ibelings, B. W., Vonk, M., Los, H. F. J., Van der Molen, D. T. & Mood, W. M. Fuzzy modeling of cyanobacterial surface waterblooms: Validation with NOAA-AVHRR satellite images. Ecol. Appl. 13, 1456–1472 (2003).
Gerten, D. & Adrian, R. Effects of climate warming, North Atlantic Oscillation, and El Nino-Southern Oscillation on thermal conditions and plankton dynamics in northern hemispheric lakes. ScientificWorldJournal 2, 586–606 (2002).
Pickett, S. In Long-Term Studies in Ecology: Approaches and Alternatives. Vol. 5. Space-for-time substitution as an alternative to long-term studies, (Springer New York, 1989).
Sommer, U. et al. Beyond the Plankton Ecology Group (PEG) model: mechanisms driving plankton succession. Annu. Rev. Ecol. Evol. Syst. 43, 429–448 (2012).
Mantzouki, E. et al. Temperature effects explain continental scale distribution of cyanobacterial toxins. Toxins 10, 156 (2018).
COST. European Cooperation in Science & Technology : Cost Actions– http://www.cost.eu/COST_Actions (2017).
CYANOCOST. COST Action ES1105: Cyanobacterial blooms and toxins in water resources: Occurrence, impacts and management http://www.cyanocost.com/ (2014).
Meriluoto, J. et al. Toxic cyanobacteria and cyanotoxins in European waters – recent progress achieved through the CYANOCOST Action and challenges for further research. AIOL 8 (2017).
NETLAKE. COST Action ES1201: Networking Lake Observatories in Europe https://www.dkit.ie/ga/node/14015 (2016).
Winslow, L. A., Read, J. S., Woolway, R. I., Brentrup, J. A., Leach, T. H. & Zwart., J. A. GLEON/rLakeAnalyzer: New algorithms and bugfixes. Zenodohttps://zenodo.org/record/1003169#.WzYwh9Uza70 (2016).
NEN. Water: photometric determination of the content of dissolved orthophosphate and the total content of phosphorous compounds by continuous flow analysis. Report No. NEN 6663 Netherlands Normalization Institute, (1986).
NEN. Water: photometric determination of the content of ammonium nitrogen and the sum of the contents of ammoniacal and organically bound nitrogen according to Kjeldahl by continuous flow analysis. Report No. NEN 6646 (Netherlands Normalization Institute, 1990).
NEN. Bepaling van het stikstofgehalte in de vorm van nitriet en in de vorm vannitraat en de som van beide met doorstroomanalyse (CFA en FIA) en spectrometrische detectie. Report No. NEN-EN-ISO 13395. Netherlands Normalization Institute, (1997).
Van der Staay, G. W. M., Brouwer, A., Baard, R. L. & van Mourik, F. Separation of photosystems I and II from the oxychlorobacterium (prochlorophyte) Prochlorothrix hollandica and association of Chlb binding antennae with PS II. Biochim. Biophys. Acta. 1102, 220–228 (1992).
Helsel, D. R. Fabricating data: How substituting values for nondetects can ruin results, and what can be done about it. Chemosphere 65, 2434–2439 (2006).
Faassen, E. J. & Lurling, M. Occurrence of the microcystins MC-LW and MC-LF in Dutch surface waters and their contribution to total microcystin toxicity. Mar. Drugs 11, 2643–2654 (2013).
Lurling, M. & Faassen, E. J. Dog poisonings associated with a Microcystis aeruginosa bloom in the Netherlands. Toxins 5, 556–567 (2013).
Fastner, J. et al. Fatal neurotoxicosis in dogs associated with tychoplanktic, anatoxin-a producing Tychonema sp. in mesotrophic lake Tegel, Berlin. Toxins 10, 60 (2018).
R Foundation for Statistical Computing, Vienna, Austria https://www.R-project.org/ (2016).
Data Citations
Mantzouki, E. et al. Environmental Data Initiative https://doi.org/10.6073/pasta/dabc352040fa58284f78883fa9debe37 (2018)
Acknowledgements
The authors acknowledge COST Action ES 1105 “CYANOCOST – Cyanobacterial blooms and toxins in water resources: Occurrence impacts and management” and COST Action ES 1201 “NETLAKE – Networking Lake Observatories in Europe” for contributing to this study through networking and knowledge sharing with European experts in the field. Evanthia Mantzouki was supported by a grant from the Swiss State Secretariat for Education, Research and Innovation (SERI) to Bas Ibelings and by supplementary funding from University of Geneva. We thank Wendy Beekman for the nutrient analysis and the University of Wageningen for covering the costs of this analysis from the personal funding of dr. Miquel Lürling. We thank Pieter Slot for assisting with the pigment analysis and the University of Amsterdam for covering the costs of the analysis through funding from the group of Prof. Jef Huisman and dr. Petra Visser (IBED). We would like to thank the Environmental Data Initiative for covering the cost of archiving the EMLS dataset. We would like to thank the Leibniz Institute of Freshwater Ecology and Inland Fisheries, Dept. of Experimental Limnology and the Aquatic Microbial Ecology Group for logistic and technical support of J. Fonvielle and H.-P. Grossart; and the Leibniz Association for financial support. The collection of data for Lough Erne and Lough Neagh were funded by the Department of Agriculture, Environment and Rural Affairs, Northern Ireland.
Author information
Authors and Affiliations
Contributions
Evanthia Mantzouki coordinated the EMLS, collected data, curated the dataset, analysed the pigment data and wrote the manuscript. Bas Ibelings conceived the idea for the EMLS, contributed to discussions throughout the study and to writing of the manuscript. James Campbell created the curation scripts, curated the dataset and wrote the manuscript. Emiel E. Van Loon and Petra Visser supervised the work of James Campbell and helped writing the manuscript. Iosif Constantinou developed the GeoNode platform, created the QGIS layers, uploaded and edited them in the platform. Maria G. Antoniou and Gregory Giuliani were involved in supervising Iosif Constantinou and brainstorming about developing the GeoNode platform. Jutta Fastner, Miquel Lürling and Elisabeth J. Faassen performed the toxin analyses. Sarah O’Leary did the graphic design and illustration of the infographic, Eilish Beirne did the graphic design of the infographic. The rest of the coauthors were responsible for data collection in lakes in their respective countries, for providing invaluable feedback and for finalizing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
ISA-Tab metadata
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Mantzouki, E., Campbell, J., van Loon, E. et al. A European Multi Lake Survey dataset of environmental variables, phytoplankton pigments and cyanotoxins. Sci Data 5, 180226 (2018). https://doi.org/10.1038/sdata.2018.226
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2018.226
This article is cited by
-
Chlorophyll soft-sensor based on machine learning models for algal bloom predictions
Scientific Reports (2022)
-
The effects of secondary bacterial metabolites on photosynthesis in microalgae cells
Biophysical Reviews (2022)
-
First occurrence of Cylindrospermopsin in the Azores (Lake São Brás, S. Miguel Island)
Limnology (2021)