Abstract
Sustained observations of microbial dynamics are rare, especially in southern hemisphere waters. The Australian Marine Microbial Biodiversity Initiative (AMMBI) provides methodologically standardized, continental scale, temporal phylogenetic amplicon sequencing data describing Bacteria, Archaea and microbial Eukarya assemblages. Sequence data is linked to extensive physical, biological and chemical oceanographic contextual information. Samples are collected monthly to seasonally from multiple depths at seven sites: Darwin Harbour (Northern Territory), Yongala (Queensland), North Stradbroke Island (Queensland), Port Hacking (New South Wales), Maria Island (Tasmania), Kangaroo Island (South Australia), Rottnest Island (Western Australia). These sites span ~30° of latitude and ~38° longitude, range from tropical to cold temperate zones, and are influenced by both local and globally significant oceanographic and climatic features. All sequence datasets are provided in both raw and processed fashion. Currently 952 samples are publically available for bacteria and archaea which include 88,951,761 bacterial (72,435 unique) and 70,463,079 archaeal (24,205 unique) 16 S rRNA v1-3 gene sequences, and 388 samples are available for eukaryotes which include 39,801,050 (78,463 unique) 18 S rRNA v4 gene sequences.
Design Type(s) | observation design • time series design • taxonomic diversity assessment by targeted gene survey • biodiversity assessment objective |
Measurement Type(s) | rRNA_16S • rRNA_18S |
Technology Type(s) | amplicon sequencing |
Factor Type(s) | temporal_interval • sampling depth • Superkingdom |
Sample Characteristic(s) | marine metagenome • Maria Island • Kangaroo Island • Sydney • Rottnest Island • North Stradbroke Island • Townsville • Darwin • marine biome |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Bacteria, Archaea and microbial eukaryotes dominate ocean ecosystems, composing 90% of marine biomass1. They display a vast phylogenetic and functional diversity, and their combined metabolic activities control the biogeochemical cycles that drive critical ecosystem services. But our oceans are changing rapidly and are under threat from both natural and man-made stressors, including rising seawater temperatures, ocean acidification, over-exploitation of resources and increasing impacts from a growing tourism industry and higher population densities. Baseline census data concerning the identity and function of marine microbes provide a benchmark against which we can assess assemblage changes.
Australia has the 7th longest coastline in the world, stretching ~36 000 km. Over the last decade, sustained temporal observations of many oceanographic parameters around Australia’s coastline have been made possible by the government funded Integrated Marine Observing System (IMOS). IMOS was established in 2007, and operates a network of national reference stations (NRS) that consist of moored sensors and regular vessel-based sampling2 (for a description of the rationale, design and implementation of IMOS NRS see http://imos.org.au/fileadmin/user_upload/shared/ANMN/NRS_rationale_and_implementation_100811.pdf).
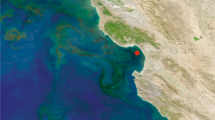
Marine microbial ecologists from a number of research institutions and universities have come together under the umbrella of the Australian Marine Microbial Biodiversity Initiative (AMMBI) to facilitate inclusion of microbial monitoring into the IMOS NRS infrastructure. Samples are collected at seven IMOS NRS: Port Hacking (site code: PHB), North Stradbroke Island (NSI), Maria Island (MAI) Rottnest Island (ROT), Yongala (YON), Darwin Harbour (DAR) and Kangaroo Island (KAI) (Fig. 1).
These IMOS NRS locations span over 30° of latitude and have been designed to maximize representation of the important marine biomes that are present around the continent2. Climatically they vary from tropical and monsoonal to cool temperate and highly seasonal habitats. All sites are affected to various degrees by basin scale events including the El Niño Southern Oscillation (ENSO) and the Indian Ocean Dipole (IOD), which are known to alter local oceanography, such as by enhancing upwelling3, and local biology, including fisheries4. Furthermore, these inter-annual events have a large influence on Australia’s two southerly flowing boundary currents, the Leeuwin Current (LC) on the Indian Ocean bounding west coast, and the East Australia Current (EAC) on the Pacific Ocean bounding east coast5. Both these currents transport warm, oligotrophic waters from the tropics to temperate latitudes. Changes in the extent and timing of southerly flow from these currents are having a large impact on both water temperatures6 and species ranges in southern Australia7,8, influencing fisheries9 and endangering native populations. However, little is known concerning the movements, adaption or range expansion of marine microbes.
Methods
Microbial sample collection
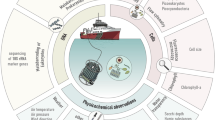
IMOS NRS sampling occurs using a variety of ocean going vessels dependent on local conditions. At all NRS, water samples are collected from between three and six pre-determined depths (depending on the water column depth) using Niskin bottles on either lines or a rosette. For each depth, a funnel and dark, sterile bottle are rinsed with 0.5 L of the retrieved seawater, and the bottle filled. In cases where filtration is conducted on shore (within 2 hours of collection) bottles are stored in a cooler with ice, out of direct sunlight. Microbial cells are collected by filtration of 2 L seawater through 0.2 μm pore Sterivex GP filter (Millipore, Massachusetts. Cat. # SVGPL10RC), using a Masterflex L/S Compact peristaltic pump fitted with an L/S 8 channel head (Cole Parmer). Pump tubing is rinsed with ~200 ml seawater from the appropriate depth prior to cell collection. Pumping continues for 1 min after sample has cleared the filter to dry. Both ends of the filter are capped and placed in individual snap-lock bags and kept on ice until being stored at −80 °C.
All AMMBI samples are collected in conjunction with IMOS NRS sampling (Fig. 1), so they are embedded within an extensive, quantitative, physical, chemical and biological oceanographic context. Contextual datasets from NRS are available for public search and download through the IMOS curated Australian Ocean Data Network Portal (https://portal.aodn.org.au/). A snapshot of IMOS data that corresponds to samples presented in this manuscript is available on Figshare (Data Citation 1). Parameters measured include conductivity, temperature, density, turbidity, fluorescence, PAR, Secchi depth, total alkalinity, total dissolved inorganic carbon, zooplankton dry weight, zooplankton size class, phytoplankton biomass, flow cytometry phytoplankton counts, phyto- and zooplankton microscopy identification and counts, HPLC pigment analysis, dissolved oxygen and nutrients including nitrite+nitrate, phosphate, silicate and ammonia. Further, in-situ moored sensor packages provide high-resolution data including acoustic doppler current profilers (ADCP), enabling the oceanographic context at the time of sampling to be accurately described. In addition, at Maria Island, Darwin, Yongala and North Stradbroke Island, surface expressions provide real-time meteorology, including wind-speed, direction, barometric pressure, humidity, air temperature and precipitation.
Sampling sites
An overview of the locations, depths and bioregions sampled and data produced by this project is provided in Table 1. All sites are located on the Australian continental shelf.
Darwin NRS is located 4 nautical miles (nm) offshore, at a depth of 20 m (full tide) over sandy substrate in Darwin Harbour, a large inverse estuary near the city of Darwin, Northern Territory (population 116,215). It is in the tropical north with a monsoonal climate where the wet season occurs between December and April. This, the northernmost NRS in the network, occupies a broad, shallow, shelf-sea region, part of the Arafura-Timor Sea. During the Austral winter or “dry season”, oceanic waters predominate and the site is vertically well mixed. During the Austral summer or ‘wet season”, due to rainfall and terrestrial inputs, the water column can become highly stratified across periods from a few days up to a few weeks. The site has a large diurnal tidal movement of up to 8 m and significant amounts of suspended sediments. Samples are collected over a 12 h period during sampling days and mid tide samples used for microbial diversity and associated biogeochemical analyses.
Based on the Australian government bioregionalisation studies, Australia has six phytoplankton provinces10. Darwin NRS is located in phytoplankton province 1: diatom dominated shelf waters of the north-west Australia, the gulf of Carpentaria, Arafura Sea and Timor Seas.
Yongala NRS is located 11 nm offshore at a depth of 28 m over sandy substrate, mid-shelf between the mainland and the coral reef in the tropical Great Barrier Reef (GBR) Lagoon. The mooring is located near the Yongala Wreck, out from Cape Bowling Green, in the central GBR. The closest major city is Townsville (population 171 824), approximately 100 km to the north. The site is generally well mixed with competing influences of the south-eastward lagoonal branch of the EAC and the opposing south-easterly trade wind forced coastal current. However, the monsoonal climate brings rain during December to April, and may result in river plume impingement and stratification. This is a site where significant ocean acidification projects are being carried out.
Phytoplankton province 3: fast growing nanoplankton diatom dominated shallow waters of the Great Barrier Reef Lagoon.
North Stradbroke Island NRS is located 6.6 nm north east of North Stradbroke Island at a depth of 60 m over sandy substrate. It is 30 km southeast of the major city of Brisbane, Queensland (population 2.099 million), at the opening to large, shallow, Moreton Bay. The site is impacted by the southerly flowing EAC and its eddies, which may cause periodic nutrient enrichment through upwelling. This latitude is the biogeographic boundary for many tropical and subtropical species. The water column is well mixed between May-August and stratified for the remainder of the year and salinity may at times be affected by floodwaters from the nearby Brisbane River outflow.
Phytoplankton province 3: fast growing nanoplankton diatom dominated shallow waters of the Great Barrier Reef Lagoon
Port Hacking NRS is located 3 nm offshore at 100 m depth over fine muddy sand, near the major city of Sydney, New South Wales (population 4.3 million). This is a sub-tropical to temperate location with strong seasonality. The site is just downstream of the EAC separation zone and is impacted by the dynamics of flow of EAC and its eddy field. The water column is very well mixed between May and Sept (although the duration of this mixing has decreased in recent years (Ingleton, T. unpublished observation)) and highly stratified between December and March. Upwelling can occur via eddies or wind driven slope water intrusions. There is a long historical oceanographic dataset from this site (with intermittent gaps in certain parameters) dating back to 1953.
Phytoplankton province 4: the productive temperate neritic province comprising coastal waters of New South Wales, Tasmania, Victoria and South Australia.
Maria Island NRS is located 4 nm offshore from Maria Island at 80 m depth over muddy sand, on the eastern coast of Tasmania. This is the southernmost NRS. The nearest population centers are Orford (population 485) and Triabunna (population 796). The site is seasonally impacted by the most southerly extent of the EAC, which has been increasing over the past 60 years, resulting in rapid ocean warming in this region6. MAI may also be influenced seasonally by the LC. The water column is well mixed year-round with some slight stratification between November and March. There is a long historical oceanographic dataset from this site (with intermittent gaps in certain parameters) dating back to 1944.
Phytoplankton province 4: the productive temperate neritic province comprising coastal waters of New South Wales, Tasmania, Victoria and South Australia
Kangaroo Island NRS is located 4.5 nm west of Kangaroo Island, at 110 m depth over medium- to coarse-grained carbonate sand with conspicuous large bryozoan and gastropod skeletons, on a broad shelf in the Great Australian Bight, 112 km southwest of Adelaide (population 1.3 million). The NRS site is subject to episodic upwelling in the summer/autumn and is impacted by the eastward flow of the warm LC and outflow of dense Spencer Gulf water during the winter. Summer waters are stratified, with warm surface waters overlying a pool of cool enriched water upwelled from the Flinders Current (a northern boundary current) which flows westward along the continental slope. A significant volume of cooler water is forced above the critical depth, promoting high productivity. Winter waters are impacted by a significant outflow of dense Spencer Gulf water, so are well mixed, downwelling dominates, and productivity is significantly reduced.
Phytoplankton province 4: the productive temperate neritic province comprising coastal waters of New South Wales, Tasmania, Victoria and South Australia
Rottnest Island NRS is located 18 nm offshore and 2.5 nm from Rottnest Island, at a depth of 50 m over sand but surrounded by low relief limestone reef which is predominately covered with macro-algae, mainly Ecklonia spp, on the Western Australia coast, near Perth (population 1.83 million). This is the only NRS situated on the Indian Ocean coastline. This station is heavily impacted by the LC, which delivers warm but less saline waters southward. The LC is strongly affected by ENSO and IOD events. The prevailing winds and current direction leads to coastal suppression of upwelling along the Western Australian coast throughout most of the year, and consequently sea surface temperatures of up to 4–5 °C warmer than upwelling systems at similar latitudes elsewhere on the globe2. There is an historical oceanographic dataset from this site (with intermittent gaps in certain parameters) dating back to 1951.
Phytoplankton province 2: the tropical neritic communities carried southwards by the Leeuwin Current
DNA extraction
All samples are couriered on dry ice, or in dewars filled with liquid nitrogen, to the Commonwealth Scientific and Industrial Research Organisation Oceans & Atmosphere (CSIRO O&A) laboratories in Hobart, Tasmania. DNA is extracted and purified using the PowerWater® Sterivex™ DNA Isolation Kit (MOBIO laboratories, Carlsbad, CA), following a slightly modified version of the manufacturer’s instructions11. The quality and quantity of DNA is checked using a NanoDrop™ 8000 Spectrophotometer (Thermo Scientific™) and DNA aliquot into multiple plates, vacuum dried and archived at -80 °C.
Further documentation outlining the standard operating procedures for sampling and DNA extraction is available at https://data.bioplatforms.com/organization/pages/bpa-marine-microbes/methods
DNA sequence generation
Generation of sequence data for this project is supported by Bioplatforms Australia (BPA). All DNA amplification and sequencing is carried out at the Ramaciotti Centre for Genomics (UNSW Sydney, Australia).
Bacterial, archaeal and eukaryotic assemblages are surveyed using small-subunit ribosomal RNA gene amplicon sequencing methods. Amplicons are prepared using Bacterial 16 S rRNA gene primers 27 F – 519 R (refs 12,13), archaeal 16 S rRNA gene primers A2F/Arch21f – 519 R (refs 12,14) and eukaryotic 18 S rRNA gene primers TAReuk454FWD1 and TAReuk- Rev3 (ref. 15) (Table 2). (We note that the reverse primer used for the 18 S V4 region (TAReuk- Rev3) may discriminate against Haptophytes, a bias which should be considered when using the data, and which has been corrected in primer sets described by Piredda et al.16).
Amplicons are sequenced using the duel indexed paired end approach on the illumina MiSeq platform according to the manufactures instructions. Further documentation outlining the standard operating procedures for generating and sequencing amplicons is available at https://data.bioplatforms.com/organization/pages/bpa-marine-microbes/methods.
DNA sequence processing
To allow for the highest possible phylogenetic resolution, data are provided as single nucleotide variants. Sequences are analyzed in a strictly standardized fashion alongside other Australian microbial biodiversity initiatives including the Biomes of Australian Soils (BASE) project17 enabling direct comparison and integration.
For each amplicon, each plate of illumina data is submitted to the following workflow separately, to identify correct biological sequences and build sample-by-read abundance matrices for samples on that plate18.
- 1
-
2
FASTA format sequences are extracted from FASTQ files and those<400 base pairs (bp) in length (or<350 bp for the 18 S amplicon) or containing N's or homopolymer runs of>8 bp are removed using MOTHUR, (v1.34.1)20,21.
-
3
Using the USEARCH 64 bit v10.0.240 (ref. 22) package, sequences are de-replicated (-derep_fulllength command), ordered by abundance and sequences with<4 representatives removed (-sortbysize command)
-
4
Chimeras are removed and biologically correct, zero-radius operational taxonomic units (zOTUs) identified (unoise3 command).
-
5
All quality-filtered sequences (from (2)) are then mapped to chimera-free zOTUs and a sample by read abundance table created using (-usearch_global)
-
6
Sample-by-read abundance matrices from each plate are merged based on the unique zOTU sequence, providing a sample-by-read abundance matrix for all samples in the dataset.
-
7
A final sample-by-read abundance data matrix is created by discarding sequences that are either not identified or not identified correctly for the given assay at the kingdom and phylum level (note this does not remove chloroplasts or mitochondrion from the Bacteria assay).
-
8
zOTUs are taxonomically classified with the SILVA v132 (ref. 23) database using MOTHUR’s implementation of the Wang classifier24 and a 60% bayesian probability cut-off. Eukaryotic zOTUs are also taxonomically classified using the Protist Ribosomal Reference database (PR2)25.
-
9
zOTUs can be readily clustered into OTUs of any similarity of the researchers choosing for comparison to other studies using a number of algorithms e.g. USEARCH cluster_fast26.
Data Records
Each data record represents a molecular description of the relative abundance taxonomic profile of the microbial assemblage at a particular site, depth and time. Each record is associated with a sample and each sample is provided with a unique ID from the Australian National Data Service. All sequence data are made publically available through the National Center for Biotechnology Information (Data Citation 2). Currently datasets for 952 samples are publically available, which in total include 88,951,761 bacterial (representing 72,435 unique zOTUs) and 70,463,079 archaeal (24,205 zOTUs) small subunit rRNA gene sequences covering variable regions 1-3, and 39,801,050 (78,463 zOTUs) eukaryote small subunit rRNA gene sequences covering variable regions 4.
Paired end read (R1, R2) and indexed read (I1, I2) data in.fastq format are also available to download at https://data.bioplatforms.com/bpa/marine_microbes/amplicon.
All contextual datasets from IMOS NRSs are available for public search and download through the Australian Ocean Data Network Portal (https://portal.aodn.org.au/).
Files relating to Bacterial (labelled AMMBI_B16S_zotus_table_Silvav132_datarelease_20032018.txt), Archaeal (AMMBI_A16S_zotus_table_Silvav132_datarelease_20032018.txt) and Eukaryote (AMMBI_18 Sv4_zotus_table_Silvav132_PR2_datarelease_20032018.txt) datasets, which include zOTU sequence, sequentially indexed unique identifiers for each zOTU, sum of zOTU abundance in the dataset, zOTU sample by abundance table, SILVA v132 taxonomy string and PR2 taxonomy string (eukaryotes only) as well as a contextual data file (labelled AMMBI_Contextual.Data.Master.sheet20180131_NSD_submission) providing a snapshot of IMOS NRS environmental data harvested from the AODN at the time of manuscript submission are available on Figshare (Data Citation 1).
A search function enabling taxonomic and contextual data informed interrogation of, and download of data from, the latest sequence read abundance table is available from
https://data.bioplatforms.com/organization/about/australian-microbiome.
Technical Validation
AMMBI represents, to our knowledge, the first methodologically standardized, continental-scale, temporal microbial ocean-observing network. We have attempted to minimize biases in the data generation pipeline by using the same simple sampling protocol at all stations and centralizing each step of DNA extraction (CSIRO), sequencing (Ramaciotti Centre for Genomics) and bioinformatics (CSIRO) in core facilities. That is, apart from sampling, all steps of the project are carried out concurrently on samples from all stations. The congruence of our phylogenetic characterization of samples with what is known concerning marine microbial assemblages globally reveals the suitability of our approach.
The alphaproteobacterial family the Pelagibacteraceae (SAR11 clade) compose ~30% of the sequences from surface waters at each station27, while the SAR86 clade is the most common and abundant gammaproteobacterial taxa (Fig. 2a). The marine cyanobacteria Prochlorococcus and Synechococcus constitute the bulk of the bacterial phototrophic assemblage and vary in their relative abundance, with Prochlorococcus relatively more prevalent at stations which display seasonal oligotrophic conditions, such as NSI and YON (Fig. 2a)28. Similarly, the Euryarchaeota Marine Groups II and Thaumarchaeota are the most abundant archaeal taxa29 (Fig. 2c).
Profiles display the relative abundance of a) bacterial, b) plastid and c) archaeal taxa contributing to>0.1% of combined 16 S rRNA reads in samples collected at 0 m and 10 m depths over the course of the study. One two litre sample per depth was collected during each NRS sample trip (except Darwin where three samples per depth were taken at three hour intervals on two of the quarterly trips) and one DNA extraction and amplicon PCR performed per sample (MAI n= 81, PHB=87, NSI=90, YON=21, KAI=10, ROT=20, DAR=21). For this analysis, bacterial and archaeal taxonomic assignments were made using the GreenGenes database30 (release 13.5) and reads corresponding to chloroplast sequences were removed from the bacterial dataset and analysed independently with taxonomy assigned using the PhytoRef database based on 6,490 plastid 16 S rRNA gene sequences. Scripts used to generate Figs. 2 and 3 are available on Github at https://github.com/martinostrowski/marinemicrobes.
18S rRNA gene relative abundance data provides information concerning heterotrophic, symbiotic and parasitic eukaryotes including Metazoa and Amoebozoa, as well as mixotrophic and phototrophic taxa including Dinophyceae, Stramenopiles, Hacrobia (cryptomonads and haptophytes) and Archaeplastida (red and green algae) (Fig. 3). The overall diversity revealed within eukaryote taxa is exceptional (2,066 taxa identified). Analyses of taxa in sub-groups (Fig. 3b–f) reveals regional trends in species abundance and distribution. For example, several diatom taxa present distinct biogeographical preferences at individual NRS, particularly Chaetoceros, Thalassiosira and Leptocylindrus, while other species display a more cosmopolitan distribution, e.g. Pseudo-nitzschia. Noctiluca scintillans, a mixotrophic dinoflagellate linked to bright red and bioluminescent coastal blooms, is detected at stations along the East Coast (MAI, PHB and NSI; Fig. 3e). A number of species associated with harmful algal blooms are detected within the data (Alexandrium, Noctiluca and Gymnodinium) which highlights the potential of this initiative to enhance understanding of bloom dynamics within the context of a holistic record of microbial community structure.
Profiles display the relative abundance of a) all taxa contributing to>0.1% of combined 18 S rRNA reads in samples collected at 0 m and 10 m depths over the course of the study. Sample details as per Fig. 2. More detailed taxonomic resolution is provided for b) the Archeaplastida and Hacrobia, (c) the Diatoms (Bacilliarophyta), (d) the Opisthokonts, (e) the Dinophyceae and (f) the Stramenopiles (other than diatoms). Note the scale differences for plots B-F which reflect the abundance of sequences for each subgroup in the entire dataset.
Data on phototrophs is further enhanced by specific analysis of plastid 16 S rRNA genes that are retrieved using the bacterial 16 S gene assay (Fig. 2b). Plastid 16 S rRNA sequences comprise between 0.02 and 33.6% of total bacterial sequences in surface samples (<25 m depth). The plastid 16 S data is highly complementary with 18 S data and provides a proxy for the relative abundances of prokaryotic and eukaryotic phototrophs which can be used to fine-tune our understanding of regional phytoplankton provinces. For example, picoeukaryote species, Ostreococcus and Micromonas contribute a significant proportion of phytoplankton sequences, particularly at mid-latitude stations (MAI, KAI, ROT, PHB) while Cyanobacteria represent a greater proportion of 16 S sequences in low-latitude stations (DAR, YON, NSI).
Usage Notes
There is considerable redundancy in the microbial taxa sampled across the seven NRS, enabling us to collect baseline data about the seasonal dynamics of a wide variety of organisms across the breadth of their habitat range. Hence the scope of this project enables us to effectively elucidate microbial niche conditions and describe microbial phenology. Further, the data provide a critical baseline against which to measure, and from which to predict, changes in microbial assemblages under future global change scenarios. Indeed, AMMBI data is being used to hindcast abundances of individual microbial taxa to long-term historic oceanographic datasets, to infer decadal scale trends in microbial composition, and to forecast taxa response to future climate scenarios. As observations are ongoing we will reveal any shifts in organismal dynamics and distributions which result from basin-scale climatic events such as the ENSO in the Pacific regions and the IOD which has its greatest affect along Australia’s western coast. We are also using this data to test the phylogenetic breadth of bioregionalisation provinces currently assigned using microscopy counts of phytoplankton.
We encourage users to download data from the Australian Microbiome searchable data portal at https://data.bioplatforms.com/organization/about/australian-microbiome. Full usage requires free registration. To search the marine dataset, navigate to “Tools/Search Facility”. Under the “Contextual Filters” box on the right use the “Environment” dropdown menu to choose “Marine”. Additional contextual filters (e.g. Temperature) can be “added” in this box, including using “NRS location code/ Voyage Code” to choose samples from specific NRS stations (e.g. text includes “MAI”). Users are also able to “Filter on Amplicon” (27f519r_bacteria, A2f519r_arcahea or 18Sv4F18Sv4R_v4_eukaryote) and/or taxonomy based on SILVAv132 6 string format.
DNA from all samples is archived in multiple aliquots at -80C at CSRIO Oceans and Atmosphere in Hobart, Tasmania and is accessible for re-analysis should the proposed methods be deemed to provide a substantial improvement or progress over prior results.
Additional information
How to cite this article: Brown, M. V. et al. Continental scale monitoring of marine microbiota by the Australian Marine Microbial Biodiversity Initiative. Sci. Data 5:180130 doi: 10.1038/sdata.2018.130 (2018).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Azam, F. Microbial control of oceanic carbon flux: The plot thickens. Science 280, 694–696 (1998).
Lynch, T. P. et al. IMOS National Reference Stations: a continental-wide physical, chemical and biological coastal observing system. PLoS ONE 9, e113652 (2014).
Middleton, J. F. et al. El Nino effects and upwelling off South Australia. J Phys Oceanogr 37, 2458–2477 (2007).
Ward, T. M. et al. Pelagic ecology of a northern boundary current system: effects of upwelling on the production and distribution of sardine (Sardinops sagax), anchovy (Engraulis australis) and southern bluefin tuna (Thunnus maccoyii) in the Great Australian Bight. Fish Oceanogr 15, 191–207 (2006).
Feng, M., Weller, E., Hill, K. The Leeuwin Current in: (eds Poloczanska, E. S., Hobday, A. J. & Richardson, A. J. ) A Marine Climate Change Impacts and Adaptation Responses in Australia Report Card. NCCARF Publication, 05/09 (2009).
Ridgway, K. R. Seasonal circulation around Tasmania: An interface between eastern and western boundary dynamics. Journal of Geophysical Research-Oceans 112 (2007).
Poloczanska, E. S. et al. Climate change and Australian marine life. Oceanogr Mar Biol 45, 407–478 (2007).
McLeod, D. J., Hallegraeff, G. M., Hosie, G. W. & Richardson, A. J. Climate-driven range expansion of the red-tide dinoflagellate Noctiluca scintillans into the Southern Ocean. Journal of Plankton Research 34, 332–337 (2012).
Figueira, W. F. & Booth, D. J. Increasing ocean temperatures allow tropical fishes to survive overwinter in temperate waters. Global Change Biology 16, 506–516 (2010).
Hayes, D. et al. Collation and Analysis of Oceanographic Datasets for National Marine Bioregionalisation. A report to the Australian Government, National Oceans Office https://www.environment.gov.au/resource/collation-and-analysis-oceanographic-datasets-national-marine-bioregionalisationDepartment of the Environment and Heritage (2005).
Appleyard, S., Abell, G. & Watson, R. Tackling microbial related issues in cultured shellfish via integrated molecular and water chemistry approaches. Seafood CRC report No. 2011/729 (2011).
Lane, D. J. et al. Rapid-Determination of 16 s Ribosomal-Rna Sequences for Phylogenetic Analyses. Proc Nat Acad Sci USA 82, 6955–6959 (1985).
Lane, D. J. 16S/23S rRNA sequencingin: Nucleic Acid Techniques In Bacterial Systematics 115–175 (John Wiley and Sons, 1991).
Delong, E. F Archaea in coastal marine environments. Proc Nat Acad Sci USA 89, 5685–5689 (1992).
Stoeck, T. et al. Multiple marker parallel tag environmental DNA sequencing reveals a highly complex eukaryotic community in marine anoxic water. Mol Ecol 19, 21–31 (2010).
Piredda, R. et al. Diversity and temporal patterns of planktonic protist assemblages at a Mediterranean Long Term Ecological Research site. FEMS Microbiol Ecol 93, fiw200 (2017).
Bissett, A. et al. Introducing BASE: the biomes of Australian soil environments soil microbial diversity database. Gigascience 5, 21 (2016).
Andrews, S. Fastqc a quality control tool for high throughput sequence datahttp://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2014).
Magoc, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75, 7537–7541 (2009).
Rideout, J. R. et al. Subsampled open-reference clustering creates consistent, comprehensive OTU definitions and scales to billions of sequences. Peerj 2, e545 (2014).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Yilmaz, P. et al. The SILVA and "All-species Living Tree Project (LTP)" taxonomic frameworks. Nucleic Acids Res 42, D643–D648 (2014).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73, 5261–5267 (2007).
Guillou, L. et al. The Protist Ribosomal Reference database (PR2): a catalog of unicellulareukaryote Small Sub-Unit rRNA sequences with curated taxonomy. Nucleic Acids Res 41, D597–D604 (2013).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nature Methods 10, 996 (2013).
Morris, R. M. et al. SAR11 clade dominates ocean surface bacterioplankton communities. Nature 420, 806–810 (2002).
Biller, S. J., Berube, P. M., Lindell, D. & Chisholm, S. W. Prochlorococcus: the structure and function of collective diversity. Nature Rev Microbiol 13, 13–27 (2015).
Karner, M. B., DeLong, E. F. & Karl, D. M. Archaeal dominance in the mesopelagic zone of the Pacific Ocean. Nature 409, 507–510 (2001).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. Isme j 6, 610–618 (2012).
Data Citations
Brown, M. V. Figshare https://doi.org/10.6084/m9.figshare.5808864.v3 (2018)
NCBI Sequence Read Archive SRP114822 (2018)
Acknowledgements
The first three years of this project and part of the activities afterwards was funded by Australian Research Council awards DP0988002 to M.V.B. & JA.F., DP120102764 to J.S., M.V.B. & L.B., DP150102326 to M.V.B., M.O., JA.F. & L.B., the Environmental Genomics Project from CSIRO Oceans and Atmosphere and a CSIRO OCE Science Leader Fellowship to L.B.
Data sourced from and sample collection funded by the Integrated Marine Observing System (IMOS) — IMOS is a national collaborative research infrastructure, supported by the Australian Government. Sequencing and bioinformatics of this program is funded by Bioplatforms Australia (BPA). We would like to acknowledge the contribution of the Marine Microbes Project consortium (https://data.bioplatforms.com/organization/pages/bpa-marine-microbes/consortium) in the generation of data used in this publication. The Marine Microbes Project is supported by funding from Bioplatforms Australia through the Australian Government National Collaborative Research Infrastructure Strategy (NCRIS).
M.V.B., J.v.d.K., A.B., M.O., J.R.S. and L.B. had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Author information
Authors and Affiliations
Contributions
Designed the Project: L.B., M.V.B., J.v.d.K., M.O., J.R.S., T.M., A.F., A.B., J.A.F. Collected and prepared samples and contextual data: T.I., M.O., M.B., T.N., F.C., C.D., L.M., B.L., N.S., T.J., J.B.I., D.F., S.R., B.H., G.A., P.C., T.K., S.L.S.S., N.P., P.M., P.v.R., R.C., M.S., K.M., J.W., Prepared DNA: J.v.d.K., S.R., M.O., T.K., P.C., B.R., K.G., M.B., L.M., JAS Managed project logistics: L.B., J.v.d.K., M.V.B., M.O., J.R.S., A.B., C.S., I.P., P.v.R., A.F., A.R., T.I., J.K. Performed bioinformatics analysis: A.B., J.v.d.K., M.B., M.O. Performed data analysis: A.B., J.v.d.K., M.V.B., M.O., L.B. Wrote the paper: M.V.B., J.v.d.K., M.O., J.R.S., A.B., L.B. All authors have read and approved the manuscript
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
ISA-Tab metadata
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/ The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files made available in this article.
About this article
Cite this article
Brown, M., van de Kamp, J., Ostrowski, M. et al. Systematic, continental scale temporal monitoring of marine pelagic microbiota by the Australian Marine Microbial Biodiversity Initiative. Sci Data 5, 180130 (2018). https://doi.org/10.1038/sdata.2018.130
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2018.130
This article is cited by
-
A marine heatwave drives significant shifts in pelagic microbiology
Communications Biology (2024)
-
Exploring the status of global terrestrial and aquatic microbial diversity through ‘Biodiversity Informatics’
Environment, Development and Sustainability (2023)
-
Predicting global distributions of eukaryotic plankton communities from satellite data
ISME Communications (2023)
-
Biogeographical and seasonal dynamics of the marine Roseobacter community and ecological links to DMSP-producing phytoplankton
ISME Communications (2022)
-
Seasonal bacterial niche structures and chemolithoautotrophic ecotypes in a North Atlantic fjord
Scientific Reports (2022)