Abstract
Human mobility continues to increase in terms of volumes and reach, producing growing global connectivity. This connectivity hampers efforts to eliminate infectious diseases such as malaria through reintroductions of pathogens, and thus accounting for it becomes important in designing global, continental, regional, and national strategies. Recent works have shown that census-derived migration data provides a good proxy for internal connectivity, in terms of relative strengths of movement between administrative units, across temporal scales. To support global malaria eradication strategy efforts, here we describe the construction of an open access archive of estimated internal migration flows in endemic countries built through pooling of census microdata. These connectivity datasets, described here along with the approaches and methods used to create and validate them, are available both through the WorldPop website and the WorldPop Dataverse Repository.
Design Type(s) | data integration objective • observation design |
Measurement Type(s) | Human Migration |
Technology Type(s) | digital curation |
Factor Type(s) | |
Sample Characteristic(s) | Homo sapiens • Angola • Burundi • Benin • Burkina Faso • Botswana • Central African Republic • Cote d'Ivoire • Cameroon • Democratic Republic of the Congo • Republic of Congo • Comoros • Djibouti • Eritrea • Ethiopia • Gabon • Ghana • Guinea • Gambia • Guinea-Bissau • Equatorial Guinea • Kenya • Liberia • Madagascar • Mali • Mozambique • Mauritania • Malawi • Mayotte • Namibia • Niger • Nigeria • Rwanda • Sudan • Senegal • Sierra Leone • Somalia • South Sudan • Sao Tome and Principe • Swaziland • Chad • Togo • Tanzania • Uganda • Republic of South Africa • Zambia • Zimbabwe • Afghanistan • Azerbaijan • Bangladesh • Bhutan • China • Georgia • Indonesia • India • Iran • Cambodia • South Korea • Laos • Sri Lanka • Myanmar • Malaysia • Nepal • Pakistan • The Philippines • Papua New Guinea • North Korea • Saudi Arabia • Solomon Islands • Thailand • Tajikistan • Timor-Leste • Viet Nam • Vanuatu • Yemen • Argentina • Belize • Bolivia • Brazil • Colombia • Costa Rica • Dominican Republic • Ecuador • Guatemala • Guyane • Guyana • Honduras • Haiti • Mexico • Nicaragua • Panama • Peru • Paraguay • El Salvador • Suriname • Venezuela • anthropogenic habitat |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
According to the International Organization for Migration1 and The World Bank2, without accounting for seasonal and temporary migrants, more than 1 billion people are currently living outside their places of origin, with about 740 million of them classified as internal migrants. Additionally, in 2014 around 67 million passengers travelled on international and domestic flights every week3 and hundreds of millions are estimated to commute daily by public transport and private vehicles4. Human mobility is expected to continue rising in volume and reach, producing increasing global connectivity that has a range of impacts, including rising numbers of invasive species, the spread of drug resistance, and disease pandemics. In this context, quantifying human mobility across multiple temporal and spatial scales, becomes crucial for quantifying its effects on society5–7, evaluating its relationship with the environment8,9, better understanding human-related processes such as urbanization and land use change10–13, and providing a strong evidence base to support both development14–16 and public health17–19 applications and policies.
In public health, the role of human mobility in the spread of infectious diseases is exemplified by the presence of HIV/AIDS in areas outside where it first emerged at the beginning of the twentieth century20–22, the 2003 SARS epidemic23, the 2007 Chikungunya outbreaks in Italy and France24,25, the 2009 H1N1 pandemic26, the 2014 Ebola outbreak in Western Africa27, the resurgence of malaria cases in areas where the disease was once eliminated28, and the worldwide spread of drug resistant pathogens29. Consequently, it is clear that to provide better informed guidelines, both at the national and international level, the effects of human mobility and connectivity in driving disease dynamics need to be better understood and accounted for refs 30–33.
Local malaria elimination and global malaria eradication are rising up the international agenda34–36. Evidence from the previous global malaria eradication program37, as well as from recent studies, control campaigns, and elimination efforts38–41 highlight the importance of accounting for human mobility in designing elimination plans. Infected people may unknowingly transport malaria parasites (potentially including antimalarial-resistant strains42) into new areas. Parasites can be imported either from other countries43 or from other areas within the same country44. Thus, because of the flow of imported cases from high- to low-transmission settings, the latter will face difficulties in achieving elimination and maintaining malaria-free status if it is achieved43. Nevertheless, despite the importance of these dynamics being long recognized45,46, attempts to translate human mobility model outputs into malaria policy are still rare47.
As detailed in Tatem7, sources of human mobility data potentially useful for modelling pathogen movements include: air and sea travel data records (including open access modelled versions of them); census migration data; travel history and displacement surveys; GPS tracking data and volunteered geographic information (with the latter including geolocated social media data), and even satellite night-time light data. In particular, patient travel history data, containing detailed demographic information and travel motivations, are traditionally used to understand malaria parasite importation patterns48–50. Recently, mobile phone call detail records (CDRs) have been increasingly used for measuring short-term human movements51,52 and thus, either alone38,53,54 or in combination with travel history data55 and malaria case data, for supporting malaria control and elimination strategic planning.
However, because of difficulties in sharing and accessing CDRs (mostly due to commercial and privacy concerns)7,56,57, alternative datasets are required in order to quantify and map internal connectivity across continental scales. To this end, using CDRs, Wesolowski et al.56 and Ruktanonchai et al.58 demonstrated that widely-available and easy-to-obtain census-derived internal migration flow data can serve as reliable proxies for the relative strength of within-country human connectivity across multiple temporal scales.
Within the framework of the WorldPop Project (www.worldpop.org), and following the approaches described in Henry et al.59 and Garcia et al.60 (Fig. 1), internal census-based migration microdata available through the online IPUMS-International (IPUMSI) database61, along with a number of other ancillary datasets, were assembled and processed to produce an open access archive of estimated 5-year (2005–2010) internal human migration flows for every Plasmodium falciparum and Plasmodium vivax (hereafter simply referred as Pf and Pv, respectively) endemic country62,63 (Supplementary Table 1).
Methods
Estimating internal migration flows between administrative units
Following Garcia et al.60 a gravity model-based approach was used to estimate the total number of people migrating from one administrative unit to any other administrative unit, between 2005 and 2010, within each malaria endemic country located in Africa, Asia, Latin America and the Caribbean62,63 (Supplementary Table 1).
The simplest gravity-type spatial interaction model, proposed by Zipf64, considers the total population in a location of origin i and in a location of destination j (henceforth simply indicated as i and j), and the distance between the two locations to predict the migration flow (MIGij) between them. Thus, migration flows between administrative units can be estimated using the following function:
where and represent the populations in the location of origin i and of destination j, respectively, and represents the distance between i and j; with α, β, and γ being parameters, used to indicate the magnitude of the effect for each covariate, that are typically estimated in the statistical modelling framework.
In this study, following the notation from Henry et al.59 and Garcia et al.60, the basic gravity-type spatial interaction in equation (1) was extended in order to include additional geographical and socioeconomic factors described in detail in the Data collection and preparation subsection below. Since the census-based migration microdata extracted from the IPUMSI database61 represent only a sample of the total census, a logistic regression was used to model the proportion of people migrating between administrative units65. In particular, the logistic regression was used to model the proportion of people residing in j in the census year who were in i ‘n’ years prior to the census. Thus, the proportion of migrants in j in the census year that were previously residing in i was estimated using the following logistic regression function:
where ; with MIGij and TOTj representing the number of people residing in j in the census year that were in i ‘n’ years prior to the census and the total population residing in j in the census year, respectively.
Initially, a separate vector β=(β0, β1, …, βn) of coefficients was used in the linear predictor of the gravity model for each country (including malaria non-endemic countries), in Africa, Asia, Latin America and the Caribbean, for which migration data were available in the IPUMSI database61 (hereafter referred as IPUMSI countries; Table 1).
However, since the main aim of this study was to estimate internal human migration flows for malaria endemic countries for which migration data are not available, ultimately, models where the linear predictors were common across all countries located in the same continent were constructed (under the assumption of homogeneity of the process along the space). To investigate possible nonlinear relationships, models where linear predictors were replaced by additive predictors, using a Generalized Additive Modelling (GAM) framework66, were also explored.
GAM is a type of regression that, while preserving the functionality of using linear terms, allows covariates to have different and possibly opposite effects on the response variable by incorporating regression coefficients with smooth nonlinear form (Fig. 2).
Thus, all possible combinations of covariates (listed in Table 2 and Supplementary Table 2) were tested in a logistic regression model and then only the linear predictors of all continuous covariates of the best predictive logistic regression model were also modelled using a GAM.
For each continent, the overall combinations of covariates and model types were explored using a multi-step approach to identify the model with the greatest predictive power in countries for which migration data were not available. The best model was then selected using a leave-one-out cross-validation approach67 in which the observed proportion of migrants in j previously residing in i for all countries except one were used for fitting models, that were subsequently used to predict the proportion of migrants in j previously residing in i in the withheld country. The correlation coefficient (R2) was selected to measure the variance explained after verifying homoscedasticity and testing overdispersion using a chi-squared test. This process was then repeated through iteratively withholding one country at the time. For each model, the R2 values for all withheld countries were averaged and used to rank each models according to their predictive power averaged across all withheld countries (Fig. 3). The overall best predictive model for each continent (Supplementary Table 3) was then used to predict the proportion of migrants residing in j who were previously residing in i for every malaria endemic country located in the corresponding continent (refer to Supplementary Table 4a,b and c for summary statistics of each best predictive model for Africa, Asia, and Latin America and the Caribbean, respectively).
The red lines represent the best averaged R2 values used to select the best predictive model for each continent (Supplementary Table 3) while the red dots represent the R2 values, for all withheld countries, calculated using the best predictive model referring to the continent in which they are located.
Finally, in order to estimate the total number of people that migrated from i to j between 2005 and 2010 (Figs 4,5,6), for each country the predicted proportion of migrants residing in j was multiplied by the 2010 total population in j; with the latter calculated using either the corresponding WorldPop68–70 or the Gridded Population of the World version 4 (GPWv4)71 dataset adjusted to match United Nations Population Division (UNPD) estimates for 2010 (ref. 72). Refer to the Data collection and preparation subsection section below for a detailed description of how the population datasets mentioned above were identified and used.
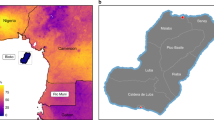
Coordinates for all three panels refer to GCS WGS 1984. For illustrative purposes, subnational unit boundaries are shown only in the insets and the colour ranges used to represent the flows are country-specific (refer to Supplementary Fig. 1 for additional close-up views of internal migration flows in Africa).
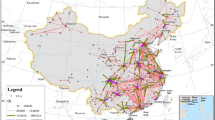
Coordinates for all three panels refer to GCS WGS 1984. For illustrative purposes, subnational unit boundaries are shown only in the insets and the colour ranges used to represent the flows are country-specific (refer to Supplementary Fig. 2a,b for additional close-up views of internal migration flows in Asia).
Coordinates for all three panels refer to GCS WGS 1984. For illustrative purposes, subnational unit boundaries are shown only in the insets and the colour ranges used to represent the flows are country-specific (refer to Supplementary Fig. 3 for additional close-up views of internal migration flows in Latin America and Caribbean).
Both model selection and prediction were performed using an R73 script contained in the WorldPop-InternalMigration-v1 code74 briefly described in the Code availability subsection below.
Data collection and preparation
In most of the countries available through the online IPUMSI database, internal migration variables were recorded by asking respondents either their administrative unit of residence 15, 5, or 1 prior to the census, or their previous residence and the number of years they are residing in the current locality. Considering that 5-year was the temporal interval available for most of the countries in the IPUMSI database and the fact that it has been demonstrated that both 1- and 5-year census-based internal migration data generally align well with shorter-term population movements in terms of relative strength of connections56,58, the 5-year migration data were used in this study. This maximised the amount of data that could be used to fit the gravity models subsequently used for predicting internal migration flows for every malaria endemic country. Thus, for each country listed in Table 1, harmonized, census-based 5-year internal migration data were extracted from the most recent census microdata available through the IPUMSI database61, downloaded locally, and eventually uploaded into a PostgreSQL database using a Microsoft Visual Studio 2010 user interface. The IPUMSI data stored in the PostgreSQL database were subsequently queried, using SQL, to quantify the number of people that migrated from each subnational administrative unit i to every other subnational administrative unit j during the 5-year timespan. These numbers were then matched to the corresponding country administrative unit spatial dataset, extracted from either the Global Administrative Areas (GADM)75 or the Global Administrative Unit Layers (GAUL)76 database, in a GIS environment. This was done by manually adding a unique ‘ID’ to each spatial unit corresponding to the one in the PostgreSQL database (hereafter referred as ‘IPUMSID’). In some cases, depending on the country, either the spatial detail of the IPUMSI migration data had to be reduced to match the lower spatial detail of the corresponding administrative unit dataset or spatially contiguous units in the administrative unit dataset had to be merged together to match the lower spatial detail of the IPUMSI migration data. In some other cases, ‘IPUMSIDs’ had to be edited or spatially contiguous units in the administrative unit dataset had to be merged together to match the reorganisation of the administrative units during the 5 years prior to the census. Finally, before calculating the migration flows between administrative units, another SQL query was used to classify each person in the census sample as either an internal migrant (1) or not (0). Examples of SQL queries used to perform the tasks described above are included in the WorldPop-InternalMigration-v1 code74 briefly described in the Code availability subsection below.
Response variable and covariates
For each country, the response variable, or the proportion of migrants residing in j in the census year that were residing in i 5 years prior to the census, was obtained by dividing the number of migrants residing in j in the census year that were residing in i 5 years prior the census by the total population residing in j in the census year; with both numbers based only on the information contained in IPUMSI census samples.
The administrative units spatially matching the IPUMSI migration microdata were used to calculate the distance between each pair of administrative units, their area, total population, and proportion of urban population. These main covariates (Table 2), along with other covariates derived from them (Supplementary Table 2), represent the pull and push migration factors, known to influence internal migration59,60,77, that were used to extend the basic gravity model proposed by Zipf64.
Other factors, including environmental factors59,60, and country-specific factors, such as literacy and percentage of male population59 or infrastructure and transportation78, were not used because (i) the factors listed in the previous paragraph alone proved to be able to explain most of the variance in the gravity models of Garcia et al.59, and (ii) only globally available datasets were explored in order to consistently model internal migration across all countries.
Calculating response variable and covariates
For each country, the total population in each administrative unit was calculated using the corresponding WorldPop79 (Data Citation 1) or GPWv4 (ref. 80) population count raster dataset adjusted to match UNPD estimates for 201072. The GPWv4 datasets were resampled to the spatial resolution of the WorlpPop datasets and used only for countries for which the WorldPop datasets were not available (Supplementary Table 1).
The area of each unit was calculated using each country vector administrative unit dataset projected to the most appropriate country-specific projected coordinate system, in order to minimize areal distortion, and ultimately reprojected to GCS WGS84.
The proportion of people in urbanized areas in each unit was calculated using the MODIS 500 m Global Urban Extent raster dataset81,82. The latter was converted to vector polygons, using the ArcGIS ‘Raster to Polygon’ tool83, and intersected with the reprojected country vector administrative unit dataset using the ArcGIS ‘Intersect’ tool83. Then, both the intersect output (containing polygons representing the total urban area within each unit uniquely identified by its ‘IPUMSID’) and the country vector administrative unit dataset were rasterized, at the resolution of the corresponding raster population dataset (i.e., 3 arc seconds 3 arc equals to approximately 100 m at the equator), and co-registered with it.
The two raster outputs, along with the population count raster dataset, were then input to the ArcGIS ‘Zonal Statistics as Table’ tool83 to generate two tables containing the total population and urban population in each unit (with the rasterized administrative units and thus their ‘IPUMSIDs’ used to define the zones). Subsequently, both tables were joined to the attribute table of the vector administrative unit dataset, using the ‘IPUMSID’ field to perform the join operation, and the proportion of urban population in each unit was calculated simply dividing its urban population by its total population.
The geodesic distance between each pair of administrative units, with the latter represented by their centroids, was calculated using the ArcGIS ‘Generate Near Table (Analysis)’ tool83. The ‘IN_FID’ and ‘NEAR_FID’ fields (identifying the administrative unit of origin and destination, respectively) in the output ‘distance’ table were then used for joining twice the ‘centroid attribute’ table using the centroid ‘ID’ field to perform the join operation. Since the ‘centroid attribute’ table contains the attributes of each administrative unit represented by the corresponding centroid, the join operation allowed to generate a ‘distance’ table containing all pairs of origin and destination administrative units along with their ‘IPUMSIDs’ and attributes including the unit’s area, total population, and proportion of urban population. Origin and destination ‘IPUMSID’ fields were then renamed ‘NODEI’ and ‘NODEJ’, respectively.
A ‘contiguity’ table containing information about spatial contiguity of administrative units (defined based on polygons sharing an edge) was generated using the ArcGIS ‘Generate Spatial Weights Matrix’ tool83 and subsequently joined with the ‘distance’ table to obtain a new table containing all main covariates, listed in Table 2, calculated at the unit level. This join operation (based on both the ‘NODEI’ and ‘NODEJ’ field the in the ‘distance’ table and the corresponding ‘IPUMSID’ and ‘NID’ field in the ‘contiguity’ table) was performed through two different R scripts depending on whether the country is an IPUMSI or a non-IPUMSI countries. In particular, the R script for the IPUMSI countries added to the new table a ‘MIGIJ’ field containing the number of people that migrated from each ‘NODEI’ to each other ‘NODEJ’ according to the IPUMSI migration microdata and calculated the response variable.
Finally, on a continent basis, all IPUMSI country tables were merged together and input to an R73 script that generated the additional covariates listed in Supplementary Table 2, identified the best predictive model for each continent, as described in the previous section, and was used to estimate the 5-year (2005–2010) internal human migration flows for every malaria endemic country using the best predictive model selected for the corresponding continent.
All operations described above, excluding the reprojection of the vector administrative unit datasets and the calculation of their surface areas, for all IPUMSI and non-IPUMSI countries, were performed using the WorldPop-InternalMigration-v1 code74 briefly described in the Code availability subsection below.
Code availability
The WorldPop-InternalMigration-v1 code74, used to produce the open access archive of estimated 5-year (2005–2010) internal human migration flows described in this article, is publicly available through Figshare. It consists of 1) a Microsoft Visual Studio 2010 user interface allowing users to upload the IPUMSI census microdata to a PostgreSQL database; 2) example SQL queries that were used to match the spatial detail of the IPUMSI migration data to spatial detail of the corresponding administrative unit dataset and to identify internal migrants within the IPUMSI census samples 3) an ArcToolbox geoprocessing tool82 that assigns a unique ID to each administrative unit and calculates the corresponding total population and proportion of urban population; 4) a Python84/ArcPy83 script that creates two tables, one containing spatial contiguity information between each pair of administrative units (‘contiguity.csv’) and another one containing the ISO country code, the continent in which the country is located, the distance between each pair of administrative units, their total population, proportion of urban population, surface area, and the geographic coordinates (GCS WGS84) of their centroid (‘distance.csv’); 5) two R73 scripts, one for the IPUMSI countries used to query the IPUMSI migration microdata loaded in the PostgreSQL database, calculate the response variable, and join the query result with the two output tables of the python script, and another one for the non-IPUMSI countries used just to join together the two output tables of the python script; and 6) an R73 script that performs the model selection and estimates the 5-year (2005–2010) internal human migration flows between subnational administrative units.
All available sets of code are named progressively and must be run sequentially according to the order in which they are presented above. They are also internally documented in order to both briefly explain their purpose and, when required, guide the user through their customization.
Data Records
All datasets described in this article, referring to all Pf and Pv endemic countries listed in Supplementary Table 1, are publicly and freely available both through the WorldPop Dataverse Repository (Data Citation 2) and the WorldPop website (http://www.worldpop.org.uk/data/data_sources/). However, it is important to note that while the datasets stored in the Dataverse Repository represent the datasets produced at the time of writing, and will be preserved in their published form, the datasets stored on the WorldPop website may be updated as more recent IPUMSI migration data for the countries listed in Table 1, become available. Similarly, the datasets stored on the WorldPop website may be updated as IPUMSI census-based migration microdata become available for additional malaria endemic and non-endemic countries located in Africa, Asia, Latin America and the Caribbean. Indeed, the availability of migration data for additional countries may enable further improvements of the predictive power of the gravity models used to estimate the internal migration flows. For each county, the corresponding internal migration dataset, along with a point dataset showing the nodes of the migration network, (Table 3) can be obtained by downloading the corresponding zipped archive associated with the continent in which the country of interest is located.
Technical Validation
Goodness of fit and error p-value
All countries available in the IPUMSI database were used to assess the accuracy of the predicted proportion of migrants in j in the census year that were previously residing in i 5 years prior to the census by comparing them with the corresponding observed values from the IPUMSI migration microdata. For each country, the goodness of fit (R2) between predicted and observed values and the corresponding error P-value, representing the average probability that predicted migration values lay outside the distribution of the observed values, are reported in Table 4 below. Both metrics were derived using (i) the observed IPUMSI migration flows from each i to any other j and (ii) the predicted IPUMSI-based migration flows calculated by multiplying the predicted proportion of migrants residing in j in the census year by the IPUMSI-based total number of people residing in j in the census year.
Usage Notes
The estimated internal human migration flows between subnational administrative units can be used to support a range of applications from planning interventions, to measuring progress, designing strategies, and predicting response variables that are intrinsically dependent on migration flows and internal connectivity.
Ongoing work involves the integration of these datasets with malaria prevalence raster datasets85–87 in order to inform local elimination and global eradication planning by identifying subnational communities of malaria movement and sources and sinks of transmission within them36,43,58. Similarly, these datasets could be used to better model the spread and improve understanding of the drivers of the distributions of other infectious diseases, such as West Nile Virus, schistosomiasis, river blindness, and yellow fever, which are endemic in some of the countries listed in Supplementary Table 1. Additionally there are many uses of these data beyond infectious disease dynamics, in the fields of trade, demography, transportation and economics, for example.
There are a number of limitations, caveats, and assumptions inherent in the approach that should be considered when using the datasets outlined here. For consistency, internal migration flows were estimated using a fixed set of pull and push factors common to all countries and thus only a limited number of covariates were used to fit the gravity-type spatial interaction models and to create predictions. For this reason, as is a trade-off in the production of generalizable models, the model fit varied between countries and for some of them, such as Malawi, China, Cambodia, India, and Venezuela (Table 4), poor fits could be improved by considering additional, locally-specific migration drivers that could help to increase the percentage of variance explained60,78. Other limitations are the fact that migration models were fitted using only a small sample (ranging between 0.07 and 10%) of the full census for each country, and that in each sample a small number of households were swapped across administrative units. Moreover, the spatial detail at which migration is captured and summarized varies by country. Because of this, for some countries, the modelled role of some of the pull and push factors, may not have been captured at the spatial level at which they influence migration as recorded in the census. It is also important to consider that the underlying migration data are based only on permanent movements captured by the census and other types of migrations, such as seasonal movements and forced displacements, may be not captured by the model88–90.
The two main assumptions behind the approach presented here are that for each country (i) the census samples are considered to be representative at the administrative unit level at which migration was recorded and (ii) the percentage of people migrating between administrative units is considered to be constant over time. Regarding the second assumption, it is important to highlight that the use of census data from many years ago for some countries may have generated inaccurate estimates for the period considered in this study (i.e., 2005–2010), for example because of major changes in the countries’ socio-economic conditions from the time period covered by the census (e.g., the rapid economic development and urbanization that has occurred in China during the last two decades91,92). Similarly, in some other countries, either the presence of conflicts93 or the occurrence of natural disasters88,89 during the specific time period covered by the census may have produced fluctuations in the number of internal migrants and consequently biased results for the period considered in this study.
Finally, the estimated internal flows represent modelling outputs generated using ancillary covariate datasets, and thus, to avoid circularity they should not be used to make predictions or explore relationships with any of these ancillary datasets. It is also important to note that these ancillary datasets are modelling outputs in themselves and thus they have a degree of uncertainty that will carry over into the migration estimates.
Additional Information
How to cite this article: Sorichetta, A. et al. Mapping internal connectivity through human migration in malaria endemic countries. Sci. Data 3:160066 doi: 10.1038/sdata.2016.66 (2016).
References
References
International Organization for Migration. Global Migration Trends: an overview. Available at http://missingmigrants.iom.int/sites/default/files/documents/Global_Migration_Trends_PDF_FinalVH_with%20References.pdf (2014).
The World Bank. International Migration at All-Time High. Available at http://www.worldbank.org/en/news/press-release/2015/12/18/international-migrants-and-remittances-continue-to-grow-as-people-search-for-better-opportunities-new-report-finds (2015).
The World Bank. Air transport, passengers carried. Available at http://data.worldbank.org/indicator/IS.AIR.PSGR/countries?display=graph (2016).
Brockmann, D., David, V. & Gallardo, A. M. Human mobility and spatial disease dynamics. Reviews of nonlinear dynamics and complexity (ed. Schuster, H. G.) (Wiley-VCH, 2009).
Undie, C. C., Johannes, J. L. & Kimani, E. Overcoming Barriers: human Mobility and Development. Human Development Report. Available at http://hdr.undp.org/sites/default/files/reports/269/hdr_2009_en_complete.pdf (United Nations Development Programme, 2009).
Antman, F. M. The impact of migration on family left behind. International Handbook on the Economics of Migration (eds Constant, A. F. & Zimmermann, K. F.) (Edward Elgar Publishing Limited, 2013).
Tatem, A. J. Mapping population and pathogen movements. Int. Health 6, 5–11 (2014).
Bremner, J. & Hunter, L. M. Migration and the Environment. Popul. Bull 69. Available at http://www.prb.org/pdf14/migration-and-environment.pdf (Population Reference Bureau, 2014).
Morrissey, J. W. Understanding the relationship between environmental change and migration: the development of an effects framework based on the case of northern Ethiopia. Global Environ. Chang 23, 1501–1510 (2013).
Potts, D. Debates about African urbanisation, migration and economic growth: what can we learn from Zimbabwe and Zambia? The Geographical Journal, doi: 10.1111/geoj.12139 (2015).
Jones, G. W. Migration and Urbanization in China, India and Indonesia: an Overview (Springer International Publishing, 2016).
Hecht, S., Yang, A. L., Basnett, B. S., Padoch, C. & Peluso, N. L. People in motion, forests in transition: trends in migration, urbanization, and remittances and their effects on tropical forests (Center for International Forestry Research, 2015).
Walters, B. B. Migration, land use and forest change in St Lucia, West Indies. Land Use Policy 51, 290–300 (2016).
Chan, K. W. Migration and development in China: trends, geography and current issues. Migration and Development 1, 187–205 (2012).
Skeldon, R. Migration and development: a global perspective (ed. Triadafyllidou, A.) (Routledge, 2014).
Delgado-Wise, R. Migration and development in Latin America. Routledge Handbook of Immigration and Refugee Studies (ed. Triadafyllidou A.) (Routledge, 2015).
Cabieses, B., Tunstall, H., Pickett, K. E. & Gideon, J. Changing patterns of migration in Latin America: how can research develop intelligence for public health? Revista panamericana de salud pública 34, 68–74 (2013).
Mou, J., Griffiths, S. M., Fong, H. F. & Dawes, M. G. Defining migration and its health impact in China. Public Health 129, 1326–1334 (2014).
Vearey, J. Healthy migration: a public health and development imperative for South (ern) Africa. SAMJ: S. Afr. Med. J 104, 663–664 (2014).
Tebit, D. M. & Arts, E. J. Tracking a century of global expansion and evolution of HIV to drive understanding and to combat disease. Lancet Infect. Dis. 11, 45–56 (2011).
Tatem, A. J., Hemelaar, J., Gray, R. R. & Salemi, M. Spatial accessibility and the spread of HIV-1 subtypes and recombinants. Aids 26, 2351–2360 (2012).
Faria, N. R. et al. The early spread and epidemic ignition of HIV-1 in human populations. Science 346, 56–61 (2014).
Colizza, V., Barrat, A., Barthélemy, M. & Vespignani, A. Predictability and epidemic pathways in global outbreaks of infectious diseases: the sars case study. BMC Medicine 5, 34 (2007).
Lines, J. Chikungunya in Italy. Brit. Med. J. 335, 576 (2007).
Grandadam, M. et al. Chikungunya virus, southeastern France. Emerg. Infect. Dis. 17, 910–913 (2011).
Balcan, D. et al. Seasonal transmission potential and activity peaks of the new influenza A (H1N1): a Monte Carlo likelihood analysis based on human mobility. BMC Medicine 7, 45 (2009).
Gomes, M. F. C. et al. Assessing the international spreading risk associated with the 2014 west African Ebola Outbreak. PLOS Currents Outbreaks 1, doi: 10.1371/currents.outbreaks.cd818f63d40e24aef769dda7df9e0da5 (2014).
Cohen, J. M. et al. Malaria resurgence: a systematic review and assessment of its causes. Malar. J. 11, 122 (2012).
MacPherson, D. W. et al. Population mobility, globalization, and antimicrobial drug resistance. Emerg. Infect. Dis. 15, 1727 (2009).
Tatem, A. J. & Hay, S. I. Climatic similarity and biological exchange in the worldwide airline transportation network. P. Roy. Soc. Lond. B: Bio 274, 1489–1496 (2007).
Huang, Z. & Tatem, A. J. Global malaria connectivity through air travel. Malar. J. 12, 269 (2013).
Perra, N. & Gonçalves, B. Modeling and predicting human infectious diseases. Social Phenomena (eds Perra, N. & Gonçalves, B.) (Springer International Publishing, 2015).
Pybus, O. G., Tatem, A. J. & Lemey, P. Virus evolution and transmission in an ever more connected world. P. Roy. Soc. Lond. B: Bio 282, 20142878 (2015).
The Roll Back Malaria Partnership. The global malaria action plan—For a malaria free world. Available at http://archiverbm.rollbackmalaria.org/gmap/gmap.pdf (The Roll Back Malaria Partnership, 2008).
World Health Organization (WHO). World Malaria Report 2015 (WHO Document Production Services, 2015).
Gates, B. & Chambers, R. From aspiration to action—What will it take to end malaria?. Available at http://endmalaria2040.org/assets/Aspiration-to-Action.pdf (2015).
Nájera, J. A., González-Silva, M. & Alonso, P. L. Some lessons for the future from the Global Malaria Eradication Programme (1955-1969). PLoS Med. 8, e1000412 (2011).
Tatem, A. J. et al. The use of mobile phone data for the estimation of the travel patterns and imported Plasmodium falciparum rates among Zanzibar residents. Malar. J. 8, 287 (2009).
Pindolia, D. K. et al. Quantifying cross-border movements and migrations for guiding the strategic planning of malaria control and elimination. Malar. J. 13, 169 (2014).
Bradley, J. et al. Infection importation: a key challenge to malaria elimination on Bioko Island, Equatorial Guinea. Malar. J. 14, 46 (2015).
Lynch, C. A. et al. Association between recent internal travel and malaria in Ugandan highland and highland fringe areas. Trop. Med. & Int. Health 20, 773–780 (2015).
Lynch, C. & Roper, C. The Transit Phase of Migration: Circulation of Malaria and Its Multidrug. PLoS Med. 8, e1001040 (2011).
Tatem, A. J. & Smith, D. L. International population movements and regional Plasmodium falciparum malaria elimination strategies. Proc. Natl. Acad. Sci 107, 12222–12227 (2010).
Wesolowski, A. et al. Quantifying the impact of human mobility on malaria. Science 338, 267–270 (2012).
Prothero, R. M. Population movements and problems of malaria eradication in Africa. B. World Health Organ 24, 405–425 (1961).
Prothero, R. M. Disease and mobility: a neglected factor in epidemiology. Int. J. Epidemiol. 6, 259 (1977).
Whittaker, M. & Smith, C. Findings of the literature review on mobility, infectious diseases and malaria. Malar. J. 11, S1 P101 (2012).
Somboon, P., Aramrattana, A., Lines, J. & Webber, R. Entomological and epidemiological investigations of malaria transmission in relation to population movements in forest areas of north-west Thailand. Southeast Asian J. Trop. Med. Public Health 29, 3–9 (1998).
Osorio, L., Todd, J. & Bradley, D. J. Travel histories as risk factors in the analysis of urban malaria in Colombia. Am. J. Trop. Med. Hyg. 71, 380–386 (2004).
Yukich, J. O. et al. Travel history and malaria infection risk in a low-transmission setting in Ethiopia: a case control study. Malar. J. 12, 33 (2013).
Song, C., Qu, Z., Blumm, N. & Barabási, A. L. Limits of predictability in human mobility. Science 327, 1018–1021 (2010).
Lu, X., Wetter, E., Bharti, N., Tatem, A. J. & Bengtsson, L. Approaching the limit of predictability in human mobility. Sci. Rep 3 doi:10.1038/srep02923 (2013).
Buckee, C. O., Wesolowski, A., Eagle, N. N., Hansena, E. & Snow, R. W. Mobile phones and malaria: Modeling human and parasite travel. Travel. Med. Infect. Dis 11, 15–22 (2013).
Tatem, A. J. et al. Integrating rapid risk mapping and mobile phone call record data for strategic malaria elimination planning. Malar. J. 13, 52 (2014).
Wesolowski, A. et al. Quantifying travel behavior for infectious disease research: a comparison of data from surveys and mobile phones. Scientific Reports 4, 5678 (2014).
Wesolowski, A. et al. The use of census migration data to approximate human movement patterns across temporal scales. PLoS ONE 8, e52971 (2013).
Flowminder Foundation. Where We Work. http://www.flowminder.org/where-we-work (2016).
Ruktanonchai, N. W. et al. Census-derived migration data as a tool for informing malaria elimination policy. Malar. J. 15, 273 (2016).
Henry, S., Boyle, P. & Lambin, E. F. Modelling inter-provincial migration in Burkina Faso, West Africa: the role of socio-demographic and environmental factors. Appl. Geogr. 23, 115–136 (2003).
Garcia, A. J., Pindolia, D. K., Lopiano, K. K. & Tatem, A. J. Modeling internal migration flows in sub-Saharan Africa using census microdata. Migrat. Stud. doi:10.1093/migration/mnu036 (2014).
Minnesota Population Center. Integrated Public Use Microdata Series, International: Version 6.4 [Machine-readable database]. Available at https://international.ipums.org/international (University of Minnesota, 2015).
World Health Organization (WHO). Malaria Coun try profiles. Available at http://www.who.int/malaria/publications/country-profiles/en/ (2015).
Howes, R. E. et al. Plasmodium vivax Transmission in Africa. PLoS Negl. Trop. Dis. 9, e0004222 (2015).
Zipf, G. K. The P1P2/D hypothesis: on intercity movement of persons. Am. Sociol. Rev. 11, 677–686 (1946).
Zhao, L., Chen, Y. & Schaffner, D. W. Comparison of logistic regression and linear regression in modeling percentage data. Appl. Environ. Microb 67, 2129–2135 (2001).
Hastie, T. J. & R. J. Generalized Additive Models (Chapman and Hall, 1990).
Zhang, P. Model Selection Via Multifold Cross Validation. Ann. Stat 21, 299–313 (1993).
Linard, C., Gilbert, M., Snow, R. W., Noor, A. M. & Tatem, A. J. Population Distribution, Settlement Patterns and Accessibility across Africa in 2010. PLoS ONE 7, e31743 (2012).
Gaughan, A. E., Stevens, F. R., Linard, C., Jia, P. & Tatem, A. J. High Resolution Population Distribution Maps for Southeast Asia in 2010 and 2015. PLoS ONE 8, e55882 (2013).
Sorichetta, A. et al. High-resolution gridded population datasets for Latin America and the Caribbean in 2010, 2015, and 2020. Sci. Data 2, 150045 (2015).
Doxsey-Whitfield, E. et al. Taking Advantage of the Improved Availability of Census Data: A First Look at the Gridded Population of the World, Version 4. Appl. Geogr. 1, 226–234 (2015).
United Nations Department of Economic and Social Affairs Population Division (UNPD). World Urbanization Prospects: The 2014 Revision. CD-ROM Edition. Available at http://esa.un.org/unpd/wup/CD-ROM/ (2014).
Core Team., R. R: a language and environment for statistical computing. Available at http://www.R-project.org/ (R Foundation for Statistical Computing, 2015).
Bird, T. J. et al. Source code for: Mapping internal connectivity through human migration in malaria endemic countries. Figshare. Available at https://dx.doi.org/10.6084/m9.figshare.3394729.v2 (2016).
GADM. Database of Global Administrative Areas. Available at http://www.gadm.org/ (2012).
GAUL. Global Administrative Unit Layers. Available at http://www.fao.org/geonetwork/srv/en/metadata.show?id=12691 (2015).
Dennet, A. Estimating flows between geographical locations: ‘get me started in’ saptial interaction modelling. UCL working paper series 181, 1–24 (2012).
Flahaux, M.-L. & De Haas, H. African migration: trends, patterns, drivers. Comp. Migr. Stud 4, doi:10.1186/s40878-015-0015-6 (2016).
WorldPop. Population—indivi dual countries. Available at http://www.worldpop.org.uk/data/data_sources/ (2015).
Center for International Earth Science Information Network, Columbia University. Gridded Population of the World, Version 4 (GPWv4): Population Count Adjusted to Match 2015 Revision of UN WPP Country Totals. Available at http://sedac.ciesin.columbia.edu/data/set/gpw-v4-population-count-adjusted-to-2015-unwpp-country-totals (NASA Socioeconomic Data and Applications Center, 2015).
Schneider, A., Friedl, M. & Potere, D. MODIS 500m Global Urban Extent. Available at http://nelson.wisc.edu/sage/data-and-models/schneider.php (2009).
Schneider, A., Friedl, M. A. & Potere, D. A new map of global urban extent from MODIS satellite data. Environ. Res. Lett. 4, 044003 (2009).
ESRI. ArcGIS Desktop: Release 10.1 (ESRI, 2012).
van Rossum, G. & de Boer, J. Interactively Testing Remote Servers Using the Python Programming Language. CWI Quarterly 4, 283–303 (1991).
Gething, P. W. et al. A new world malaria map: Plasmodium falciparum endemicity in 2010. Malar. J 10, 1–16 (2011).
Gething, P. W. et al. A long neglected world malaria map: Plasmodium vivax endemicity in 2010. PLoS Negl. Trop. Dis. 6, e1814 (2012).
Bhatt, S. et al. The effect of malaria control on Plasmodium falciparum in Africa between 2000 and 2015. Nature 526, 207–211 (2015).
Wilson, R. et al. Rapid and near Realtime Assessments of Population Displacement Using Mobile Phone Data Following Disasters: the 2015 Nepal Earthquake. PLOS Currents Disasters 1, doi: 10.1371/currents.dis.d073fbece328e4c39087bc086d694b5c (2016).
Bengtsson, L., Lu, X., Thorson, A., Garfield, R. & Von Schreeb, J. Improved response to disasters and outbreaks by tracking population movements with mobile phone network data: a post-earthquake geospatial study in Haiti. PLoS Med. 8, e1001083 (2011).
Bharti, N. et al. Explaining seasonal fluctuations of measles in Niger using nighttime lights imagery. Science 334, 1424–1427 (2011).
Schneider, A. et al. A new urban landscape in East-Southeast Asia, 2000-2010. Environ. Res. Lett. 10, 034002 (2015).
Gaughan, A. E. et al. Spatiotemporal patterns of population in mainland China, 1990 to 2010. Sci. Data 3, 160005 (2016).
Lozano-Gracia, N., Piras, G., Ibáñez, A. M. & Hewings, G. J. The journey to safety: conflict-driven migration flows in Colombia. Int. Regional Sci. Rev 33, 157–180 (2010).
International Organization for Standardization. 3166-1 alpha-3 (three-letter) country codes. Available at http://www.iso.org/iso/catalogue_detail.htm?csnumber=63545 (2015).
Data Citations
Sorichetta, A Harvard Dataverse https://doi.org/10.7910/DVN/PUGPVR (2015)
Sorichetta, A Harvard Dataverse https://doi.org/10.7910/DVN/RUWQQK (2016)
Acknowledgements
The authors wishes to acknowledge the statistical offices that provided the underlying data making this research possible: Central Bureau of Census and Population Studies, Cameroon; Ghana Statistical Services; National Statistics Directorate, Guinea; National Statistical Office, Malawi; National Directorate of Statistics and Informatics, Mali; National Agency of Statistics and Demography, Senegal; Statistics South Africa; Bureau of Statistics, Uganda; Central Statistics Office, Zambia; Central Agency for Public Mobilization and Statistics, Egypt; Department of Statistics, Morocco; National Statistical Service, Armenia; National Statistical Committee, Kyrgyzstan; Ministry of Statistics and Programme Implementation, India; BPS Statistics Indonesia; National Statistical Office, Thailand; National Institute of Statistics, Cambodia; National Bureau of Statistics, China; Department of Statistics, Malaysia; National Statistics Office, Philippines; General Statistics Office, Vietnam; National Statistical Office, Mongolia; Bureau of Statistics, Fiji; National Institute of Statistics and Censuses, Argentina; National Institute of Statistics, Bolivia; Institute of Geography and Statistics, Brazil; National Administrative Department of Statistics, Colombia; National Institute of Statistics and Censuses, Costa Rica; National Statistics Office, Dominican Republic; National Institute of Statistics and Censuses, Ecuador; Department of Statistics and Censuses, El Salvador; Institute of Statistics and Informatics, Haiti; National Institute of Statistics, Geography, and Informatics, Mexico; National Institute of Information Development, Nicaragua; National Institute of Statistics and Informatics, Peru; National Institute of Statistics, Venezuela; Office of National Statistics, Cuba; Statistical Institute, Jamaica; National Institute of Statistics, Uruguay. A.S. is supported by funding from the Bill & Melinda Gates Foundation (OPP1106427, 1032350). A.J.T. is supported by funding from NIH/NIAID (U19AI089674), the Bill & Melinda Gates Foundation (OPP1106427, 1032350, 1134076), the Clinton Health Access Initiative, National Institutes of Health, and a Wellcome Trust Sustaining Health Grant (106866/Z/15/Z). This work forms part of the outputs of WorldPop (www.worldpop.org) and the Flowminder Foundation (www.flowminder.org). The funders had no role in study design, data collection and analysis, decision to publish, and preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
A.S. coordinated the study, undertook data collection and assembly, and drafted the manuscript. A.S. and T.J.B. undertook data analyses, generated the internal migration datasets, and prepared the tables and figures. A.S. and L.S. performed the technical validation of the internal migration datasets. A.S., T.J.B., N.R., and A.J.T. conceptualized the study. A.S., T.J.B., N.R., E.z.E.S., C.P., N.T., I.C.W., J.D.S., A.J.G., and L.S. developed the WorldPop-InternalMigration-v1 code. T.J.B. implemented the R script used for model selection and prediction. C.P., A.J.G., I.C.W., and J.D.S. aided with data assembly. A.J.T. aided drafting the manuscript and conceived the study. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Sorichetta, A., Bird, T., Ruktanonchai, N. et al. Mapping internal connectivity through human migration in malaria endemic countries. Sci Data 3, 160066 (2016). https://doi.org/10.1038/sdata.2016.66
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2016.66
This article is cited by
-
Global holiday datasets for understanding seasonal human mobility and population dynamics
Scientific Data (2022)
-
Quantifying efficient information exchange in real network flows
Communications Physics (2021)
-
Critical behavior in interdependent spatial spreading processes with distinct characteristic time scales
Communications Physics (2021)
-
Mapping the travel patterns of people with malaria in Bangladesh
BMC Medicine (2020)
-
Utilizing general human movement models to predict the spread of emerging infectious diseases in resource poor settings
Scientific Reports (2019)