Abstract
Emerging evidence suggests Jumonji domain-containing proteins are epigenetic regulators in diverse biological processes including cellular differentiation and proliferation. RNA interference-based analyses combined with gene expression profiling can effectively characterize the cellular functions of these enzymes. We found that the depletion of Jumonji domain-containing protein 6 (JMJD6) and its paralog protein Jumonji domain-containing protein 4 (JMJD4) individually by small hairpin RNAs (shRNAs) slowed cell proliferation of mouse NIH3T3 fibroblasts. We subsequently performed gene expression profiling on both JMJD6- and JMJD4-depleted mouse NIH3T3 fibroblasts using the Affymetrix GeneChip Mouse Exon 1.0 ST Array. Here we report the gene profiling datasets along with the experimental procedures. The information can be used to further investigate how JMJD6 and JMJD4 affect gene expression and cellular physiology.
Design Type(s) | parallel group design • RNA Interference |
Measurement Type(s) | transcription profiling assay |
Technology Type(s) | DNA microarray |
Factor Type(s) | biological replicate role • shRNA |
Sample Characteristic(s) | Mus musculus • NIH-3T3 cell |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
Jumonji domain-containing proteins are iron- and 2-oxoglutarate-dependent oxygenases that act on diverse substrates including proteins, nucleic acids and small molecules1,2. These enzymes either hydroxylate or demethylate their substrates in an oxygen-dependent manner. Many JmjC domain proteins play a key role in the epigenetic regulation of mammalian development and of diseases such as cancer2,3.
Phylogenetic analysis classifies the Jumonji domain containing proteins into several subgroups3. One subgroup comprises the asparaginyl hydroxylase FIH4,5, the ribosomal hydroxylases MINA53 and NO66 (ref. 6) , the lysyl hydroxylase JMJD4 (ref. 7), the lysine demethylase/hydroxylase JMJD5 (refs 8,9) , the lysyl hydroxylase/arginine demethylase JMJD6 (refs 10,11) , the tRNA hydroxylase TYW5 (ref. 12), and the enzymatically uncharacterized proteins JMJD8 and HSPBAP1. Although these enzymes share the conserved JmjC domain with the histone lysine demethylases, they generally lack chromatin-binding domain, indicating potential functions other than modifying histones.
Among the members of this functionally diverse subgroup, JMJD6 was recently characterized as a crucial regulator for gene expression at the level of histone modification10,13, transcriptional elongation14 and RNA splicing11,15,16. Jmjd6-deficient mice show various developmental defects and perinatal lethality17,18. Moreover, JMJD6 is required for angiogenesis15, adipocyte differentiation19 and T cell proliferation20. Elevation of JMJD6 expression has been observed in breast21,22, lung23, colon24, and oral cancer25. Depletion of JMJD6 by RNA interference reduced the proliferation of human cancer cell lines21,24. Therefore, JMJD6 is essential for cellular proliferation and differentiation. In contrast, the paralog protein JMJD4 that shares 34% sequence identity with JMJD6 was shown to be involved in translational termination by hydroxylating the translational termination factor eRF1 (ref. 7). Other biological functions of JMJD4 have not been demonstrated.
We conducted a loss-of-function study using an RNA interference approach to reduce the levels of endogenous JMJD6 and JMJD4 in proliferating mouse NIH3T3 fibroblasts. We found that depletion of either JMJD6 or JMJD4 alone significantly reduced cell proliferation. In order to determine whether and how these JmjdC domain proteins affect gene expression on a global scale, we performed gene expression profiling on the JMJD6 and JMJD4 knockdown NIH3T3 cells using the Affymetrix Exon Arrays. In this report, we provide a detailed description of the gene expression profiling datasets.
Methods
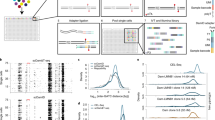
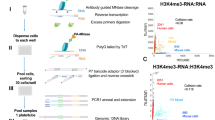
The scheme of the experimental procedures is presented in Fig. 1.
Cell culture
The mouse NIH3T3 fibroblast cell line was maintained in DMEM high glucose medium (Invitrogen) containing 10% calf serum (Sigma) and 100 U ml−1 penicillin/streptomycin (Invitrogen). Human 293T cells were maintained in DMEM high glucose medium containing 10% fetal calf serum (Sigma) and 100 U ml−1 penicillin/streptomycin (Invitrogen). To evaluate cell proliferation, 1×105 cells were seeded in 6-well plates, and the number of viable cells was counted every other day with a hemocytometer.
Plasmid construction and virus transduction
The preparation of small hairpin RNA (shRNA) lentiviral constructs was performed as previously described (Campeau, 2009). The shJMJD6 target sequence is 5′-GCTGACACCCAGAGAACAA-3′. The shJMJD4 target sequence is GAAGAATTCTGGAGAGCAT. The control shRNA sequence is TGGCGGCGAGTGAAGTACGTGATAA. The shRNA lentiviral constructs were co-transfected with pLP1, pLP2 and pVSVG packaging vectors into 293T cells with Lipofectamine 2000 reagent (Invitrogen). The viral supernatant was harvested after 48 h incubation and filtered through a 0.45 μm syringe filter (Millipore). To infect NIH3T3 cells, one million cells were incubated with 2 ml of the filtered viral supernatant supplemented with 4 μg ml−1 of polybrene (Sigma) for 48 h. The infected cells were subsequently selected in media containing 2.5 μg ml−1 puromycin (Invitrogen) for 7 days.
RNA isolation and real-time qPCR analysis
One million cells were plated overnight in 10 cm culture dishes. The cells were washed with 5 ml cold PBS and then lysed directly in the culture dish by adding 1 ml of TRIzol reagent (Invitrogen). Total RNA was isolated from TRIzol lysates using RNeasy Mini Kit (Qiagen) according to the manufacturer’s instructions. An on-column DNAse digestion step is included in this protocol. cDNA was prepared from 1 μg of total RNA by Superscript III reverse transcriptase kit (Invitrogen). Quantitative PCR was performed on a StepOne Plus real-time PCR machine with Fast SYBR Green Master mix (Applied Biosystems). The relative gene expression levels were calculated as 2^ (CtEef1a1−Ctgene) and were normalized to the scrambled shRNA control as indicated. The primers are listed below:
Eef1a1 (forward): 5′-AGCTTCTCTGACTACCCTCCACTT-3′
Eef1a1 (reverse): 5′-GACCGTTCTTCCACCACTGATT-3′
Jmjd6 (forward): 5′-GTTCCAGCTCGTCAGACTCG-3′
Jmjd6 (reverse): 5′-TGCCCCTAAGACATGACCAC-3′
Jmjd4 (forward): 5′-TCCTGCTGGAATGTCGCACCTGT-3′
Jmjd4 (reverse): 5′-ACCCCAAATAGGGACCGGAGGC-3′
Microarray analysis
The RNA samples were processed at the Genomics Core Facility of UMass Medical School. RNA quantity and quality were assessed using an Agilent Bioanalyzer 2100. cDNA was synthesized from 500 ng RNA using the Ambion WT kit (Life Technologies) and the single stranded cDNA was prepared using the Affymetrix GeneChip WT Terminal Labeling Kit (Affymetrix) according to the manufacturer’s instructions. The labeled mix was hybridized to GeneChip Mouse Exon 1.0 ST Array (Affymetrix) in the Gene Chip Hybridization Oven 640 overnight. Probe intensities were measured using the Affymetrix GeneChip Scanner 3000 7G. The probe cell intensity data (CEL) from GeneChip Mouse Exon 1.0 ST Arrays was analyzed in the Affymetrix Expression Console software to generate CHP files using the Robust Multichip Analysis (RMA)-sketch algorithm workflow. The transcript structure confidence levels for both gene and exon level analyses were set as Core, which limits analysis to exon-level probe sets that map to BLAT alignments of mRNA with annotated full-length coding sequence (CDS) regions. Differentially expressed genes and exons were identified by Transcriptome Analysis Console (TAC) 3.0 software (Affymetrix).
Data Records
Gene expression profiling on RNA samples collected from the control shRNA-treated (shCtrl_R1 and shCtrl_R2), JMJD6 knockdown (shJMJD6_R1 and shJMJD6_R2), JMJD4 knockdown (shJMJD4_R1 and shJMJD4_R2) cells was performed using GeneChip Mouse Exon 1.0 ST Arrays. Two biological replicates were performed. Both gene level and exon level expression were analyzed. All samples and datasets are described in Table 1. The primary data are available at the NCBI Gene Expression Omnibus (GEO) under the accession numbers(Data Citation 1).
Technical Validation
Confirmation of the shRNA-mediated knockdown
The shRNA-mediated knockdown reduced cell proliferation is shown in Fig. 2a. The signal intensity values of expression from the Jmjd6 and Jmjd4 genes in the microarray datasets are shown in Fig. 2b. The expression levels from the endogenous Jmjd6 and Jmjd4 genes were confirmed by real-time qPCR and are shown in Fig. 2c.
(a) The cell proliferation curve of the control shRNA-treated (shCtrl), JMJD6 knockdown (shJMJD6), and JMJD4 knockdown (shJMJD4) mouse NIH3T3 fibroblasts. (b) The signal intensity of expression from the Jmjd6 and Jmdj4 genes from each individual array. (c) The relative expression levels from the endogenous Jmjd6 and Jmdj4 genes in each sample were determined by real-time qPCR analysis. The value of the shCtrl_R1 sample was set as 1.
Quality control of microarray data
The probe cell intensity values in each individual array were generated from the Affymetrix Expression Console software and are presented in the box plot (Fig. 3a). The data indicate that the probe cell intensities from the individual arrays were similar. The reproducibility of the microarray results is shown by the correlation analyses on the biological replicates. The correlation between the samples at the gene level (Fig. 3b) and at the exon level (Fig. 3c) are presented in the scatter plots with the squared Pearson correlation coefficient (R2). All pairs of biological replicates have very high correlation (R2≥0.98 for gene level analysis; R2≥0.96 for exon level analysis). Knockdown of JMJD4 has more profound impact on gene expression than knockdown of JMJD6 (R2=0.97 versus R2=0.99 for gene level analysis; R2=0.96 versus R2=0.98 for exon level analysis).
(a) The box plot of feature probe cell intensity values from each individual array. (b) The scatter plots of signal intensity from each sample pair at the level of gene expression. (c) The scatter plots of signal intensity from each sample pair at the level of exon expression. R2: the squared Pearson correlation coefficient.
Usage Notes
The full quality control report can be accessed using the Affymetrix Expression Console software. The differentially expressed genes and exons can be identified using the Affymetrix Transcriptome Analysis Console software.
Additional Information
How to cite this article: Hu, Y.-J. & Imbalzano, A. N. Global gene expression profiling of JMJD6- and JMJD4-depleted mouse NIH3T3 fibroblasts. Sci. Data 3:160022 doi: 10.1038/sdata.2016.22 (2016).
References
References
Klose, R. J., Kallin, E. M. & Zhang, Y. JmjC-domain-containing proteins and histone demethylation. Nat. Rev. Genet. 7, 715–727 (2006).
Loenarz, C. & Schofield, C. J. Physiological and biochemical aspects of hydroxylations and demethylations catalyzed by human 2-oxoglutarate oxygenases. Trends. Biochem. Sci. 36, 7–18 (2011).
Johansson, C. et al. The roles of Jumonji-type oxygenases in human disease. Epigenomics 6, 89–120 (2014).
Hewitson, K. S. et al. Hypoxia-inducible factor (HIF) asparagine hydroxylase is identical to factor inhibiting HIF (FIH) and is related to the cupin structural family. J. Biol. Chem. 277, 26351–26355 (2002).
Lando, D. et al. FIH-1 is an asparaginyl hydroxylase enzyme that regulates the transcriptional activity of hypoxia-inducible factor. Genes. Dev. 16, 1466–1471 (2002).
Ge, W. et al. Oxygenase-catalyzed ribosome hydroxylation occurs in prokaryotes and humans. Nat. Chem. Biol. 8, 960–962 (2012).
Feng, T. et al. Optimal translational termination requires C4 lysyl hydroxylation of eRF1. Mol. Cell 53, 645–654 (2014).
Hsia, D. A. et al. KDM8, a H3K36me2 histone demethylase that acts in the cyclin A1 coding region to regulate cancer cell proliferation. Proc. Natl Acad Sci. USA 107, 9671–9676 (2010).
Youn, M. Y. et al. JMJD5, a Jumonji C (JmjC) domain-containing protein, negatively regulates osteoclastogenesis by facilitating NFATc1 protein degradation. J. Biol. Chem. 287, 12994–13004 (2012).
Chang, B., Chen, Y., Zhao, Y. & Bruick, R. K. JMJD6 is a histone arginine demethylase. Science 318, 444–447 (2007).
Webby, C. J. et al. Jmjd6 catalyses lysyl-hydroxylation of U2AF65, a protein associated with RNA splicing. Science 325, 90–93 (2009).
Noma, A. et al. Expanding role of the jumonji C domain as an RNA hydroxylase. J. Biol. Chem. 285, 34503–34507 (2010).
Unoki, M. et al. Lysyl 5-hydroxylation, a novel histone modification, by Jumonji domain containing 6 (JMJD6). J. Biol. Chem. 288, 6053–6062 (2013).
Liu, W. et al. Brd4 and JMJD6-associated anti-pause enhancers in regulation of transcriptional pause release. Cell 155, 1581–1595 (2013).
Boeckel, J. N. et al. Jumonji domain-containing protein 6 (Jmjd6) is required for angiogenic sprouting and regulates splicing of VEGF-receptor 1. Proc. Natl Acad Sci. USA 108, 3276–3281 (2011).
Heim, A. et al. Jumonji domain containing protein 6 (Jmjd6) modulates splicing and specifically interacts with arginine-serine-rich (RS) domains of SR- and SR-like proteins. Nucleic Acids Res. 42, 7833–7850 (2014).
Kunisaki, Y. et al. Defective fetal liver erythropoiesis and T lymphopoiesis in mice lacking the phosphatidylserine receptor. Blood 103, 3362–3364 (2004).
Bose, J. et al. The phosphatidylserine receptor has essential functions during embryogenesis but not in apoptotic cell removal. J. Biol. 3, 15 (2004).
Hu, Y. J. et al. Transcriptional and post-transcriptional control of adipocyte differentiation by Jumonji domain-containing protein 6. Nucleic Acids Res. 43, 7790–7804 (2015).
Chen, C. F. et al. Regulation of T cell proliferation by JMJD6 and PDGF-BB during chronic hepatitis B infection. Sci. Rep. 4, 6359 (2014).
Lee, Y. F. et al. JMJD6 is a driver of cellular proliferation and motility and a marker of poor prognosis in breast cancer. Breast Cancer Res. 14, R85 (2012).
Poulard, C. et al. Role of JMJD6 in Breast Tumourigenesis. PLoS One 10, e0126181 (2015).
Zhang, J., Ni, S. S., Zhao, W. L., Dong, X. C. & Wang, J. L. High expression of JMJD6 predicts unfavorable survival in lung adenocarcinoma. Tumour Biol. 34, 2397–2401 (2013).
Wang, F. et al. JMJD6 promotes colon carcinogenesis through negative regulation of p53 by hydroxylation. PLoS Biol. 12, e1001819 (2014).
Lee, C. R. et al. Elevated expression of JMJD6 is associated with oral carcinogenesis and maintains cancer stemness properties. Carcinogenesis 37, 119–128 (2015).
Data Citations
Hu, Y. J., & Imbalzano, A. N. Gene Expression Omnibus GSE76758 (2016)
Acknowledgements
We thank P. Spatrick and R. Barutcu for technical assistance. This work was supported by funding from NIH grant GM56244 to A.I.
Author information
Authors and Affiliations
Contributions
Y.H. and A.I. designed the experiments and wrote the manuscript. Y.H. generated RNA samples and performed data analyses.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
ISA-Tab metadata
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Hu, YJ., Imbalzano, A. Global gene expression profiling of JMJD6- and JMJD4-depleted mouse NIH3T3 fibroblasts. Sci Data 3, 160022 (2016). https://doi.org/10.1038/sdata.2016.22
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2016.22