Abstract
Large-scale reconstructions of neuronal populations are critical for structural analyses of neuronal cell types and circuits. Dense reconstructions of neurons from image data require ultrastructural resolution throughout large volumes, which can be achieved by automated volumetric electron microscopy (EM) techniques. We used serial block face scanning EM (SBEM) and conductive sample embedding to acquire an image stack from an olfactory bulb (OB) of a zebrafish larva at a voxel resolution of 9.25×9.25×25 nm3. Skeletons of 1,022 neurons, 98% of all neurons in the OB, were reconstructed by manual tracing and efficient error correction procedures. An ergonomic software package, PyKNOSSOS, was created in Python for data browsing, neuron tracing, synapse annotation, and visualization. The reconstructions allow for detailed analyses of morphology, projections and subcellular features of different neuron types. The high density of reconstructions enables geometrical and topological analyses of the OB circuitry. Image data can be accessed and viewed through the neurodata web services (http://www.neurodata.io). Raw data and reconstructions can be visualized in PyKNOSSOS.
Design Type(s) | observation design |
Measurement Type(s) | neuronal morphology phenotype |
Technology Type(s) | electron microscopy imaging assay |
Factor Type(s) | |
Sample Characteristic(s) | Danio rerio • olfactory bulb |
Machine-accessible metadata file describing the reported data (ISA-Tab format)
Similar content being viewed by others
Background & Summary
The dense anatomical reconstruction of neuronal populations is a major goal in neuroscience because it provides detailed structural information about neuronal cell types, circuits and wiring diagrams. Such information is important to understand the diversity of neuron types and the organization of neuronal networks1–17. Given that neuronal processes can be long, geometrically complex, and <100 nm in diameter, imaging methods for dense neuronal reconstructions need to provide ultrastructural resolution throughout large volumes. These goals can be achieved by volumetric electron microscopy (EM) approaches based on serial sectioning1,13,18–20. We used serial block face EM (SBEM), an approach that is based on the operation of an automated ultramicrotome inside the vacuum chamber of a scanning electron microscope1,6,18. Stacks of images are acquired by repeated removal of ultrathin sections (typically <30 nm) from a sample block and imaging of the block face. Brain tissue is impregnated with heavy metals to generate contrast of lipid membranes and other structures including synaptic densities21,22. To alleviate problems caused by charging of the sample18,23–25 we introduced conductive silver particles into the resin surrounding the sample. This approach increased the signal to noise ratio because images could be acquired in high vacuum26.
Tracing of neuronal processes in 3D image data is necessary to obtain geometrical representations of neurons. This process is challenging and requires substantial human input because fully automated image segmentation methods do not yet achieve sufficiently high accuracy. We traced neurons manually by placing connected nodes onto cross-sections of neurites in the image data. Tracing was performed in KNOSSOS27 or PyKNOSSOS26, two software tools designed specifically for high-throughput 3D image annotation and neuron reconstruction. The bulk of skeleton tracing tasks was outsourced to a company (www.ariadne-service.ch). Each neuron was initially reconstructed by three different individuals (‘tracers’). Skeletons were then compared and corrected iteratively by ‘COnvergence by Redundancy and Experts’ (CORE), a procedure that involves local re-tracing around mismatch points and focused input from a scientific expert. This workflow is efficient and achieves high accuracy26.
We analyzed the organization of neurons in the olfactory bulb (OB), which receives sensory input through an array of discrete neuropil units, the glomeruli. In adult vertebrates, each glomerulus receives input from olfactory sensory neurons expressing the same odorant receptor28,29. Principal neurons of the OB, the mitral/tufted cells, receive input from olfactory sensory neurons and provide output to multiple target areas of the OB. Within the OB, mitral/tufted cells interact via different types of GABAergic interneurons (INs)30. In the adult OB, the most prominent IN types are periglomerular and short axon cells in the upper layers and granule cells in the deep layers. INs are critical for neuronal computations such as the equalization and decorrelation of odor-evoked activity patterns30–33 but the topological organization of IN networks has been poorly understood. Moreover, most INs arise late in development34,35 but it has been unclear how IN networks change as the OB matures.
To address these and additional questions we acquired a stack of SBEM images of the OB in a zebrafish larva and reconstructed approximately 98% of all OB neurons (n=1,022; Figs 1 and 2). The results identified novel rare cell types and indicate that granule cells develop later than other IN types. Inter-glomerular projections were mediated primarily by INs and formed a specific pattern that is governed by glomerular identity, as defined by the associated odorant receptors. The datasets allow for multiple additional analyses including further morphological analyses of individual neurons, geometrical analyses of glomeruli and neuronal populations, ultrastructural analyses of synapses in different cell types, other ultrastructural analyses, and analyses of neuronal subpopulations associated with specific glomeruli. Moreover, the high density and accuracy of skeleton reconstructions may be useful for the development and validation of procedures for automated segmentation of volumetric EM image data7,8,36–39. Data are accessible through the neurodata web services of the open connectome project (http://neurodata.io)40.
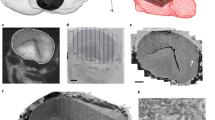
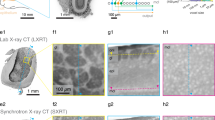
(a) EM image showing cross-section through the OB. ON, olfactory nerve; S, silver particles. (b) Reslices through a subvolume. (c) Examples of synapses. (d) Orientation of image plane in (a) is illustrated relative to the reconstruction of all 1,022 neurons in the OB (Dataset 2). Colors depict the group of the parent glomerulus (longest neurite length). (e) 3D outlines of the 17 glomeruli in the OB (Dataset 3). Colors depict different groups of glomeruli as defined by Braubach and colleagues47,48. 3D rendering in (d,e) was done in PyKNOSSOS. Parts of this figure have been modified from illustrations in a previous publication26.
Methods
Most methods have been described in detail in a previous publication26.
Electron microscopy
Dissection
All animal procedures were approved by the Veterinary Department of the Canton Basel-Stadt (Switzerland). Fish were housed and bred in an animal facility under standard conditions. The sample was prepared from a zebrafish larva 4.5 days post fertilization (dpf) that expressed the genetically encoded calcium indicator GCaMP5 under the control of the panneuronal elavl3 promoter41. The sex is not yet determined at this developmental stage. Prior to sample preparation, the larva was paralyzed with Mivacron, mounted in low-melting agarose (Sigma Aldrich A9414), and immersed in standard E3 zebrafish medium42 (5 mM NaCl, 0.17 mM KCl, 0.33 mM CaCl2, 0.33 mM MgSO4, pH 7.4). Responses of olfactory bulb neurons to odor stimulation were then measured by multiphoton microscopy as described43 over a period of approximately 4 h. Subsequently, the larva was anesthetized in tricaine methanesulfonate (MS-222) and a small craniotomy was made above the contralateral olfactory bulb using a glass pipette to facilitate the penetration of reagents into the brain. Immediately afterwards the larva was transferred into EM fixative (Table 1).
Fixation and staining
The sample was fixed and stained en bloc using an established protocol with minor modifications21,22. The procedure includes successive exposures of the sample to reduced OsO4, thiocarbohydrazide (TCH), OsO4, uranyl acetate and lead aspartate. Reagents and solutions are described in Table 1. Tissue was initially fixed in EM fixative for 1 h at room temperature and 1 h on ice. After five washes (always 3 min each) with ice-cold cacodylate buffer, samples were postfixed in RedOs solution for 1 h on ice, washed 5 times in bidistilled H20, and incubated in fresh TCH solution for 20 min at room temperature. In a second postfixation step, samples were incubated in Os solution for 30 min at room temperature and washed five times in bidistilled H20. The sample was then incubated in UA solution and left in a refrigerator (4 °C) overnight, washed five times in bidistilled H20, incubated at 60 °C for 30 min in bidistilled H20, incubated in Walton’s lead aspartate for 20 min at 60 °C, and washed five times in bidistilled H20 at room temperature. The sample was then dehydrated in an ethanol series (20%, 50%, 70%, 90%, 100%, 100%; 5 min each), incubated in 50% ethanol and 50% Epon resin for 30 min, and incubated in 100% Epon resin for 1 h at room temperature. Epon resin was exchanged and samples were again incubated at room temperature for 4–12 h (overnight).
Embedding and mounting of the sample
After fixation and staining the sample was embedded in a silver-filled epoxy to minimize charging of the sample block during exposure to the electron beam. The silver-filled epoxy was made from a commercially available 2-component glue that contains elongated silver particles with a size up to 45 μm (EE129-4; Epo-Tek). Deviating slightly from the instructions for normal use, the two components A and B were mixed in a ratio of 1.25:1. This ratio yielded conductive sample blocks with mechanical properties that allowed for reliable thin sectioning (25 nm). The sample was transferred to the conductive medium and moved gently to ensure that silver particles contact the surface of the tissue. The epoxy was then cured by incubation at 60 °C for 48 h. The resistance across the sample block was measured with an Ohm-meter and found to be <1 Ohm. The sample block was glued on an aluminum stub for SBEM (Gatan) using cyanoacrylate glue and trimmed to a pyramid with a block face area of approximately 300 μm×200 μm.
Electron microscopy
Images were acquired using a scanning electron microscope (QuantaFEG 200; FEI) equipped with an automated ultramicrotome inside the vacuum chamber (3View; Gatan). The ultramicrotome cut successive sections at a thickness of 25 nm. After each section, the sample block-face was scanned. The microscope, the stage and the ultramicrotome were controlled using DigitalMicrograph software (Gatan). Images were generated by detection of backscattered electrons with a silicon diode detector (Opto Diode Corp., USA). The signal was preamplified and further amplified by standard components of the 3View system (Gatan) before digitizing at 16 bit. The sample was scanned in high vacuum with a landing energy of 2 kV. Due to conductive embedding, artifacts arising from sample charging were negligible26. Pixel size was 9.25×9.25 nm2, the electron dose was 17.5 e−nm−2, and the pixel acquisition rate was 200 kHz. To cover the entire cross-section of the OB, multiple images (tiles) were acquired with a size up to 4,096×4,096 pixels and an overlap of 5–8%. Choosing a relatively small tile size allowed us to effectively avoid image distortions. The size and arrangement of tiles was adjusted to changes in the cross-section of the OB during acquisition of the stack. 4,746 successive sections were acquired over a period of 35 days. One section was lost due to technical problems (between slices 1301 and 1302). The focus was readjusted on average every 14.5 h. The final stack was cropped to a size of 72.2×107.8×118.6 μm3.
Image processing
Because distortions of raw images were minimized by conductive embedding and small tile size, registration of images in the stack could be achieved by simple translational alignment procedures. Pre-processing, registration and stitching of images was performed using custom software tools written in MATLAB that allowed for parallel batch processing of large datasets. For image registration, translational offsets between neighboring image tiles were calculated using a custom optimized cross-correlation procedure in MATLAB. Image columns were standardized by subtracting the mean and dividing by the standard deviation of the pixel intensities. The same standardization was subsequently applied to the rows. Translational offsets between neighboring images were calculated by determining the maximum 2D cross-correlation of the standardized images in the Fourier domain. The second inverse Fourier transform was restricted to the central region of the 2D cross-correlogram reflecting a maximal expected offset between overlapping image parts of 256 pixels. Using this procedure the offsets between two 1024×1024 pixel images with translational offsets of dX=69 and dY=66 pixels could be calculated in <0.36 s on a laptop computer (Intel Core i7-3520M CPU @ 2.90 GHz×4, 7.5 GB RAM, Ubuntu 12.10). Offsets were used to optimize the tile positions in a global total least square displacement sense. Image contrast was normalized by fitting a Gaussian distribution to the pixel intensity histogram and thresholding at 1.5–3 standard deviations around the peak of the Gaussian distribution to convert the images to 8 bit. Stacks were then divided into cubes of 128×128×128 voxels for dynamic data loading in KNOSSOS or PyKNOSSOS.
Software: PyKNOSSOS
PyKNOSSOS: general description
We developed PyKNOSSOS, a software package for ergonomic manual skeleton tracing, visualization and annotation of neurons in 3D image datasets (Figs 3 and 4). Other publicly available applications for similar purposes include KNOSSOS (www.knossostool.org)27, CATMAID (http://catmaid.org/)44 and ilastik (www.ilastik.org)45. PyKNOSSOS was inspired by KNOSSOS but is fully implemented in Python, which enables rapid and efficient code development, modification and extension. Efficient real-time 3D visualization of large numbers of reconstructed neurons together with raw image data is achieved using the VTK-Python bindings in combination with custom image data loading procedures written in C++. The user interface of PyKNOSSOS has been designed by close interactions between programmers and users to integrate user feedback into the application as directly as possible. PyKNOSSOS is written in Python (version 2.7) and runs on various operating systems including Linux and Windows 7. The main graphical user interface is based on the PyQT4 library. PyKNOSSOS can be run as stand-alone executable or directly from script using a Python interpreter. All source code and detailed installation instructions can be found on https://github.com/adwanner/PyKNOSSOS.
PyKNOSSOS running on a local machine streams cubed image data from a web server (data store; NeuroData) and stores cubes on disk. Cubes are generated by NeuroData and dynamically loaded into PyKNOSSOS. Annotations (e.g., skeleton reconstructions) are downloaded from a repository (label store) and may be modified locally.
(a) Screenshot of PyKNOSSOS in tracing mode. (b) Screenshot of PyKNOSSOS using the NeuronLibrary plugin for visualization of specific neurons. The table on the left shows part of Dataset 4 (neurite length of each neuron in each glomerulus). Specific neurons were extracted by filtering, color-coded, and visualized together with 3D outlines of glomeruli (Dataset 3).
PyKNOSSOS: multi-scale dynamic data loading
Similar to KNOSSOS, PyKNOSSOS accesses a cubed version of the dataset in which the imaging data is divided into 8-bit cubes (Fig. 3). The default cube size is 128 pixel edge length but arbitrary cube dimensions can be processed. While navigating through the dataset, a custom C++ routine dynamically loads the image data in a pre-defined neighborhood around the current focal point into RAM. If multiple versions of the dataset at different zoom levels exist, PyKNOSSOS supports instantaneous seamless browsing and multi-scale zooming. At a given zoom level li, a cubed neighborhood of typically 320–576 pixels around the current focal point is loaded into memory for all stored consecutive zoom levels li-1, li and li+1. This permits navigation through large datasets (>4 TB) with minimal RAM requirements (<4.5 GB). The data can be loaded from local hard disk storage, via a hybrid streaming pipeline from HTTP accessible servers, or via the JPEG stack service of the data API of neurodata (http://www.neurodata.io; Fig. 3).
PyKNOSSOS: built-in Python console and plugin interface
In development mode, PyKNOSSOS allows direct interaction and access to its core variables and functions via a built-in Python console. In addition, PyKNOSSOS features an easy-to-use plugin interface that allows arbitrary custom Python scripts to interact with PyKNOSSOS at runtime.
PyKNOSSOS: Orthogonal tracing mode and arbitrary reslices
In the default configuration, PyKNOSSOS has five viewports (Fig. 4a). Image data is displayed in four viewports: the YX viewport (imaging plane) and three mutually orthogonal viewports of arbitrary orientation. In tracing mode, one of the latter is perpendicular to the current tracing direction. We find that this ‘auto-orthogonal’ view increases tracing speed and facilitates the identification of branch points and synapses. The reslice views are generated by a tri-linear interpolation procedure to efficiently extract reslices at arbitrary orientations and zoom levels. The fifth viewport displays the skeleton reconstructions.
PyKNOSSOS: synapse annotation mode
Synapses can be annotated by three successive clicks to place connected nodes onto the first neuron, the synaptic cleft, and the second neuron. If skeletons of the two neurons are available, the first and the last click connect to existing skeleton nodes. The first click can be omitted if a skeleton node is active. Synapses can be assigned to user-defined classes and scored with a confidence level.
PyKNOSSOS: visualization
All visualizations are done using the VTK-Python bindings. Skeleton nodes can be rendered either as points or spheres and skeleton edges can be rendered as lines or tubes. For simple volume representation of convex objects such as somata or olfactory glomeruli, convex hulls can be rendered from sets of nodes.
Neuron reconstruction
Manual tracing
Skeletons of neurons were traced manually using KNOSSOS or PyKNOSSOS. Tracings of OB cells were initiated from seed points close to somata. When cells showed obvious features of glia such as sheet-like processes, tracings were abandoned. A small number of cells was excluded because their morphology was not neuron-like (usually no or only minor processes) or because they did not extend processes into neuropil regions of the OB. All reconstructions of OB cells with neuron-like morphology were completed and included. Starting from seed points, tracers followed neuronal processes by placing successive nodes onto cross-sections of neurites in the original image data, resulting in a skeleton representation of each neuron27. Most skeletons were reconstructed by a professional tracing service (www.ariadne-service.ch).
Consolidation of skeletons and error correction
Each neuron was initially reconstructed three times by different individuals. Multiple tracings were then combined into a ‘consolidated’ skeleton by CORE (‘COnvergence by Redundancy and Experts’), a procedure based on redundancy (multiple independent tracings of each neuron) and focused input by an expert. During the CORE procedure, errors were identified and corrected. The procedure is described in detail elsewhere26. Briefly, individual skeletons were resampled to a node density of approximately 100 nm. Starting from seeds, nodes of skeletons were iteratively combined into ‘cliques’ based on their distance and connectedness. The resulting cliques thus represented the agreement between independent tracings and formed the basis of the consolidated skeletons. Disagreements between one skeleton and the others resulted in nodes that were not part of a clique. To resolve such disagreements, the point of divergence (‘mismatch point’) between the skeletons was inspected by a scientific expert. The expert usually recruited two additional tracers to generate additional tracings of local segments around the mismatch point. In rare cases, particularly when errors were obvious, the expert intervened directly and overruled the tracers. A new consolidated skeleton was then computed and the procedure was repeated until all mismatch points were resolved26.
Annotation of glomeruli
Glomeruli were defined as distinct neuropil regions containing axons of olfactory sensory neurons, which were identified by their characteristic dark cytoplasmic staining46. Seventeen glomeruli were outlined manually in 3D by placing vertices around glomerular boundaries in PyKNOSSOS (Fig. 2e). Because glomerular boundaries were sometimes difficult to delineate when adjacent glomeruli were not separated by somata the outlines may not be fully precise. Sixteen glomerular outlines corresponded in size, shape and position to a previous light microscopic description of glomeruli in the larval zebrafish OB at similar or slightly earlier developmental stages47,48. These glomeruli were subdivided into seven groups and named according to the nomenclature of Braubach et al.47,48 (dG group: dG; dlG group: dlG; vmG/vaG group: vmG, vaG; vpG group: vpG1, vpG2; lG group: lG3, lG4, lGx; mdG group: mdG1, mdG2, mdG3, mdG4, mdG5, mdG6; maG group: maG). The seventeenth glomerulus was assigned to the mdG group and named mdGx based on its position. In addition, we identified an elongated neuropil volume below lG3 and between dlG and vpG2 that contained sensory axons. This volume was not included in the set of glomeruli because it was less distinct and smaller than other glomeruli. Furthermore, the central OB contained a neuropil region that was not classified as a glomerulus because we did not detect axons of olfactory sensory neurons. This volume may correspond to the plexiform layer of the adult OB.
Code availability
The code for PyKNOSSOS and a manual can be found at https://github.com/adwanner/PyKNOSSOS. See Usage Notes for instructions to install and run PyKNOSSOS.
Data Records
Dataset 1: EM image stack
Dataset 1 (Data Citation 1) contains 4,746 serial EM images in 8-bit TIF format covering the OB of a zebrafish larva at 4.5 dpf and parts of the adjacent telencephalon (374 GB; Fig. 2a–c). The voxel size is 9.25×9.25×25 nm3. Images have been acquired, stitched and aligned as described in Methods.
Completeness
The volume contains >99.96% of an entire OB and parts of the adjacent telencephalon. The remaining 0.04% of the OB volume is located in glomerulus mdG6 and accounts for <3.97% of the volume of mdG6. One section was lost (between slices 1301 and 1302).
Data access
The image data is available under http://doi.org/10.7281/T1MS3QN7 and can be accessed through the neurodata web services (NeuroData; http://neurodata.io/wanner16). Data can either be viewed interactively through the NeuroDataViz web interface (http://viz.neurodata.io/WannerAA201605/) or using PyKNOSSOS (Fig. 3). NeuroData provides various APIs to extract and download specific subvolumes (‘cutouts’) for external analysis. A detailed description of the services is available at http://neurodata.io/wanner16.
Data viewing in PyKNOSSOS
The repository containing PyKNOSSOS code (https://github.com/adwanner/PyKNOSSOS) includes a dataset configuration file (in folder Datasets/wanner16) to access EM image data hosted by neurodata web services from a local PyKNOSSOS installation. After installation of PyKNOSSOS the dataset can be accessed by loading the configuration file from the menu entry ‘Datasets’ in the ‘Settings’ window. The program automatically fetches the cubed image data around the current location using the online JPEG stack service of NeuroData. By default, PyKNOSSOS uses one download stream but the user can increase the number of parallel download streams in the ‘Loader’ tab of the ‘Settings’ window. PyKNOSSOS uses a hybrid approach for online image data streaming. Any downloaded image cube is stored permanently on the local disk in the same folder as the configuration file (Fig. 3). This minimizes the data streaming traffic because each image cube has only to be downloaded once. In addition, the permanent caching enables offline browsing of already visited locations.
Dataset 2: Skeleton reconstructions of neurons in the OB
Dataset 2 (Data Citation 2) contains manual skeleton reconstructions and soma outlines of 1,022 neurons in the larval zebrafish OB (101 MB; Fig. 2d). Each reconstructed neuron has a unique ID and is provided in the native PyKNOSSOS file format *.nmx. An NMX-file is a zip-archive container in which annotations and skeletons are saved in individual NML-files, the native KNOSSOS file format, which is XML-based27. Each skeleton consists of 3D nodes (vertices) connected by edges. Soma outlines are computed from convex sets of 3D points.
Completeness
The 1,022 reconstructed neurons in the dataset represent approximately 98% of all neurons in the OB26.
Data access
Data can be downloaded from https://doi.org/10.5281/zenodo.58985. Further information including a link to the download site is given on a website hosted by NeuroData (http://neurodata.io/wanner16).
Data viewing using PyKNOSSOS
Download and extract the WannerAA201605_SkeletonsGlomeruli.zip from the location specified above. Each NMX-file with the file name pattern Neuron_id*.nmx contains the soma outline and reconstructed skeleton of the neuron with the indicated unique ID. To load one or multiple of these NMX files, start PyKNOSSOS, click on the ‘Open’ command in the ‘File’ menu of the ‘Settings’ window, and select one or multiple files. Multiple skeletons can be loaded simultaneously or sequentially by appending new skeletons to the already loaded ones. Load image data as described above (Dataset 1) to overlay skeletons onto raw data. Various display options for skeletons and somata can be found in the ‘Skeleton’ and ‘Soma’ tab of the ‘Visualization’ tab of the ‘Settings’ window, respectively.
Dataset 3: Manually outlined volumes of glomeruli in the OB
Dataset 3 (Data Citation 2) contains the outlines of 17 glomeruli that are represented as convex hulls around manually placed 3D points (Fig. 2e). The point sets are provided in the native PyKNOSSOS file format *.nmx. Each glomerulus has an unique ID and a label according to the nomenclature of Braubach et al.47,48.
Completeness
The 17 glomeruli include all glomeruli annotated by Braubach et al.47,48 at a similar developmental stage and one additional glomerulus (mdGX)26. In addition, we noticed an elongated neuropil volume below lG3 between dlG and vpG2 that contained sensory axons. This volume was not included in the set of glomeruli because it was less distinct and smaller than other glomeruli.
Data access
Data can be downloaded from https://doi.org/10.5281/zenodo.58985. Further information including a link to the download site is given on a website hosted by NeuroData (http://neurodata.io/wanner16).
Data viewing using PyKNOSSOS
Download and extract the WannerAA201605_SkeletonsGlomeruli.zip from the location specified above. The file Glomeruli.nmx contains the outlines of all glomeruli. Use the ‘Open’ command in the ‘File’ menu of the ‘Settings’ window to load the file in PyKNOSSOS. To view glomeruli together with skeletons and image data we recommend to adjust the glomerular region opacity in the ‘Region’ tab of the ‘Visualization’ tab in the ‘Settings’ window.
Dataset 4: Glomerular innervation pattern of OB neurons
Dataset 4 (Data Citation 2) is a spreadsheet that contains a table in *.csv format that lists the neurite length of each neuron in each glomerulus, generated from the information in Datasets 2 and 3. In addition, the table contains information about the identity (ID) and type of each neuron. The following cell types are distinguished: (1) mitral cells (principal neurons of the OB), (2) class 1 INs (small INs with local processes), (3) class 2 INs (larger INs, often with polarized neurites innervating multiple glomeruli), (4) class 3 INs (usually large INs, often with extensive neurites innervating multiple glomeruli), (5) unusual projection neurons (UPNs; four neurons with IN-like neurites that project out of the OB), (6) large olfactory bulb cells (LOCs; two very large cells with neuron-like morphology but unusual ultrastructure), and (7) other cells that could not be classified unambiguously. IN classes 1–3 were distinguished by clustering of morphological features. For more information see ref. 26.
Data access
Data can be downloaded from https://doi.org/10.5281/zenodo.58985. Further information including a link to the download site is given on a website hosted by NeuroData (http://neurodata.io/wanner16).
Data viewing using PyKNOSSOS
Download and extract the WannerAA201605_SkeletonsGlomeruli.zip from the location specified above. The file WannerAA201605.csv contains a table that lists the neurite length of each neuron in each glomerulus. The csv-file can be loaded into PyKNOSSOS using the NeuronLibrary plugin (Fig. 4b). This plugin facilitates various operations such as filtering of neurons by cell type or glomerulus (Fig. 5).
(a) Mitral cells with >50 μm neurite length in glomerulus mdGX. (b) INs of classes 1, 2, and 3 with >50 μm neurite length in lG3 and lGX. (c) Unusual projection neurons (UPNs) and large olfactory bulb cells (LOCs). (d) Class 1 INs with >30 μm neurite length in dG. (e) Class 2 INs with >100 μm neurite length in dG. (f) Class 3 INs with >100 μm neurite length in dG. (g) Examples of mitral cells innervating different subcompartments of dG, viewed in three different orientations. Scale bar: 20 μm.
Technical Validation
The signal-to-noise ratio and resolution of image data is clearly sufficient to trace neurites and identify synapses. Conductive sample embedding enabled data acquisition in high vacuum. In tests performed with comparable samples, high vacuum conditions increased the signal-to-noise ratio of EM images by a factor of three or more compared to low vacuum conditions (14 or 20 Pa water pressure)26, which have been used in most previous SBEM applications6,18. No noticeable image distortions were seen throughout most of the dataset because tile sizes were small. Occasionally, small local distortions occurred in regions where the density of conductive material is low. However, these distortions do not affect image interpretation. High quality of image alignment throughout the volume was confirmed by orthogonal reslices and specific inspection of stitching boundaries26.
Neuron reconstruction is error-prone and requires error correction procedures to obtain high accuracy. In order to achieve efficient error correction we initially generated three independent reconstructions of each neuron. Errors were then identified and corrected by CORE, a semi-automated method that involves inspection of mismatch points by an expert and local re-annotation around mismatch points26. The final reconstruction accuracy was quantified by different measures: (1) Recall, a measure for errors caused by missed processes; (2) precision, a measure for errors caused by wrongly traced processes; and (3) the ‘relative length error’, a measure that takes into account both types of errors26. Accuracy was evaluated against a ground truth generated by highly redundant reconstruction and expert error correction, and by comparing independent reconstructions to each other. Error quantification showed that accuracy was high: typically, the total length of missing or incorrectly annotated processes was <5% of the total neurite length26. Additional tracing of ‘orphan’ profiles showed that missed processes were usually short terminal branches, <3.5 μm in length, with few synapses26.
Usage Notes
Only a small amount of the information contained in the image data (Dataset 1) and in the skeleton reconstructions (Dataset 2) has been extracted and analyzed so far26. These datasets are unique because they represent the near-complete 3D ultrastructure of an entire brain region and a very dense morphological reconstruction of the neurons contained in this brain region. To facilitate further annotations and analyses of these datasets we describe basic procedures to access, visualize and organize these datasets.
Installation of PyKNOSSOS and data browsing
-
1
Download and extract the zip-archive of the latest release of PyKNOSSOS from https://github.com/adwanner/PyKNOSSOS.
-
2
Run PyKNOSSOS
-
a
from the Python script ‘PyKnosssos.py’ in a Python interpreter: Make sure that you have installed the necessary Python modules listed at the beginning of the script. Some modules such as QxtSpanSlider and images2gif are non-standard and might be missing in some Python distributions.
-
b
using the stand-alone binary PyKnossos.exe.
-
a
-
3
To load the image data (Dataset 1), click on ‘Load new dataset’ in the ‘Dataset’ menu of the ‘Settings’ window.
-
4
Browse to the folder ‘Datasets/wanner16’ that was included in the release zip-archive and select the dataset configuration file ‘wanner16.conf’.
-
5
If you are in ‘online’ mode, i.e. you are connected to the internet and the option ‘Working offline’ in the ‘Dataset’ menu of the ‘Settings window’ is not checked, the hybrid streaming of image dataset cubes should start immediately.
-
6
Load skeletons and glomerular outlines as described above (Data Records, Datasets 2 and 3). Reconstructions will be shown together with image data.
-
7
See the included PyKNOSSOS manual for detailed instructions on how to use PyKNOSSOS.
Basic navigation:
-
Panning: Keep the left mouse button pressed and move the mouse cursor
-
Move perpendicular to the currently active viewport: F/D keys or UP/DOWN keys or scroll with the mouse wheel
-
Zoom in/out:
-
○ with mouse: Keep CTRL+right mouse button pressed while moving the mouse cursor up/down
-
○ keyboard: +/− keys
-
○ reset zoom to 0 (native resolution) Space-key
-
Filtering and visualization of reconstructed neurons
-
1
Install and run PyKNOSSOS
-
2
Download and extract the zip-file ‘SkeletonsGlomeruli.zip’ that contains the datasets 2, 3 and 4.
-
3
Click on ‘Load plugin’ in the ‘Plugins’ menu of the ‘Settings’ window.
-
4
Browse to the folder ‘plugins’ that was included in the release zip-archive and select the Python script ‘NeuronLibrary.py’.
-
5
An ‘Open file…’ dialog will be displayed that allows you to load the desired neuron library file.
-
6
Browse to the folder where SkeletonsGlomeruli.zip was extracted to and select the csv-file ‘WannerAA201605.csv’.
-
7
A new dockable window containing a table view of all reconstructed neurons will be displayed.
-
8
Click on the button ‘Data source’ and browse to the folder where SkeletonsGlomeruli.zip was extracted to.
Basic usage:
-
Double-clicking on any row will load and display the corresponding neuron in PyKNOSSOS.
-
By left-clicking on the table header, you can choose different filter options in order to filter the table.
-
The different glomeruli have the two filter options ‘>’ and ‘<=’. Depending on the threshold value in the ‘Inn. thres’ spinbox, these filters show those neurons that have more/less neurite length in the corresponding glomerulus.
-
Clicking on the ‘Show’ button will load and show all currently filtered/displayed neurons.
-
In order to display the glomerular outlines together with the neurons, check ‘Include glomeruli’.
-
Right-clicking on any row allows you to change the display color of the corresponding neuron. If multiple rows are selected (partially), the new color will be assigned to all (partially) selected rows.
-
Various display options for skeletons, somas and glomerular regions can be found in the ‘Skeleton’, ‘Soma’ and ‘Region’ tabs of the ‘Visualization’ tab of the ‘Settings’ window, respectively.
Future perspectives
Our datasets can be further mined to address a wide range of questions in cellular, developmental and systems neuroscience. An obvious next step is to annotate the synapses between reconstructed neurons in order to reconstruct the full wiring diagram, an effort that is underway. Exhaustive dense reconstructions of wiring diagrams have so far been achieved only in C. elegans2,3 and in representative subvolumes of very few other circuits5,7–11,14. The topological analysis of wiring diagrams is particularly important to understand the function of neuronal circuits whose connectivity cannot be approximated based on topological relationships or other means, as in the OB.
Future work may generate dense volumetric reconstructions of neurons in the zebrafish OB by combining automated segmentation with skeleton reconstructions7,8,36–39. This approach uses machine learning methods to over-segment EM image data in 3D, resulting in ‘supervoxels’ representing subvolumes of neurons. Skeletons are then used to merge supervoxels from the same neurons. The generation of skeletons is typically a bottleneck in this procedure because it involves substantial manual labor. The skeletons provided along with our EM image data should therefore greatly facilitate volumetric neuron reconstructions.
Previous studies have mapped glomeruli in the developing zebrafish OB, reported expression patterns of various marker genes, determined projection patterns of mitral cells, and characterized odor responses of glomeruli and OB neurons43,47–56. Anatomical information about neurons in the zebrafish OB is, however, incomplete. Most anatomical studies focused on mitral cells43,49,50,57,58 but only few IN types have been characterized in detail31,59. The detailed morphological information contained in our datasets may therefore be exploited to generate a comprehensive atlas of the zebrafish OB that integrates molecular, anatomical and functional information.
The OB of zebrafish and other species contains glomeruli responding to common odorants, as well as subsets of glomeruli that detect specific odorants with a defined biological function such as pheromones52,53. Our datasets provide a unique opportunity to explore whether these classes of glomeruli differ in their cellular composition and microcircuit organization, and to study how these glomeruli interact via intra-bulbar projections. Moreover, the high density of our reconstructions allows for a detailed analysis of the sub-glomerular organization of neuronal microcircuits. Preliminary observations indicate that individual mitral cells do not always innervate entire glomeruli but can be restricted to distinct subcompartments (Fig. 5g)60. It may thus be interesting to examine whether these subcompartments reflect distinct microcircuits associated with the same glomerulus.
Datasets described in this paper should be particularly valuable to link the morphology of neurons to their ultrastructure. The high resolution of the EM data allows for the quantitative analysis of subcellular components such as mitochondria, endoplasmatic reticulum (ER), primary cilia and synapses. Features of these organelles may then be mapped onto the 3D reconstructions of individual neurons and compared across neurons. A quantitative analysis of synapses, for example, would provide the opportunity to compare the size and other ultrastructural features of synapses within the same neuron, across neurons of the same type, and across neurons of different types. Additional analyses may then ask whether synapses of the same neuron vary systematically depending on the identity of the synaptic partner. Moreover, it is possible to determine whether mitochondria, ER or other organelles are associated preferentially with specific types of synapses. A combined morphological and ultrastructural analysis of neurons has major potential to discover unknown rules governing the organization of neuronal microcircuits12,13.
The development of automated procedures for the reconstruction of neurons from volumetric EM data is an active field that requires ground truth datasets. Such datasets are usually obtained by labor-intensive manual reconstruction and often contain only parts of neurons, which complicates the quantification of reconstruction performance at the level of entire neurons. Our skeleton reconstructions may serve as valuable ground truth because they comprise a large number of neurons that were fully reconstructed with high accuracy. In the future, automated reconstruction may also incorporate statistical information about the geometry of neurons. Such information needs to be extracted from large, high-resolution morphological datasets such as our skeleton reconstructions. Our datasets can therefore serve a resource for the future development of automated reconstruction procedures.
Additional Information
How to cite this article: Wanner, A. A. et al. 3-dimensional electron microscopic imaging of the zebrafish olfactory bulb and dense reconstruction of neurons. Sci. Data 3:160100 doi: 10.1038/sdata.2016.100 (2016).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Denk, W., Briggman, K. L. & Helmstaedter, M. Structural neurobiology: missing link to a mechanistic understanding of neural computation. Nat. Rev. Neurosci. 13, 351–358 (2012).
White, J. G., Southgate, E., Thomson, J. N. & Brenner, S. The structure of the nervous system of the nematode Caenorhabditis elegans. Philosophical transactions of the Royal Society of London 314, 1–340 (1986).
Varshney, L. R., Chen, B. L., Paniagua, E., Hall, D. H. & Chklovskii, D. B. Structural properties of the Caenorhabditis elegans neuronal network. PLoS computational biology 7, e1001066 (2011).
Jarrell, T. A. et al. The connectome of a decision-making neural network. Science 337, 437–444 (2012).
Ohyama, T. et al. A multilevel multimodal circuit enhances action selection in Drosophila. Nature 520, 633–639 (2015).
Briggman, K. L., Helmstaedter, M. & Denk, W. Wiring specificity in the direction-selectivity circuit of the retina. Nature 471, 183–188 (2011).
Helmstaedter, M. et al. Connectomic reconstruction of the inner plexiform layer in the mouse retina. Nature 500, 168–174 (2013).
Kim, J. S. et al. Space-time wiring specificity supports direction selectivity in the retina. Nature 509, 331–336 (2014).
Lee, W. C. et al. Anatomy and function of an excitatory network in the visual cortex. Nature 532, 370–374 (2016).
Berck, M. E. et al. The wiring diagram of a glomerular olfactory system. eLife 5 (2016).
Takemura, S. Y. et al. A visual motion detection circuit suggested by Drosophila connectomics. Nature 500, 175–181 (2013).
Morgan, J. L., Berger, D. R., Wetzel, A. W. & Lichtman, J. W. The Fuzzy Logic of Network Connectivity in Mouse Visual Thalamus. Cell 165, 192–206 (2016).
Kasthuri, N. et al. Saturated Reconstruction of a Volume of Neocortex. Cell 162, 648–661 (2015).
Randel, N. et al. Neuronal connectome of a sensory-motor circuit for visual navigation. eLife 3 (2014).
Zwart, M. F. et al. Selective Inhibition Mediates the Sequential Recruitment of Motor Pools. Neuron 91, 615–628 (2016).
Ding, H., Smith, R. G., Poleg-Polsky, A., Diamond, J. S. & Briggman, K. L. Species-specific wiring for direction selectivity in the mammalian retina. Nature 535, 105–110 (2016).
Schneider-Mizell, C. M. et al. Quantitative neuroanatomy for connectomics in Drosophila. eLife 5 (2016).
Denk, W. & Horstmann, H. Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue nanostructure. PLoS Biol 2, e329 (2004).
Briggman, K. L. & Bock, D. D. Volume electron microscopy for neuronal circuit reconstruction. Curr. Opin. Neurobiol. 22, 154–161 (2012).
Lichtman, J. W. & Denk, W. The big and the small: challenges of imaging the brain's circuits. Science 334, 618–623 (2011).
Deerinck, T. J. et al. Enhancing serial block-face scanning electron microscopy to enable high resolution 3D nanohistology of cells and tissues. Microsc. Microanal. 16, 1138–1139 (2010).
Tapia, J. C. et al. High-contrast en bloc staining of neuronal tissue for field emission scanning electron microscopy. Nat Protoc. 7, 193–206 (2012).
Titze, B. & Denk, W. Automated in-chamber specimen coating for serial block-face electron microscopy. Journal of microscopy 250, 101–110 (2013).
Robinson, V. N. E. The elimination of charging artefacts in the scanning electron microscope. J. Phys. E: Sci. Instrum 8, 638–640 (1975).
Mathieu, C. Beam-gas and signal-gas interactions in the variable pressure scanning electron microscope. Scan Microsc 13, 23–41 (1999).
Wanner, A. A., Genoud, C., Masudi, T., Siksou, L. & Friedrich, R. W. Dense EM-based reconstruction of the interglomerular projectome in the zebrafish olfactory bulb. Nat Neurosci 19, 816–825 (2016).
Helmstaedter, M., Briggman, K. L. & Denk, W. High-accuracy neurite reconstruction for high-throughput neuroanatomy. Nat Neurosci 14, 1081–1088 (2011).
Axel, R. The molecular logic of smell. Sci. Am. 273, 130–137 (1995).
Yoshihara, Y. Molecular genetic dissection of the zebrafish olfactory system. Results and problems in cell differentiation 47, 97–120 (2009).
Wilson, R. I. & Mainen, Z. F. Early events in olfactory processing. Annu. Rev. Neurosci. 29, 163–201 (2006).
Zhu, P., Frank, T. & Friedrich, R. W. Equalization of odor representations by a network of electrically coupled inhibitory interneurons. Nature Neurosci. 16, 1678–1686 (2013).
Friedrich, R. W. Information processing in the olfactory system of zebrafish. Annu. Rev. Neurosci. 36, 383–402 (2013).
Banerjee, A. et al. An Interglomerular Circuit Gates Glomerular Output and Implements Gain Control in the Mouse Olfactory Bulb. Neuron 87, 193–207 (2015).
Rosselli-Austin, L. & Altman, J. The postnatal development of the main olfactory bulb of the rat. J. Dev. Physiol. 1, 295–313 (1979).
Mack-Bucher, J. A., Li, J. & Friedrich, R. W. Early functional development of interneurons in the zebrafish olfactory bulb. Eur. J. Neurosci. 25, 460–470 (2007).
Berning, M., Boergens, K. M. & Helmstaedter, M. SegEM: Efficient Image Analysis for High-Resolution Connectomics. Neuron 87, 1193–1206 (2015).
Chklovskii, D. B., Vitaladevuni, S. & Scheffer, L. K. Semi-automated reconstruction of neural circuits using electron microscopy. Curr Opin Neurobiol 20, 667–675 (2010).
Jain, V., Seung, H. S. & Turaga, S. C. Machines that learn to segment images: a crucial technology for connectomics. Curr. Opin. Neurobiol. 20, 653–666 (2010).
Kaynig, V. et al. Large-scale automatic reconstruction of neuronal processes from electron microscopy images. Med. Image. Anal. 22, 77–88 (2015).
Burns, R. et al. in Proceedings of the 25th International Conference on Scientific and Statistical Database Management (eds Szalay, A., Budavari, T., Balazinska, M., Meliou, A. & Sacan, A.) (Scientific and Statistical Database Management, 2013).
Ahrens, M. B., Orger, M. B., Robson, D. N., Li, J. M. & Keller, P. J. Whole-brain functional imaging at cellular resolution using light-sheet microscopy. Nat Methods 10, 413–420 (2013).
Westerfield, M . The zebrafish book. A guide for the laboratory use of zebrafish (Danio rerio). 4 edn, (University of Oregon Press, 2000).
Li, J. et al. Early development of functional spatial maps in the zebrafish olfactory bulb. J. Neurosci. 25, 5784–5795 (2005).
Saalfeld, S., Cardona, A., Hartenstein, V. & Tomancak, P. CATMAID: collaborative annotation toolkit for massive amounts of image data. Bioinformatics 25, 1984–1986 (2009).
Sommer, C., Straehle, C., Koethe, U. & Hamprecht, F. A. ilastik: Interactive learning and segmentation toolkit. 8th Internat. Symp. Biomed. Imaging (ISBI) Proceedings 230–233, doi:10.1109/ISBI.2011.5872394 (2011).
Pinching, A. J. & Powell, T. P. The neuropil of the glomeruli of the olfactory bulb. J. Cell. Sci. 9, 347–377 (1971).
Braubach, O. R., Fine, A. & Croll, R. P. Distribution and functional organization of glomeruli in the olfactory bulbs of zebrafish (Danio rerio). J Comp Neurol 520, 2317–2339 (2012).
Braubach, O. R. et al. Experience-dependent versus experience-independent postembryonic development of distinct groups of zebrafish olfactory glomeruli. J Neurosci 33, 6905–6916 (2013).
Miyasaka, N. et al. From the olfactory bulb to higher brain centers: genetic visualization of secondary olfactory pathways in zebrafish. J. Neurosci. 29, 4756–4767 (2009).
Miyasaka, N. et al. Olfactory projectome in the zebrafish forebrain revealed by genetic single-neuron labelling. Nat. Commun. 5, 3639 (2014).
Dynes, J. L. & Ngai, J. Pathfinding of olfactory neuron axons to stereotyped glomerular targets revealed by dynamic imaging in living zebrafish embryos. Neuron. 20, 1081–1091 (1998).
Yabuki, Y. et al. Olfactory receptor for prostaglandin F2alpha mediates male fish courtship behavior. Nat. Neurosci. 19, 897–904 (2016).
Friedrich, R. W. & Korsching, S. I. Chemotopic, combinatorial and noncombinatorial odorant representations in the olfactory bulb revealed using a voltage-sensitive axon tracer. J. Neurosci. 18, 9977–9988 (1998).
Friedrich, R. W. & Korsching, S. I. Combinatorial and chemotopic odorant coding in the zebrafish olfactory bulb visualized by optical imaging. Neuron 18, 737–752 (1997).
Yaksi, E., von Saint Paul, F., Niessing, J., Bundschuh, S. T. & Friedrich, R. W. Transformation of odor representations in target areas of the olfactory bulb. Nature Neurosci. 12, 474–482 (2009).
Yaksi, E., Judkewitz, B. & Friedrich, R. W. Topological reorganization of odor representations in the olfactory bulb. PLoS Biol. 5, e178 (2007).
Byrd, C. A. & Brunjes, P. C. Organization of the olfactory system in the adult zebrafish: histological, immunohistochemical, and quantitative analysis. J. Comp. Neurol. 358, 247–259 (1995).
Fuller, C. L., Yettaw, H. K. & Byrd, C. A. Mitral cells in the olfactory bulb of adult zebrafish (Danio rerio): Morphology and distribution. J. Comp. Neurol. 499, 218–230 (2006).
Bundschuh, S. T., Zhu, P., Zhang Schärer, Y.-P. & Friedrich, R. W. Dopaminergic modulation of mitral cells and odor responses in the zebrafish olfactory bulb. J. Neurosci. 32, 6830–6840 (2012).
Miyasaka, N. et al. Functional development of the olfactory system in zebrafish. Mech. Dev. 130, 336–346 (2013).
Walton, J. Lead asparate, an en bloc contrast stain particularly useful for ultrastructural enzymology. J. Histochem. Cytochem. 27, 1337–1342 (1979).
Data Citations
Wanner, A. A., Genoud, C., & Friedrich, R. W. NeuroData http://doi.org/10.7281/T1MS3QN7 (2016)
Wanner, A. A., Genoud, C., & Friedrich, R. W. Zenodo https://doi.org/10.5281/zenodo.58985 (2016)
Acknowledgements
We thank C. Bleck, B. Anderson and B. Titze for scientific input, W. Ong, L. Ong and R. Ong for help with tracer training and management and A. Baden, A. Eusman, K. Lillaney and J. Vogelstein for help with data ingestion on neurodata.io. The work was supported by the Novartis Research Foundation, the Human Frontiers Science Program (HFSP) and the Swiss National Science Foundation (SNF).
Author information
Authors and Affiliations
Contributions
A.A.W. participated in all tasks. He acquired and processed image data, developed, supervised and participated in neuron tracing and error correction procedures, created PyKNOSSOS, analyzed data, and participated in writing the manuscript. C.G. participated in sample preparation, methods development, and image acquisition. R.W.F. participated in data analysis and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Part of the results disclosed herein have been included in European patent application EP14736451 and US patent application US14897514. A.A.W. is the founder and owner of ariadne-service.
ISA-Tab metadata
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0 Metadata associated with this Data Descriptor is available at http://www.nature.com/sdata/ and is released under the CC0 waiver to maximize reuse.
About this article
Cite this article
Wanner, A., Genoud, C. & Friedrich, R. 3-dimensional electron microscopic imaging of the zebrafish olfactory bulb and dense reconstruction of neurons. Sci Data 3, 160100 (2016). https://doi.org/10.1038/sdata.2016.100
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/sdata.2016.100
This article is cited by
-
RoboEM: automated 3D flight tracing for synaptic-resolution connectomics
Nature Methods (2024)
-
Volume EM: a quiet revolution takes shape
Nature Methods (2023)
-
Landmark-based retrieval of inflamed skin vessels enabled by 3D correlative intravital light and volume electron microscopy
Histochemistry and Cell Biology (2022)
-
The glomerular network of the zebrafish olfactory bulb
Cell and Tissue Research (2021)
-
Serial blockface SEM suggests that stem cells may participate in adult notochord growth in an invertebrate chordate, the Bahamas lancelet
EvoDevo (2020)