Abstract
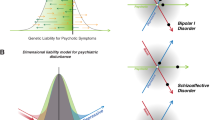
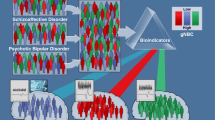
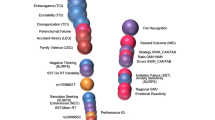
Every common psychiatric syndrome is genetically, neurobiologically and clinically heterogeneous. This heterogeneity may reflect the dimensional variability of a single illness with common etiology, varying in severity. Alternatively, as in many areas of medicine, it may reflect different discrete types of illness leading to similar alterations of mood or behavior. Resolving uncertainty as to the nature of heterogeneity in psychiatric illness is crucial for advancing the development of novel therapeutics and improving the precision with which pharmacological interventions are chosen for specific patients. Recent work resolving illness heterogeneity has shown promise in identifying biologically distinct patient subgroups within common psychiatric syndromes. This progress offers hope for the longer-term aim of enhancing our understanding of biological alterations associated with psychiatric syndromes, so that drug development and pharmacological interventions can shift towards altering specific targeted biological processes, instead of working to change complex behavioral features that may represent the final common pathway of different biological illness mechanisms. Using approaches such as neuroimaging and peripheral immune markers, studies have identified subgroups with different patterns of biological features that provide translational targets for novel drug development programmes, and may, in the longer term, together with psychological and social perspectives, more closely link diagnostics and therapeutics in psychiatry. So far, multiple discrete subgroups have been identified, and these have been associated with clinical features. Work is now needed to improve the validity and reliability of biologically derived subtypes, and better characterize their clinical, developmental and psychosocial features. This is required to establish their clinical utility for predicting illness course and response to different therapies, and to determine how biologically distinct features of patient subgroups can guide the development of novel therapies targeting those alterations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$59.00 per year
only $4.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Chaste, P. et al. A genome-wide association study of autism using the Simons Simplex Collection: does reducing phenotypic heterogeneity in autism increase genetic homogeneity? Biol. Psychiatry 77, 775–784 (2015).
Kapur, S., Phillips, A. G. & Insel, T. R. Why has it taken so long for biological psychiatry to develop clinical tests and what to do about it? Mol. Psychiatry 17, 1174–1179 (2012).
Jablensky, A. Subtyping schizophrenia: implications for genetic research. Mol. Psychiatry 11, 815–836 (2006).
Jablensky, A. The diagnostic concept of schizophrenia: its history, evolution and future prospects. Dialogues Clin. Neurosci. 12, 271–287 (2010).
Sagvolden, T., Johansen, E. B., Aase, H. & Russell, V. A. A dynamic developmental theory of attention-deficit/hyperactivity disorder (ADHD) predominantly hyperactive/impulsive and combined subtypes. Behav. Brain Sci. 28, 397–419 (2005). Discussion 419–368.
Beijers, L. et al. Biomarker-based subtyping of depression and anxiety disorders using latent class analysis. A NESDA study. Psychol. Med. 49, 617–627 (2019).
Clementz, B. A. et al. Psychosis biotypes: replication and validation from the B-SNIP Consortium. Schizophr. Bull. 48, 56–68 (2022).
Hermens, D. F., Lagopoulos, J., Naismith, S. L., Tobias-Webb, J. & Hickie, I. B. Distinct neurometabolic profiles are evident in the anterior cingulate of young people with major psychiatric disorders. Transl. Psychiatry 2, e110 (2012).
Lizano, P. et al. Multivariate relationships between peripheral inflammatory marker subtypes and cognitive and brain structural measures in psychosis. Mol. Psychiatry 26, 3430–3443 (2021).
Stefanik, L. et al. Brain-behavior participant similarity networks among youth and emerging adults with schizophrenia spectrum, autism spectrum, or bipolar disorder and matched controls. Neuropsychopharmacology 43, 1180–1188 (2018).
Insel, T. R. & Cuthbert, B. N. Medicine. Brain disorders? Precisely. Science 348, 499–500 (2015).
Cuthbert, B. N. & Insel, T. R. Toward the future of psychiatric diagnosis: the seven pillars of RDoC. BMC Med. 11, 126 (2013).
Cuthbert, B. N. The RDoC framework: facilitating transition from ICD/DSM to dimensional approaches that integrate neuroscience and psychopathology. World Psychiatry 13, 28–35 (2014).
Normanno, N. & Cree, I. A. Genomics driven-oncology: challenges and perspectives. BMC Cancer 15, 141 (2015).
Kalia, M. Biomarkers for personalized oncology: recent advances and future challenges. Metabolism 64, S16–S21 (2015).
El Achkar, C. M., Olson, H. E., Poduri, A. & Pearl, P. L. The genetics of the epilepsies. Curr. Neurol. Neurosci. Rep. 15, 39 (2015).
Sweeney, J. A. et al. Mixture analysis of pursuit eye-tracking dysfunction in schizophrenia. Biol. Psychiatry 34, 331–340 (1993).
Kaur, M. et al. Mismatch negativity/P3a complex in young people with psychiatric disorders: a cluster analysis. PLoS ONE 7, e51871 (2012).
Zhang, W. et al. Discrete patterns of cortical thickness in youth with bipolar disorder differentially predict treatment response to quetiapine but not lithium. Neuropsychopharmacology 43, 2256–2263 (2018).
Luo, C. et al. Subtypes of schizophrenia identified by multi-omic measures associated with dysregulated immune function. Mol. Psychiatry 26, 6926–6936 (2021).
Ivleva, E. I., Turkozer, H. B. & Sweeney, J. A. Imaging-based subtyping for psychiatric syndromes. Neuroimaging Clin. N. Am. 30, 35–44 (2020).
Bishop, J. R., Zhang, L. & Lizano, P. Inflammation subtypes and translating inflammation-related genetic findings in schizophrenia and related psychoses: a perspective on pathways for treatment stratification and novel therapies. Harv. Rev. Psychiatry 30, 59–70 (2022).
Beijers, L., Wardenaar, K. J., van Loo, H. M. & Schoevers, R. A. Data-driven biological subtypes of depression: systematic review of biological approaches to depression subtyping. Mol. Psychiatry 24, 888–900 (2019).
Sahin, M. et al. Discovering translational biomarkers in neurodevelopmental disorders. Nat. Rev. Drug Discov. 18, 235–236 (2019).
Clementz, B. A. et al. Identification of distinct psychosis biotypes using brain-based biomarkers. Am. J. Psychiatry 173, 373–384 (2016).
McGinty, E. E. & Eisenberg, M. D. Mental health treatment gap—the implementation problem as a research problem. JAMA Psychiatry 79, 746–747 (2022).
Paykel, E. S. Classification of depressed patients: a cluster analysis derived grouping. Br. J. Psychiatry 118, 275–288 (1971).
Goldstein, G. Neuropsychological heterogeneity in schizophrenia: a consideration of abstraction and problem-solving abilities. Arch. Clin. Neuropsychol. 5, 251–264 (1990).
Heinrichs, R. W. & Awad, A. G. Neurocognitive subtypes of chronic schizophrenia. Schizophr. Res. 9, 49–58 (1993).
Heinrichs, R. W., Ruttan, L., Zakzanis, K. K. & Case, D. Parsing schizophrenia with neurocognitive tests: evidence of stability and validity. Brain Cogn. 35, 207–224 (1997).
Goldstein, G., Allen, D. N. & Seaton, B. E. A comparison of clustering solutions for cognitive heterogeneity in schizophrenia. J. Int. Neuropsychol. Soc. 4, 353–362 (1998).
van Hulst, B. M., de Zeeuw, P. & Durston, S. Distinct neuropsychological profiles within ADHD: a latent class analysis of cognitive control, reward sensitivity and timing. Psychol. Med. 45, 735–745 (2015).
Mostert, J. C. et al. Similar subgroups based on cognitive performance parse heterogeneity in adults with ADHD and healthy controls. J. Atten. Disord. 22, 281–292 (2018).
Fair, D. A., Bathula, D., Nikolas, M. A. & Nigg, J. T. Distinct neuropsychological subgroups in typically developing youth inform heterogeneity in children with ADHD. Proc. Natl Acad. Sci. USA 109, 6769–6774 (2012).
Dawes, S. E., Jeste, D. V. & Palmer, B. W. Cognitive profiles in persons with chronic schizophrenia. J. Clin. Exp. Neuropsychol. 33, 929–936 (2011).
Lewandowski, K. E., Sperry, S. H., Cohen, B. M. & Ongür, D. Cognitive variability in psychotic disorders: a cross-diagnostic cluster analysis. Psychol. Med. 44, 3239–3248 (2014).
Geisler, D. et al. Brain structure and function correlates of cognitive subtypes in schizophrenia. Psychiatry Res. 234, 74–83 (2015).
Van Dam, N. T. et al. Data-driven phenotypic categorization for neurobiological analyses: beyond DSM-5 labels. Biol. Psychiatry 81, 484–494 (2017).
Wu, M. J. et al. Identification and individualized prediction of clinical phenotypes in bipolar disorders using neurocognitive data, neuroimaging scans and machine learning. Neuroimage 145, 254–264 (2017).
Weinberg, D. et al. Cognitive subtypes of schizophrenia characterized by differential brain volumetric reductions and cognitive decline. JAMA Psychiatry 73, 1251–1259 (2016).
Wessman, J. et al. Mixture model clustering of phenotype features reveals evidence for association of DTNBP1 to a specific subtype of schizophrenia. Biol. Psychiatry 66, 990–996 (2009).
Hallmayer, J. F. et al. Genetic evidence for a distinct subtype of schizophrenia characterized by pervasive cognitive deficit. Am. J. Hum. Genet. 77, 468–476 (2005).
Fanous, A. H. & Kendler, K. S. Genetic heterogeneity, modifier genes, and quantitative phenotypes in psychiatric illness: searching for a framework. Mol. Psychiatry 10, 6–13 (2005).
Fanous, A. H. & Kendler, K. S. Genetics of clinical features and subtypes of schizophrenia: a review of the recent literature. Curr. Psychiatry Rep. 10, 164–170 (2008).
Hochberger, W. C. et al. Unitary construct of generalized cognitive ability underlying BACS performance across psychotic disorders and in their first-degree relatives. Schizophr. Res. 170, 156–161 (2016).
Reilly, J. L. & Sweeney, J. A. Generalized and specific neurocognitive deficits in psychotic disorders: utility for evaluating pharmacological treatment effects and as intermediate phenotypes for gene discovery. Schizophr. Bull. 40, 516–522 (2014).
Voineskos, A. N., Jacobs, G. R. & Ameis, S. H. Neuroimaging heterogeneity in psychosis: neurobiological underpinnings and opportunities for prognostic and therapeutic innovation. Biol. Psychiatry 88, 95–102 (2020).
Chahal, R., Gotlib, I. H. & Guyer, A. E. Research review: brain network connectivity and the heterogeneity of depression in adolescence - a precision mental health perspective. J. Child Psychol. Psychiatry 61, 1282–1298 (2020).
Sun, H. et al. Two patterns of white matter abnormalities in medication-naive patients with first-episode schizophrenia revealed by diffusion tensor imaging and cluster analysis. JAMA Psychiatry 72, 678–686 (2015).
Dwyer, D. B. et al. Brain subtyping enhances the neuroanatomical discrimination of schizophrenia. Schizophr. Bull. 44, 1060–1069 (2018).
Wen, J. et al. Characterizing heterogeneity in neuroimaging, cognition, clinical symptoms and genetics among patients with late-life depression. JAMA Psychiatry 79, 464–474 (2022).
Xiao, Y. et al. Subtyping schizophrenia patients based on patterns of structural brain alterations. Schizophr. Bull. 48, 241–250 (2022).
Zhao, Q. et al. A subtype of institutionalized patients with schizophrenia characterized by pronounced subcortical and cognitive deficits. Neuropsychopharmacology 47, 2024–2032 (2022).
Brodersen, K. H. et al. Dissecting psychiatric spectrum disorders by generative embedding. Neuroimage Clin. 4, 98–111 (2014).
Price, R. B. et al. Parsing heterogeneity in the brain connectivity of depressed and healthy adults during positive mood. Biol. Psychiatry 81, 347–357 (2017).
Hawco, C. et al. Separable and replicable neural strategies during social brain function in people with and without severe mental illness. Am. J. Psychiatry 176, 521–530 (2019).
Costa Dias, T. G. et al. Characterizing heterogeneity in children with and without ADHD based on reward system connectivity. Dev. Cogn. Neurosci. 11, 155–174 (2015).
Feder, S. et al. Sample heterogeneity in unipolar depression as assessed by functional connectivity analyses is dominated by general disease effects. J. Affect. Disord. 222, 79–87 (2017).
Gates, K. M., Molenaar, P. C., Iyer, S. P., Nigg, J. T. & Fair, D. A. Organizing heterogeneous samples using community detection of GIMME-derived resting state functional networks. PLoS ONE 9, e91322 (2014).
Liang, S. et al. Aberrant triple-network connectivity patterns discriminate biotypes of first-episode medication-naive schizophrenia in two large independent cohorts. Neuropsychopharmacology 46, 1502–1509 (2021).
Price, R. B., Gates, K., Kraynak, T. E., Thase, M. E. & Siegle, G. J. Data-driven subgroups in depression derived from directed functional connectivity paths at rest. Neuropsychopharmacology 42, 2623–2632 (2017).
Volkow, N. D., Wang, G. J., Fowler, J. S., Tomasi, D. & Telang, F. Addiction: beyond dopamine reward circuitry. Proc. Natl Acad. Sci. USA 108, 15037–15042 (2011).
Drysdale, A. T. et al. Resting-state connectivity biomarkers define neurophysiological subtypes of depression. Nat. Med. 23, 28–38 (2017).
Wang, Y. et al. Data-driven clustering differentiates subtypes of major depressive disorder with distinct brain connectivity and symptom features. Br. J. Psychiatry 219, 606–613 (2021).
Ferenczi, E. A. et al. Prefrontal cortical regulation of brainwide circuit dynamics and reward-related behavior. Science 351, aac9698 (2016).
Knutson, B., Bhanji, J. P., Cooney, R. E., Atlas, L. Y. & Gotlib, I. H. Neural responses to monetary incentives in major depression. Biol. Psychiatry 63, 686–692 (2008).
Graybiel, A. M., Aosaki, T., Flaherty, A. W. & Kimura, M. The basal ganglia and adaptive motor control. Science 265, 1826–1831 (1994).
Chand, G. B. et al. Two distinct neuroanatomical subtypes of schizophrenia revealed using machine learning. Brain 143, 1027–1038 (2020).
Pan, Y. et al. Morphological profiling of schizophrenia: cluster analysis of MRI-based cortical thickness data. Schizophr. Bull. 46, 623–632 (2020).
Sugihara, G. et al. Distinct patterns of cerebral cortical thinning in schizophrenia: a neuroimaging data-driven approach. Schizophr. Bull. 43, 900–906 (2017).
Honnorat, N., Dong, A., Meisenzahl-Lechner, E., Koutsouleris, N. & Davatzikos, C. Neuroanatomical heterogeneity of schizophrenia revealed by semi-supervised machine learning methods. Schizophr. Res. 214, 43–50 (2019).
Hermens, D. F. et al. Cluster analysis reveals abnormal hippocampal neurometabolic profiles in young people with mood disorders. Eur. Neuropsychopharmacol. 25, 836–845 (2015).
Tamminga, C. A. et al. Clinical phenotypes of psychosis in the Bipolar-Schizophrenia Network on Intermediate Phenotypes (B-SNIP). Am. J. Psychiatry 170, 1263–1274 (2013).
Ivleva, E. I. et al. Brain structure biomarkers in the psychosis biotypes: findings from the Bipolar-Schizophrenia Network for Intermediate Phenotypes. Biol. Psychiatry 82, 26–39 (2017).
Guimond, S. et al. A diagnosis and biotype comparison across the psychosis spectrum: investigating volume and shape amygdala-hippocampal differences from the B-SNIP study. Schizophr. Bull. 47, 1706–1717 (2021).
Kelly, S. et al. White matter microstructure across brain-based biotypes for psychosis—findings from the bipolar-schizophrenia network for intermediate phenotypes. Psychiatry Res. Neuroimaging 308, 111234 (2021).
Ji, L. et al. Characterizing functional regional homogeneity (ReHo) as a B-SNIP psychosis biomarker using traditional and machine learning approaches. Schizophr. Res. 215, 430–438 (2020).
Meda, S. A. et al. Examining functional resting-state connectivity in psychosis and its subgroups in the Bipolar-Schizophrenia Network on Intermediate Phenotypes cohort. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 1, 488–497 (2016).
Zhang, W. et al. Brain gray matter network organization in psychotic disorders. Neuropsychopharmacology 45, 666–674 (2020).
Ewen, J. B., Sweeney, J. A. & Potter, W. Z. Conceptual, regulatory and strategic imperatives in the early days of EEG-based biomarker validation for neurodevelopmental disabilities. Front. Integr. Neurosci. 13, 45 (2019).
Xiao, Y. et al. Altered cortical thickness related to clinical severity but not the untreated disease duration in schizophrenia. Schizophr. Bull. 41, 201–210 (2015).
Zhang, W. et al. Brain structural abnormalities in a group of never-medicated patients with long-term schizophrenia. Am. J. Psychiatry 172, 995–1003 (2015).
Cole, V. T., Apud, J. A., Weinberger, D. R. & Dickinson, D. Using latent class growth analysis to form trajectories of premorbid adjustment in schizophrenia. J. Abnorm. Psychol. 121, 388–395 (2012).
McGorry, P. & Nelson, B. Why we need a transdiagnostic staging approach to emerging psychopathology, early diagnosis and treatment. JAMA Psychiatry 73, 191–192 (2016).
Seaton, B. E., Goldstein, G. & Allen, D. N. Sources of heterogeneity in schizophrenia: the role of neuropsychological functioning. Neuropsychol. Rev. 11, 45–67 (2001).
Marquand, A. F., Wolfers, T., Mennes, M., Buitelaar, J. & Beckmann, C. F. Beyond lumping and splitting: a review of computational approaches for stratifying psychiatric disorders. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 1, 433–447 (2016).
Kokkeler, K. J. E. et al. Subtyping late-life depression according to inflammatory and metabolic dysregulation: a prospective study. Psychol. Med. 52, 515–525 (2022).
Chaudhury, D. et al. Rapid regulation of depression-related behaviours by control of midbrain dopamine neurons. Nature 493, 532–536 (2013).
Tye, K. M. et al. Amygdala circuitry mediating reversible and bidirectional control of anxiety. Nature 471, 358–362 (2011).
Krishnan, V. et al. Molecular adaptations underlying susceptibility and resistance to social defeat in brain reward regions. Cell 131, 391–404 (2007).
Yanagi, M. et al. Kv3.1-containing K+ channels are reduced in untreated schizophrenia and normalized with antipsychotic drugs. Mol. Psychiatry 19, 573–579 (2014).
Egerton, A. et al. Glutamate in schizophrenia: neurodevelopmental perspectives and drug development. Schizophr. Res. 223, 59–70 (2020).
Liu, Z. et al. Resolving heterogeneity in schizophrenia through a novel systems approach to brain structure: individualized structural covariance network analysis. Mol. Psychiatry 26, 7719–7731 (2021).
Demchenko, I., Tassone, V. K., Kennedy, S. H., Dunlop, K. & Bhat, V. Intrinsic connectivity networks of glutamate-mediated antidepressant response: a neuroimaging review. Front. Psychiatry 13, 864902 (2022).
Kamburov, A., Stelzl, U., Lehrach, H. & Herwig, R. The ConsensusPathDB interaction database: 2013 update. Nucleic Acids Res. 41, D793–D800 (2013).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Jeppesen, R. et al. Efficacy and safety of anti-inflammatory agents in treatment of psychotic disorders - a comprehensive systematic review and meta-analysis. Brain Behav. Immun. 90, 364–380 (2020).
Köhler-Forsberg, O. et al. Efficacy of anti-inflammatory treatment on major depressive disorder or depressive symptoms: meta-analysis of clinical trials. Acta Psychiatr. Scand. 139, 404–419 (2019).
Ehrenreich, H. et al. Improvement of cognitive functions in chronic schizophrenic patients by recombinant human erythropoietin. Mol. Psychiatry 12, 206–220 (2007).
Pape, K., Tamouza, R., Leboyer, M. & Zipp, F. Immunoneuropsychiatry—novel perspectives on brain disorders. Nat. Rev. Neurol. 15, 317–328 (2019).
Asslih, S., Damri, O. & Agam, G. Neuroinflammation as a common denominator of complex diseases (cancer, diabetes type 2 and neuropsychiatric disorders). Int. J. Mol. Sci. 22, 6138 (2021).
Meyer, J. H. et al. Neuroinflammation in psychiatric disorders: PET imaging and promising new targets. Lancet Psychiatry 7, 1064–1074 (2020).
Yuan, N., Chen, Y., Xia, Y., Dai, J. & Liu, C. Inflammation-related biomarkers in major psychiatric disorders: a cross-disorder assessment of reproducibility and specificity in 43 meta-analyses. Transl. Psychiatry 9, 233 (2019).
Fillman, S. G. et al. Increased inflammatory markers identified in the dorsolateral prefrontal cortex of individuals with schizophrenia. Mol. Psychiatry 18, 206–214 (2013).
Fillman, S. G., Sinclair, D., Fung, S. J., Webster, M. J. & Shannon Weickert, C. Markers of inflammation and stress distinguish subsets of individuals with schizophrenia and bipolar disorder. Transl. Psychiatry 4, e365 (2014).
Fillman, S. G. et al. Elevated peripheral cytokines characterize a subgroup of people with schizophrenia displaying poor verbal fluency and reduced Broca’s area volume. Mol. Psychiatry 21, 1090–1098 (2016).
Schwarz, E. et al. Identification of subgroups of schizophrenia patients with changes in either immune or growth factor and hormonal pathways. Schizophr. Bull. 40, 787–795 (2014).
Boerrigter, D. et al. Using blood cytokine measures to define high inflammatory biotype of schizophrenia and schizoaffective disorder. J. Neuroinflammation 14, 188 (2017).
Hoang, D. et al. Inflammatory subtypes in antipsychotic-naïve first-episode schizophrenia are associated with altered brain morphology and topological organization. Brain Behav. Immun. 100, 297–308 (2022).
McIntyre, R. S. et al. Efficacy of adjunctive Infliximab vs placebo in the treatment of adults with bipolar I/II depression: a randomized clinical trial. JAMA Psychiatry 76, 783–790 (2019).
Elad, D. et al. Improving the predictive potential of diffusion MRI in schizophrenia using normative models—towards subject-level classification. Hum. Brain Mapp. 42, 4658–4670 (2021).
Tokuda, T. et al. Identification of depression subtypes and relevant brain regions using a data-driven approach. Sci. Rep. 8, 14082 (2018).
Hein, A. M. et al. Sustained hippocampal IL-1β overexpression impairs contextual and spatial memory in transgenic mice. Brain Behav. Immun. 24, 243–253 (2010).
Gonzalez, P. V., Schiöth, H. B., Lasaga, M. & Scimonelli, T. N. Memory impairment induced by IL-1β is reversed by α-MSH through central melanocortin-4 receptors. Brain Behav. Immun. 23, 817–822 (2009).
Zhang, L. et al. Inflammation subtypes in psychosis and their relationships with genetic risk for psychiatric and cardiometabolic disorders. Brain Behav. Immun. Health 22, 100459 (2022).
Kindler, J. et al. Dysregulation of kynurenine metabolism is related to proinflammatory cytokines, attention, and prefrontal cortex volume in schizophrenia. Mol. Psychiatry 25, 2860–2872 (2020).
Schwarcz, R., Bruno, J. P., Muchowski, P. J. & Wu, H. Q. Kynurenines in the mammalian brain: when physiology meets pathology. Nat. Rev. Neurosci. 13, 465–477 (2012).
Jones, K. A. & Thomsen, C. The role of the innate immune system in psychiatric disorders. Mol. Cell Neurosci. 53, 52–62 (2013).
Barichello, T. et al. Exposure to perinatal infections and bipolar disorder: a systematic review. Curr. Mol. Med. 16, 106–118 (2016).
Girgis, R. R. et al. A randomized, double-blind, placebo-controlled clinical trial of tocilizumab, an interleukin-6 receptor antibody, for residual symptoms in schizophrenia. Neuropsychopharmacology 43, 1317–1323 (2018).
Raison, C. L. et al. A randomized controlled trial of the tumor necrosis factor antagonist infliximab for treatment-resistant depression: the role of baseline inflammatory biomarkers. JAMA Psychiatry 70, 31–41 (2013).
Kalman, B. Autoimmune encephalitides: a broadening field of treatable conditions. Neurologist 22, 1–13 (2017).
Bowen, E. F. W., Burgess, J. L., Granger, R., Kleinman, J. E. & Rhodes, C. H. DLPFC transcriptome defines two molecular subtypes of schizophrenia. Transl. Psychiatry 9, 147 (2019).
Yu, C., Arcos-Burgos, M., Licinio, J. & Wong, M. L. A latent genetic subtype of major depression identified by whole-exome genotyping data in a Mexican-American cohort. Transl. Psychiatry 7, e1134 (2017).
Arnedo, J. et al. Uncovering the hidden risk architecture of the schizophrenias: confirmation in three independent genome-wide association studies. Am. J. Psychiatry 172, 139–153 (2015).
Yin, L. et al. Leveraging genome-wide association and clinical data in revealing schizophrenia subgroups. J. Psychiatr. Res. 106, 106–117 (2018).
Zhang, J. P. et al. Schizophrenia polygenic risk score as a predictor of antipsychotic efficacy in first-episode psychosis. Am. J. Psychiatry 176, 21–28 (2019).
Pardiñas, A. F. et al. Interaction testing and polygenic risk scoring to estimate the association of common genetic variants with treatment resistance in schizophrenia. JAMA Psychiatry 79, 260–269 (2022).
Meijs, H. et al. A polygenic-informed approach to a predictive EEG signature empowers antidepressant treatment prediction: a proof-of-concept study. Eur. Neuropsychopharmacol. 62, 49–60 (2022).
Vita, A. et al. Treatment-resistant schizophrenia: genetic and neuroimaging correlates. Front. Pharmacol. 10, 402 (2019).
Eum, S., Lee, A. M. & Bishop, J. R. Pharmacogenetic tests for antipsychotic medications: clinical implications and considerations. Dialogues Clin. Neurosci. 18, 323–337 (2016).
Milosavljevic, F. et al. Association of CYP2C19 and CYP2D6 poor and intermediate metabolizer status with antidepressant and antipsychotic exposure: a systematic review and meta-analysis. JAMA Psychiatry 78, 270–280 (2021).
Dinga, R. et al. Evaluating the evidence for biotypes of depression: methodological replication and extension of. Neuroimage Clin. 22, 101796 (2019).
Cheng, Y. et al. Delineation of early and later adult onset depression by diffusion tensor imaging. PLoS ONE 9, e112307 (2014).
Hall, M. H. et al. Patterns of deficits in brain function in bipolar disorder and schizophrenia: a cluster analytic study. Psychiatry Res. 200, 272–280 (2012).
Kleinman, A. et al. Attention-based classification pattern, a research domain criteria framework, in youths with bipolar disorder and attention-deficit/hyperactivity disorder. Aust. N. Z. J. Psychiatry 49, 255–265 (2015).
Marek, S. et al. Reproducible brain-wide association studies require thousands of individuals. Nature 603, 654–660 (2022).
Lombardo, M. V., Lai, M. C. & Baron-Cohen, S. Big data approaches to decomposing heterogeneity across the autism spectrum. Mol. Psychiatry 24, 1435–1450 (2019).
Tamminga, C. A. et al. Biotyping in psychosis: using multiple computational approaches with one data set. Neuropsychopharmacology 46, 143–155 (2021).
Bijsterbosch, J. et al. Challenges and future directions for representations of functional brain organization. Nat. Neurosci. 23, 1484–1495 (2020).
Botvinik-Nezer, R. et al. Variability in the analysis of a single neuroimaging dataset by many teams. Nature 582, 84–88 (2020).
Gordon, E. M. et al. Precision functional mapping of individual human brains. Neuron 95, 791–807 (2017).
Wang, D. et al. Parcellating cortical functional networks in individuals. Nat. Neurosci. 18, 1853–1860 (2015).
Wolfers, T., Buitelaar, J. K., Beckmann, C. F., Franke, B. & Marquand, A. F. From estimating activation locality to predicting disorder: a review of pattern recognition for neuroimaging-based psychiatric diagnostics. Neurosci. Biobehav. Rev. 57, 328–349 (2015).
Eitel, F., Schulz, M. A., Seiler, M., Walter, H. & Ritter, K. Promises and pitfalls of deep neural networks in neuroimaging-based psychiatric research. Exp. Neurol. 339, 113608 (2021).
Kim, Y. K. & Na, K. S. Application of machine learning classification for structural brain MRI in mood disorders: critical review from a clinical perspective. Prog. Neuropsychopharmacol. Biol. Psychiatry 80, 71–80 (2018).
Zhang, L., Wang, M., Liu, M. & Zhang, D. A survey on deep learning for neuroimaging-based brain disorder analysis. Front. Neurosci. 14, 779 (2020).
Varol, E., Sotiras, A. & Davatzikos, C. HYDRA: revealing heterogeneity of imaging and genetic patterns through a multiple max-margin discriminative analysis framework. Neuroimage 145, 346–364 (2017).
Dong, A., Honnorat, N., Gaonkar, B. & Davatzikos, C. CHIMERA: clustering of heterogeneous disease effects via distribution matching of imaging patterns. IEEE Trans. Med. Imaging 35, 612–621 (2016).
Yao, Y. et al. Variational Bayesian inversion for hierarchical unsupervised generative embedding (HUGE). Neuroimage 179, 604–619 (2018).
Raman, S., Deserno, L., Schlagenhauf, F. & Stephan, K. E. A hierarchical model for integrating unsupervised generative embedding and empirical Bayes. J. Neurosci. Methods 269, 6–20 (2016).
Powers, A. R. III, Kelley, M. & Corlett, P. R. Hallucinations as top-down effects on perception. Biol. Psychiatry Cogn. Neurosci. Neuroimaging 1, 393–400 (2016).
Sheldon, A. D. et al. Perceptual pathways to hallucinogenesis. Schizophr. Res. 245, 77–89 (2022).
Gershon, E. S. & Guroff, J. J. Information from relatives. Diagnosis of affective disorders. Arch. Gen. Psychiatry 41, 173–180 (1984).
Holleran, L. et al. The relationship between white matter microstructure and general cognitive ability in patients with schizophrenia and healthy participants in the ENIGMA consortium. Am. J. Psychiatry 177, 537–547 (2020).
McPartland, J. C. et al. The Autism Biomarkers Consortium for Clinical Trials (ABC-CT): scientific context, study design and progress toward biomarker qualification. Front. Integr. Neurosci. 14, 16 (2020).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grants 82101998, 8212018014, 82071908, 81820108018 and 81621003), the National Key R&D Program of China (grants 2022YFC2009901 and 2022YFC2009900), Sichuan Science and Technology Program (grant 2021JDTD0002) and the Post-Doctor Research Project, West China Hospital, Sichuan University (grant 2020HXBH005). J.A.S. acknowledges support from the University of Cincinnati Schizophrenia Research Fund. S.L. acknowledges support from the Humboldt Foundation Friedrich Wilhelm Bessel Research Award and Chang Jiang Scholars (programme T2019069).
Author information
Authors and Affiliations
Contributions
J.A.S., Q.G. and S.L. conceived the topic of this Review. W.Z. and J.A.S. conceived the structure of the Review. W.Z. and J.A.S. wrote the manuscript, with input from J.R.B., Q.G. and S.L. All authors took part in extensive discussions to refine the arguments presented in this manuscript and gave approval of the final version to be published.
Corresponding authors
Ethics declarations
Competing interests
W.Z. and J.A.S. are consultants to VeraSci. J.R.B. has served as a consultant to OptumRx. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Mental Health thanks Albert R. Powers, Michael Moutoussis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, W., Sweeney, J.A., Bishop, J.R. et al. Biological subtyping of psychiatric syndromes as a pathway for advances in drug discovery and personalized medicine. Nat. Mental Health 1, 88–99 (2023). https://doi.org/10.1038/s44220-023-00019-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44220-023-00019-x
This article is cited by
-
Choroid plexus volume enlargement in first-episode antipsychotic-naïve schizophrenia
Schizophrenia (2024)
-
Classifying and clustering mood disorder patients using smartphone data from a feasibility study
npj Digital Medicine (2023)