Abstract
Histopathological studies have revealed key processes of atherosclerotic plaque thrombosis. However, the diversity and complexity of lesion types highlight the need for improved subphenotyping. Here, we analyzed the gene expression profiles of 654 advanced human carotid plaques. The unsupervised, transcriptome-driven clustering revealed five dominant plaque types. These plaque phenotypes were associated with clinical presentation and showed differences in cellular compositions. Validation in coronary segments showed that the molecular signature of these plaques was linked to coronary ischemia. One of the plaque types with the most severe clinical symptoms pointed to both inflammatory and fibrotic cell lineages. Furthermore, we did a preliminary analysis of potential circulating biomarkers that mark the different plaque phenotypes. In conclusion, the definition of the plaque at risk for a thrombotic event can be fine-tuned by in-depth transcriptomic-based phenotyping. These differential plaque phenotypes prove clinically relevant for both carotid and coronary artery plaques and point to distinct underlying biology of symptomatic lesions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Raw RNA-seq data from the Athero-Express cohort are not publicly available owing to research participant privacy and consent. Data can be accessed via DataverseNL at this address: https://doi.org/10.34894/D1MDKL. There are restrictions on use by commercial parties and on sharing openly based on (inter)national laws and regulations and written informed consent. Therefore, these data (and additional clinical data) are available only upon discussion and upon signing a data sharing agreement (see Terms of Access in DataverseNL) and within a specially designed environment provided by UMC Utrecht.
The processed Athero-Express RNA-seq dataset can be accessed using the PlaqView portal (https://www.plaqview.com/).
Raw and processed bulk RNA-seq data from the coronary artery tissues will be made available on the Gene Expression Omnibus as well as on https://www.plaqview.com/ upon publication of the primary manuscript describing these data. In the meantime, requests to access these data can be addressed to C.L.M. (clintm@virginia.edu).
Code availability
The core scripts used for the analysis can be found at https://github.com/CirculatoryHealth/PlaqueCluster.
References
Falk, E. Plaque rupture with severe pre-existing stenosis precipitating coronary thrombosis. Characteristics of coronary atherosclerotic plaques underlying fatal occlusive thrombi. Br. Heart J. 50, 127–134 (1983).
Davies, M. J. & Thomas, A. C. Plaque fissuring—the cause of acute myocardial infarction, sudden ischaemic death, and crescendo angina. Br. Heart J. 53, 363–373 (1985).
Franzén, O. et al. Cardiometabolic risk loci share downstream cis- and trans-gene regulation across tissues and diseases. Science 353, 827–830 (2016).
Hartman, R. J. G. et al. Sex-stratified gene regulatory networks reveal female key driver genes of atherosclerosis involved in smooth muscle cell phenotype switching. Circulation 143, 713–726 (2021).
Perisic, L. et al. Gene expression signatures, pathways and networks in carotid atherosclerosis. J. Intern. Med. 279, 293–308 (2016).
Shakhtshneider, E. V. et al. RNA-SEQ transcriptome analysis of stable and unstable atherosclerotic plaques. Atherosclerosis 315, e133 (2020).
Aure, M. R. et al. Integrative clustering reveals a novel split in the luminal A subtype of breast cancer with impact on outcome. Breast Cancer Res. 19, 44 (2017).
Zhao, L., Zhao, H. & Yan, H. Gene expression profiling of 1200 pancreatic ductal adenocarcinoma reveals novel subtypes. BMC Cancer 18, 603 (2018).
Lapointe, J. et al. Gene expression profiling identifies clinically relevant subtypes of prostate cancer. Proc. Natl Acad. Sci. USA 101, 811–816 (2004).
Bertucci, F. et al. Gene expression profiling identifies molecular subtypes of inflammatory breast cancer. Cancer Res. 65, 2170–2178 (2005).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
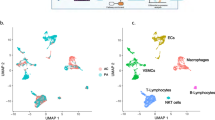
Depuydt, M. A. C. et al. Microanatomy of the human atherosclerotic plaque by single-cell transcriptomics. Circ. Res. 127, 1437–1455 (2020).
Costales, P. et al. K domain CR9 of low density lipoprotein (LDL) receptor-related protein 1 (LRP1) is critical for aggregated LDL-induced foam cell formation from human vascular smooth muscle cells. J. Biol. Chem. 290, 14852–14865 (2015).
Erdmann, J., Kessler, T., Munoz Venegas, L. & Schunkert, H. A decade of genome-wide association studies for coronary artery disease: the challenges ahead. Cardiovasc. Res. 114, 1241–1257 (2018).
Timmerman, N. et al. Pre-operative plasma extracellular vesicle proteins are associated with a high risk of long term secondary major cardiovascular events in patients undergoing carotid endarterectomy. Eur. J. Vasc. Endovasc. Surg. 62, 705–715 (2021).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Hartiala, J. A. et al. Genome-wide analysis identifies novel susceptibility loci for myocardial infarction. Eur. Heart J. 42, 919–933 (2021).
Turner, A. W. et al. Single-nucleus chromatin accessibility profiling highlights regulatory mechanisms of coronary artery disease risk. Nat. Genet. 54, 804–816 (2022).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Khoukaz, H. B. et al. Drug targeting of plasminogen activator inhibitor-1 inhibits metabolic dysfunction and atherosclerosis in a murine model of metabolic syndrome. Arterioscler. Thromb. Vasc. Biol. 40, 1479–1490 (2020).
Josefs, T. et al. Neutrophil extracellular traps promote macrophage inflammation and impair atherosclerosis resolution in diabetic mice. JCI Insight 5, e134796 (2020).
Schumski, A. et al. Endotoxinemia accelerates atherosclerosis through electrostatic charge-mediated monocyte adhesion. Circulation 143, 254–266 (2021).
Döring, Y., Libby, P. & Soehnlein, O. Neutrophil extracellular traps participate in cardiovascular diseases: recent experimental and clinical insights. Circ. Res. 126, 1228–1241 (2020).
Silvestre-Roig, C. et al. Externalized histone H4 orchestrates chronic inflammation by inducing lytic cell death. Nature 569, 236–240 (2019).
Gomez, I. et al. Neutrophil microvesicles drive atherosclerosis by delivering miR-155 to atheroprone endothelium. Nat. Commun. 11, 214 (2020).
Tomas, L. et al. Altered metabolism distinguishes high-risk from stable carotid atherosclerotic plaques. Eur. Heart J. 39, 2301–2310 (2018).
Newman, A. A. C. et al. Multiple cell types contribute to the atherosclerotic lesion fibrous cap by PDGFRβ and bioenergetic mechanisms. Nat. Metab. 3, 166–181 (2021).
Shi, J., Yang, Y., Cheng, A., Xu, G. & He, F. Metabolism of vascular smooth muscle cells in vascular diseases. Am. J. Physiol. Heart Circ. Physiol. 319, H613–H631 (2020).
De Bock, K. et al. Role of PFKFB3-driven glycolysis in vessel sprouting. Cell 154, 651–663 (2013).
Kang, H., Kim, H., Lee, S., Youn, H. & Youn, B. Role of metabolic reprogramming in epithelial–mesenchymal transition (EMT). Int. J. Mol. Sci. 20, 2042 (2019).
Cornelissen, A., Guo, L., Sakamoto, A., Virmani, R. & Finn, A. V. New insights into the role of iron in inflammation and atherosclerosis. EBioMedicine 47, 598–606 (2019).
Sakamoto, A. et al. CD163+ alternative macrophage inhibits atherosclerotic calcification via enhancement of hyaluronan production in vascular smooth muscle cells. Circulation 140, A13413 (2019).
Mega, J. L. et al. Genetic risk, coronary heart disease events, and the clinical benefit of statin therapy: an analysis of primary and secondary prevention trials. Lancet 385, 2264–2271 (2015).
Fateh-Moghadam, S. et al. Platelet degranulation is associated with progression of intima-media thickness of the common carotid artery in patients with diabetes mellitus type 2. Arterioscler. Thromb. Vasc. Biol. 25, 1299–1303 (2005).
Li, X. et al. The regulation of exosome-derived miRNA on heterogeneity of macrophages in atherosclerotic plaques. Front. Immunol. 11, 2175 (2020).
Bhatia, V. K. et al. Complement C1q reduces early atherosclerosis in low-density lipoprotein receptor-deficient mice. Am. J. Pathol. 170, 416–426 (2007).
Tomaniak, M. et al. Vulnerable plaques and patients: state-of-the-art. Eur. Heart J. 41, 2997–3004 (2020).
Lind, L. et al. Plasma protein profile of carotid artery atherosclerosis and atherosclerotic outcomes. Arterioscler. Thromb. Vasc. Biol. 41, 1777–1788 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Nurmohamed, N. S. et al. Targeted proteomics improves cardiovascular risk prediction in secondary prevention. Eur. Heart J. 43, 1569–1577 (2022).
Verhoeven, B. A. N. et al. Athero-express: differential atherosclerotic plaque expression of mRNA and protein in relation to cardiovascular events and patient characteristics. Rationale and design. Eur. J. Epidemiol. 19, 1127–1133 (2004).
Hellings, W. E. et al. Composition of carotid atherosclerotic plaque is associated with cardiovascular outcome. Circulation 121, 1941–1950 (2010).
Hellings, W. E. et al. Atherosclerotic plaque composition and occurrence of restenosis after carotid endarterectomy. JAMA 299, 547–554 (2008).
van Lammeren, G. W. et al. Time-dependent changes in atherosclerotic plaque composition in patients undergoing carotid surgery. Circulation 129, 2269–2276 (2014).
Laurie, C. C. et al. Quality control and quality assurance in genotypic data for genome-wide association studies. Genet. Epidemiol. 34, 591–602 (2010).
Hashimshony, T. et al. CEL-Seq2: sensitive highly-multiplexed single-cell RNA-Seq. Genome Biol. 17, 77 (2016).
Waltman, L. & van Eck, N. J. A smart local moving algorithm for large-scale modularity-based community detection. Eur. Phys. J. B 86, 471 (2013).
satijalab/seurat: R toolkit for single cell genomics. https://github.com/satijalab/seurat.
Yu, G. & He, Q.-Y. ReactomePA: an R/Bioconductor package for reactome pathway analysis and visualization. Mol. Biosyst. 12, 477–479 (2016).
Bennett, B. J. et al. Genetic architecture of atherosclerosis in mice: a systems genetics analysis of common inbred strains. PLoS Genet. 11, e1005711 (2015).
Dai, X. et al. Betaine supplementation attenuates S-adenosylhomocysteine hydrolase-deficiency-accelerated atherosclerosis in apolipoprotein E-deficient mice. Nutrients 14, 718 (2022).
Ikeda, J. et al. Radiation impacts early atherosclerosis by suppressing intimal LDL accumulation. Circ. Res. 128, 530–543 (2021).
Gullberg, M. et al. Cytokine detection by antibody-based proximity ligation. Proc. Natl Acad. Sci. USA 101, 8420–8424 (2004).
van der Laan, S. W. et al. Genetic susceptibility loci for cardiovascular disease and their impact on atherosclerotic plaques. Circ. Genom. Precis. Med. 11, e002115 (2018).
Delaneau, O., Marchini, J. & Zagury, J.-F. A linear complexity phasing method for thousands of genomes. Nat. Methods 9, 179–181 (2011).
Inouye, M. et al. Genomic risk prediction of coronary artery disease in 480,000 adults: implications for primary prevention. J. Am. Coll. Cardiol. 72, 1883–1893 (2018).
Choi, S. W. & O’Reilly, P. F. PRSice-2: polygenic risk score software for biobank-scale data. Gigascience 8, giz082 (2019).
Acknowledgements
This work was supported by the Dutch Heart Foundation (CVON2017-20: Generating the best evidence-based pharmaceutical targets and drugs for atherosclerosis (GENIUS II) to G.P., S.W.v.d.L. and M.P.J.d.W. and Targeting macrophages in atherosclerotic disease (T-MAD) to M.P.J.d.W.); Fondation Leducq (Transatlantic Network Grant PlaqOmics) to N.J.L., M.C., G.K.O., A.V.F., J.L.M.B., C.L.M. and G.P.; Transatlantic Network LEAN to M.P.J.d.W.; EU 755320 Taxinomisis grant to E.P., E.A., G.J.d.B., A.B. and G.P.; and the Dutch Research Council VENI grant (VI.VENI.212.196) to K.H.M.P. We acknowledge the European Research Area Network on Cardiovascular Diseases (grant no. 01KL1802 to F.W.A. and S.W.v.d.L.); the ERA-Endless Consortium (Dutch Heart Foundation, grant no. 2017/T099 to H.M.d.R. and G.P.); a European Research Council Consolidator Grant (grant no. 866478 UCARE to H.M.d.R.); and National Institutes of Health grant R01HL148239 to C.L.M. F.W.A. is supported by UCL Hospitals NIHR Biomedical Research Centre. The authors would like to thank the Utrecht Sequencing Facility for continuous support and patience.
Author information
Authors and Affiliations
Contributions
M.M. and A.B. analyzed and integrated the data. S.W.v.d.L. and J.M. provided PRS calculations. A.B. and S.W.v.d.L. performed patient selection, randomization and sample handling. G.J.d.B. performed CEA procedures. N.A.M.v.d.D., N.L. and E.M. tested library preparation strategies and processed coronary samples for sequencing. M.A.C.D., K.H.M.P., L.S., M.P.J.W. and J.P. provided, analyzed and interpreted single-cell sequencing data. K.C. and G.B.-B. performed deconvolution of bulk RNA-seq data. J.M., N.T., F.W. and M.C.V. recruited the patients, coordinated by D.P.V.d.K. D.P.v.d.K. and E.S.G.S. provided and coordinated Olink measurements. L.S. and S.W.v.d.L. performed MAGMA analysis. A.B., N.T., F.W., D.P.v.d.K., H.M.d.R., F.W.A., J.L.M.B. and S.W.v.d.L. participated in conceptualization and data interpretation and provided critical feedback on the article. E.D.B., R.J.G.H., E.P., E.A., H.S., G.K.O., C.M., J.C.K., A.V.F., R.V., N.J.L. and M.C. participated in data interpretation and provided critical feedback on the article. A.W.T., M.D.K., C.J.H., J.C.K., J.L.M.B. and C.L.M. recruited, processed and analyzed coronary samples. E.A., F.W.A., S.W.v.d.L., C.L.M., M.M. and G.P. provided funding. M.M., C.L.M. and G.P. participated in the conceptualization and supervision of the project and the finalization of the article. M.M. prepared the figures. M.M. and G.P. drafted the manuscript. All authors provided feedback on the research, analyses and article.
Corresponding authors
Ethics declarations
Competing interests
C.L.M. has received funding support from AstraZeneca for work unrelated to this study. G.P. received funding support from Roche to partly cover the generation of biomarker data. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Muredach Reilly, Dennis Wolf and Anders Malarstig for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Unsupervised clustering of plaques based on transcriptomics data.
a) PCA plot and b) tSNE projection of the 654 plaque samples based on RNA-seq dataset. The color indicates the cluster corresponding to the plaque type cluster from the SNN modularity optimization based clustering algorithm. c) Distribution of non-ribosomal protein-coding genes with annotated HGNC name; reads mapping to mitochondrial genes and mean Pearson correlation of samples per cluster
Extended Data Fig. 2 Robustness of the clustering.
a) UMAP projection of the 654 plaque samples based on RNA-seq dataset. The color indicates the cluster corresponding to the plaque type cluster from the SNN modularity optimization-based clustering algorithm after batch and hospital correction. b) Heatmap depicts relative gene expression levels of selected plaque type enriched genes in individual samples and plaque clusters. c) Correspondence of clusters derived from batch corrected and original dataset. Numbers indicate sample counts in the intersection of the corresponding clusters. d) UMAP projection of the 654 plaque samples based on RNA-seq dataset. The color indicates the cluster corresponding to the plaque type cluster from the SNN modularity optimization-based clustering algorithm after batch and hospital correction using Harmony integration. e) Heatmap depicts relative gene expression levels of selected plaque type enriched genes in individual samples and plaque clusters. f) Correspondence of clusters derived from batch corrected (using Harmony) and original dataset. Numbers indicate sample counts in the intersection of the corresponding clusters. g) UMAP projection of the 654 plaque samples based on RNA-seq dataset. The color indicates the frequency of the sample being assigned in other than the original cluster in permutation analysis. h) Distribution of the frequencies of the sample being assigned in other than the original cluster in permutation analysis.
Extended Data Fig. 3 Integrative analysis of coronary and carotid dataset.
a) Heatmaps depicting relative gene expression levels of selected plaque type enriched genes in individual samples and plaque clusters (upper panel - combined coronary and carotid dataset, lower panel – original carotid dataset). b) UMAP projection of the coronary and carotid sample based on RNA-seq data. The color indicates the cluster corresponding to the plaque type cluster from the SNN modularity optimization based clustering algorithm
Extended Data Fig. 4 Plaque transcriptomics.
a) Bioanalyzer profiles of total RNA isolated from 10 random samples of advanced atherosclerotic lesions. b) Distribution of sequencing reads between samples using four different library preparation strategies. (n = 12 per library preparation method, in the case of SMARTer n = 6) c) Percentage of sequenced reads mapped to annotated genes using four different library preparation strategies. (n = 12 per library preparation method, in the case of SMARTer n = 6) Boxplot’s top, middle, and bottom lines in c) and d) represent values at 25th, 50th, and 75th percentile. Whiskers extend up to 1.5 times the interquartile range from the top (bottom) of the box to the furthest data point within that distance. d) Number of annotated genes identified per sample with at least one mapped read. Samples with less than 9000 genes were excluded from the analysis e) Number of non-ribosomal protein-coding genes with annotated HGNC name per sample used in the analysis.
Supplementary information
Supplementary Table 1
Baseline table
Supplementary Table 2
Cluster-specific genes
Supplementary Table 3
Pathway enrichment analysis
Supplementary Table 4
Scaled gene expression values per cluster
Supplementary Table 5
Histological parameters
Supplementary Table 6
Circulating biomarkers
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mokry, M., Boltjes, A., Slenders, L. et al. Transcriptomic-based clustering of human atherosclerotic plaques identifies subgroups with different underlying biology and clinical presentation. Nat Cardiovasc Res 1, 1140–1155 (2022). https://doi.org/10.1038/s44161-022-00171-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-022-00171-0
This article is cited by
-
Uncovering atherosclerotic cardiovascular disease by PET imaging
Nature Reviews Cardiology (2024)
-
X chromosome inactivation skewing is common in advanced carotid atherosclerotic lesions in females and predicts secondary peripheral artery events
Biology of Sex Differences (2023)
-
Loss-of-function mutations in Dnmt3a and Tet2 lead to accelerated atherosclerosis and concordant macrophage phenotypes
Nature Cardiovascular Research (2023)
-
A new era in understanding atherosclerotic plaques
Nature Cardiovascular Research (2022)