Abstract
Pulmonary hypertension (PH) is a severe cardiopulmonary disease characterized by pulmonary vascular remodeling. Immunoglobulin E (IgE) is known to participate in aortic vascular remodeling, but whether IgE mediates pulmonary vascular disease is unknown. In the present study, we found serum IgE elevation in pulmonary arterial hypertension (PAH) patients, hypoxia-induced PH mice and monocrotaline-induced PH rats. Neutralizing IgE with an anti-IgE antibody was effective in preventing PH development in mice and rat models. The IgE receptor FcεRIα was also upregulated in PH lung tissues and Fcer1a deficiency prevented the development of PH. Single-cell RNA-sequencing revealed that FcεRIα was mostly expressed in mast cells (MCs) and MC-specific Fcer1a knockout protected against PH in mice. IgE-activated MCs produced interleukin (IL)-6 and IL-13, which subsequently promoted vascular muscularization. Clinically approved IgE antibody omalizumab alleviated the progression of established PH in rats. Using genetic and pharmacological approaches, we have demonstrated that blocking IgE–FcεRIα signaling may hold potential for PAH treatment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Sequencing data generated in the present study have been deposited at the Genome Sequence Archive (https://ngdc.cncb.ac.cn/gsa) and are publicly available at the date of publication. The accession nos. of scRNA-seq and RNA-seq data are CRA007001 and CRA007019, respectively. Additional data supporting the findings in the present study are included in the main article and associated files. Source data are provided with this paper.
References
Sakao, S., Voelkel, N. F. & Tatsumi, K. The vascular bed in COPD: pulmonary hypertension and pulmonary vascular alterations. Eur. Respir. Rev. 23, 350–355 (2014).
Humbert, M. et al. Pathology and pathobiology of pulmonary hypertension: state of the art and research perspectives. Eur. Respir. J. 53, 1801887 (2019).
Colvin, K. L. et al. Bronchus-associated lymphoid tissue in pulmonary hypertension produces pathologic autoantibodies. Am. J. Respir. Crit. Care Med. 188, 1126–1136 (2013).
Frid, M. G. et al. Immunoglobulin-driven complement activation regulates proinflammatory remodeling in pulmonary hypertension. Am. J. Respir. Crit. Care Med. 201, 224–239 (2020).
Terrier, B. et al. Identification of target antigens of antifibroblast antibodies in pulmonary arterial hypertension. Am. J. Respir. Crit. Care Med. 177, 1128–1134 (2008).
Arends, S. J. et al. Prevalence of anti-endothelial cell antibodies in idiopathic pulmonary arterial hypertension. Eur. Respir. J. 35, 923–925 (2010).
Dib, H. et al. Targets of anti-endothelial cell antibodies in pulmonary hypertension and scleroderma. Eur. Respir. J. 39, 1405–1414 (2012).
Kashiwada, M. et al. IL-4-induced transcription factor NFIL3/E4BP4 controls IgE class switching. Proc. Natl Acad. Sci. USA 107, 821–826 (2010).
Srikakulapu, P. et al. Artery tertiary lymphoid organs control multilayered territorialized atherosclerosis B-cell responses in aged ApoE–/– mice. Arterioscler. Thromb. Vasc. Biol. 36, 1174–1185 (2016).
Wang, J. et al. IgE stimulates human and mouse arterial cell apoptosis and cytokine expression and promotes atherogenesis in Apoe–/– mice. J. Clin. Invest. 121, 3564–3577 (2011).
Wang, J. et al. IgE actions on CD4+ T cells, mast cells, and macrophages participate in the pathogenesis of experimental abdominal aortic aneurysms. EMBO Mol. Med. 6, 952–969 (2014).
Wu, L. C. & Zarrin, A. A. The production and regulation of IgE by the immune system. Nat. Rev. Immunol. 14, 247–259 (2014).
Dahal, B. K. et al. Involvement of mast cells in monocrotaline-induced pulmonary hypertension in rats. Respir. Res. 12, 60 (2011).
Hoffmann, J. et al. Mast cells promote lung vascular remodelling in pulmonary hypertension. Eur. Respir. J. 37, 1400–1410 (2011).
Bartelds, B. et al. Mast cell inhibition improves pulmonary vascular remodeling in pulmonary hypertension. Chest 141, 651–660 (2012).
Shu, T., Xing, Y. & Wang, J. Autoimmunity in pulmonary arterial hypertension: evidence for local immunoglobulin production. Front. Cardiovasc. Med. 8, 680109 (2021).
Stearman, R. S. et al. Systems analysis of the human pulmonary arterial hypertension lung transcriptome. Am. J. Respir. Cell Mol. Biol. 60, 637–649 (2019).
Travaglini, K. J. et al. A molecular cell atlas of the human lung from single-cell RNA sequencing. Nature 587, 619–625 (2020).
Huang, J. et al. Progressive endothelial cell damage in an inflammatory model of pulmonary hypertension. Exp. Lung Res. 36, 57–66 (2010).
Daley, E. et al. Pulmonary arterial remodeling induced by a Th2 immune response. J. Exp. Med. 205, 361–372 (2008).
Chen, G. et al. Inhibition of CRTH2-mediated Th2 activation attenuates pulmonary hypertension in mice. J. Exp. Med. 215, 2175–2195 (2018).
Zhang, Y. & Michelakis, E. D. A phase-2 NIH-sponsored randomized clinical trial of rituximab in scleroderma-associated pulmonary arterial hypertension did not reach significance for its endpoints: end of story? Not so fast! Am. J. Respir. Crit. Care Med. 204, 123–125 (2021).
de Bourcy, C. F. A., et al., Dynamics of the human antibody repertoire after B cell depletion in systemic sclerosis. Sci. Immunol. 2, eaan8289 (2017).
Lambrecht, B. N. & Hammad, H. The immunology of asthma. Nat. Immunol. 16, 45–56 (2015).
Xu, Z. et al. The relationship of serum antigen-specific and total immunoglobulin E with adult cardiovascular diseases. Int. J. Med. Sci. 15, 1098–1104 (2018).
Andersson, C. K. et al. Novel site-specific mast cell subpopulations in the human lung. Thorax 64, 297–305 (2009).
Dahlin, J. S. & Hallgren, J. Mast cell progenitors: origin, development and migration to tissues. Mol. Immunol. 63, 9–17 (2015).
Platzer, B., Stout, M. & Fiebiger, E. Functions of dendritic-cell-bound IgE in allergy. Mol. Immunol. 68, 116–119 (2015).
Perros, F. et al. Dendritic cell recruitment in lesions of human and experimental pulmonary hypertension. Eur. Respir. J. 29, 462–468 (2007).
Montani, D. et al. C-kit-positive cells accumulate in remodeled vessels of idiopathic pulmonary arterial hypertension. Am. J. Respir. Crit. Care Med. 184, 116–123 (2011).
Chaouat, A. et al. Role for interleukin-6 in COPD-related pulmonary hypertension. Chest 136, 678–687 (2009).
Prins, K. W. et al. Interleukin-6 is independently associated with right ventricular function in pulmonary arterial hypertension. J. Heart Lung Transplant. 37, 376–384 (2018).
Breitling, S. et al. The mast cell-B cell axis in lung vascular remodeling and pulmonary hypertension. Am. J. Physiol. Lung Cell. Mol. Physiol. 312, L710–L721 (2017).
Cho, W. K. et al. IL-13 receptor alpha2-arginase 2 pathway mediates IL-13-induced pulmonary hypertension. Am. J. Physiol. Lung Cell. Mol. Physiol. 304, L112–L124 (2013).
Takagi, K. et al. IL-13 enhances mesenchymal transition of pulmonary artery endothelial cells via down-regulation of miR-424/503 in vitro. Cell Signal. 42, 270–280 (2018).
Stone, K. D., Prussin, C. & Metcalfe, D. D. IgE, mast cells, basophils, and eosinophils. J. Allergy Clin. Immunol. 125, S73–S80 (2010).
Siracusa, M. C. et al. Basophils and allergic inflammation. J. Allergy Clin. Immunol. 132, 789–801 (2013). quiz 788.
Ramirez, G. A. et al. Eosinophils from physiology to disease: a comprehensive review. BioMed. Res. Int. 2018, 9095275 (2018).
Galiè, N. et al. 2015 ESC/ERS Guidelines for the diagnosis and treatment of pulmonary hypertension: The Joint Task Force for the Diagnosis and Treatment of Pulmonary Hypertension of the European Society of Cardiology (ESC) and the European Respiratory Society (ERS): Endorsed by: Association for European Paediatric and Congenital Cardiology (AEPC), International Society for Heart and Lung Transplantation (ISHLT). Eur. Respir. J. 46, 903–975 (2015).
Ge, W. et al. The role of immunoglobulin E and mast cells in hypertension. Cardiovasc. Res. https://doi.org/10.1093/cvr/cvac010 (2022).
Zhao, H. et al. Role of IgE-FcepsilonR1 in pathological cardiac remodeling and dysfunction. Circulation 143, 1014–1030 (2021).
Xie, T. et al. Single-cell deconvolution of fibroblast heterogeneity in mouse pulmonary fibrosis. Cell Rep. 22, 3625–3640 (2018).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Butler, A. et al. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Satija, R. et al. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Yu, G. et al. clusterProfiler: an R package for comparing biological themes among gene clusters. Omics J. Integrat. Biol. 16, 284–287 (2012).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563, 347–353 (2018).
Wang, L. et al. Targeting IL-17 attenuates hypoxia-induced pulmonary hypertension through downregulation of beta-catenin. Thorax 74, 564–578 (2019).
Kim, K. et al. Transcriptome analysis reveals nonfoamy rather than foamy plaque macrophages are proinflammatory in atherosclerotic murine models. Circ. Res. 123, 1127–1142 (2018).
Gallagher, M. P. et al. Detection of true IgE-expressing mouse B lineage cells. J. Vis. Exp. https://doi.org/10.3791/52264 (2014).
Tu, L. et al. Selective BMP-9 inhibition partially protects against experimental pulmonary hypertension. Circ. Res. 124, 846–855 (2019).
Kikuchi, N. et al. Selenoprotein P promotes the development of pulmonary arterial hypertension: possible novel therapeutic target. Circulation 138, 600–623 (2018).
Kuklina, E. M., Nekrasova, I. V. & Valieva, Y. V. Involvement of semaphorin (Sema4D) in T-dependent activation of B cells. Bull. Exp. Biol. Med. 163, 447–450 (2017).
Zhu, J. et al. GATA-3 promotes Th2 responses through three different mechanisms: induction of Th2 cytokine production, selective growth of Th2 cells and inhibition of Th1 cell-specific factors. Cell Res. 16, 3–10 (2006).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
This project was supported by grants from the National Natural Science Foundation of China (grant no. 82000063 to Y.L.), the Chinese Academy of Medical Sciences Innovation Fund for Medical Sciences (grant no. 2021-I2M-1-049 to J.W.), the Non-pit Central Research Institute Fund of Chinese Academy of Medical Sciences (grant nos. 2018JB31001 and 2019JB310001 to C.W.) and State Key Laboratory Special Fund (grant no. 2060204). We thank C. Yan (Department of Medicine, University of Rochester) for advice.
Author information
Authors and Affiliations
Contributions
T.S. carried out investigations, methodology and visualization, and wrote the original draft. Y.L. carried out investigations, methodology, visualization and formal analysis. Y.Z. carried out investigations, created software, and did a formal analysis, data curation and visualization. Z.Z. provided resources, did a formal analysis and administered the project. P.Y. supervised the writing, review and editing of the paper. J.L. carried out investigations, validation and visualization. Y.X. carried out investigations and writing (review and editing). X.S. carried out validation and visualization. B.L. and J.P. created software and curated data. X.N. and X.Q. carried out investigations and validation. C.X. provided resources and carried out a formal analysis. H.Y. and Q.C. provided resources. J.C. supervised the project and provided resources. Y.Y. supervised the project and wrote, reviewed and edited the paper. J.W. and C.W. conceived and supervised the project, wrote, reviewed and edited the paper, administered the project and acquired funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Wolfgang Kuebler and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of the hypoxia-induced PH mouse model and sample collection for scRNA-seq.
a. Schematic diagram for the hypoxia-induced PH mouse model in C57BL/6 mice and sample collection for scRNA-seq. b. RVSP of C57BL/6 mice before (H0w) and after 4 weeks of hypoxia (H4w). c. RV/LV + S of these mice. d. PA AT/ET of these mice. e. Representative image of H&E and α-SMA staining, scale bar=25μm. f. Quantification of wall thickness. g. Proportion of non-muscularized (N), partially muscularized (P), or full muscularized (F) pulmonary arterioles. All above quantitative results are shown as mean ± SEM, and difference between groups was evaluated by unpaired two-tailed t-test.
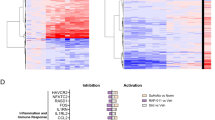
Extended Data Fig. 2 Bar plots of enriched GO pathways between pairs of time points.
Differentially enriched genes (DEGs) were calculated between two adjacent time points by Seurat and used to perform GO enrichment analysis employed by clusterProfiler package, whereas over representation analysis (ORA) was used for statistic test. P values was calculated by hypergeometric distribution and adjusted by FDR with Benjamini–Hochberg procedure controlled. The color in bar plots was coded by log-transformed P values.
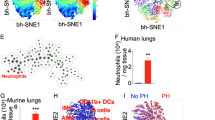
Extended Data Fig. 3 ScRNA-seq revealed Th2 response and T cell-B cell interaction in hypoxia-induced mice PH model.
a. Cell cluster identification by Cd45, Cd3d and Cd4 enrichment on t-SNE plot. b. GO pathway enrichment bar plot of DEGs in Cd4+ T cells during hypoxia exposure. c. Flow cytometry quantification of CD3+ CD4+ T cells. (n = 5 mice in 0-week hypoxia and n = 8 mice in 4-week hypoxia). d. Flow cytometry quantification of CD4+ IL4+ T cells. (n = 5 mice in 0-week hypoxia and n = 8 mice in 4-week hypoxia). e. Relative mRNA expression of Il4 and Gata3 in CD4+ T cells (n = 4). f. Dot plot of the interaction of gene pairs between B cells and Cd4+ T cells (Cd40lg_CD40, Cd28_Cd86) in four hypoxia groups. g. Dot plots of the expression of Cd86, Cd40 in B cell (left), and Cd28, Cd40lg in Cd4+ T cell (right) in four hypoxia groups. h. Relative mRNA expression of Cd28 in CD4+ T cells and relative mRNA expression of Cd40 and Cd86 in CD19+ B cells (n = 4). For b, p value was calculated by hypergeometric distribution and adjusted by FDR with Benjamini–Hochberg procedure controlled. For c-e and h, quantitative results are shown as mean ± SEM and compared by unpaired two-tailed t-test.
Extended Data Fig. 4 Proportion changes in B cells subtypes from control and hypoxic lung.
a. Flow cytometry gate strategy for germinal center B cells, memory B cells, plasma cells and plasma blasts in lung tissues. b. Relative Aicda mRNA expression in sorted germinal center B cells (GC B cells) from control and hypoxic lungs. c. The proportion of memory B cells from control and hypoxic lungs. d. The proportion of IgE+ cells in memory B cell population. e. The proportion of plasma cells from control and hypoxic lungs. f. The proportion of plasma blasts from control and hypoxic lungs. g. Immunochemistry staining of CD138 in bronchovascular space from control and hypoxic lung. h. The proportion of IgE+ cells in B cell population in spleen from control and hypoxic mice. i. The proportion of IgE+ cells in B cell population in lymph node from control and hypoxic mice. All values are presented as the mean ± SEM. Difference between groups were evaluated by unpaired two-tailed t-test. n = 5 for each group. PA, pulmonary artery; Br, bronchus.
Extended Data Fig. 5 The efficiency of anti-IgE-neutralizing antibody in mice and rats.
a. Serum IgE levels in isotype control antibody- and anti-IgE antibody-treated mice under control or hypoxia. n = 5 for control groups, n = 8 for hypoxia groups. b. The representative pressure tracing waveform of RVSP in isotype control or anti-IgE treated mice. c. Serum IgE levels in isotype control antibody- and anti-IgE antibody-treated rats after saline or MCT injection. n = 6 for saline groups, n = 6 for MCT rat treated with isotype control, n = 8 for MCT rats treated with anti-IgE. d. The representative pressure tracing waveform of PAP in isotype control or anti-IgE treated rats. All above quantitative results are shown as mean ± SEM. Difference between multiple groups was evaluated by two-way ANOVA with Bonferroni’s post hoc test.
Extended Data Fig. 6 Bar plot of FCER1A mRNA levels for individual patients’ data.
Data were analyzed from published genome-wide expression data (GSE117261) through GEO2R. Donor, n = 23; PAH, n = 54, logFc=0.338, adj. p value=0.00706 adjusted by Benjamini & Hochberg (False discovery rate).
Extended Data Fig. 7 Immunofluorescence staining of FcεRIα in human lung tissues.
a. Representative images showing staining of tryptase (green) and FcεRIα (red) in lung sections of control subjects (non-PAH) and PAH patients. Scale bar=50 μm. b. Representative images showing staining of α-SMA (green) and FcεRIα (red) in lung sections of control subjects and PAH patients. Scale bar=50 μm. c. Representative images showing staining of CC10 (green) and FcεRIα (red) in lung sections of control subjects and PAH patients. Scale bar=50 μm. The arrows indicate examples of positive-staining cells. PA, pulmonary artery; AE, airway epithelium. For a-c, the same results were observed in 4 individual samples each group.
Extended Data Fig. 8 Immune cell proportions in WT and MCKO mice at basal level.
a. The proportion of CD45+ cells from indicated lungs. b. The proportion of CD19+ cells from indicated lungs. c. The proportion of CD3+ cells from indicated lungs. d. The proportion of CD11b+ cells in CD45+ population from indicated lungs. e. The proportion of CD11c+ cells in CD45+ population from indicated lungs. f. Quantification of tryptase+ cell numbers in lung tissues from indicated mice. g. Serum histamine levels from indicated mice. h. Serum IgE levels in KO mice model. i. Serum IgE levels in MCKO mice model. All values are presented as the mean ± SEM. For a-e, n = 4 for each group. Differences between groups were evaluated by unpaired two-tailed t-test. For f and g, n = 4 for control group. n = 7 for hypoxia group, for h and i, n = 4 for control groups, n = 6 for hypoxia groups. Differences between groups were evaluated by two-way ANOVA with Bonferroni’s post hoc test.
Extended Data Fig. 9 mRNA expression of Il6 and Il13 in mouse lung tissues.
a. Relative Il6 mRNA expression from lung tissues in anti-IgE treated mouse model. b. Relative Il13 mRNA expression from lung tissues in anti-IgE treated mouse model. c. Relative Il6 mRNA expression from lung tissues in KO mouse model. d. Relative Il13 mRNA expression from lung tissues in KO mouse model. e. Relative Il6 mRNA expression from lung tissues in MCKO mouse model. f. Relative Il13 mRNA expression from lung tissues in MCKO mouse model. All values are presented as the mean ± SEM and compared by two-way ANOVA with Bonferroni’s post hoc test. n = 4 for control groups, n = 6 for hypoxia groups.
Extended Data Fig. 10 mRNA and protein levels of IL6 and IL13 in rat lung tissues.
a. Relative Il6 mRNA expression from lung tissues in anti-IgE treated rat model. b. Relative Il13 mRNA expression from lung tissues in anti-IgE treated rat model. c. IL6 protein levels from lung tissues in anti-IgE treated rat model. d. IL13 protein levels from lung tissues in anti-IgE treated rat model. n = 4 for control groups, n = 6 for MCT groups. All values are presented as the mean ± SEM and compared by two-way ANOVA with Bonferroni’s post hoc test.
Supplementary information
Supplementary Information
Supplementary Figure 1, Supplementary Table 1
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 3
Unprocessed Western Blots
Source Data Fig. 4
Statistical Source Data
Source Data Fig. 5
Statistical Source Data
Source Data Fig. 5
Unprocessed Western Blots
Source Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 5
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 8
Statistical Source Data
Source Data Extended Data Fig. 9
Statistical Source Data
Source Data Extended Data Fig. 10
Statistical Source Data
Rights and permissions
About this article
Cite this article
Shu, T., Liu, Y., Zhou, Y. et al. Inhibition of immunoglobulin E attenuates pulmonary hypertension. Nat Cardiovasc Res 1, 665–678 (2022). https://doi.org/10.1038/s44161-022-00095-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-022-00095-9
This article is cited by
-
Periodontitis exacerbates pulmonary hypertension by promoting IFNγ+ T cell infiltration in mice
International Journal of Oral Science (2024)
-
Inhibiting IgE in pulmonary hypertension
Nature Reviews Drug Discovery (2022)