Abstract
Stem and progenitor cells residing in the intestinal crypts drive the majority of colorectal cancers (CRCs), yet vascular contribution to this niche remains largely unexplored. Vascular endothelial growth factor A (VEGFA) is a key driver of physiological and tumor angiogenesis. Accordingly, current anti-angiogenic cancer therapies target the VEGFA pathway. Here we report that in CRC expansion of the stem/progenitor pool in intestinal crypts requires VEGFA-independent growth and remodeling of blood vessels. Epithelial transformation-induced expression of the endothelial peptide apelin, directs migration of distant venous endothelial cells toward progenitor niche vessels ensuring optimal perfusion. In the absence of apelin, loss of injury-inducible PROX1+ epithelial progenitors inhibited both incipient and advanced intestinal tumor growth. Our results establish fundamental principles for the reciprocal communication between vasculature and the intestinal progenitor niche and provide a mechanism for resistance to VEGFA-targeting drugs in CRCs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data generated or analyzed are included in the main article and associated files. Gene expression profiling data were deposited in a Super Series (GSE178118) including both the intestinal blood endothelial (GSE111783) and epithelial cell data (GSE178110). Source data are provided with this paper.
References
Gehart, H. & Clevers, H. Tales from the crypt: new insights into intestinal stem cells. Nat. Rev. Gastroenterol. Hepatol. 16, 19–34 (2019).
Zeuner, A., Todaro, M., Stassi, G. & De Maria, R. Colorectal cancer stem cells: from the crypt to the clinic. Cell Stem Cell 15, 692–705 (2014).
Lu, J. et al. Endothelial cells promote the colorectal cancer stem cell phenotype through a soluble form of Jagged-1. Cancer Cell 23, 171–185 (2013).
De Palma, M., Biziato, D. & Petrova, T. V. Microenvironmental regulation of tumour angiogenesis. Nat. Rev. Cancer 17, 457 (2017).
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015).
Lenz, H.-J. et al. Impact of consensus molecular subtype on survival in patients with metastatic colorectal cancer: results from CALGB/SWOG 80405 (Alliance). J. Clin. Oncol. 37, 1876–1885 (2019).
Kerr, R. S. et al. Adjuvant capecitabine plus bevacizumab versus capecitabine alone in patients with colorectal cancer (QUASAR 2): an open-label, randomised phase 3 trial. Lancet Oncol. 17, 1543–1557 (2016).
Kamba, T. et al. VEGF-dependent plasticity of fenestrated capillaries in the normal adult microvasculature. Am. J. Physiol. Heart. Circ. Physiol. 290, H560–H576 (2006).
Bernier-Latmani, J. & Petrova, T. V. Intestinal lymphatic vasculature: structure, mechanisms and functions. Nat. Rev. Gastroenterol. Hepatol. 14, 510–526 (2017).
de Sousa E Melo, F. et al. Methylation of cancer-stem-cell-associated Wnt target genes predicts poor prognosis in colorectal cancer patients. Cell Stem Cell 9, 476–485 (2011).
Ragusa, S. et al. PROX1 promotes metabolic adaptation and fuels outgrowth of Wnthigh metastatic colon cancer cells. Cell Rep. 8, 1957–1973 (2014).
Wiener, Z. et al. Prox1 promotes expansion of the colorectal cancer stem cell population to fuel tumor growth and ischemia resistance. Cell Reports 8, 1943–1956 (2014).
Petrova, T. V. et al. Transcription factor Prox1 induces colon cancer progression by promoting the transition from benign to highly dysplastic phenotype. Cancer Cell 13, 407–419 (2008).
Rocha, S. F. et al. Esm1 modulates endothelial tip cell behavior and vascular permeability by enhancing VEGF bioavailability. Circ. Res. 115, 581–590 (2014).
Yang, Y. et al. Anti-VEGF- and anti-VEGF receptor-induced vascular alteration in mouse healthy tissues. Proc. Natl Acad. Sci. USA 110, 12018–12023 (2013).
Korsisaari, N. et al. Inhibition of VEGF-A prevents the angiogenic switch and results in increased survival of Apc+/min mice. Proc. Natl Acad. Sci. USA 104, 10625–10630 (2007).
Carmeliet, P. & Jain, R. K. Molecular mechanisms and clinical applications of angiogenesis. Nature 473, 298–307 (2011).
Lundgren, O. Studies on blood flow distribution and countercurrent exchange in the small intestine. Acta Physiol. Scand. Suppl. 303, 1–42 (1967).
Witte, L. et al. Monoclonal antibodies targeting the VEGF receptor-2 (Flk1/KDR) as an anti-angiogenic therapeutic strategy. Cancer Metastasis Rev. 17, 155–161 (1998).
Sansom, O. J. et al. Loss of Apc in vivo immediately perturbs Wnt signaling, differentiation, and migration. Genes Dev. 18, 1385–1390 (2004).
Schepers, A. G. et al. Lineage tracing reveals Lgr5+ stem cell activity in mouse intestinal adenomas. Science 337, 730–735 (2012).
Corbett, T. H., Griswold, D. P., Roberts, B. J., Peckham, J. C. & Schabel, F. M. Tumor induction relationships in development of transplantable cancers of the colon in mice for chemotherapy assays, with a note on carcinogen structure. Cancer Res. 35, 2434–2439 (1975).
Jinushi, M. et al. Milk fat globule epidermal growth factor–8 blockade triggers tumor destruction through coordinated cell-autonomous and immune-mediated mechanisms. J. Exp. Med. 206, 1317–1326 (2009).
Dalerba, P. et al. Phenotypic characterization of human colorectal cancer stem cells. Proc. Natl Acad. Sci. USA 104, 10158–10163 (2007).
Ragusa, S. et al. Antiangiogenic immunotherapy suppresses desmoplastic and chemoresistant intestinal tumors in mice. J. Clin. Invest. 130, 1199–1216 (2020).
McKenzie, J. A. G. et al. Apelin is required for non-neovascular remodeling in the retina. Am. J. Pathol. 180, 399–409 (2012).
Langen, U. H. et al. Cell–matrix signals specify bone endothelial cells during developmental osteogenesis. Nat. Cell Biol. 19, 189–201 (2017).
Chen, Q. et al. Apelin+ endothelial niche cells control hematopoiesis and mediate vascular regeneration after myeloablative injury. Cell Stem Cell 25, 768–783.e6 (2019).
Kidoya, H. et al. The apelin/APJ system induces maturation of the tumor vasculature and improves the efficiency of immune therapy. Oncogene 31, 3254–3264 (2012).
Kidoya, H. et al. APJ regulates parallel alignment of arteries and veins in the skin. Dev. Cell 33, 247–259 (2015).
Kidoya, H. & Takakura, N. Biology of the apelin–APJ axis in vascular formation. J. Biochem. 152, 125–131 (2012).
Tian, X. et al. Subepicardial endothelial cells invade the embryonic ventricle wall to form coronary arteries. Cell Res. 23, 1075–1090 (2013).
Lee, H.-O. et al. Lineage-dependent gene expression programs influence the immune landscape of colorectal cancer. Nat. Genet. 52, 594–603 (2020).
Kuba, K. et al. Impaired heart contractility in apelin gene–deficient mice associated with aging and pressure overload. Circ. Res. 101, e32–e42 (2007).
Baluk, P., Morikawa, S., Haskell, A., Mancuso, M. & McDonald, D. M. Abnormalities of basement membrane on blood vessels and endothelial sprouts in tumors. Am. J. Pathol. 163, 1801–1815 (2003).
Kim, K. A. et al. Mitogenic influence of human R-spondin1 on the intestinal epithelium. Science 309, 1256–1259 (2005).
Ootani, A. et al. Sustained in vitro intestinal epithelial culture within a Wnt-dependent stem cell niche. Nat. Med. 15, 701–706 (2009).
Hua, G. et al. Crypt base columnar stem cells in small intestines of mice are radioresistant. Gastroenterology 143, 1266–1276 (2012).
Yan, K. S. et al. Intestinal enteroendocrine lineage cells possess homeostatic and injury-inducible stem cell activity. Cell Stem Cell 21, 78–90 (2017).
Nakanishi, Y. et al. Dclk1 distinguishes between tumor and normal stem cells in the intestine. Nat. Genet. 45, 98–103 (2013).
Westphalen, C. B. et al. Long-lived intestinal tuft cells serve as colon cancer-initiating cells. J. Clin. Invest. 124, 1283–1295 (2014).
Frentzas, S. et al. Vessel co-option mediates resistance to anti-angiogenic therapy in liver metastases. Nat. Med. 22, 1294–1302 (2016).
Mohyeldin, A., Garzón-Muvdi, T. & Quiñones-Hinojosa, A. Oxygen in stem cell biology: a critical component of the stem cell niche. Cell Stem Cell 7, 150–161 (2010).
Sancho, P., Barneda, D. & Heeschen, C. Hallmarks of cancer stem cell metabolism. Br. J. Cancer 114, 1305 (2016).
La Vecchia, S. & Sebastián, C. Metabolic pathways regulating colorectal cancer initiation and progression. Semin. Cell Dev. Biol. 98, 63–70 (2020).
Pitulescu, M. E. et al. Dll4 and Notch signalling couples sprouting angiogenesis and artery formation. Nat. Cell Biol. 19, 915–927 (2017).
Xu, C. et al. Arteries are formed by vein-derived endothelial tip cells. Nat. Commun. 5, 5758 (2014).
Lee, H. W. et al. The role of venous endothelial cells in developmental and pathologic Angiogenesis. Circulation 144, 1308–1322 (2021).
Kasai, A. et al. Retardation of retinal vascular development in apelin-deficient mice. Arterioscl. Throm. Vas. 28, 1717–1722 (2008).
Kasai, A. et al. Apelin is a crucial factor for hypoxia-induced retinal angiogenesis. Arterioscl. Throm. Vas. 30, 2182–2187 (2010).
Eyries, M. et al. Hypoxia-induced apelin expression regulates endothelial cell proliferation and regenerative angiogenesis. Circ. Res. 103, 432–440 (2008).
Liu, Q. et al. Genetic targeting of sprouting angiogenesis using Apln-CreER. Nat. Commun. 6, 6020–6020 (2015).
Mastrella, G. et al. Targeting APLN/APLNR improves antiangiogenic efficiency and blunts proinvasive side effects of VEGFA/VEGFR2 blockade in glioblastoma. Cancer Res. 79, 2298–2313 (2019).
Uribesalgo, I. et al. Apelin inhibition prevents resistance and metastasis associated with anti-angiogenic therapy. EMBO Mol. Med. 11, e9266 (2019).
Zuurbier, L. et al. Apelin: a putative novel predictive biomarker for bevacizumab response in colorectal cancer. Oncotarget 8, 42949–42961 (2017).
Jaiprasart, P., Dogra, S., Neelakantan, D., Devapatla, B. & Woo, S. Identification of signature genes associated with therapeutic resistance to anti-VEGF therapy. Oncotarget 11, 99–114 (2020).
Zhang, L., Takara, K., Yamakawa, D., Kidoya, H. & Takakura, N. Apelin as a marker for monitoring the tumor vessel normalization window during antiangiogenic therapy. Cancer Sci. 107, 36–44 (2016).
Lathen, C. et al. ERG–APLNR axis controls pulmonary venule endothelial proliferation in pulmonary veno-occlusive disease. Circulation 130, 1179–1191 (2014).
Strohbach, A. et al. The apelin receptor influences biomechanical and morphological properties of endothelial cells. J. Cell. Physiol. 233, 6250–6261 (2018).
Sharma, B. et al. Alternative progenitor cells compensate to rebuild the coronary vasculature in Elabela- and Apj-deficient hearts. Dev. Cell 42, 655–666.e653 (2017).
Shibata, H. et al. Rapid colorectal adenoma formation initiated by conditional targeting of the Apc gene. Science 278, 120–123 (1997).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Marino, S., Vooijs, M., van der Gulden, H., Jonkers, J. & Berns, A. Induction of medulloblastomas in p53-null mutant mice by somatic inactivation of Rb in the external granular layer cells of the cerebellum. Genes Dev. 14, 994–1004 (2000).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
El Marjou, F. et al. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193 (2004).
Moser, A. R., Pitot, H. C. & Dove, W. F. A dominant mutation that predisposes to multiple intestinal neoplasia in the mouse. Science 247, 322–324 (1990).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Hooper, A. T. et al. Engraftment and reconstitution of hematopoiesis is dependent on Vegfr2-mediated regeneration of sinusoidal endothelial cells. Cell Stem Cell 4, 263–274 (2009).
Sörensen, I., Adams, R. H. & Gossler, A. DLL1-mediated Notch activation regulates endothelial identity in mouse fetal arteries. Blood 113, 5680–5688 (2009).
Harvey, N. L. et al. Lymphatic vascular defects promoted by Prox1 haploinsufficiency cause adult-onset obesity. Nat. Genet. 37, 1072–1081 (2005).
Bernier-Latmani, J. & Petrova, T. V. High-resolution 3D analysis of mouse small-intestinal stroma. Nat. Protoc. 11, 1617–1629 (2016).
Yang, B. et al. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell 158, 945–958 (2014).
Sato, T. et al. Single Lgr5 stem cells build crypt–/villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Squadrito, MarioL. et al. miR-511-3p modulates genetic programs of tumor-associated macrophages. Cell Rep. 1, 141–154 (2012).
Carvalho, B. S. & Irizarry, R. A. A framework for oligonucleotide microarray preprocessing. Bioinformatics 26, 2363–2367 (2010).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Masiero, M. et al. A core human primary tumor angiogenesis signature identifies the endothelial orphan receptor ELTD1 as a key regulator of angiogenesis. Cancer Cell 24, 229–241 (2013).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Zudaire, E., Gambardella, L., Kurcz, C. & Vermeren, S. A computational tool for quantitative analysis of vascular networks. PLoS ONE 6, e27385 (2011).
Skog, M. et al. Expression and prognostic value of transcription factor PROX1 in colorectal cancer. Br. J. Cancer 105, 1346–1351 (2011).
Acknowledgements
We thank N. Piazzon (Institute of Pathology Biobank, CHUV and University of Lausanne, Lausanne, Switzerland) for providing human tumor samples, M. Lutolf for providing Rspo1-Fc, A. Zuba, M. Bruand and M. LeCoultre for participation in the initial stages of this work, C. Beauverd for mouse genotyping and colony maintenance and S. Robine and L. Claesson-Welsh for providing Villin-CreERT2 and Kdrfl/fl; Cdh5-CreERT2 mice. The UNIL Animal, Cellular Imaging, Genomic Technologies, Mouse Pathology and Flow Cytometry Facilities and EPFL Histology Core and Protein Production facilities are gratefully acknowledged. We apologize to colleagues whose work we could not reference due to size constraints.
This work was supported by grants from the Swiss Cancer League (KLS 3406-02-2016 and KFS-4895-08-2019), Foundation MEDIC, the Emma Muschamp Foundation, the Swiss National Science Foundation (31003A-156266 to T.V.P., 31003A-166161 to S.A.L.), the European Research Council (ERC EVOLVE-725051 to M.D.P.) and the Gabriela Kummer MD-PhD fellowship and Alfred and Anneliese Sutter-Stöttner private fellowships (to C.C.).
Author information
Authors and Affiliations
Contributions
J.B.L. and T.V.P. designed the study; J.B.L., C.C., S.M., S.R., I.D., D.B., F.R., S.N., P.A., M.L.S., S.S., S.D., A.G.L. and N.F. conducted experiments and acquired and analyzed data; S.A.L., R.B., P.V., B.Z., M.D.P., M.D. and C.S. provided reagents and samples; J.B.L. and T.V.P. wrote the manuscript. All authors discussed the results.
Corresponding author
Ethics declarations
Competing interests
T.V.P. receives research funding from Roche for an unrelated project. The authors declare no other competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Hyung Chun, Gou Young Koh and the other, anonymous, reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Reduced VEGFA signaling near Wnthigh progenitors in human CRC.
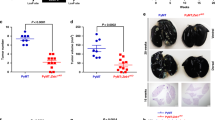
(a) Wnthigh and Wntlow human CRC tumors are in CMS2 and CMS1, respectively. Distribution of CMS assignment for Wnthigh and Wntlow human CRC tumors, n = 469. (b) Angiogenic gene expression is decreased in CMS2 human CRC tumors compared to CMS1. Experimentally-derived angiogenesis signature score based on Masiero et al. 103 classified by CMS assignment, n = 2,275. p values from one-ANOVA with post-hoc Tukey test are listed. (c) VEGF-dependent angiogenesis gene expression103 is highest in Wntlow CRC tumors, n = 469 tumors. For boxplots, minimum (Q1) and maximum (Q3) of the box are the 25th and 75th percentile, the center is the median, the whiskers minima and maxima highlight the Q1-1.5xIQR and Q3 + 1.5xIQR, where IQR is the interquartile range (Q3-Q1). Points outside of these regions are outliers. (d-h) VEGFA signaling is compartmentalized into low Wnt signaling areas of human CRC tumors. Serial paraffin section immunostaining of high (H) or low (L) areas of PROX1 staining within the same Wnthigh CRC tumors for (d) PROX1, (e) PECAM1 and (f) ESM1. (g-h) Quantification of percentage (g) vessel area (left, p = 0.0401) or (h) ESM1+ vessels (left, p = 0.0008) in Wnthigh-Progenitor (Pro) or Wnthigh-Differentiated (Diff) areas or luminal/bulk or invasive (L/B or Inv) areas of Wntlow CRC tumors (n = 11 Wnthigh tumors and n = 8 Wntlow tumors). Scale bars: 100 μm. All data are shown as mean ± SD. *P < 0.05, ***P < 0.001, 2-tailed unpaired Student’s t-test.
Extended Data Fig. 2 Compartmentalized VEGFA in the intestinal vascular unit.
(a) Filopodia are restricted to villus vessels in the small intestine. White, VEGFR2; arrowheads indicate perinuclear VEGFR2. Quantification of the percentage of filopodia observed in villi or crypts (p < 0.0001, n = 3 mice). (b-c) VEGFA expression and signaling are restricted to the villus. (b) Whole-mount immunostaining for VEGFA (red) in adult intestinal villi is restricted to villus blood vessels (green, PECAM1). (c) ESM1 staining (red) is restricted to villus vessels (green, podocalyxin). Cryosection immunostaining, DNA (blue). (d) Crypt progenitor epithelial cells are normoxic while differentiated epithelial cells are hypoxic. Small intestine and colon paraffin section immunostaining for hypoxia markers pimonidazole (Pimo) and HIF1α (red). Both hypoxia markers co-stain with the differentiated epithelial cell marker Keratin-20 (green, KRT20), DNA (blue). (e) Vessels associated with progenitor and differentiated colon epithelial cells are compartmentalized into normal and angiogenic vessels, respectively, as in the small intestine. Whole-mount immunostaining of colon blood capillaries (red, VEGFR2) with a side- and top-view. (f-h) Villus blood vessel branching is reduced in mice treated with DC101, while crypt vessel length and branching are unaffected. (f) Whole-mount immunostaining for intestinal vessels (red, VEGFR2) and quantification of the length (x100 μm) of crypt vessels and number of branchpoints per 100 μm vessel length in (g) villus (p = 0.0002) and (h) crypt vessels in mice treated with either IgG or DC101 (n = 6 mice). (i) Villus vessel filopodia are dependent on active VEGFA signaling. The number of villus vessel (black, VEGFR2/mTomato) filopodia (arrowheads) is significantly reduced in mice treated with DC101. Quantification of the number of filopodia per 100μm of villus vessel length (n = 4 IgG-treated mice and n = 5 DC101-treated mice; p < 0.0001). (j-k) VEGFA signaling blockade decreases intestinal ESM1 expression. (j) Whole-mount immunostaining for ESM1 (red) on adult intestinal villi blood vessels (green, PECAM1). (k) Total intestine Esm1 expression from adult control antibody- or DC101-treated C57BL/6 mice analyzed by RT–qPCR; Esm1 expression normalized by Cdh5 (n = 6 mice; p = 0.0224). (l) Scheme of intestinal vasculature and VEGFA-dependence. Villus blood vessel are VEGFA-dependent and associated with differentiated epithelial cells while crypt vessels are VEGFA-independent and associated with crypt cells, including stem and proliferating progenitor cells. VEGFA blockade reduces vascular density in the villi but has no impact on crypt vessels or epithelial cell proliferation. Scale bars: 50 μm: b, d, f; 20 μm: a, c, e, i, j. All data are shown as mean ± SD. *P < 0.05, ***P < 0.001, ****P < 0.0001, 2-tailed unpaired Student’s t-test. Images in b–e are representative of at least 3 independent experiments.
Extended Data Fig. 3 Stem/progenitor cells are VEGFA-independent and normoxic in mouse adenomas.
(a) Villus vessel length (green, PECAM1) is significantly increased in ApcΔIEC intestine while vessel branching is decreased. (b-c) Quantification of (b) total vessel length (x100μm; p = 0.0019) and (c) total branchpoints per 100μm of vessel length (p = 0.0003) in control or ApcΔIEC intestine (n = 6 mice). (d) Progenitor cell expansion maintains crypt/villus vessel compartmentalization. Whole-mount immunostaining in Apcmin/+; Lgr5-EGFP-CreERT2 adenomas. Blood vessel (red, VEGFR2) density and branching is higher in the luminal part of adenomas (Panel 1) compared to expanded epithelial crypts (Panel 2, Lgr5-GFP, green). Representative of 2 independent experiments. (e) VEGFA expression is not induced during progenitor expansion. qPCR for expression of Vegfa from total intestine of control and ApcΔIEC mice, n = 3 mice. (f-g) VEGFR2 blockade decreases ESM1 expression during progenitor expansion. (f) Whole-mount immunostaining of ESM1 (red) on villus blood vessels (green, PECAM1) in control antibody- or DC101-treated ApcΔIEC mice. (g) ESM1 (red) is reduced in DC101-treated Apcfl/fl; Lgr5-EGFP-CreERT2 adenoma blood vessels (green, MECA32). Paraffin section immunostaining; DNA, blue. (h) Quantification of percentage vessel area in adenomas of control antibody or DC101-treated Apcfl/fl; Lgr5-EGFP-CreERT2 mice, n = 4 mice. (i) Paraffin section immunostaining of GFP+ (green) stem cells in control antibody or DC101-treated Apcfl/fl; Lgr5-EGFP-CreERT2 adenomas (white outlines). (j–l) Quantification of control IgG or DC101-treated Apcfl/fl; Lgr5-EGFP-CreERT2 (j) adenoma area, (k) number of adenomas and (l) percentage tumor area GFP+, n = 4 tumors. (m) KRT20-negative Apcmin/+ adenoma crypts are Pimo-negative while differentiated KRT20+ lumen-adjacent epithelial cells (green) are Pimo+ (red). (n) Apcmin/+ adenoma crypt cells are HIF1α-negative (arrow) while lumen-adjacent epithelial cells are HIF1α+ (red). Scale bars: 200 μm: d, i; 50 μm: a, d (insets), f, g, m, n. All data are shown as mean ± SD, **P < 0.01, ***P < 0.0 1, 2-tailed unpaired Student’s t-test. Images in m and n are representative of 2 independent experiments.
Extended Data Fig. 4 Stem/progenitor cells are VEGFA-independent and normoxic in models of advanced CRC.
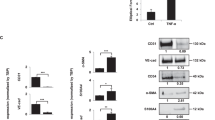
(a-b) MC38 tumor cells have low Wnt signaling. (a) qPCR for expression of Wnt target genes Axin2, Lgr5 and Prox1 in AKP organoid and MC38 tumors, n = 3 independent samples. (b) Paraffin section immunostaining for β-CATENIN in AKP organoid and MC38 tumors. DNA, blue. (c-d) VEGFR2 blockade inhibits MC38 tumor growth and proliferation. Quantification of subcutaneous MC38 tumor organoid (c) tumor volume (left, IgG vs DC101, p value: 0.0264, day 10; 0.0093, day 11; 0.0061, day 12; 0.0046, day 13; 0.0035, day 14) and weight (right, p = 0.0002; n = 7 tumors in IgG-treated mice and n = 8 tumors in DC101-treated mice). (d) Percentage of EdU+ MC38 tumor cells in control antibody- and DC101-treated mice (p < 0.0001, n = 6 tumors in IgG-treated mice and n = 7 tumors in DC101-treated mice, data from two independent experiments). (e-f) AKP organoid tumor vessels are less sensitive to VEGFR2 blockade than vessels from MC38 tumors. Quantification of tumor vessel area as a percentage of tumor area of control IgG antibody- or DC101-treated NSG mice with (e) AKP organoid tumors (p = 0.0056, n = 9 tumors) or (f) MC38 tumors (p < 0.0001, n = 14 tumors). (g-h) AKP organoid tumors have fewer vessels expressing the VEGFA signaling-dependent ESM1 than MC38 tumors. Quantification of percentage of ESM1+vessels from control antibody- or DC101-treated mice with (g) AKP organoid tumors (p = 0.0378, n = 6 tumors) or (h) MC38 tumors (p < 0.0001, n = 10 tumors). (i-j) AKP tumor blood endothelial cells (BECs) proliferation is resistant to VEGFR2 blockade. Quantification of % Ki67+ BECs from control antibody- or DC101-treated mice with (i) AKP organoid tumors (n = 9 tumors) or (j) MC38 tumors (p = 0.0039, n = 5 IgG-treated tumors and n = 7 DC101-treated tumors). (k) Endothelial-specific ablation of VEGFR2 does not alter AKP organoid tumor growth. Experimental plan for AKP organoid tumor subcutaneous injection into Kdrfl/fl and Kdrfl/fl; Cdh5-CreERT2 mice; tumor volume (mean ± SEM), dotted line indicates start of tamoxifen injections n = 6 tumors; final tumor weight (mean ± SD, n = 5 tumors in Kdrfl/fl and n = 4 tumors in Kdrfl/fl; Cdh5-CreERT2 mice). (l) VEGFR2 is efficiently depleted in Kdrfl/fl; Cdh5-CreERT2 mice without altering blood vessel density. Staining for PECAM1 (green), VEGFR2 (red) and DAPI (blue). Quantification of vessel density in AKP organoid tumors injected into Kdrfl/fl and Kdrfl/fl; Cdh5-CreERT2 mice (n = 5 tumors in Kdrfl/fl and n = 4 tumors in Kdrfl/fl; Cdh5-CreERT2 mice). (m) Endothelial-specific ablation of VEGFR2 arrests growth of MC38 tumors. Left, experimental plan for MC38 tumor subcutaneous injection into Kdrfl/fl and Kdrfl/fl; Cdh5-CreERT2 mice. Middle, tumor volume (p value: 0.0038, day 22; 0.0011, day 25; dotted line indicates start of tamoxifen injections). Right, final tumor weight (p = 0.0055, n = 4 tumors in Kdrfl/fl and n = 3 tumors in Kdrfl/fl; Cdh5-CreERT2 mice). (n) Increased endothelial VEGFA signaling in tumors with decreased Wnt signaling. Staining for ESM1 (green), PECAM1 (red) and DAPI (blue). Quantification of percentage of ESM1+ vessels in AP and APP organoid tumor vessels (p = 0.0034, n = 7 tumors). Scale bars: 50 μm: b (AKP organoids), l, n; 20 μm: b (MC38). All data (except k, middle) shown as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001, 2-tailed unpaired Student’s t-test.
Extended Data Fig. 5 Confirmation of Apln expression in villus endothelial cells.
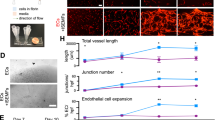
(a) Density plots showing the gating strategy for flow cytometric sorting of intestinal blood and lymphatic endothelial cell (BEC and LEC) sorting. Single, live, CD45neg, EpCAMneg cells were selected (upper plot). Among those, BEC were identified and collected as CD31+, gp38neg cells and LEC as CD31+, gp38+ cells. (b) Intestinal LEC and BEC quality control qPCR for Flt1 (p = 0.0059), Nrp1 (p = 0.0036), Lyve1 (p = 0.0051) and Prox1 (p < 0.0001, n = 3 sorted cell samples from individual mice). (c-d) Apelin expression is induced during progenitor cell expansion. qPCR for Apln expression in (c) sorted intestinal BECs (p = 0.0071, n = 3 sorted cell samples from individual mice) and (d) total intestine from control and ApcΔIEC mice (p = 0.0051, n = 4 WT and n = 3 ApcΔIEC mice). (e) Paraffin section immunostaining for blood vessels (red, PECAM1) and in situ hybridization (ISH) for Apln (green). Zoomed areas marked by 1 + 2. (f-h) The majority of apelin-expressing endothelial cells are in or near the villus venule. (f) Whole-mount immunostaining of GFP expression (green) and either all intestinal vessels (red, VEGFR2/mTomato) or MADCAM1+ vessels (blue) in Apln-CreERT2; mTmG mice 24 hours post 4-OHT injection. (g) Quantification of the percentage of GFP+ ECs that are in the MADCAM1+ villus venule or endothelial cells directly in contact with the villus venule (MADCAM1-adjacent) or crypt vessels in Apln-CreERT2; mTmG mice 24 hours post 4-OHT injection (n = 3 mice). (h) Quantification of percentage of GFP+ endothelial cells in the villus or crypt (n = 3 mice). (i) Aplnr is widely expressed in intestinal blood vessels. Paraffin section immunostaining/ISH for Aplnr (green) expressed on intestinal endothelial cells (red, PECAM1) of wild-type and Apcfl/fl; p53fl/fl Vil-CreERT2 intestine; blue, DAPI. (j) Apln expression is maintained after VEGFR2 signaling blockade. Experimental setup for Apln-CreERT2; mTmG lineage tracing combined with DC101 treatment. Whole-mount immunostaining for intestinal vessels (white, PECAM1) and GFP+ endothelial cells (green) after 24 hours of treatment with control or DC101 blocking antibodies. Red, ESM1; arrowheads, villus venule; arrows, capillary endothelial cells; asterisk, ESM1 staining comparison. Quantification of number of GFP+ endothelial cells in the villus or crypt, n = 4 IgG-treated and n = 5 DC101-treated mice). Scale bars: 50 μm: e, f, j; 20 μm: b (insets), i. All data are shown as mean ± SD. **P < 0.01, ***P < 0.001 2-tailed unpaired Student’s t-test. Images in e and i representative of 2 independent experiments.
Extended Data Fig. 6 Apelin maintains crypt venous capillary patency at steady state and during crypt expansion.
(a-b) Neither vessel density nor blood endothelial cell (BEC) proliferation are decreased in Apln-/- mice. (a) Quantification of crypt vessel density in control and Apln-/- mice (p = 0.0368, n = 3 WT mice and n = 4 Apln-/- mice). (b) Quantification of percentage of villus and crypt Ki67+ BECs in control and Apln-/- mice (n = 8 WT mice and n = 6 Apln-/- mice). (c) Quantification of the number of empty collagen sleeves / villus area of wild-type and Apln-/- mice (n = 4 WT mice and n = 3 Apln-/- mice). (d) Venous identity of regressing crypt vessels in Apln-/- mice. Whole-mount immunostaining for VEGFR2 (green), venous marker endomucin (red) and collagen IV (white). Arrowheads, empty collagen sleeves between endomucin+ vessels; arrow, regressing vessel. Quantification of number of regressed crypt vessels in Apln-/- mice sorted by endomucin status of each end of regressed vessel (n = 3 mice). (e) Quantification of crypt vessel density in ApcΔIEC and ApcΔIEC; Apln-/- mice (n = 3 ApcΔIEC and n = 4 ApcΔIEC; Apln-/- mice). (f) Quantification of percentage of villus and crypt Ki67+ BECs in ApcΔIEC and ApcΔIEC; Apln-/- mice (n = 4 mice). (g) Increased number of blind-ended crypt blood vessels in ApcΔIEC; Apln-/- mice. Whole-mount immunostaining of intestinal vessels (red, VEGFR2) from ApcΔIEC and ApcΔIEC; Apln-/- mice. Insets: blind-ended crypt vessels (arrowheads). (h) Increased empty collagen sleeves (green, collagen IV) in crypt vessels (red, VEGFR2) of ApcΔIEC; Apln-/- compared to ApcΔIEC mice. Quantification of number of empty collagen sleeves per crypt vessel length (p = 0.0002, n = 5 ApcΔIEC and n = 4 ApcΔIEC; Apln-/- mice). (i) Venous identity of regressing crypt vessels in ApcΔIEC; Apln-/- mice. Whole-mount immunostaining of crypt vessels (green, VEGFR2) and venous identity marker endomucin (red) from ApcΔIEC and ApcΔIEC; Apln-/- mice. Quantification of number of regressed crypt vessels in ApcΔIEC; Apln-/- mice sorted by endomucin status of each end of regressed vessel (n = 4 mice). Scale bars: 50 μm: g-i; 20 μm: d, g + h (insets). All data are shown as mean ± SD. *P < 0.05, ***P < 0.001, 2-tailed unpaired Student’s t-test.
Extended Data Fig. 7 Clonal analysis of Apln-CreERT2: mTmG lineage tracing.
(a) Clonal expansion of GFP+ cells in the villus, but not crypt vessels of Apln-CreERT2; mTmG mice. Analysis of venule and venule adjacent veinous GFP+ cell cluster size in the villus and crypt after 24 hr and 3 weeks of lineage tracing after 4-OHT injection, (mean ± SD, n = 4 Apln-CreERT2; mTmG 24 hr mice and n = 3 Apln-CreERT2; mTmG 3w mice). (b) Lack of sex-specific differences in crypt vessel phenotype in Apln-deficient mice. Representative whole-mount immunostainings for crypt vessels (white, VEGFR2) in male and female mice heterozygous or null for Apln. (c) Paraffin section immunostaining for crypt proliferation (green, Ki67) in mice treated with PBS or Rspo1-FC for 4 days; blue, DAPI. (d) Model for APLN-mediated control of crypt vessel patency. APLN+ endothelial cells migrate during steady state and expansion to the crypts in an APLN-dependent fashion. Scale bars: 50 μm: b; 20 μm: c. Images in a and c representative of 2 independent experiments.
Extended Data Fig. 8 Apelin promotes crypt perfusion and epithelial cell fitness.
(a) Villi are smaller in Apln-/- mice compared to controls. Whole-mount immunostaining for villus vessels (red, VEGFR2) in control and Apln-/- mice. Quantification of villus area between control and Apln-/- mice (p = 0.0221, n = 4 mice). (b) Crypt length is smaller in Apln-/- mice compared to controls. Paraffin section immunostaining; red, EdU; blue, DNA. Quantification of crypt length (p = 0.0004, n = 4 mice). (c) Decreased cypt epithelial cell proliferation in Apln-/- mice. Quantification of the number of Ki67+ crypt epithelial cells in control and Apln-/- mice (p = 0.0004, n = 4 WT mice and n = 3 Apln-/- mice). (d) Epithelial cell migration is slowed in Apln-/- mice. Paraffin immunostaining for EdU (red) after 72 h pulse; blue, DNA. Quantification of position of EdU+ epithelial cells 72 h after pulse as a percentage of villus height (p = 0.0028, n = 3 mice). (e) Increased number of EdUneg phospho-γH2A.X+ crypt epithelial cells in Apln-/- mice. Paraffin section immunostaining; green, EdU; red, phospho-γH2A.X; blue, DNA. Quantification of percentage of crypt EdUneg phospho-γH2A.X+ epithelial cells (p = 0.032, n = 3 mice). (f) Definition of crypt epithelial proliferation polarization index. EdU+ crypt intestinal epithelial cells were annotated as either on the left or right side and the percentage of each to a side was used to generate a score where 0 indicates exactly the same number of EdU+ epithelial cells on each side of the crypt and 1 indicates all EdU+ epithelial cells are on one side of the crypt. (g) Crypt EdU+ epithelial cells are polarized on one side of the crypt in Apln-/- mice. Polarization index of EdU+ crypt epithelial cells in control and Apln-/- mice (p < 0.0001, n = 4 mice). (h) Pulsed EdU+ IECs are polarized on one side of the villus in Apln-/- mice. Polarization index of EdU+ IECs after 72 h pulse in control and Apln-/- mice (p = 0.0214, n = 3 mice). (i) Decreased perfusion of crypt vessels in Apln-/- mice. Whole-mount immunostaining for VEGFR2 (green) after i.v. injection of PE-conjugated PECAM1 antibody (red). Arrowheads: non-perfused VEGFR2+PECAM1-PEneg vessels. (j) Density plots depicting the gating strategy for sorting CD44+ and CD44neg epithelial cells using flow cytometry. Single, live CD45neg CD31neg cells expressing EPCAM were selected (left plot) and then collected as either CD44+ or CD44neg epithelial cell subsets. (k) Apelin receptor is not expressed on intestinal epithelial cells (IECs). qPCR for Lgr5, Cdh5 and Aplnr expression in CD44+ IECs (progenitor cells), CD44neg IECs (differentiated IECs) and BECs (n = 2 CD44+ sorted IECs, n = 2 CD44neg sorted IECs and n = 5 sorted BECs from individual animals). (l, m) Defect of crypt epithelial progenitor survival in Apln-/- mice is cell extrinsic. Images and quantification of organoid area from either wild-type or Apln-/- mice after (l) first plating (p = 0.0013) or (m) re-plating (n = 4 organoid populations isolated from individual mice). Scale bars: 50 μm: a, b, d, i, l, m; 20 μm: e. All data are shown as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001, 2-tailed unpaired Student’s t-test.
Extended Data Fig. 9 Apelin maintains distinct populations of crypt epithelial cells.
(a, b) Secretory progenitor- and differentiated secretory-specific gene expression is decreased in Apln-/- crypt epithelial cells. (a) UMAP plots of mouse intestinal epithelial scRNAseq data55. Cell type-specific markers allowed cluster annotation (left). The top 100 cluster-specific genes were used as input for GSEA of differentially expressed genes between wild-type and Apln-/- CD44+ crypt epithelial cells. Clusters enriched for genes increased or decreased in Apln-/- crypt epithelial cells are marked in green and red, respectively (right). (b) GSEA of genes differentially regulated in Apln-/- crypt epithelial cells against top 100 genes of each cluster shown in (a). (c) Annotation of clusters used for GSEA analysis for genes up- and downregulated in wild-type vs Apln-/- crypt epithelial cells. (d) No increase in the number Lgr5-GFP stem cells (green, GFP, p = 0.0266) or their proliferation (red, Ki67, p = 0.0484) in Apln-/- crypts; blue, DAPI (n = 3 mice). (e, f) Decreased number of epithelial cells expressing tuft and enteroendocrine markers in Apln-/- crypts. Paraffin section immunostaining for (e) DCLK1 (green) or (f) CHGA (red) in wild-type and Apln-/- mice; blue, DAPI. Quantification of the number of (e) DCLK1+ (p = 0.0188, WT vs Apln-/- crypt) or (f) CHGA+ (p = 0.0055, WT vs Apln-/- crypt) in crypts and villi of wild-type and Apln-/- mice (n = 3 mice). Scale bars: 50 μm: e, f; 20 μm: d. *P < 0.05, **P < 0.01 2-tailed unpaired Student’s t-test.
Extended Data Fig. 10 Apelin maintains tumor progenitor cell fitness and proliferation.
(a-d) Apln deficiency reduces crypt expansion (a) Staining for EdU (green), Ki67 (red) and DNA (blue) in ApcΔIEC and ApcΔIEC; Apln-/- mice. (b-d) Quantification for the (b) crypt depth (p = 0.0007) and percentage of (c) Ki67+ (p = 0.0026) and (d) EdU+ crypt IECs (p = 0.0303, n = 4 mice). (e) Increased number of EdUneg phospho-γH2A.X+ crypt epithelial cells in ApcΔIEC; Apln-/- mice. Paraffin section immunostaining; green, EdU; red, phospho-γH2A.X; blue, DNA. Quantification of percentage of crypt EdUneg phospho-γH2A.X+ epithelial cells (p = 0.0004, n = 4 mice). (f) In situ hybridization for Lgr5 (green) in AKP organoid tumors implanted into wild-type or Apln-/- mice; DAPI, blue. Quantification for percentage of tumor area positive for Lgr5 (n = 6 tumors). (g) The majority of DCLK1+ cells in AKP organoid tumors also express PROX1. Quantification of percentage DCLK1+ cells that are also PROX1+ in either control or Apln-/- mice (n = 5 tumors in WT mice and n = 3 tumors in Apln-/- mice). (h) Apelin deficiency reduces vessel density in progenitor cell areas. Staining for progenitor marker PROX1 (green), PECAM1 (red) and DNA (blue) in AKP tumor grown in control and Apln-/- mice. Quantification of tumor vascularization as percentage PECAM1+ area in PROX1+ (p = 0.0082, n = 8 tumors in WT mice and n = 5 tumors in Apln-/- mice) and PROX1neg zones (n = 8 tumors in WT mice and n = 6 tumors in Apln-/- mice) of AKP tumor organoids grown in control and Apln-/- mice. Scale bars: 50 μm: a, f; 20 μm: e. All data are shown as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001, 2-tailed unpaired Student’s t-test.
Supplementary information
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 3
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 7
Statistical Source Data.
Source Data Extended Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 9
Statistical Source Data.
Source Data Extended Data Fig. 10
Statistical Source Data.
Rights and permissions
About this article
Cite this article
Bernier-Latmani, J., Cisarovsky, C., Mahfoud, S. et al. Apelin-driven endothelial cell migration sustains intestinal progenitor cells and tumor growth. Nat Cardiovasc Res 1, 476–490 (2022). https://doi.org/10.1038/s44161-022-00061-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44161-022-00061-5
This article is cited by
-
Adipo-oncology: adipocyte-derived factors govern engraftment, survival, and progression of metastatic cancers
Cell Communication and Signaling (2024)
-
Apelin-VEGF-C mRNA delivery as therapeutic for the treatment of secondary lymphedema
EMBO Molecular Medicine (2024)
-
Endothelial sensing of dietary metabolites supports barrier tissue homeostasis
Nature Cardiovascular Research (2023)
-
ADAMTS18+ villus tip telocytes maintain a polarized VEGFA signaling domain and fenestrations in nutrient-absorbing intestinal blood vessels
Nature Communications (2022)
-
Apelin drives maintenance and expansion of the vascular niche in intestinal crypts
Nature Cardiovascular Research (2022)