Abstract
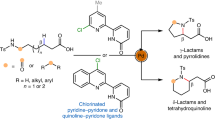
Enantioselectivity prediction in asymmetric catalysis has been a long-standing challenge in synthetic chemistry because of the high-dimensional nature of the structure–enantioselectivity relationship. A lack of understanding of the synthetic space results in laborious and time-consuming efforts in the discovery of asymmetric reactions, even if the same transformation has already been optimized on model substrates. Here we present a data-driven workflow to achieve a holistic enantioselectivity prediction of asymmetric pallada-electrocatalysed C–H activation by implementing transition state knowledge in machine learning. The vectorization of transition state knowledge allowed for an excellent description and extrapolation of the machine learning model, and enabled the quantitative evaluation of 846,720 possibilities. Model interpretation revealed the non-intuitive olefin effect on the enantioselectivity determination. Subsequent density functional theory calculations unravelled mechanistic knowledge that the rate-determining step depends on the olefin reactivity in the insertion step. Therefore, the olefin insertion step can be involved in the overall enantioselectivity determination. These results highlight the complementary features of knowledge-based machine learning with an interpretation-driven mechanistic study, which provides the opportunity to harness widely existing catalysis screening data and transition state models in molecular synthesis.

This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data related to ML details, experimental procedures, HPLC spectra and NMR spectra are available in the Supplementary Information. Source data are provided with this paper.
Code availability
Codes for target transformation, descriptor generation, model training, feature selection, feature ranking and synthetic space exploration are freely available at https://github.com/licheng-xu-echo/SyntheticSpacePrediction.
References
Noyori, R. Asymmetric catalysis: science and opportunities (Nobel Lecture). Angew. Chem. Int. Ed. 41, 2008–2022 (2002).
Trost, B. M. Asymmetric catalysis: an enabling science. Proc. Natl Acad. Sci. USA 101, 5348–5355 (2004).
Noyori, R. Synthesizing our future. Nat. Chem. 1, 5–6 (2009).
Taylor, M. S. & Jacobsen, E. N. Asymmetric catalysis in complex target synthesis. Proc. Natl Acad. Sci. USA 101, 5368–5373 (2004).
Woodard, S. S., Finn, M. G. & Sharpless, K. B. Mechanism of asymmetric epoxidation. 1. Kinetics. J. Am. Chem. Soc. 113, 106–113 (1991).
Cheong, P. H.-Y., Legault, C. Y., Um, J. M., Çelebi-Ölçüm, N. & Houk, K. N. Quantum mechanical investigations of organocatalysis: mechanisms, reactivities, and selectivities. Chem. Rev. 111, 5042–5137 (2011).
Bahmanyar, S., Houk, K. N., Martin, H. J. & List, B. Quantum mechanical predictions of the stereoselectivities of proline-catalyzed asymmetric intermolecular aldol reactions. J. Am. Chem. Soc. 125, 2475–2479 (2003).
Lam, Y.-h, Grayson, M. N., Holland, M. C., Simon, A. & Houk, K. N. Theory and modeling of asymmetric catalytic reactions. Acc. Chem. Res. 49, 750–762 (2016).
Knowles, R. R. & Jacobsen, E. N. Attractive noncovalent interactions in asymmetric catalysis: links between enzymes and small molecule catalysts. Proc. Natl Acad. Sci. USA 107, 20678–20685 (2010).
Neel, A. J., Milo, A., Sigman, M. S. & Toste, F. D. Enantiodivergent fluorination of allylic alcohols: data set design reveals structural interplay between achiral directing group and chiral Anion. J. Am. Chem. Soc. 138, 3863–3875 (2016).
Crawford, J. M., Kingston, C., Toste, F. D. & Sigman, M. S. Data science meets physical organic chemistry. Acc. Chem. Res. 54, 3136–3148 (2021).
Zahrt, A. F., Athavale, S. V. & Denmark, S. E. Quantitative structure–selectivity relationships in enantioselective catalysis: past, present, and future. Chem. Rev. 120, 1620–1689 (2020).
Oliveira, J. C. A. et al. When machine learning meets molecular synthesis. Trends Chem. 4, 863–885 (2022).
Mater, A. C. & Coote, M. L. Deep learning in chemistry. J. Chem. Inf. Model. 59, 2545–2559 (2019).
Tkatchenko, A. Machine learning for chemical discovery. Nat. Commun. 11, 4125 (2020).
Niemeyer, Z. L., Milo, A., Hickey, D. P. & Sigman, M. S. Parameterization of phosphine ligands reveals mechanistic pathways and predicts reaction outcomes. Nat. Chem. 8, 610–617 (2016).
Reid, J. P. & Sigman, M. S. Holistic prediction of enantioselectivity in asymmetric catalysis. Nature 571, 343–348 (2019).
Zahrt, A. F. et al. Prediction of higher-selectivity catalysts by computer-driven workflow and machine learning. Science 363, eaau5631 (2019).
Henle, J. J. et al. Development of a computer-guided workflow for catalyst optimization. Descriptor validation, subset selection, and training set analysis. J. Am. Chem. Soc. 142, 11578–11592 (2020).
Singh, S. et al. A unified machine-learning protocol for asymmetric catalysis as a proof of concept demonstration using asymmetric hydrogenation. Proc. Natl Acad. Sci. USA 117, 1339–1345 (2020).
Gallarati, S. et al. Reaction-based machine learning representations for predicting the enantioselectivity of organocatalysts. Chem. Sci. 12, 6879–6889 (2021).
Kutchukian, P. S. et al. Chemistry informer libraries: a chemoinformatics enabled approach to evaluate and advance synthetic methods. Chem. Sci. 7, 2604–2613 (2016).
Hase, F., Roch, L. M., Kreisbeck, C. & Aspuru-Guzik, A. Phoenics: a Bayesian optimizer for chemistry. ACS Cent. Sci. 4, 1134–1145 (2018).
Coley, C. W. Defining and exploring chemical spaces. Trends Chem. 3, 133–145 (2021).
Shields, B. J. et al. Bayesian reaction optimization as a tool for chemical synthesis. Nature 590, 89–96 (2021).
Dhawa, U. et al. Enantioselective pallada-electrocatalyzed C–H activation by transient directing groups: expedient access to helicenes. Angew. Chem. Int. Ed. 59, 13451–13457 (2020).
Moskal, M., Beker, W., Szymkuc, S. & Grzybowski, B. A. Scaffold-directed face selectivity machine-learned from vectors of non-covalent interactions. Angew. Chem. Int. Ed. 60, 15230–15235 (2021).
Jorner, K., Brinck, T., Norrby, P.-O. & Buttar, D. Machine learning meets mechanistic modelling for accurate prediction of experimental activation energies. Chem. Sci. 12, 1163–1175 (2021).
Zhang, S. Q. & Hong, X. Mechanism and selectivity control in Ni- and Pd-catalyzed cross-couplings involving carbon–oxygen bond activation. Acc. Chem. Res. 54, 2158–2171 (2021).
Tomberg, A., Johansson, M. J. & Norrby, P. O. A predictive tool for electrophilic aromatic substitutions using machine learning. J. Org. Chem. 84, 4695–4703 (2019).
Guan, Y. et al. Regio-selectivity prediction with a machine-learned reaction representation and on-the-fly quantum mechanical descriptors. Chem. Sci. 12, 2198–2208 (2020).
Li, X., Zhang, S. Q., Xu, L. C. & Hong, X. Predicting regioselectivity in radical C–H functionalization of heterocycles through machine learning. Angew. Chem. Int. Ed. 59, 13253–13259 (2020).
Gallegos, L. C., Luchini, G., St John, P. C., Kim, S. & Paton, R. S. Importance of engineered and learned molecular representations in predicting organic reactivity, selectivity, and chemical properties. Acc. Chem. Res. 54, 827–836 (2021).
Ramakrishnan, R., Dral, P. O., Rupp, M. & von Lilienfeld, O. A. Big data meets quantum chemistry approximations: the delta-machine learning approach. J. Chem. Theory Comput. 11, 2087–2096 (2015).
Xu, L. C. et al. Towards data-driven design of asymmetric hydrogenation of olefins: database and hierarchical learning. Angew. Chem. Int. Ed. 60, 22804–22811 (2021).
Martin, T. M. et al. Does rational selection of training and test sets improve the outcome of QSAR modeling? J. Chem. Inf. Model. 52, 2570–2578 (2012).
Ahneman, D. T., Estrada, J. G., Lin, S., Dreher, S. D. & Doyle, A. G. Predicting reaction performance in C–N cross-coupling using machine learning. Science 360, 186–190 (2018).
Rinehart, N. I., Zahrt, A. F., Henle, J. J. & Denmark, S. E. Dreams, false starts, dead ends, and redemption: a chronicle of the evolution of a chemoinformatic workflow for the optimization of enantioselective catalysts. Acc. Chem. Res. 54, 2041–2054 (2021).
Dewyer, A. L., Argüelles, A. J. & Zimmerman, P. M. Methods for exploring reaction space in molecular systems. WIREs Comput. Mol. Sci. 8, e1354 (2018).
Acknowledgements
Generous support by the National Natural Science Foundation of China (21873081 and 22122109, X. Hong; 22103070, S.-Q.Z.), the Starry Night Science Fund of Zhejiang University Shanghai Institute for Advanced Study (SN-ZJU-SIAS-006, X. Hong), Beijing National Laboratory for Molecular Sciences (BNLMS202102, X. Hong), CAS Youth Interdisciplinary Team (JCTD-2021-11, X. Hong), Fundamental Research Funds for the Central Universities (226-2022-00140 and 226-2022-00224, X. Hong), the Center of Chemistry for Frontier Technologies and Key Laboratory of Precise Synthesis of Functional Molecules of Zhejiang Province (PSFM 2021-01, X. Hong), the State Key Laboratory of Clean Energy Utilization (ZJUCEU2020007, X. Hong), China Scholarship Council (fellowship to X. Hou), the European Union (ERC advanced grant no. 101021358 conferred to L.A.) and the DFG (Gottfried-Wilhelm-Leibniz-Preis attributed to L.A. and SPP 2363) are gratefully acknowledged. Calculations and ML trainings were performed on the high‐performance computing system at the Department of Chemistry, Zhejiang University.
Author information
Authors and Affiliations
Contributions
X. Hong and L.A. conceived and supervised the project. X. Hong and S.-Q.Z. designed the workflow of the ML. L.-C.X. and S.-W.L. performed the ML training and analysed the training data. J.F. and X. Hou performed the experiments and analysed the experimental data. Y.-Y.L. and J.C.A.O. performed the DFT calculations for the physical organic descriptors and the mechanistic studies. X. Hong. and L.A. wrote the manuscript with input from all the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Synthesis thanks Tobias Gensch, Bartosz Grzybowski and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Peter Seavill, in collaboration with the Nature Synthesis team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Machine learning and experimental details, Supplementary Figs. 1–37 and Tables 1–17.
Supplementary Data 1
Collected data of asymmetric pallada-electrocatalysed C–H activation
Source data
Source Data Fig. 4
Data for the three regression diagrams of Fig. 4a.
Source Data Fig. 5
Importance scores for top-5 features.
Source Data Fig. 6
Data for the regression diagram of Fig. 6e.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, LC., Frey, J., Hou, X. et al. Enantioselectivity prediction of pallada-electrocatalysed C–H activation using transition state knowledge in machine learning. Nat. Synth 2, 321–330 (2023). https://doi.org/10.1038/s44160-022-00233-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s44160-022-00233-y
This article is cited by
-
Enabling late-stage drug diversification by high-throughput experimentation with geometric deep learning
Nature Chemistry (2024)
-
Data-driven design of new chiral carboxylic acid for construction of indoles with C-central and C–N axial chirality via cobalt catalysis
Nature Communications (2023)
-
Reaction performance prediction with an extrapolative and interpretable graph model based on chemical knowledge
Nature Communications (2023)