Abstract
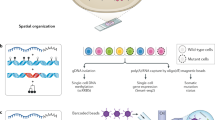
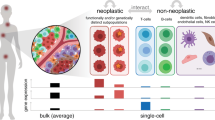
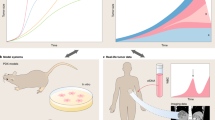
Individual cells within an otherwise genetically homogenous population constantly undergo fluctuations in their molecular state, giving rise to non-genetic heterogeneity. Such diversity is being increasingly implicated in cancer therapy resistance and metastasis. Identifying the origins of non-genetic heterogeneity is therefore crucial for making clinical breakthroughs. We discuss with examples how dynamical models and computational tools have provided critical multiscale insights into the nature and consequences of non-genetic heterogeneity in cancer. We demonstrate how mechanistic modeling has been pivotal in establishing key concepts underlying non-genetic diversity at various biological scales, from population dynamics to gene regulatory networks. We discuss advances in single-cell longitudinal profiling techniques to reveal patterns of non-genetic heterogeneity, highlighting the ongoing efforts and challenges in statistical frameworks to robustly interpret such multimodal datasets. Moving forward, we stress the need for data-driven statistical and mechanistically motivated dynamical frameworks to come together to develop predictive cancer models and inform therapeutic strategies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Welch, D. R. & Hurst, D. R. Defining the hallmarks of metastasis. Cancer Res. 79, 3011–3027 (2019).
Bell, C. C. & Gilan, O. Principles and mechanisms of non-genetic resistance in cancer. Br. J. Cancer 122, 465–472 (2020).
Ramirez, M. et al. Diverse drug-resistance mechanisms can emerge from drug-tolerant cancer persister cells. Nat. Commun. 7, 10690 (2016).
Marine, J.-C., Dawson, S.-J. & Dawson, M. A. Non-genetic mechanisms of therapeutic resistance in cancer. Nat. Rev. Cancer 20, 743–756 (2020).
Rambow, F. et al. Toward minimal residual disease-directed therapy in melanoma. Cell 174, 843–855.e19 (2018).
Sharma, S. V. et al. A chromatin-mediated reversible drug-tolerant state in cancer cell subpopulations. Cell 141, 69–80 (2010).
Gupta, P. B. et al. Stochastic state transitions give rise to phenotypic equilibrium in populations of cancer cells. Cell 146, 633–644 (2011).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Su, Y. et al. Single-cell analysis resolves the cell state transition and signaling dynamics associated with melanoma drug-induced resistance. Proc. Natl Acad. Sci. USA 114, 13679–13684 (2017).
Quinn, J. J. et al. Single-cell lineages reveal the rates, routes, and drivers of metastasis in cancer xenografts. Science 371, eabc1944 (2021).
Fennell, K. A. et al. Non-genetic determinants of malignant clonal fitness at single-cell resolution. Nature 601, 125–131 (2022).
Wouters, J. et al. Robust gene expression programs underlie recurrent cell states and phenotype switching in melanoma. Nat. Cell Biol. 22, 986–998 (2020).
Marin-Bejar, O. et al. Evolutionary predictability of genetic versus nongenetic resistance to anticancer drugs in melanoma. Cancer Cell 39, 1135–1149.e8 (2021).
Roesch, A. et al. Overcoming intrinsic multidrug resistance in melanoma by blocking the mitochondrial respiratory chain of slow-cycling JARID1B(high) cells. Cancer Cell 23, 811–825 (2013).
Goyal, Y. et al. Pre-determined diversity in resistant fates emerges from homogenous cells after anti-cancer drug treatment. Preprint at bioRxiv https://doi.org/10.1101/2021.12.08.471833 (2021).
Emert, B. L. et al. Variability within rare cell states enables multiple paths toward drug resistance. Nat. Biotechnol. 39, 865–876 (2021).
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Nevozhay, D., Adams, R. M., Van Itallie, E., Bennett, M. R. & Balázsi, G. Mapping the environmental fitness landscape of a synthetic gene circuit. PLoS Comput. Biol. 8, e1002480 (2012).
Mooney, S. M., Jolly, M. K., Levine, H. & Kulkarni, P. Phenotypic plasticity in prostate cancer: role of intrinsically disordered proteins. Asian J. Androl. 18, 704–710 (2016).
Raj, A. & van Oudenaarden, A. Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135, 216–226 (2008).
Schuh, L. et al. Gene networks with transcriptional bursting recapitulate rare transient coordinated high expression states in cancer. Cell Syst. 10, 363–378.e12 (2020).
Kim, J. & DeBerardinis, R. J. Mechanisms and implications of metabolic heterogeneity in cancer. Cell Metab. 30, 434–446 (2019).
Huh, D. & Paulsson, J. Non-genetic heterogeneity from stochastic partitioning at cell division. Nat. Genet. 43, 95–100 (2011).
Su, Y. et al. Phenotypic heterogeneity and evolution of melanoma cells associated with targeted therapy resistance. PLoS Comput. Biol. 15, e1007034 (2019).
Roux, J. et al. Fractional killing arises from cell-to-cell variability in overcoming a caspase activity threshold. Mol. Syst. Biol. 11, 803 (2015).
Huang, B. et al. Modeling the transitions between collective and solitary migration phenotypes in cancer metastasis. Sci. Rep. 5, 17379 (2015).
Huang, S., Eichler, G., Bar-Yam, Y. & Ingber, D. E. Cell fates as high-dimensional attractor states of a complex gene regulatory network. Phys. Rev. Lett. 94, 128701 (2005).
Li, Q. et al. Dynamics inside the cancer cell attractor reveal cell heterogeneity, limits of stability, and escape. Proc. Natl Acad. Sci. USA 113, 2672–2677 (2016).
Blake, W. J. et al. Phenotypic consequences of promoter-mediated transcriptional noise. Mol. Cell 24, 853–865 (2006).
Pillai, M. & Jolly, M. K. Systems-level network modeling deciphers the master regulators of phenotypic plasticity and heterogeneity in melanoma. iScience 24, 103111 (2021).
Paudel, B. B. et al. A nonquiescent ‘idling’ population state in drug-treated, BRAF-mutated melanoma. Biophys. J. 114, 1499–1511 (2018).
Udyavar, A. R. et al. Novel hybrid phenotype revealed in small cell lung cancer by a transcription factor network model that can explain tumor heterogeneity. Cancer Res. 77, 1063–1074 (2017).
Wooten, D. J. et al. Systems-level network modeling of small cell lung cancer subtypes identifies master regulators and destabilizers. PLoS Comput. Biol. 15, e1007343 (2019).
Chauhan, L., Ram, U., Hari, K. & Jolly, M. K. Topological signatures in regulatory network enable phenotypic heterogeneity in small cell lung cancer. Elife 10, e64522 (2021).
Kalluri, R. & Weinberg, R. A. The basics of epithelial–mesenchymal transition. J. Clin. Invest. 119, 1420–1428 (2009).
Hari, K., Ullanat, V., Balasubramanian, A., Gopalan, A. & Jolly, M. K. Landscape of epithelial-mesenchymal plasticity as an emergent property of coordinated teams in regulatory networks. Elife https://doi.org/10.7554/eLife.76535 (2022).
Celià-Terrassa, T. et al. Hysteresis control of epithelial-mesenchymal transition dynamics conveys a distinct program with enhanced metastatic ability. Nat. Commun. 9, 5005 (2018).
Pastushenko, I. & Blanpain, C. EMT transition states during tumor progression and metastasis. Trends Cell Biol. 29, 212–226 (2019).
Lee, J. et al. Network of mutually repressive metastasis regulators can promote cell heterogeneity and metastatic transitions. Proc. Natl Acad. Sci. USA 111, E364–E373 (2014).
Bhang, H.-E. C. et al. Studying clonal dynamics in response to cancer therapy using high-complexity barcoding. Nat. Med. 21, 440–448 (2015).
Turke, A. B. et al. Preexistence and clonal selection of MET amplification in EGFR mutant NSCLC. Cancer Cell 17, 77–88 (2010).
Oren, Y. et al. Cycling cancer persister cells arise from lineages with distinct programs. Nature 596, 576–582 (2021).
Russo, M. et al. A modified fluctuation-test framework characterizes the population dynamics and mutation rate of colorectal cancer persister cells. Nat. Genet. 54, 976–984 (2022).
Fröhlich, F., Gerosa, L., Muhlich, J. & Sorger, P. K. Mechanistic model of MAPK signaling reveals how allostery and rewiring contribute to drug resistance. Mol. Syst. Biol. 19, e10988 (2023).
Goldman, A. et al. Temporally sequenced anticancer drugs overcome adaptive resistance by targeting a vulnerable chemotherapy-induced phenotypic transition. Nat. Commun. 6, 6139 (2015).
Risom, T. et al. Differentiation-state plasticity is a targetable resistance mechanism in basal-like breast cancer. Nat. Commun. 9, 3815 (2018).
Sahoo, S. et al. A mechanistic model captures the emergence and implications of non-genetic heterogeneity and reversible drug resistance in ER+ breast cancer cells. NAR Cancer 3, zcab027 (2021).
Rehman, S. K. et al. Colorectal cancer cells enter a diapause-like DTP state to survive chemotherapy. Cell 184, 226–242.e21 (2021).
Pisco, A. O. et al. Non-Darwinian dynamics in therapy-induced cancer drug resistance. Nat. Commun. 4, 2467 (2013).
Farquhar, K. S. et al. Role of network-mediated stochasticity in mammalian drug resistance. Nat. Commun. 10, 2766 (2019).
Hayford, C. E. et al. An in vitro model of tumor heterogeneity resolves genetic, epigenetic, and stochastic sources of cell state variability. PLoS Biol. 19, e3000797 (2021).
Vipparthi, K. et al. Emergence of hybrid states of stem-like cancer cells correlates with poor prognosis in oral cancer. iScience 25, 104317 (2022).
Gillespie, D. T. Exact stochastic simulation of coupled chemical reactions. J. Phys. Chem. 81, 2340–2361 (1977).
Hari, K. et al. Identifying inhibitors of epithelial-mesenchymal plasticity using a network topology-based approach. npj Syst. Biol. Appl. 6, 15 (2020).
Nevozhay, D., Adams, R. M., Murphy, K. F., Josic, K. & Balázsi, G. Negative autoregulation linearizes the dose–response and suppresses the heterogeneity of gene expression. Proc. Natl Acad. Sci. USA 106, 5123–5128 (2009).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Spencer, S. L., Gaudet, S., Albeck, J. G., Burke, J. M. & Sorger, P. K. Non-genetic origins of cell-to-cell variability in TRAIL-induced apoptosis. Nature 459, 428–432 (2009).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Linderman, G. C. et al. Zero-preserving imputation of single-cell RNA-seq data. Nat. Commun. 13, 192 (2022).
Hie, B., Bryson, B. & Berger, B. Efficient integration of heterogeneous single-cell transcriptomes using Scanorama. Nat. Biotechnol. 37, 685–691 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Chen, W. S. et al. Uncovering axes of variation among single-cell cancer specimens. Nat. Methods 17, 302–310 (2020).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Xi, N. M. & Li, J. J. Benchmarking computational doublet-detection methods for single-cell RNA sequencing data. Cell Syst. 12, 176–194.e6 (2021).
van Dijk, D. et al. Recovering gene interactions from single-cell data using data diffusion. Cell 174, 716–729.e27 (2018).
Azizi, E. et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell 174, 1293–1308.e36 (2018).
Shaffer, S. M. et al. Memory sequencing reveals heritable single-cell gene expression programs associated with distinct cellular behaviors. Cell 182, 947–959.e17 (2020).
Karacosta, L. G. et al. Mapping lung cancer epithelial–mesenchymal transition states and trajectories with single-cell resolution. Nat. Commun. 10, 5587 (2019).
Wagner, J. et al. A single-cell atlas of the tumor and immune ecosystem of human breast cancer. Cell 177, 1330–1345.e18 (2019).
Luria, S. E. & Delbrück, M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics 28, 491–511 (1943).
Kaur, A. et al. Metastatic potential in clonal melanoma cells is driven by a rare, early-invading subpopulation. Preprint at bioRxiv https://doi.org/10.1101/2022.04.17.488591 (2022).
Mold, J. E. et al. Clonally heritable gene expression imparts a layer of diversity within cell types. Preprint at bioRxiv https://doi.org/10.1101/2022.02.14.480352 (2022).
Banerji, C. R. S. et al. Cellular network entropy as the energy potential in Waddington’s differentiation landscape. Sci. Rep. 3, 3039 (2013).
Park, Y., Lim, S., Nam, J.-W. & Kim, S. Measuring intratumor heterogeneity by network entropy using RNA-seq data. Sci. Rep. 6, 37767 (2016).
Rigau, J., Feixas, M., Sbert, M., Bardera, A. & Boada, I. Medical image segmentation based on mutual information maximization. In Medical Image Computing and Computer-Assisted Intervention 135–142 (Springer, 2004).
Margolin, A. A. et al. ARACNE: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinformatics 7, S7 (2006).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Qiu, X. et al. Mapping transcriptomic vector fields of single cells. Cell 185, 690–711.e45 (2022).
Couturier, C. P. et al. Single-cell RNA-seq reveals that glioblastoma recapitulates a normal neurodevelopmental hierarchy. Nat. Commun. 11, 3406 (2020).
Weinreb, C., Wolock, S., Tusi, B. K., Socolovsky, M. & Klein, A. M. Fundamental limits on dynamic inference from single-cell snapshots. Proc. Natl Acad. Sci. USA 115, E2467–E2476 (2018).
Chaligne, R. et al. Epigenetic encoding, heritability and plasticity of glioma transcriptional cell states. Nat. Genet. https://doi.org/10.1038/s41588-021-00927-7 (2021).
Bergen, V., Soldatov, R. A., Kharchenko, P. V. & Theis, F. J. RNA velocity—current challenges and future perspectives. Mol. Syst. Biol. 17, e10282 (2021).
Gorin, G., Fang, M., Chari, T. & Pachter, L. RNA velocity unraveled. PLoS Comput. Biol. 18, e1010492 (2022).
Zheng, S. C., Stein-O’Brien, G., Boukas, L., Goff, L. A. & Hansen, K. D. Pumping the brakes on RNA velocity—understanding and interpreting RNA velocity estimates. Preprint at bioRxiv https://doi.org/10.1101/2022.06.19.494717 (2022).
Biddy, B. A. et al. Single-cell mapping of lineage and identity in direct reprogramming. Nature 564, 219–224 (2018).
Gutierrez, C. et al. Multifunctional barcoding with ClonMapper enables high-resolution study of clonal dynamics during tumor evolution and treatment. Nat. Cancer 2, 758–772 (2021).
Umkehrer, C. et al. Isolating live cell clones from barcoded populations using CRISPRa-inducible reporters. Nat. Biotechnol. 39, 174–178 (2021).
Leighton, J., Hu, M., Sei, E., Meric-Bernstam, F. & Navin, N. E. Reconstructing mutational lineages in breast cancer by multi-patient-targeted single-cell DNA sequencing. Cell Genom. 3, 100215 (2023).
Weinreb, C., Rodriguez-Fraticelli, A., Camargo, F. D. & Klein, A. M. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science 367, eaaw3381 (2020).
McKenna, A. et al. Whole-organism lineage tracing by combinatorial and cumulative genome editing. Science 353, aaf7907 (2016).
Jiang, C. L. et al. Cell type determination for cardiac differentiation occurs soon after seeding of human-induced pluripotent stem cells. Genome Biol. 23, 90 (2022).
Weinreb, C. & Klein, A. M. Lineage reconstruction from clonal correlations. Proc. Natl Acad. Sci. USA 117, 17041–17048 (2020).
Richman, L. P., Goyal, Y., Jiang, C. L. & Raj, A. ClonoCluster: a method for using clonal origin to inform transcriptome clustering. Cell Genom. 3, 100247 (2023).
Wang, S.-W., Herriges, M. J., Hurley, K., Kotton, D. N. & Klein, A. M. CoSpar identifies early cell fate biases from single-cell transcriptomic and lineage information. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01209-1 (2022).
Wagner, D. E. & Klein, A. M. Lineage tracing meets single-cell omics: opportunities and challenges. Nat. Rev. Genet. 21, 410–427 (2020).
Gerosa, L. et al. Receptor-driven ERK pulses reconfigure MAPK signaling and enable persistence of drug-adapted BRAF-mutant melanoma cells. Cell Syst. 11, 478–494.e9 (2020).
Yang, C., Tian, C., Hoffman, T. E., Jacobsen, N. K. & Spencer, S. L. Melanoma subpopulations that rapidly escape MAPK pathway inhibition incur DNA damage and rely on stress signalling. Nat. Commun. 12, 1747 (2021).
Stichel, D. et al. An individual-based model for collective cancer cell migration explains speed dynamics and phenotype variability in response to growth factors. npj Syst. Biol. Appl. 3, 5 (2017).
Meyer, M. et al. Profiling the non-genetic origins of cancer drug resistance with a single-cell functional genomics approach using predictive cell dynamics. Cell Syst. 11, 367–374.e5 (2020).
You, L. et al. Linking the genotypes and phenotypes of cancer cells in heterogenous populations via real-time optical tagging and image analysis. Nat. Biomed. Eng. https://doi.org/10.1038/s41551-022-00853-x (2022).
Bao, F. et al. Integrative spatial analysis of cell morphologies and transcriptional states with MUSE. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01251-z (2022).
Tian, C., Yang, C. & Spencer, S. L. EllipTrack: a global-local cell-tracking pipeline for 2D fluorescence time-lapse microscopy. Cell Rep. 32, 107984 (2020).
Bagheri, N., Carpenter, A. E., Lundberg, E., Plant, A. L. & Horwitz, R. The new era of quantitative cell imaging-challenges and opportunities. Mol. Cell 82, 241–247 (2022).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Su, Y. et al. Multi-omic single-cell snapshots reveal multiple independent trajectories to drug tolerance in a melanoma cell line. Nat. Commun. 11, 2345 (2020).
Sammut, S.-J. et al. Multi-omic machine learning predictor of breast cancer therapy response. Nature 601, 623–629 (2022).
Rao, A., Barkley, D., França, G. S. & Yanai, I. Exploring tissue architecture using spatial transcriptomics. Nature 596, 211–220 (2021).
Atta, L. & Fan, J. Computational challenges and opportunities in spatially resolved transcriptomic data analysis. Nat. Commun. 12, 5283 (2021).
Nam, A. S., Chaligne, R. & Landau, D. A. Integrating genetic and non-genetic determinants of cancer evolution by single-cell multi-omics. Nat. Rev. Genet. 22, 3–18 (2021).
Salgia, R. & Kulkarni, P. The genetic/non-genetic duality of drug ‘resistance’ in cancer. Trends Cancer Res. 4, 110–118 (2018).
Liang, W. et al. Development and validation of a nomogram for predicting survival in patients with resected non-small-cell lung cancer. J. Clin. Oncol. 33, 861–869 (2015).
Karakiewicz, P. I. et al. Multi-institutional validation of a new renal cancer-specific survival nomogram. J. Clin. Oncol. 25, 1316–1322 (2007).
Huang, C., Zhao, J. & Zhu, Z. Prognostic nomogram of prognosis-related genes and clinicopathological characteristics to predict the 5-year survival rate of colon cancer patients. Front Surg. 8, 681721 (2021).
Phillips, R. Theory in biology: Fig. 1 or Figure 7? Trends Cell Biol. 25, 723–729 (2015).
Theorists and experimentalists must join forces. Nat. Comput. Sci. 1, 299–299 (2021).
Cassidy, T., Nichol, D., Robertson-Tessi, M., Craig, M. & Anderson, A. R. A. The role of memory in non-genetic inheritance and its impact on cancer treatment resistance. PLoS Comput. Biol. 17, e1009348 (2021).
Mellis, I. A. & Raj, A. Half dozen of one, six billion of the other: what can small- and large-scale molecular systems biology learn from one another? Genome Res. 25, 1466–1472 (2015).
Bauer, A. L., Jackson, T. L. & Jiang, Y. A cell-based model exhibiting branching and anastomosis during tumor-induced angiogenesis. Biophys. J. 92, 3105–3121 (2007).
Zhang, L., Strouthos, C. G., Wang, Z. & Deisboeck, T. S. Simulating brain tumor heterogeneity with a multiscale agent-based model: linking molecular signatures, phenotypes and expansion rate. Math. Comput. Model. 49, 307–319 (2009).
Saez-Rodriguez, J. et al. Comparing signaling networks between normal and transformed hepatocytes using discrete logical models. Cancer Res. 71, 5400–5411 (2011).
Macklin, P., Edgerton, M. E., Thompson, A. M. & Cristini, V. Patient-calibrated agent-based modelling of ductal carcinoma in situ (DCIS): from microscopic measurements to macroscopic predictions of clinical progression. J. Theor. Biol. 301, 122–140 (2012).
Fumiã, H. F. & Martins, M. L. Boolean network model for cancer pathways: predicting carcinogenesis and targeted therapy outcomes. PLoS ONE 8, e69008 (2013).
Steinway, S. N. et al. Network modeling of TGFβ signaling in hepatocellular carcinoma epithelial-to-mesenchymal transition reveals joint sonic hedgehog and Wnt pathway activation. Cancer Res. 74, 5963–5977 (2014).
Wang, Z., Zhang, L., Sagotsky, J. & Deisboeck, T. S. Simulating non-small cell lung cancer with a multiscale agent-based model. Theor. Biol. Med. Model. 4, 50 (2007).
Andasari, V., Roper, R. T., Swat, M. H. & Chaplain, M. A. J. Integrating intracellular dynamics using CompuCell3D and Bionetsolver: applications to multiscale modelling of cancer cell growth and invasion. PLoS ONE 7, e33726 (2012).
Swat, M. H. et al. in Methods in Cell Biology Vol. 110 (eds Asthagiri, A. R. & Arkin, A. P.) 325–366 (Academic Press, 2012).
Lu, M., Jolly, M. K., Levine, H., Onuchic, J. N. & Ben-Jacob, E. MicroRNA-based regulation of epithelial-hybrid-mesenchymal fate determination. Proc. Natl Acad. Sci. USA 110, 18144–18149 (2013).
Wang, W. et al. Dynamics between cancer cell subpopulations reveals a model coordinating with both hierarchical and stochastic concepts. PLoS ONE 9, e84654 (2014).
Jain, P., Bhatia, S., Thompson, E. W. & Jolly, M. K. Population dynamics of epithelial–mesenchymal heterogeneity in cancer cells. Biomolecules 12, 348 (2022).
Di Filippo, M. et al. Zooming-in on cancer metabolic rewiring with tissue specific constraint-based models. Comput. Biol. Chem. 62, 60–69 (2016).
Ghaffarizadeh, A., Heiland, R., Friedman, S. H., Mumenthaler, S. M. & Macklin, P. PhysiCell: an open source physics-based cell simulator for 3-D multicellular systems. PLoS Comput. Biol. 14, e1005991 (2018).
Damiani, C. et al. A metabolic core model elucidates how enhanced utilization of glucose and glutamine, with enhanced glutamine-dependent lactate production, promotes cancer cell growth: the WarburQ effect. PLoS Comput. Biol. 13, e1005758 (2017).
Gatto, F., Miess, H., Schulze, A. & Nielsen, J. Flux balance analysis predicts essential genes in clear cell renal cell carcinoma metabolism. Sci. Rep. 5, 10738 (2015).
Zhou, J. X., Pisco, A. O., Qian, H. & Huang, S. Nonequilibrium population dynamics of phenotype conversion of cancer cells. PLoS ONE 9, e110714 (2014).
Angel, P., Hattori, K., Smeal, T. & Karin, M. The jun proto-oncogene is positively autoregulated by its product, Jun/AP-1. Cell 55, 875–885 (1988).
Castles, C. G., Oesterreich, S., Hansen, R. & Fuqua, S. A. Auto-regulation of the estrogen receptor promoter. J. Steroid Biochem. Mol. Biol. 62, 155–163 (1997).
Gardner, T. S., Cantor, C. R. & Collins, J. J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Cherry, J. L. & Adler, F. R. How to make a biological switch. J. Theor. Biol. 203, 117–133 (2000).
Burk, U. et al. A reciprocal repression between ZEB1 and members of the miR-200 family promotes EMT and invasion in cancer cells. EMBO Rep. 9, 582–589 (2008).
Kim, Y., Roh, S., Lawler, S. & Friedman, A. miR451 and AMPK mutual antagonism in glioma cell migration and proliferation: a mathematical model. PLoS ONE 6, e28293 (2011).
Lu, M. et al. Tristability in cancer-associated microRNA-TF chimera toggle switch. J. Phys. Chem. B 117, 13164–13174 (2013).
Novák, B. & Tyson, J. J. Design principles of biochemical oscillators. Nat. Rev. Mol. Cell Biol. 9, 981–991 (2008).
Ma, L. et al. A plausible model for the digital response of p53 to DNA damage. Proc. Natl Acad. Sci. USA 102, 14266–14271 (2005).
Purvis, J. E. et al. p53 dynamics control cell fate. Science 336, 1440–1444 (2012).
Acknowledgements
We thank the members of the Goyal and Jolly labs for helpful discussions. We thank K. Kiani and I. Mellis for their critical reading of the manuscript. The Goyal lab thanks R. Valadka at Northwestern University for his prompt and unwavering support in setting up the lab space. M.P. was supported by grants to Y.G. including the Career Award at the Scientific Interface from BWF (1020614.01) and start-up funds from Northwestern University, and KVPY Fellowship (Department of Science and Technology, Government of India). E.H. acknowledges support from the Career Award at the Scientific Interface from BWF (1020614.01) and Northwestern University’s Biomedical Engineering Department for the BME Summer Undergraduate Research Grant Award (SURA). E.H. thanks E. Hojel and M. Hojel for the support and encouragement to pursue her passions. M.K.J. acknowledges support from Ramanujan Fellowship awarded by the Science and Engineering Research Board, Department of Science and Technology, Government of India (SB/S2/RJN-049/2018), and from the InfoSys Foundation, Bangalore. Y.G. acknowledges support from the Career Award at the Scientific Interface from BWF (1020614.01), start-up funds from Northwestern University, and a grant (10063150.01) from Research Catalyst Program from the McCormick School of Engineering at Northwestern University.
Author information
Authors and Affiliations
Contributions
M.K.J. and Y.G. conceived the project, wrote the first draft of the manuscript and contributed to figure design. M.P. revised the initial draft, contributed to figure design, and prepared the boxes and the table. E.H. contributed to the figure, box and table design and literature survey. All authors contributed to revisions.
Corresponding authors
Ethics declarations
Competing interests
Y.G. received consultancy fee from the Schmidt Science Fellows and the Rhodes Trust. The other authors declare no competing interests.
Peer review
Peer review information
Nature Computational Science thanks Gábor Balázsi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Ananya Rastogi and Kaitlin McCardle, in collaboration with the Nature Computational Science team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pillai, M., Hojel, E., Jolly, M.K. et al. Unraveling non-genetic heterogeneity in cancer with dynamical models and computational tools. Nat Comput Sci 3, 301–313 (2023). https://doi.org/10.1038/s43588-023-00427-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-023-00427-0