Abstract
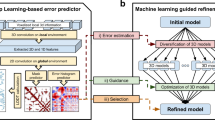
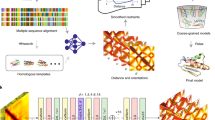
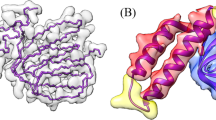
Protein model refinement is the last step applied to improve the quality of a predicted protein model. Currently, the most successful refinement methods rely on extensive conformational sampling and thus take hours or days to refine even a single protein model. Here, we propose a fast and effective model refinement method that applies graph neural networks (GNNs) to predict a refined inter-atom distance probability distribution from an initial model and then rebuilds three-dimensional models from the predicted distance distribution. Tested on the Critical Assessment of Structure Prediction refinement targets, our method has an accuracy that is comparable to those of two leading human groups (FEIG and BAKER), but runs substantially faster. Our method may refine one protein model within ~11 min on one CPU, whereas BAKER needs ~30 h on 60 CPUs and FEIG needs ~16 h on one GPU. Finally, our study shows that GNN outperforms ResNet (convolutional residual neural networks) for model refinement when very limited conformational sampling is allowed.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Our in-house data are available at http://raptorx.uchicago.edu/download/. Click on this link and fill in your name, email address and organization name to obtain a data link, through which you will find a text file 0README.Data4GNNRefine.txt that specifies the names of the data files to be downloaded. The data are also available at Zenodo38. The DeepAccNet data are available at https://github.com/hiranumn/DeepAccNet. The CASP13 and CASP14 models for refinement are available at https://predictioncenter.org/. The CAMEO models are available at https://www.cameo3d.org/modeling/. The CAMEO dataset includes 208 starting models for all the CAMEO hard targets released between 1 May 2018 and 1 May 2020. We keep only the targets with sequence length in [50, 500] and native structures containing at least 80% of sequence residues. Following CASPs, we select the best-predicted models (in terms of GDT-HA) for each target as the starting models, and only keep the starting models with lDDT > 50. For the CASP13 FM (free-modeling) dataset, there are 28 test targets corresponding to 32 official FM domains. For each target we build ~150 decoys as its starting models using our in-house template-free modeling software RaptorX-Contact. Source data are provided with this paper.
Code availability
The source code is available at Code Ocean39.
References
Wang, S., Sun, S., Li, Z., Zhang, R. & Xu, J. Accurate de novo prediction of protein contact map by ultra-deep learning model. PLoS Comput. Biol. 13, e1005324 (2017).
Xu, J. Distance-based protein folding powered by deep learning. Proc. Natl. Acad. Sci. USA 116, 16856–16865 (2019).
Senior, A. W. et al. Improved protein structure prediction using potentials from deep learning. Nature 577, 706–710 (2020).
Yang, J. et al. Improved protein structure prediction using predicted interresidue orientations. Proc. Natl Acad. Sci. USA 117, 1496–1503 (2020).
Read, R. J., Sammito, M. D., Kryshtafovych, A. & Croll, T. I. Evaluation of model refinement in CASP13. Proteins Struct. Funct. Bioinf. 87, 1249–1262 (2019).
Heo, L., Arbour, C. F. & Feig, M. Driven to near-experimental accuracy by refinement via molecular dynamics simulations. Proteins Struct. Funct. Bioinf. 87, 1263–1275 (2019).
Park, H. et al. High-accuracy refinement using Rosetta in CASP13. Proteins Struct. Funct. Bioinf. 87, 1276–1282 (2019).
Xu, D. & Zhang, Y. Improving the physical realism and structural accuracy of protein models by a two-step atomic-level energy minimization. Biophys. J. 101, 2525–2534 (2011).
Heo, L., Park, H. & Seok, C. GalaxyRefine: protein structure refinement driven by side-chain repacking. Nucleic Acids Res. 41, W384–W388 (2013).
Bhattacharya, D., Nowotny, J., Cao, R. & Cheng, J. 3Drefine: an interactive web server for efficient protein structure refinement. Nucleic Acids Res. 44, W406–W409 (2016).
Bhattacharya, D. refineD: improved protein structure refinement using machine learning based restrained relaxation. Bioinformatics 35, 3320–3328 (2019).
Lee, G. R., Won, J., Heo, L. & Seok, C. GalaxyRefine2: simultaneous refinement of inaccurate local regions and overall protein structure. Nucleic Acids Res. 47, W451–W455 (2019).
Hiranuma, N. et al. Improved protein structure refinement guided by deep learning based accuracy estimation. Nat. Commun. 12, 1340 (2021).
Mirjalili, V., Noyes, K. & Feig, M. Physics-based protein structure refinement through multiple molecular dynamics trajectories and structure averaging. Proteins Struct. Funct. Bioinf. 82, 196–207 (2014).
Sanyal, S., Anishchenko, I., Dagar, A., Baker, D. & Talukdar, P. ProteinGCN: protein model quality assessment using graph convolutional networks. Preprint at bioRxiv https://doi.org/10.1101/2020.04.06.028266 (2020).
Baldassarre, F., Hurtado, D. M., Elofsson, A. & Azizpour, H. GraphQA: protein model quality assessment using graph convolutional networks. Bioinformatics 37, 360–366 (2021).
Chaudhury, S., Lyskov, S. & Gray, J. J. PyRosetta: a script-based interface for implementing molecular modeling algorithms using Rosetta. Bioinformatics 26, 689–691 (2010).
Conway, P., Tyka, M. D., DiMaio, F., Konerding, D. E. & Baker, D. Relaxation of backbone bond geometry improves protein energy landscape modeling. Protein Sci. 23, 47–55 (2014).
Mariani, V., Biasini, M., Barbato, A. & Schwede, T. lDDT: a local superposition-free score for comparing protein structures and models using distance difference tests. Bioinformatics 29, 2722–2728 (2013).
Critical Assessment of Techniques for Protein Structure Prediction Thirteenth Round—Abstract Book (Prediction Center, 2018); https://predictioncenter.org/casp13/doc/CASP13_Abstracts.pdf
Critical Assessment of Techniques for Protein Structure Prediction Fourteenth Round—Abstract Book (Prediction Center, 2020); https://predictioncenter.org/casp14/doc/CASP14_Abstracts.pdf
Heo, L., Arbour, C. F., Janson, G. & Feig, M. Improved sampling strategies for protein model refinement based on molecular dynamics simulation. J. Chem. Theory Comput. 17, 1931–1943 (2021).
Shuid, A. N., Kempster, R. & McGuffin, L. J. ReFOLD: a server for the refinement of 3D protein models guided by accurate quality estimates. Nucleic Acids Res. 45, W422–W428 (2017).
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983).
Igashov, I., Olechnovič, L., Kadukova, M., Venclovas, Č. & Grudinin, S. VoroCNN: deep convolutional neural network built on 3D Voronoi tessellation of protein structures. Bioinformatics https://doi.org/10.1093/bioinformatics/btab118 (2021).
Zhang, J. & Zhang, Y. A novel side-chain orientation dependent potential derived from random-walk reference state for protein fold selection and structure prediction. PLoS ONE 5, e15386 (2010).
Won, J., Baek, M., Monastyrskyy, B., Kryshtafovych, A. & Seok, C. Assessment of protein model structure accuracy estimation in CASP13: challenges in the era of deep learning. Proteins Struct. Funct. Bioinf. 87, 1351–1360 (2019).
Rives, A. et al. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc. Natl Acad. Sci. USA 118, e2016239118 (2021).
Rao, R. et al. MSA transformer. Preprint at bioRxiv https://doi.org/10.1101/2021.02.12.430858 (2021).
Dawson, N. L. et al. CATH: an expanded resource to predict protein function through structure and sequence. Nucleic Acids Res. 45, D289–D295 (2017).
Wang, G. & Dunbrack, R. L. PISCES: a protein sequence culling server. Bioinformatics 19, 1589–1591 (2003).
Thomas, N. et al. Tensor field networks: rotation- and translation-equivariant neural networks for 3D point clouds. Preprint at https://arxiv.org/pdf/1802.08219.pdf (2018).
Huang, B. & Carley, K. M. Residual or gate? Towards deeper graph neural networks for inductive graph representation learning. Preprint at https://arxiv.org/pdf/1904.08035.pdf (2019).
Wang, M. et al. Deep Graph Library: a graph-centric, highly-performant package for graph neural networks. Preprint at https://arxiv.org/pdf/1909.01315.pdf (2020).
Paszke, A. et al. PyTorch: an imperative style, high-performance deep learning library. In Advances in Neural Information Processing Systems 32 (eds Wallach, H. et al.) 8026–8037 (Curran Associates, 2019).
Zhou, H. & Zhou, Y. Distance-scaled, finite ideal-gas reference state improves structure-derived potentials of mean force for structure selection and stability prediction. Protein Sci. 11, 2714–2726 (2002).
Park, H. et al. Simultaneous optimization of biomolecular energy functions on features from small molecules and macromolecules. J. Chem. Theory Comput. 12, 6201–6212 (2016).
Xu, J. Data for protein model refinement and model quality assessment. Zenodo https://doi.org/10.5281/zenodo.4635356
Jing, X. GNNRefine: fast and effective protein model refinement by deep graph neural networks (Code Ocean, 2021); https://doi.org/10.24433/CO.8813669.v1
Acknowledgements
We thank D. Baker’s team, including H. Park, who provided us with the DeepAccNet training data and helpful comments on our manuscript. We are also grateful to L. Heo for explaining FEIG and FEIG-S to us. This work is supported by National Institutes of Health grant no. R01GM089753 to J.X. and National Science Foundation grant no. DBI1564955 to J.X. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
X.J. conceived the research, developed the GNNRefine and carried out the benchmarking experiments. J.X. built the in-house training data and guided the research. X.J. and J.X. analyzed the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Computational Science thanks Hahnbeom Park, Lim Heo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Handling editor: Jie Pan, in collaboration with the Nature Computational Science team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Extended Data Fig. 1 Quality improvement by different methods on the CASP13 refinement targets.
Boxplot of the distribution of ΔGDT-HA, ΔGDT-TS, and ΔlDDT on the CASP13 refinement targets. The five lines in each boxplot from top to bottom in turn mean: Maximum (Q3 + 1.5IQR), Third quartile (Q3, 75th percentile), Median (50th percentile), First quartile (Q1, 25th percentile), and Minimum (Q1-1.5IQR), where IQR is Q3-Q1. The precision is 2.
Extended Data Fig. 2 Quality improvement by different methods on the CASP14 refinement targets.
Box plot of the distribution of ΔGDT-HA, ΔGDT-TS, and ΔlDDT on the CASP14 refinement targets.The five lines in each boxplot from top to bottom in turn mean: Maximum (Q3 + 1.5IQR), Third quartile (Q3, 75th percentile), Median (50th percentile), First quartile (Q1, 25th percentile), and Minimum (Q1-1.5IQR), where IQR is Q3-Q1. The precision is 2.
Supplementary information
Supplementary Information
Supplementary sections 1–12, Figs. 1–6 and Tables 1–23.
Source data
Source Data Fig. 2
The PDB structure files for Fig. 2.
Source Data Fig. 3
The running time data for GNNRefine and DeepAccNet.
Source Data Extended Data Fig. 1
The source data used to draw the boxplot of Extended Data Fig. 1.
Source Data Extended Data Fig. 2
The source data used to draw the boxplot of Extended Data Fig. 2.
Rights and permissions
About this article
Cite this article
Jing, X., Xu, J. Fast and effective protein model refinement using deep graph neural networks. Nat Comput Sci 1, 462–469 (2021). https://doi.org/10.1038/s43588-021-00098-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43588-021-00098-9
This article is cited by
-
ScanNet: an interpretable geometric deep learning model for structure-based protein binding site prediction
Nature Methods (2022)
-
Rapid protein model refinement by deep learning
Nature Computational Science (2021)